Genome Browsers UCSC Santa Cruz California and Ensembl

Genome Browsers UCSC (Santa Cruz, California) and Ensembl (EBI, UK) http: //genome. ucsc. edu/ http: //www. ensembl. org/

Eukaryotic Genomes: Not only collections of genes • Protein coding genes • RNA genes (r. RNA, sno. RNA, mi. RNA, t. RNA) • Structural DNA (centromeres, telomeres) • Regulation-related sequences (promoters, enhancers, silencers, insulators) • Parasite sequences (transposons) • Pseudogenes (non-functional gene-like sequences) • Simple sequence repeats
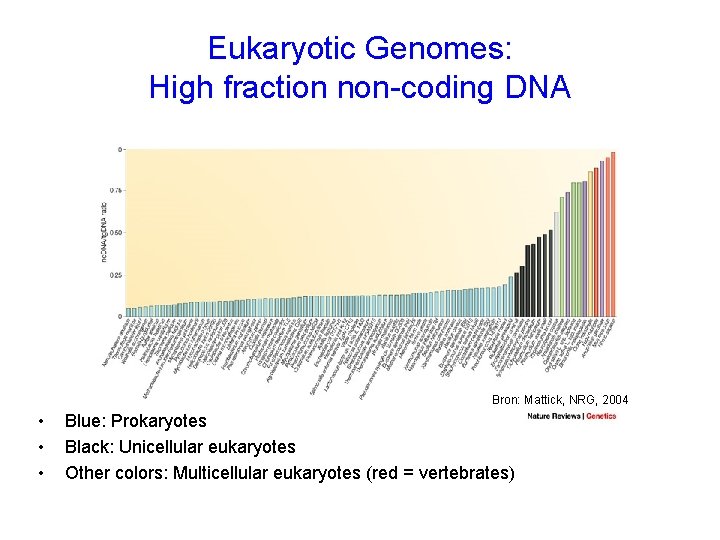
Eukaryotic Genomes: High fraction non-coding DNA Bron: Mattick, NRG, 2004 • • • Blue: Prokaryotes Black: Unicellular eukaryotes Other colors: Multicellular eukaryotes (red = vertebrates)

Human Genome • 3 billion basepairs (3 Gb) • 22 chromosome pairs + X en Y chromosomes • Chromosome length varies from ~50 Mb to ~250 Mb • About 22000 protein-coding genes – compare with ~14000 for fruitfly en ~19000 for Nematode C. elegans

Human genome Bron: Molecular Biology of the Cell (4 th edition) (Alberts et al. , 2002) • • Only 1. 2% codes for proteins, 3. 5 -5% is under selection Long introns, short exons Large spaces between genes More than half exists of repetitive DNA
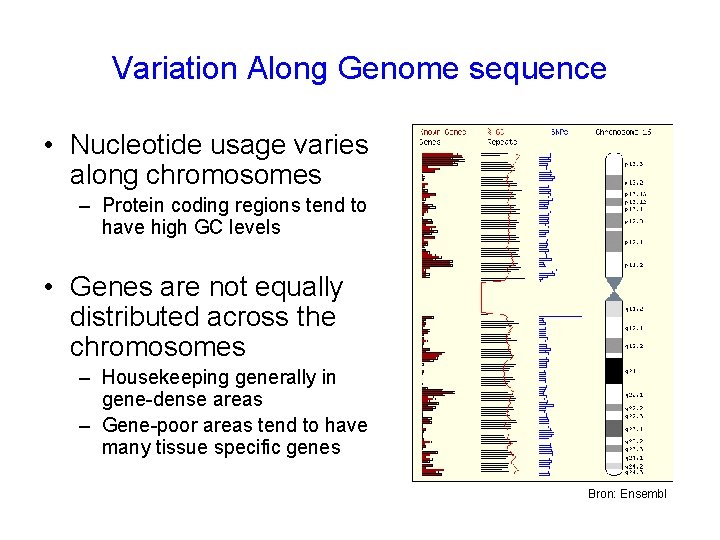
Variation Along Genome sequence • Nucleotide usage varies along chromosomes – Protein coding regions tend to have high GC levels • Genes are not equally distributed across the chromosomes – Housekeeping generally in gene-dense areas – Gene-poor areas tend to have many tissue specific genes Bron: Ensembl
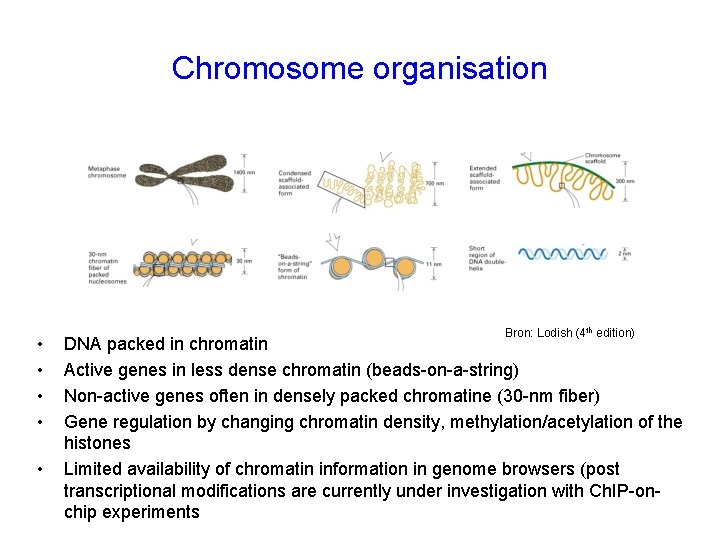
Chromosome organisation • • • Bron: Lodish (4 th edition) DNA packed in chromatin Active genes in less dense chromatin (beads-on-a-string) Non-active genes often in densely packed chromatine (30 -nm fiber) Gene regulation by changing chromatin density, methylation/acetylation of the histones Limited availability of chromatin information in genome browsers (post transcriptional modifications are currently under investigation with Ch. IP-onchip experiments

