HDF Basics Tools Dataset Datatypes 11252020 1 Tutorial

HDF Basics Tools, Dataset, Datatypes 11/25/2020 1

Tutorial Topics • Tools • Groups and Links • Datasets • Partial I/O • Storage properties • Filters and compression in HDF 5 11/25/2020 2

HDF 5 File An HDF 5 file is a container that holds data objects. 11/25/2020 lat | lon | temp ----|----12 | 23 Se|r. Exper 3. 1 D ial im Nu ent Co ate| 15 | 24 m N nf : 3/1 4. 2 ig 3 be ote ur /0 r: s: a 9 99 17 | 21 |tion 3. 6 37 : S tan 89 da 3 rd 3 20
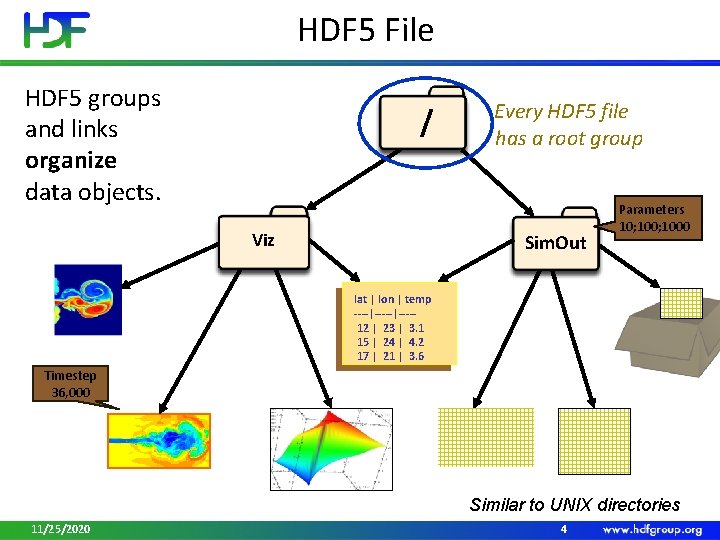
HDF 5 File HDF 5 groups and links organize data objects. / Viz Every HDF 5 file has a root group Sim. Out Parameters 10; 1000 lat | lon | temp ----|----12 | 23 | 3. 1 15 | 24 | 4. 2 17 | 21 | 3. 6 Timestep 36, 000 Similar to UNIX directories 11/25/2020 4

TOOLS 11/25/2020 5

Tools • HDFView • Command-line tools • • h 5 dump, h 5 ls h 5 diff h 5 repack h 5 copy HDF 5 for Python or h 5 py http: //www. h 5 py. org/ MATLAB, Mathematica, IDL R package For more information visit https: //support. hdfgroup. org/products/hdf 5_tools/ 11/25/2020 6
Demo • HDFView SS_01499_STD_F 1262. h 5 • h 5 dump gz 6_SCRIS_npp_d 20140522_t 0754579_e 0802557_b 13293__noaa_pop. h 5 11/25/2020 7

GROUPS AND LINKS 11/25/2020 8

Groups and Links • Groups are containers for links (graph edges) • Warning: Many APIs in H 5 G interface are obsolete - use H 5 L interfaces to discover and manipulate file structure 11/25/2020 9
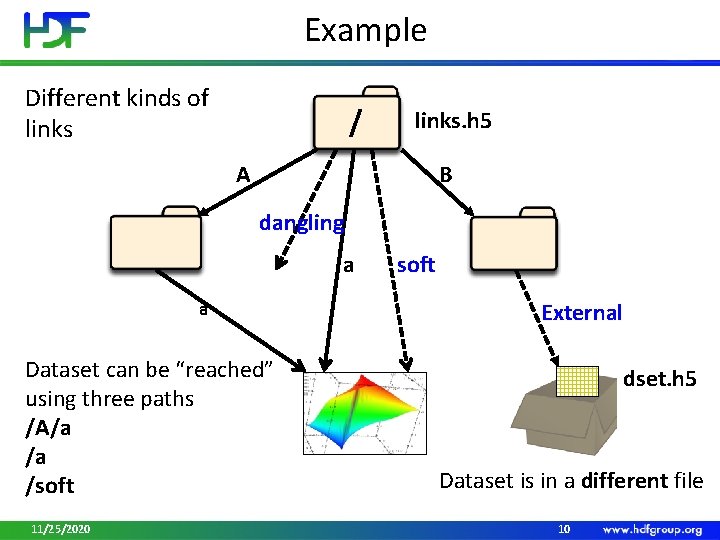
Example Different kinds of links / links. h 5 A B dangling a a Dataset can be “reached” using three paths /A/a /a /soft 11/25/2020 soft External dset. h 5 Dataset is in a different file 10

Links • Name • Example: “A”, “B”, “a”, “dangling”, “soft” • Unique within a group; “/” are not allowed in names • Type • Hard Link • Value is object’s address in a file • Created automatically when object is created • Can be added to point to existing object • Soft Link • Value is a string , for example, “/A/a”, but can be anything • Use to create aliases 11/25/2020 11

Links (cont. ) • Type • External Link • Value is a pair of strings , for example, (“dset. h 5”, “dset” ) • Use to access data in stored other HDF 5 files through a particular file • Enable file caching for better performance H 5 Pset_elink_file_cache_size 11/25/2020 12

Links Properties • ASCII or UTF-8 encoding for names • Create intermediate groups • Saves programming effort • C example lcpl_id = H 5 Pcreate(H 5 P_LINK_CREATE); H 5 Pset_create_intermediate_group( lcpl_id, 1 ); H 5 Gcreate (fid, "A/B", lcpl_id, H 5 P_DEFAULT); • Group “A” will be created if it doesn’t exist 11/25/2020 13

Operations on Links • • • See H 5 L interface in Reference Manual Create Delete Copy Iterate Check if exists 11/25/2020 14

Groups Properties • Creation properties • Type of links storage • Compact • Used with a few members (default under 8) • Saves space in the file, open/close times • Dense (default behavior) • Used with many (>16) members (default) • Tunable size for a local heap • Saves space by providing estimate for size of the storage required for links names • Can be compressed • Many links with similar names (XXX-abc, XXX-d, XXX-efgh, etc. ) • Requires more time to compress/uncompress data 11/25/2020 15

Groups Properties • Creation properties • Links may have creation order tracked and indexed • Indexing by name (default) • A, B, a, dangling, soft • Indexing by creation order (has to be enabled) • A, B, a, soft, dangling 11/25/2020 16