Genome browsers UCSC NCBI http: //genome. ucsc. edu/ Ensembl http: //www. ensembl. org/

Genome Browsing With the UCSC Genome Browser http: //genome. ucsc. edu/
UCSC Genome browser
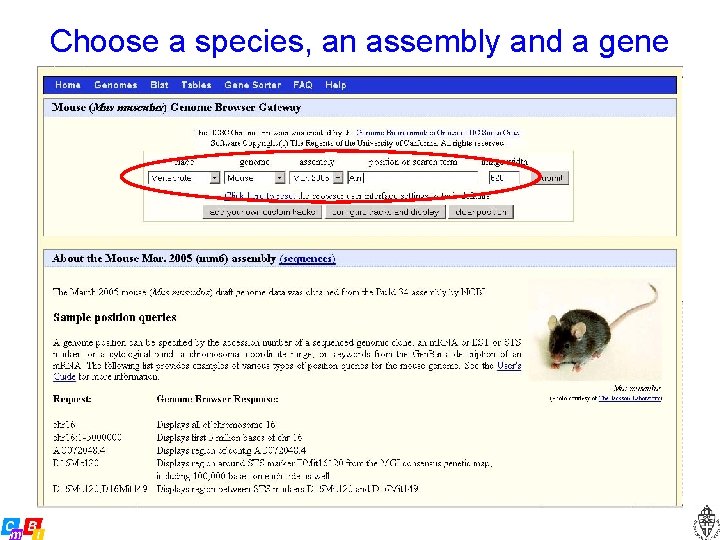
Choose a species, an assembly and a gene
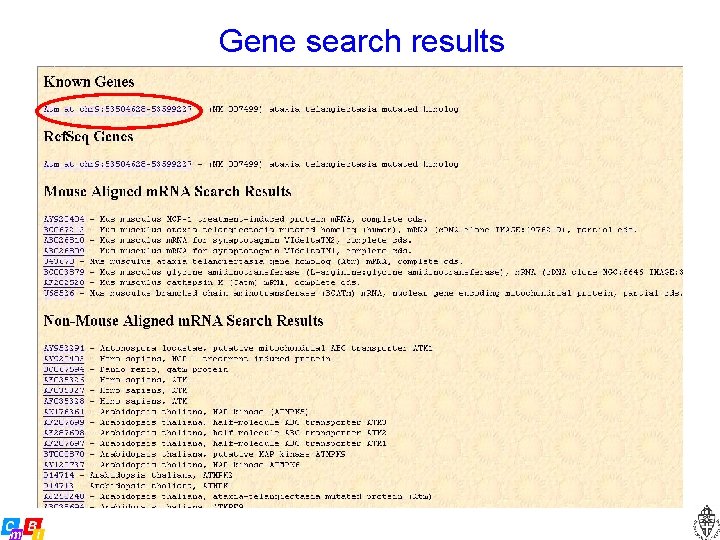
Gene search results
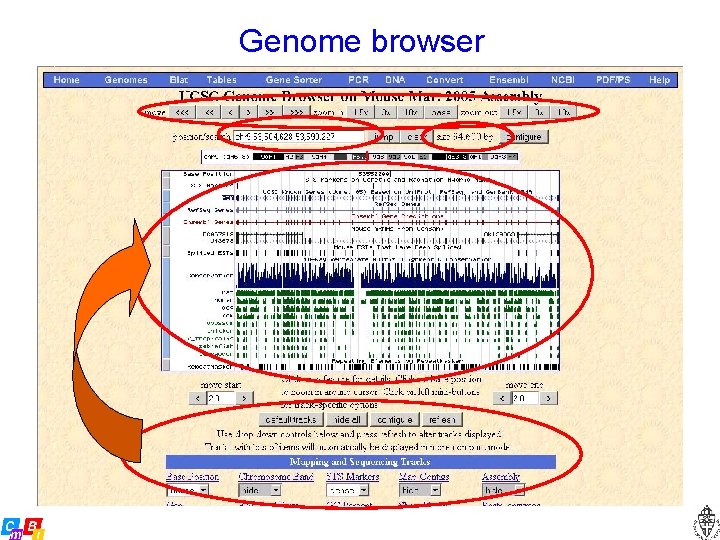
Genome browser
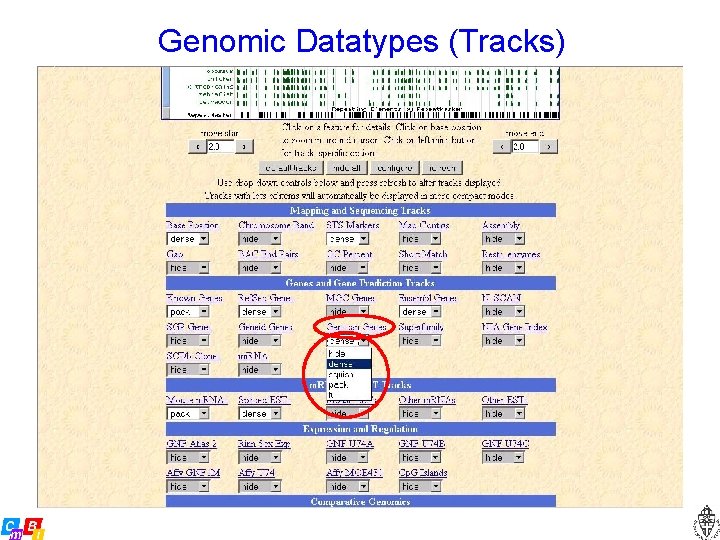
Genomic Datatypes (Tracks)