Discovering HDF 5 file’s structure • HDF 5 provides APIs for recursive and nonrecursive iterations over the groups and attributes • H 5 Ovisit and H 5 Literate (H 5 Giterate) • Life is much easier with H 5 Py import h 5 py def print_info(name, obj): print name for name, value in obj. attrs. iteritems(): print name+": ", value f = h 5 py. File('GATMO-SATMS-npp. h 5', 'r+') f. visititems(print_info) f. close() 11/25/2020 17
Demo h 5 ex_g_iterate. c 11/25/2020 18
Demo h 5 ex_g_visit. h 5 11/25/2020 19

Hints • Use latest file format (see H 5 Pset_libver_bound function in RM) to set up file access property list • Save space when creating a lot of groups in a file • Save time when accessing many objects (>1000) 11/25/2020 20

Informal Benchmark • Create a file and a group in a file • Create up to 10^6 groups with one dataset in each group • Compare files sizes and performance of HDF 5 using the latest group format with the performance of default format • For old versions performance became very slow after 700000 groups 11/25/2020 21
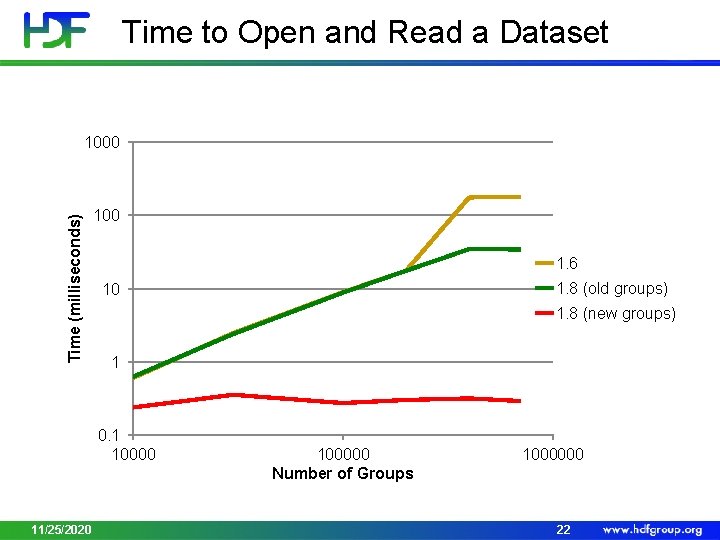
Time to Open and Read a Dataset Time (milliseconds) 1000 1. 6 1. 8 (new groups) 1 0. 1 10000 11/25/2020 1. 8 (old groups) 10 100000 Number of Groups 1000000 22
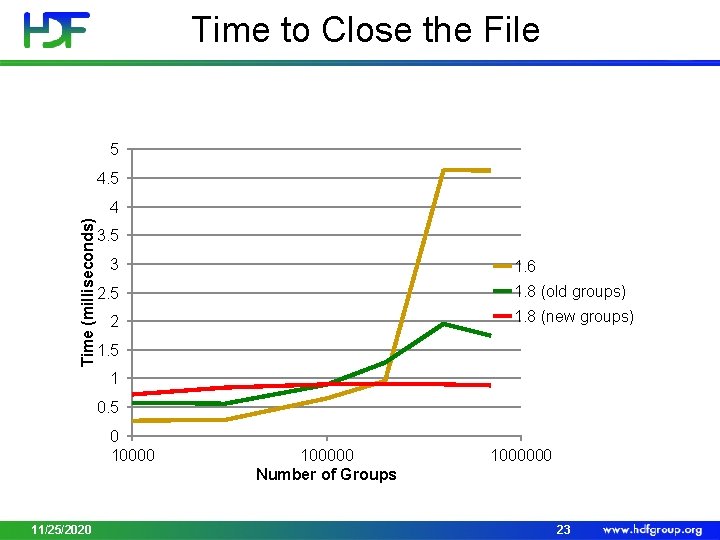
Time to Close the File 5 4. 5 Time (milliseconds) 4 3. 5 3 1. 6 1. 8 (old groups) 2. 5 1. 8 (new groups) 2 1. 5 1 0. 5 0 10000 11/25/2020 100000 Number of Groups 1000000 23
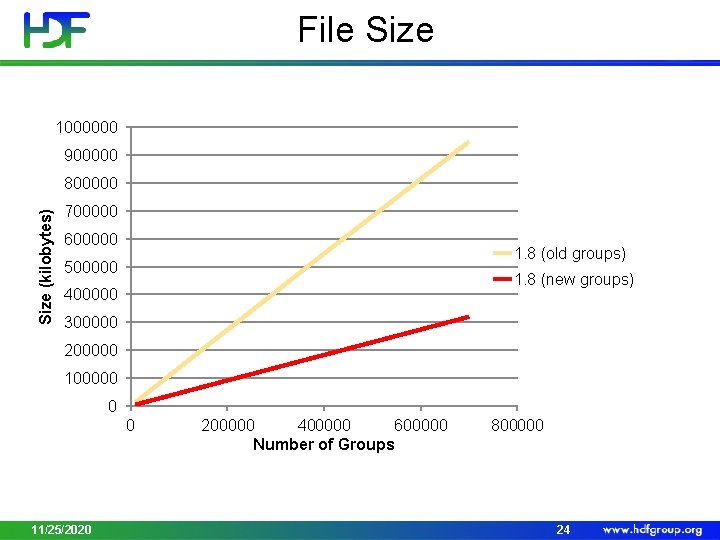
File Size 1000000 900000 Size (kilobytes) 800000 700000 600000 1. 8 (old groups) 500000 1. 8 (new groups) 400000 300000 200000 100000 0 0 11/25/2020 200000 400000 600000 Number of Groups 800000 24