Transcription data rather complicated
Browser → Gene record
Gene record
Gene record (2)
Gene record (3)
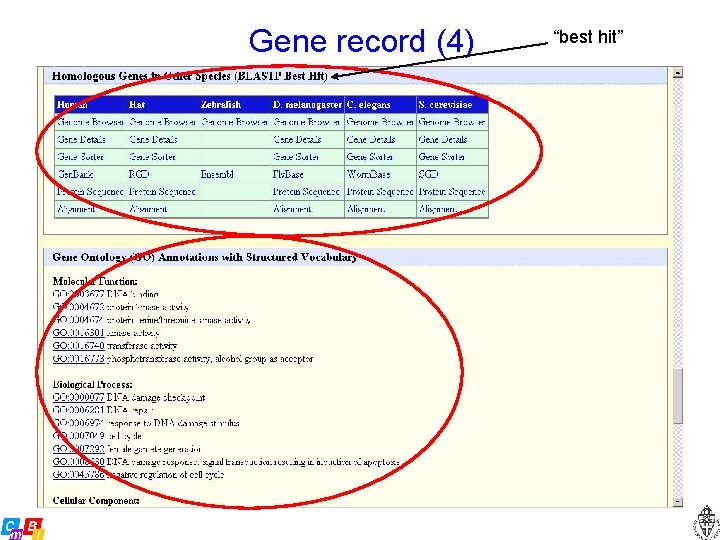
Gene record (4) “best hit”
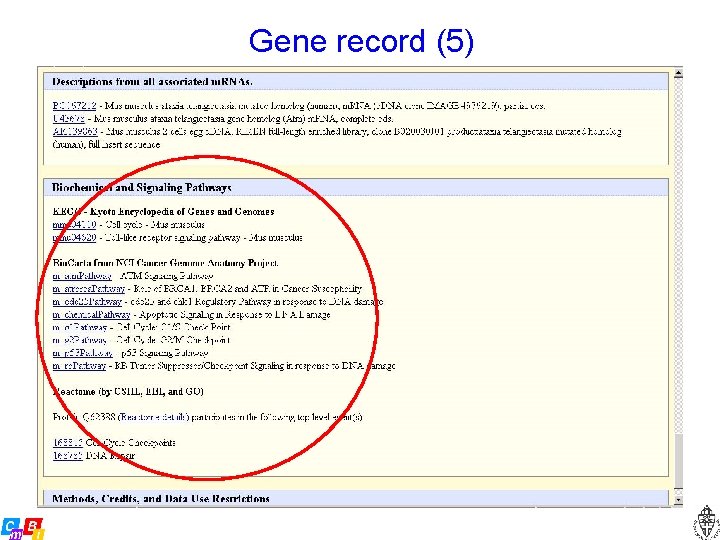
Gene record (5)