DATASETS 11/25/2020 25

Section Topics • Introduction to HDF 5 hyperslab selections • Introduction to HDF 5 partial I/O • Introduction to HDF 5 dataset storage 11/25/2020 26
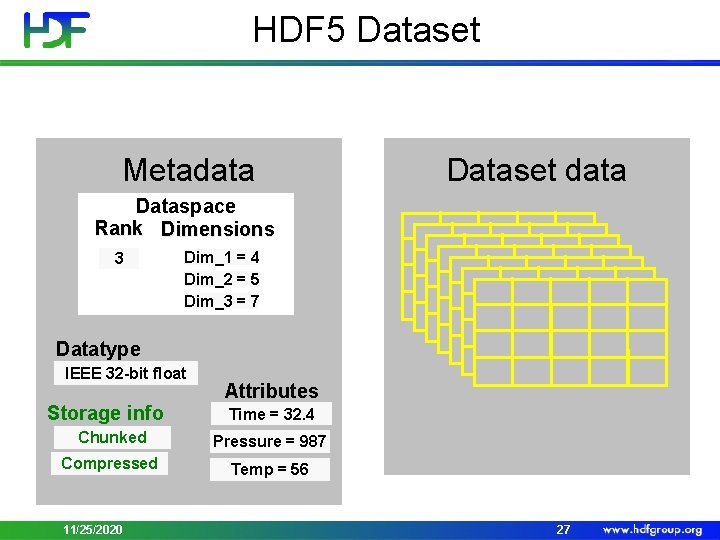
HDF 5 Dataset Metadata Dataset data Dataspace Rank Dimensions 3 Dim_1 = 4 Dim_2 = 5 Dim_3 = 7 Datatype IEEE 32 -bit float Storage info Attributes Time = 32. 4 Chunked Pressure = 987 Compressed Temp = 56 11/25/2020 27
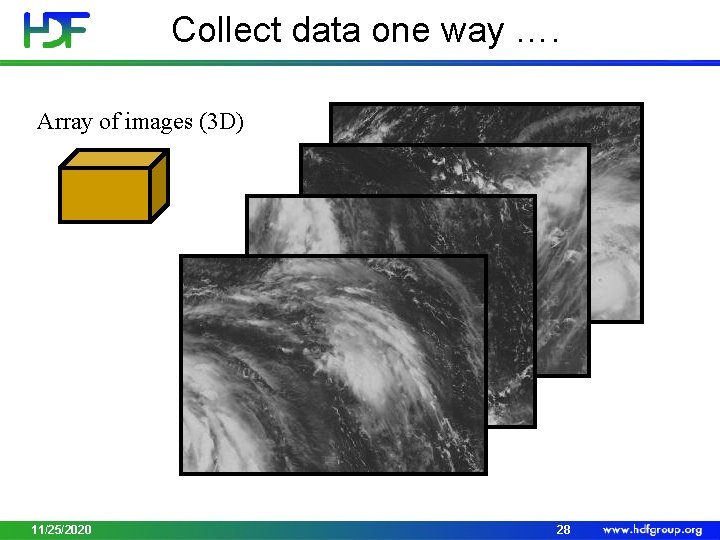
Collect data one way …. Array of images (3 D) 11/25/2020 28

Display data another way … Stitched image (2 D array) 11/25/2020 29

Data is too big to read…. 11/25/2020 30

HDF 5 Library Features • HDF 5 Library provides capabilities to • Describe subsets of data and perform write/read operations on subsets • Hyperslab selections and partial I/O • Store descriptions of the data subsets in a file • Object references • Region references • Use efficient storage mechanism to achieve good performance while writing/reading subsets of data • Chunking, compression 11/25/2020 31

PARTIAL I/O IN HDF 5 11/25/2020 32

How to write a row? $ h 5 dump file. h 5 HDF 5 "file. h 5" { GROUP "/" { DATASET "A" { DATATYPE H 5 T_STD_I 32 BE DATASPACE SIMPLE { ( 4, 6 ) / ( 4, 6 ) } DATA { (0, 0): 0, 0, 0, (1, 0): 1, 2, 3, 4, 5, 6, (2, 0): 0, 0, 0, (3, 0): 0, 0, 0, 0 } } GROUP "B" { } } } 11/25/2020 33

How to Describe a Subset in HDF 5? • Before writing and reading a subset of data one has to describe it to the HDF 5 Library. • HDF 5 APIs and documentation refer to a subset as a “selection” or “hyperslab selection”. • If specified, HDF 5 Library will perform I/O on a selection only and not on all elements of a dataset. 11/25/2020 34

Types of Selections in HDF 5 • Two types of selections • Hyperslab selection • Regular hyperslab • Simple hyperslab • Result of set operations on hyperslabs (union, difference, …) • Point selection • Hyperslab selection is especially important for doing parallel I/O in HDF 5 (See Parallel HDF 5 Tutorial) 11/25/2020 35
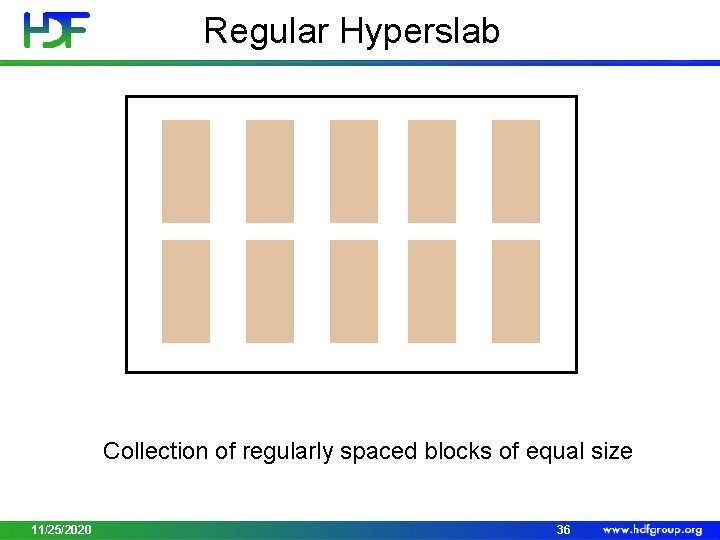
Regular Hyperslab Collection of regularly spaced blocks of equal size 11/25/2020 36
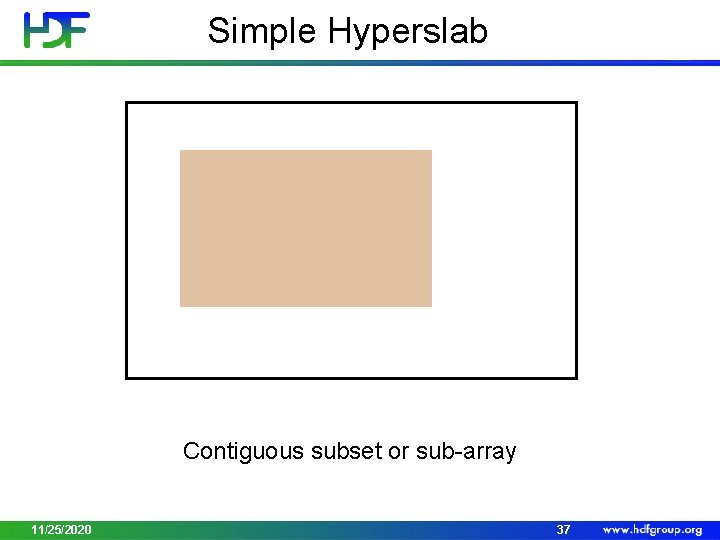
Simple Hyperslab Contiguous subset or sub-array 11/25/2020 37
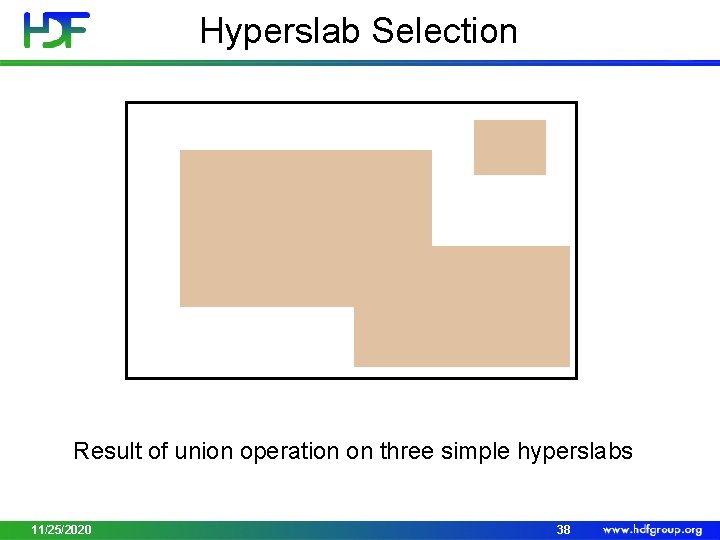
Hyperslab Selection Result of union operation on three simple hyperslabs 11/25/2020 38

HDF 5 Hyperslab Description • • • Everything is “measured” in number of elements Start - starting location of a hyperslab (1, 1) Stride - number of elements that separate each block (3, 2) Count - number of blocks (2, 6) Block - block size (2, 1) 11/25/2020 39

Simple Hyperslab Description • Two ways to describe a simple hyperslab • As several blocks • Stride – (1, 1) • Count – (2, 6) • Block – (2, 1) • As one block • Stride – (1, 1) • Count – (1, 1) • Block – (4, 6) No performance penalty for one way or another 11/25/2020 40

Writing a row • Memory space selection is 1 -dim array of size 6 • File space selection start = {1, 0}, stride = {1, 1}, count = {1, 6}, block = {1, 1} Number of elements selected in memory should be the same as selected in the file 11/25/2020 41
![Demo: Writing a row (h 5_wrow. c) hid_t mspace_id, fspace_id; hsize_t dims[1] = {6}; Demo: Writing a row (h 5_wrow. c) hid_t mspace_id, fspace_id; hsize_t dims[1] = {6};](http://slidetodoc.com/presentation_image_h/99bd80d275a8699c0c8ea57042376157/image-42.jpg)
Demo: Writing a row (h 5_wrow. c) hid_t mspace_id, fspace_id; hsize_t dims[1] = {6}; hsize_t start[2], count[2]; …. . /* Create memory dataspace */ mspace_id = H 5 Screate_simple(RANK, dims, NULL); /* Get file space identifier from the dataset */ fspace_id = H 5 Dget_space(dataset_id); /* Select hyperslab in the dataset to write too */ start[0] = 1; start[1] = 0; count[0] = 1; count[1] = 6; status = H 5 Sselect_hyperslab(fspace_id, H 5 S_SELECT_SET, start, NULL, count, NULL); H 5 Dwrite(dataset_id, H 5 T_NATIVE_INT, mspace_id, fspace_id, H 5 P_DEFAULT, wdata); 11/25/2020 42

Writing a subset h 5_subset. c • Creates an 8 x 10 integer dataset • Selects and writes to a 3 x 4 subset, offset by 1 x 2 11/25/2020 43

Demo • h 5_wrow. c • h 5_subset. c 11/25/2020 44

DATASET STORAGE 11/25/2020 45
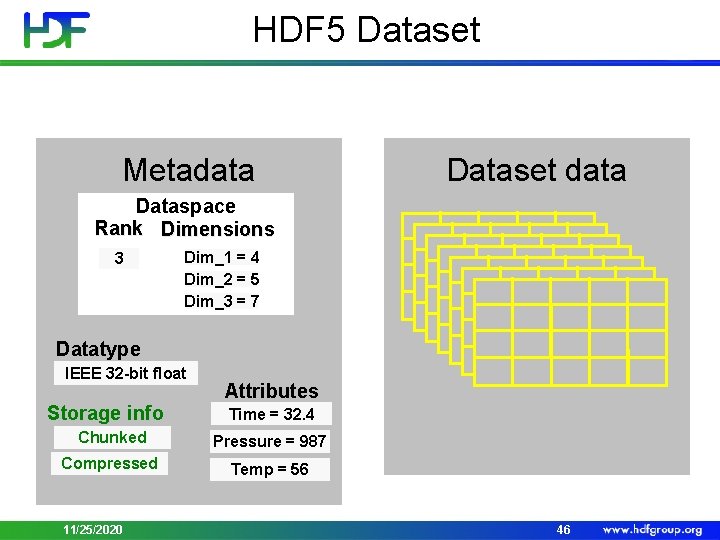
HDF 5 Dataset Metadata Dataset data Dataspace Rank Dimensions 3 Dim_1 = 4 Dim_2 = 5 Dim_3 = 7 Datatype IEEE 32 -bit float Storage info Attributes Time = 32. 4 Chunked Pressure = 987 Compressed Temp = 56 11/25/2020 46