Genomic elements • Genome browsers can be used to examine other things – Genomic sequence conservation – Pseudogenes – Duplications en deletions of pieces chromosome (Copy Number Variations, CNVs)

Genomic Sequence Conservation • Not only protein coding parts are conserved in evolution • Conserved non-coding genomic sequences can be involved in gene regulation (enhancers, silencers, insulators) • With the UCSC browser one can examine genomic conservation

Genomic Conservation (UCSC)

Pseudogenes • Pseudogenes “look” like (are homologous to) proteincoding genes, but are non-functional • Two types: – Unprocessed pseudogenes (loss of function) – Processed pseudogenes (m. RNAs that are retrotranscribed onto the genome they miss introns and sometimes have a poly. A) • The UCSC contains various databases of pseudogenes: – Yale pseudogenes (both types pseudogenes) – Vega pseudogenes (both types pseudogenes) – Retroposed genes (only processed pseudogenes)

Pseudogenes (UCSC)

Copy Number Variation • People do not only vary at the nucleotide level (SNPs); short pieces genome can be present in varying number of copies (Copy Number Polymorphisms (CNPs) or Copy Number Variants (CNVs) • When there are genes in the CNV areas, this can lead to variations in the number of gene copies between individuals • With the UCSC browser CNVs can be examined

Copy Number Variation (UCSC)
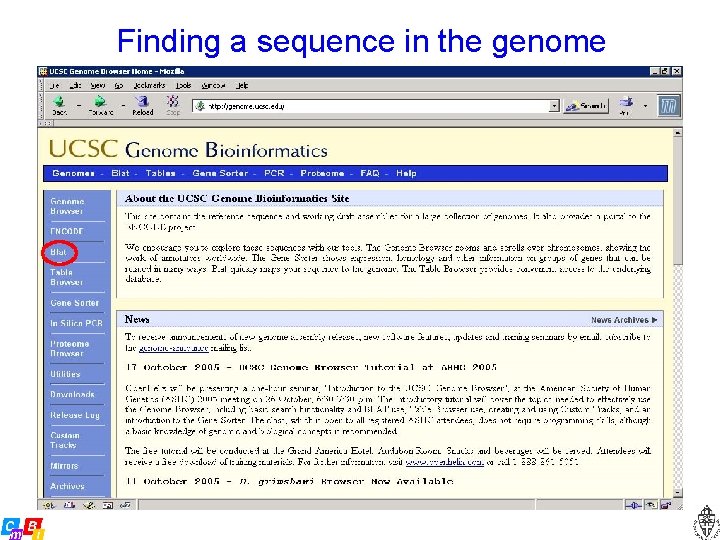
Finding a sequence in the genome
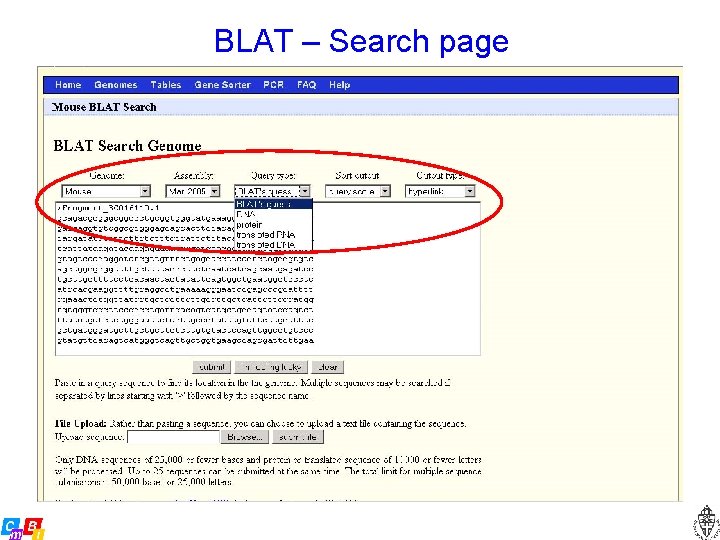
BLAT – Search page
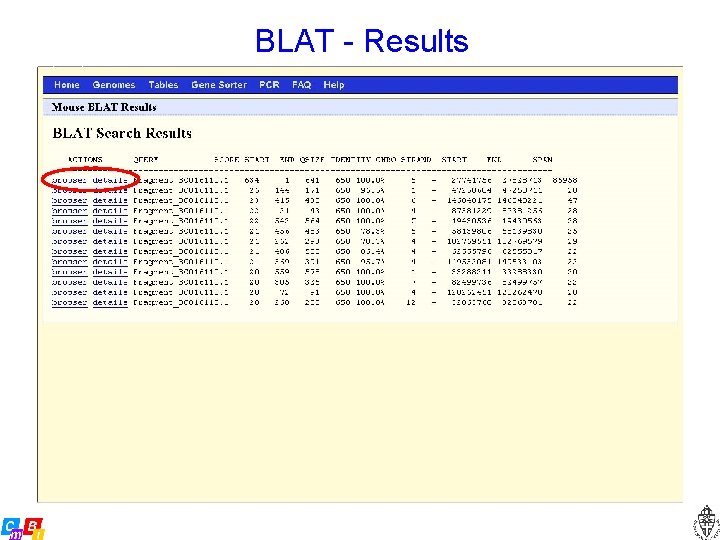
BLAT - Results
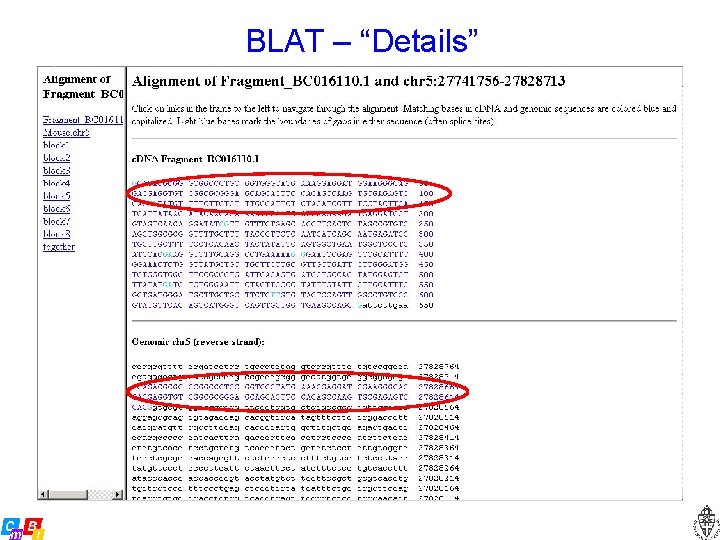
BLAT – “Details”

BLAT – “Browser”

Genome browsers UCSC http: //genome. ucsc. edu/ Ensembl http: //www. ensembl. org/
Genome Browsing With the Ensembl Genome browser http: //www. ensembl. org/
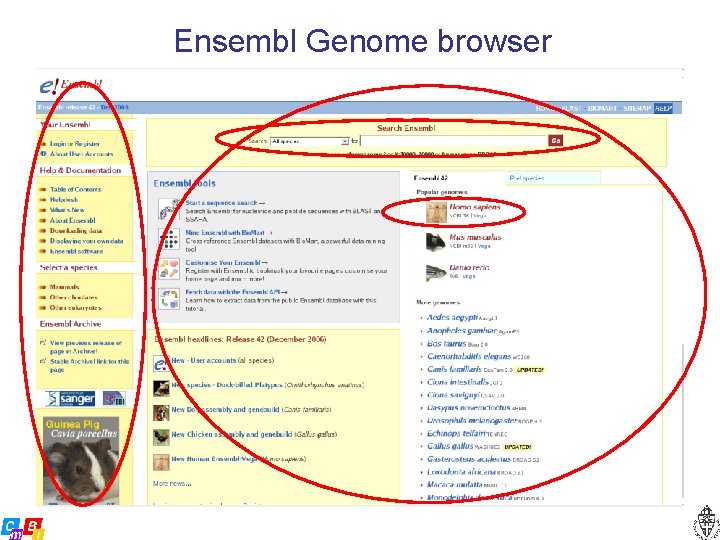
Ensembl Genome browser
Het Human Genome
Map. View – Overview chromosome