CONTIGUOUS 11/25/2020 47
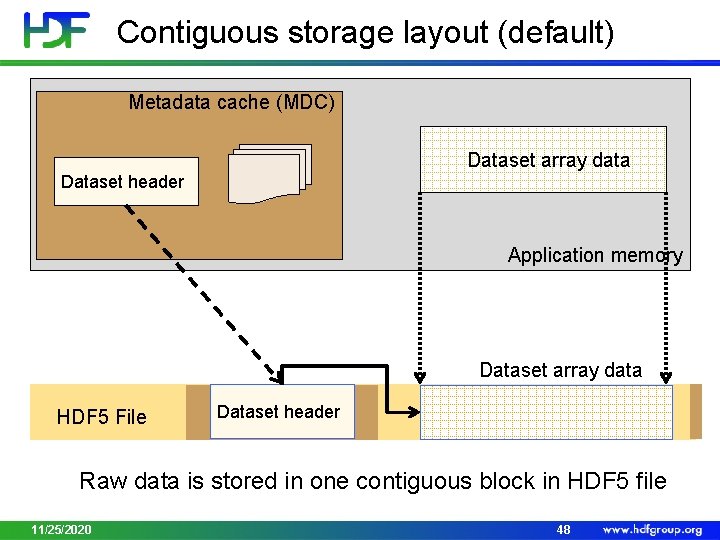
Contiguous storage layout (default) Metadata cache (MDC) Dataset array data Dataset header Application memory Dataset array data HDF 5 File Dataset header Raw data is stored in one contiguous block in HDF 5 file 11/25/2020 48

Contiguous Storage • Pros: • Default storage mechanism for HDF 5 dataset • Allows sub-setting • Efficient access to the whole dataset or to contiguous (in the file) subset • Can be easily located in the HDF 5 file • Cons: • No compression • Data cannot be added 11/25/2020 49

CHUNKED 11/25/2020 50

Chunked storage layout • Chunking – storage layout where a dataset is partitioned in fixed-size multi-dimensional tiles or chunks • Each chunk is stored as contiguous block • HDF 5 library treats each chunk as atomic object for I/O • Greatly affects performance and file sizes • Use for extendible datasets and datasets with filters applied (checksum, compression) • Use for sub-setting of big datasets 11/25/2020 51
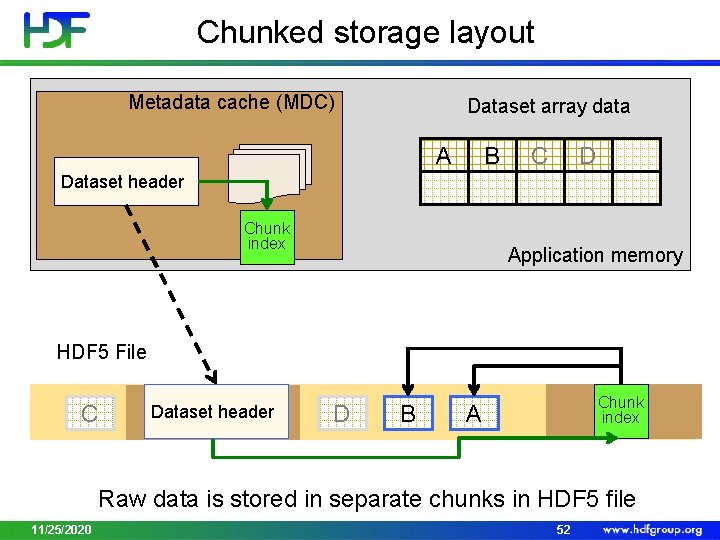
Chunked storage layout Metadata cache (MDC) Dataset array data B A C D Dataset header Chunk index Application memory HDF 5 File C Dataset header D B Chunk index A Raw data is stored in separate chunks in HDF 5 file 11/25/2020 52
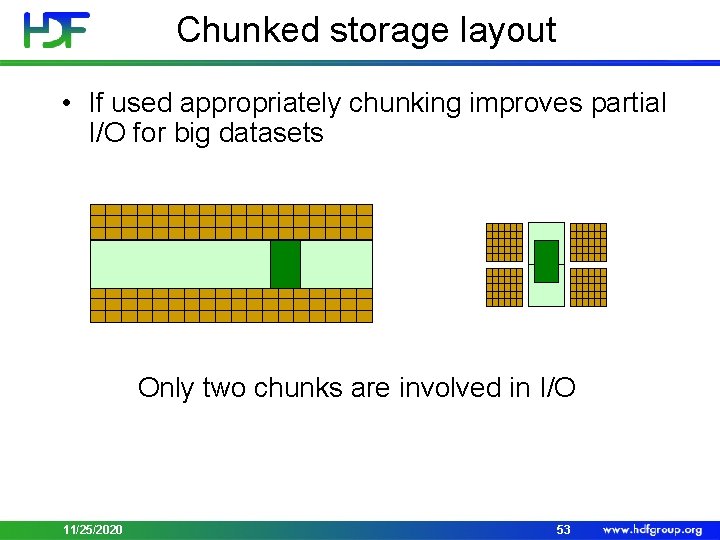
Chunked storage layout • If used appropriately chunking improves partial I/O for big datasets Only two chunks are involved in I/O 11/25/2020 53

Chunked Storage • Pros: • Allows adding/deleting data and compression • Cons: • Wrong chunking parameters may hurt performance and memory footprint 11/25/2020 54

Creating Chunked Dataset 1. 2. 3. Create a dataset creation property list. Set property list to use chunked storage layout. Create dataset with the above property list. dcpl_id = H 5 Pcreate(H 5 P_DATASET_CREATE); rank = 2; ch_dims[0] = 100; ch_dims[1] = 200; H 5 Pset_chunk(dcpl_id, rank, ch_dims); dset_id = H 5 Dcreate (…, dcpl_id); H 5 Pclose(dcpl_id); 11/25/2020 55
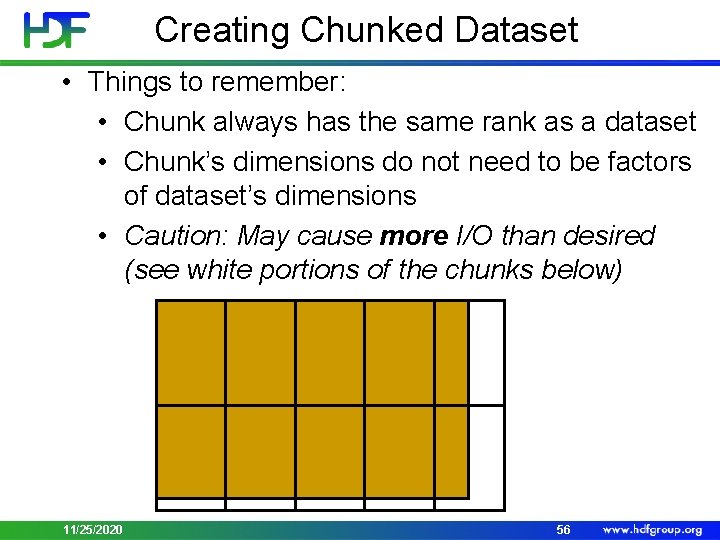
Creating Chunked Dataset • Things to remember: • Chunk always has the same rank as a dataset • Chunk’s dimensions do not need to be factors of dataset’s dimensions • Caution: May cause more I/O than desired (see white portions of the chunks below) 11/25/2020 56