Contig. View – Zooming in (compare UCSD)

Contig. View (2)
Gene. View – Gene record

Trans. View - m. RNA Transcript
Trans. View - m. RNA Transcript (2)
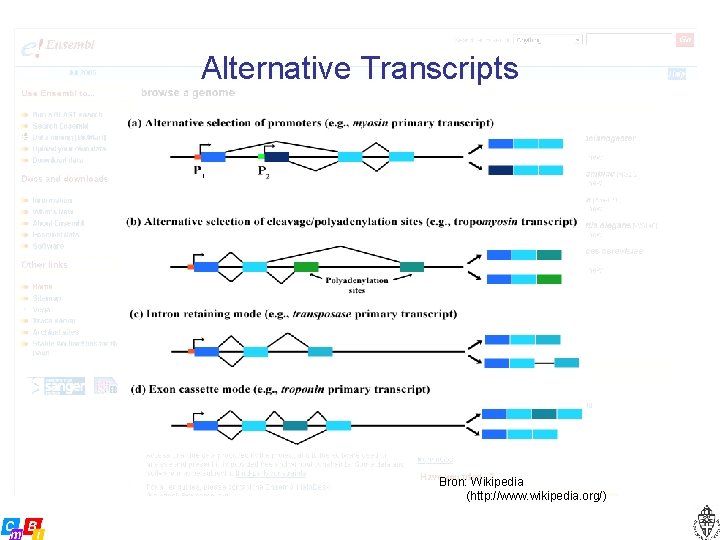
Alternative Transcripts Bron: Wikipedia (http: //www. wikipedia. org/)

Gene. View - Show Alternative Transcripts

Gene. Splice. View - Alternative Transcripts

Single Nucleotide Polymorphisms (SNPs) • Sequence variations within a species • Similar to mutations, but are simultaneously present in the population, and generaly have little effect • Are being used as genetic markers (a genetic disease is e. g. associated with a SNP) • ENSEMBL offers a nice SNP view
Gene. View - Show SNPs
Gene. SNPView - SNPs
Gene. View - Show Protein
Prot. View - Protein
Prot. View - Protein Sequence

Prot. View – Search proteins with the same domains

Domain. View – Proteins with a certain domain (Interpro = SMART + PFAM + others)
Prot. View - Find Proteins In the Same Protein Family

Family. View – Alignments of homologous proteins
Finding Human Genes

Finding a human gene (2)

Blast
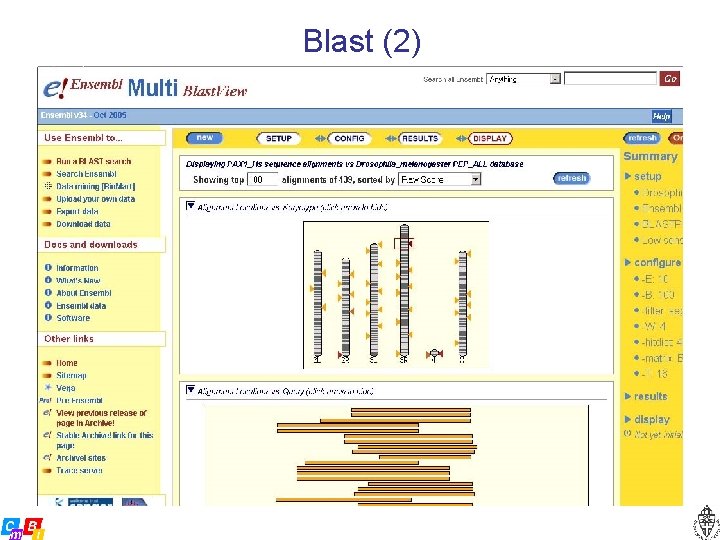
Blast (2)

UCSC vs Ensembl: Which is better ? • They more or less contain the same information • UCSC is a bit easier in use • Ensembl gives more detailed information and more flexible data export • Other small differences in data (e. g. UCSC has more extensive genomic conservation data) • Whatever your are familiar with !!
- Slides: 61