Chunking Limitations • Chunk dimensions cannot be bigger than dataset dimensions • Number of elements a chunk is limited to 4 GB • H 5 Pset_chunk fails otherwise • Total size chunk is limited to 4 GB • Total size = (number of elements) * (size of the datatype) • H 5 Dwrite fails later on 11/25/2020 57

Writing or Reading Chunked Dataset 1. 2. Chunking is transparent to the application. Use the same set of operations as for contiguous dataset, for example, H 5 Dopen(…); H 5 Sselect_hyperslab (…); H 5 Dread(…); 3. Selections do not need to coincide precisely with the chunk boundaries. 11/25/2020 58

Creating Compressed Dataset 1. 2. 3. 4. Create a dataset creation property list Set property list to use chunked storage layout Set property list to use filters Create dataset with the above property list dcpl_id = H 5 Pcreate(H 5 P_DATASET_CREATE); rank = 2; ch_dims[0] = 100; ch_dims[1] = 100; H 5 Pset_chunk(dcpl_id, rank, ch_dims); H 5 Pset_deflate(dcpl_id, 9); dset_id = H 5 Dcreate (…, dcpl_id); H 5 Pclose(dcpl_id); Example: h 5_d_unlimgzip. c 11/25/2020 59

h 5 d_unlimgzip. h 5 [ 0 -1 -2 -3 -4 -5 -6 7 8 9 ] [ 0 0 7 8 9 ] [ 0 1 2 3 4 5 6 7 8 9 ] [ 0 2 4 6 8 10 12 7 8 9 ] [ 0 1 2 3 4 5 6 7 8 9 ] 11/25/2020 First write Second write 60

Demo • h 5 d_unlimgzip. c 11/25/2020 61

VIRTUAL DATASET (VDS) 11/25/2020 62

Challenge • How to view data stored across the HDF 5 files as an HDF 5 dataset on which normal operations can be performed? • High-level approach • Native HDF 5 implementation • Transparent to applications 11/25/2020 63

TWO SIMPLE USE CASES 11/25/2020 64

Collect data one way …. File: a. h 5 Dataset /A File: c. h 5 Dataset /C 11/25/2020 File: b. h 5 Dataset /B File: d. h 5 Dataset /D 65

Present it in a different way… Whole image File: F. h 5 Dataset /D 11/25/2020 66

Present it in a different way… Subset of data File: F. h 5 Dataset /F 11/25/2020 67
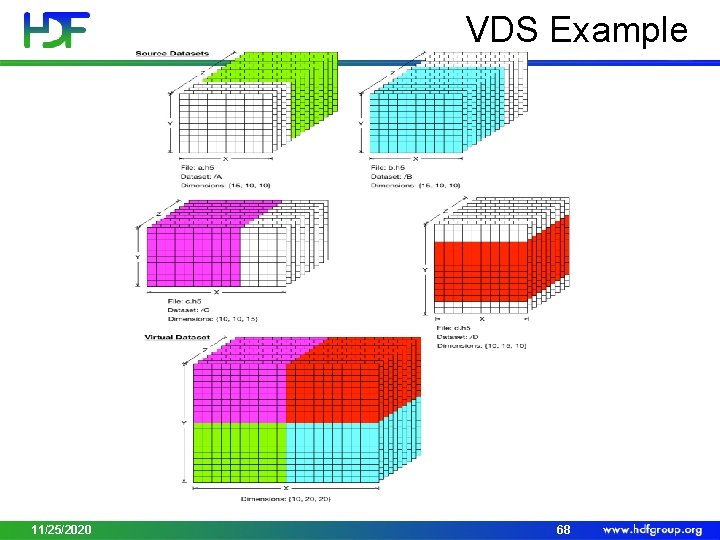
VDS Example 11/25/2020 68

VDS Storage • Pros: • Allows data aggregation transparent to user • Allows to have multiple parallel data streams • Compatible with Single Writer/Multiple Reader access • Cons: • Performance may not be as good as when data stored in regular HDF 5 dataset 11/25/2020 69

PROGRAMMING MODEL AND EXAMPLES OF MAPPING 11/25/2020 70

VDS Programming Model • Create datasets that comprise the VDS (the source datasets) (optional) • Create the VDS • Define VDS storage • Map elements from the source datasets to the elements of the VDS • Call H 5 Dcreate using the properties defined above • Access the VDS as a regular HDF 5 dataset • Close the VDS when finished 11/25/2020 71
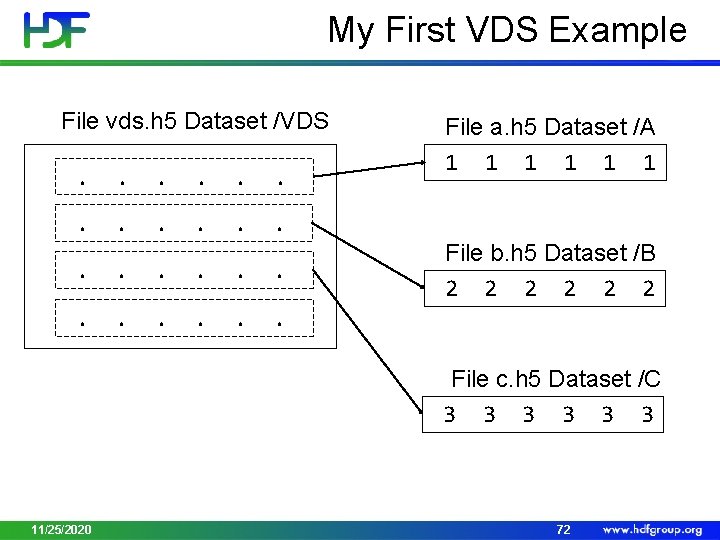
My First VDS Example File vds. h 5 Dataset /VDS . . . File a. h 5 Dataset /A 1 1 1 File b. h 5 Dataset /B 2 2 2 File c. h 5 Dataset /C 3 3 3 11/25/2020 72

My First VDS Example File vds. h 5 Dataset /VDS 1 1 1 2 2 2 3 3 3 -1 -1 -1 File a. h 5 Dataset /A 1 1 1 File b. h 5 Dataset /B 2 2 2 File c. h 5 Dataset /C 3 3 3 Data the application will see when reading /VDS dataset from file vds. h 5 The last row is filled with the fill value 11/25/2020 74

Demo • h 5_vds. c 11/25/2020 76

EXTERNAL 11/25/2020 77
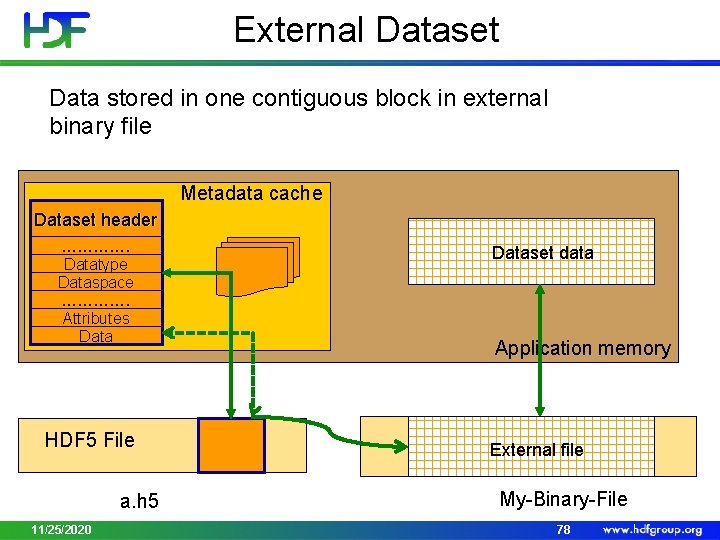
External Dataset Data stored in one contiguous block in external binary file Metadata cache Dataset header …………. Datatype Dataspace …………. Attributes Data HDF 5 File a. h 5 11/25/2020 Dataset data Application memory External file My-Binary-File 78

External Storage • Pros: • Mechanism to reference data stored in a non-HDF 5 binary file • Can be easily “imported” to HDF 5 with h 5 repack • Allows sub-setting • Efficient access to the whole dataset or to contiguous (in the file) subset • Cons: • Two or more files • No compression • Data cannot be added 11/25/2020 79

External Example and Demo HDF 5 "h 5 ex_d_extern-new. h 5" { GROUP "/" { DATASET "DSExternal" { DATATYPE H 5 T_STD_I 32 BE DATASPACE SIMPLE { ( 28 ) / ( 28 ) } STORAGE_LAYOUT { CONTIGUOUS EXTERNAL { FILENAME h 5 ex_d_extern. data SIZE 112 OFFSET 0 } } FILTERS { NONE } FILLVALUE { FILL_TIME H 5 D_FILL_TIME_IFSET VALUE 0 } ALLOCATION_TIME { H 5 D_ALLOC_TIME_LATE } DATA { (0): 0, 0, 1, 1, 2, 2, 3, (22): 3, 3, 3, 3 } …. 11/25/2020 80

COMPACT 11/25/2020 81

Compact storage layout • Raw data is stored in a dataset object header • Raw data read/written with the header • Use for small (few K) datasets to minimize small I/O operations 11/25/2020 82
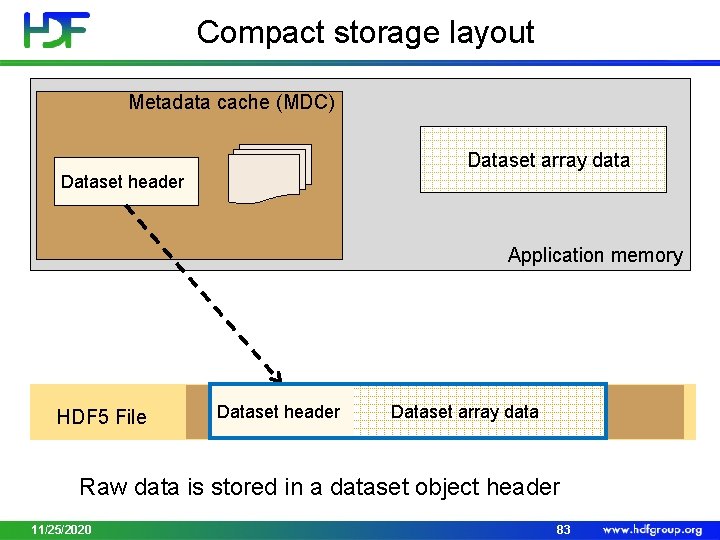
Compact storage layout Metadata cache (MDC) Dataset array data Dataset header Application memory HDF 5 File Dataset header Dataset array data Raw data is stored in a dataset object header 11/25/2020 83

DATATYPES 11/25/2020 84

An HDF 5 Datatype is… • A description of dataset element type • Grouped into “classes”: • • • Atomic – integers, floating-point values Enumerated Compound – like C structs Array Opaque References • Object – similar to soft link • Region – similar to soft link to dataset + selection • Variable-length • Strings – fixed and variable-length • Sequences – similar to Standard C++ vector class 11/25/2020 85

HDF 5 Datatypes • HDF 5 has a rich set of pre-defined datatypes and supports the creation of an unlimited variety of complex user-defined datatypes. • Self-describing: • Datatype definitions are stored in the HDF 5 file with the data. • Datatype definitions include information such as byte order (endianness), size, and floating point representation to fully describe how the data is stored and to insure portability across platforms. 11/25/2020 86

Datatype Conversion • Datatypes that are compatible, but not identical are converted automatically when I/O is performed • Compatible datatypes: • All atomic datatypes are compatible • Identically structured array, variable-length and compound datatypes whose base type or fields are compatible • Enumerated datatype values on a “by name” basis • Make datatypes identical for best performance 11/25/2020 87

Datatypes Used in JPSS files • In h 5 dump output search for “DATATYPE” keyword; those correspond to (…) DATATYPE H 5 T_STRING C string DATATYPE DATATYPE H 5 T_STD_U 8 BE unsigned char H 5 T_STD_U 16 BE short (big-endian) H 5 T_STD_I 32 BE int (big-endian) H 5 T_IEEE_F 32 BE float (big-endian) H 5 T_STD_U 64 BE unsigned long H 5 T_REFERENCE { H 5 T_STD_REF_OBJECT } H 5 T_REFERENCE { H 5 T_STD_REF_DSETREG } Native types from each language are mapped to HDF 5 pre-defined datatypes https: //www. hdfgroup. org/HDF 5/doc/RM/Predef. DTypes. html 11/25/2020 88
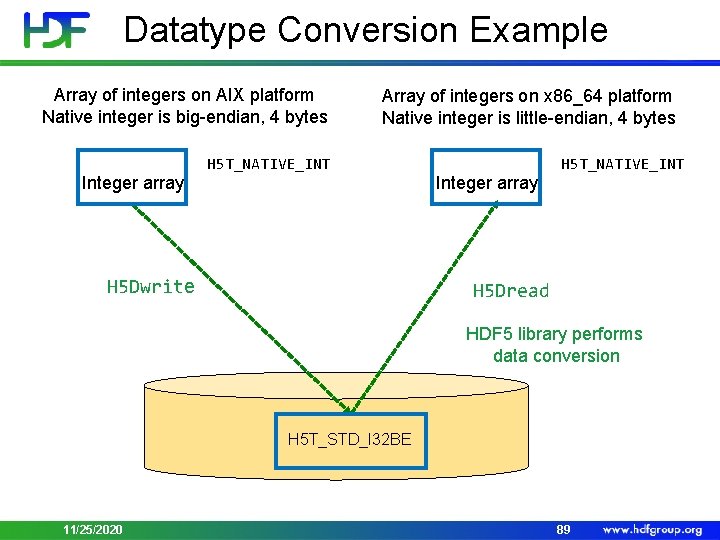
Datatype Conversion Example Array of integers on AIX platform Native integer is big-endian, 4 bytes Integer array Array of integers on x 86_64 platform Native integer is little-endian, 4 bytes H 5 T_NATIVE_INT H 5 Dwrite Integer array H 5 T_NATIVE_INT H 5 Dread HDF 5 library performs data conversion H 5 T_STD_I 32 BE 11/25/2020 89

Datatype Conversion Datatype of data in the file dataset = H 5 Dcreate(file, DATASETNAME, H 5 T_STD_I 32 LE, space, H 5 P_DEFAULT); Datatype of data in memory buffer H 5 Dwrite(dataset, H 5 T_NATIVE_INT, H 5 S_ALL, H 5 P_DEFAULT, buf); By specifying 4 -bytes little-endian in the file, one will avoid data conversion when reading on little-endian machines 11/25/2020 90

STRINGS 11/25/2020 91

Storing Strings in HDF 5 • Array of characters (Array datatype or extra dimension in dataset) • Quick access to each character • Extra work to access and interpret each string • Fixed length string_id = H 5 Tcopy(H 5 T_C_S 1); H 5 Tset_size(string_id, size); • Wasted space in shorter strings • Can be compressed if used in datasets • Variable length string_id = H 5 Tcopy(H 5 T_C_S 1); H 5 Tset_size(string_id, H 5 T_VARIABLE); • Overhead as for all VL datatypes • Compression will not be applied to actual data in datasets 11/25/2020 92

Storing Strings in HDF 5 • Store the data in two datasets: • 1 D char dataset stores concatenated strings. • 2 D dataset stores (start, end) indices of each string in the first dataset. • Essentially a "hand-constructed" version of how we store variable length data. • Since the data are stored in datasets, they can be chunked, compressed, etc. • Slower than other access methods. • When using attribute, use VL string H 5_SCALAR dataspace instead H 5 S_SIMPLE to save some programming effort • See CGNS example; uses array of chars confusing HDFView 11/25/2020 93

USING DATATYPE TO REFERENCE DATA 11/25/2020 94

Reference Datatypes • Object Reference • Pointer to an object in a file • Predefined datatype H 5 T_STD_REF_OBJ • Dataset Region Reference • Pointer to a dataset + dataspace selection • Predefined datatype H 5 T_STD_REF_DSETREG 11/25/2020 95
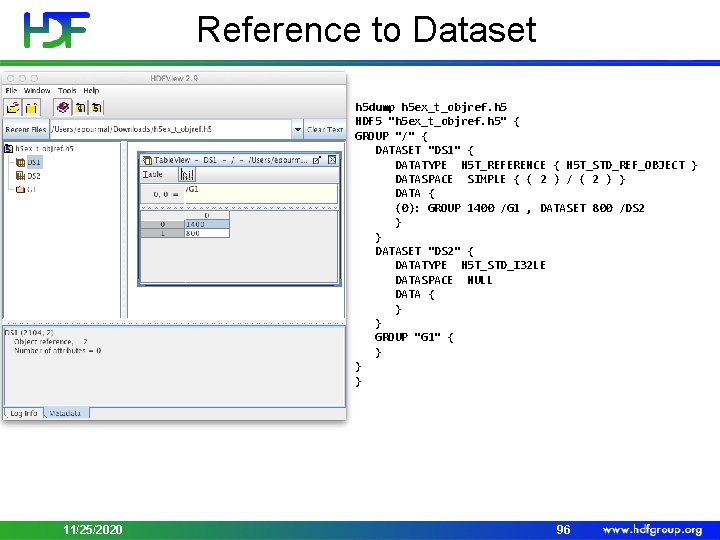
Reference to Dataset h 5 dump h 5 ex_t_objref. h 5 HDF 5 "h 5 ex_t_objref. h 5" { GROUP "/" { DATASET "DS 1" { DATATYPE H 5 T_REFERENCE { H 5 T_STD_REF_OBJECT } DATASPACE SIMPLE { ( 2 ) / ( 2 ) } DATA { (0): GROUP 1400 /G 1 , DATASET 800 /DS 2 } } DATASET "DS 2" { DATATYPE H 5 T_STD_I 32 LE DATASPACE NULL DATA { } } GROUP "G 1" { } } } 11/25/2020 96

Example: h 5 ex_t_objref. c HDF 5 "h 5 ex_t_objref. h 5" { GROUP "/" { DATASET "DS 1" { DATATYPE H 5 T_REFERENCE { H 5 T_STD_REF_OBJECT } DATASPACE SIMPLE { ( 2 ) / ( 2 ) } DATA { (0): GROUP 1400 /G 1 , DATASET 800 /DS 2 } } DATASET "DS 2" { DATATYPE H 5 T_STD_I 32 LE DATASPACE NULL DATA { } } GROUP "G 1" { } } } 11/25/2020 97

Saving Selected Region in a File Need to select and access the same elements of a dataset 11/25/2020 98
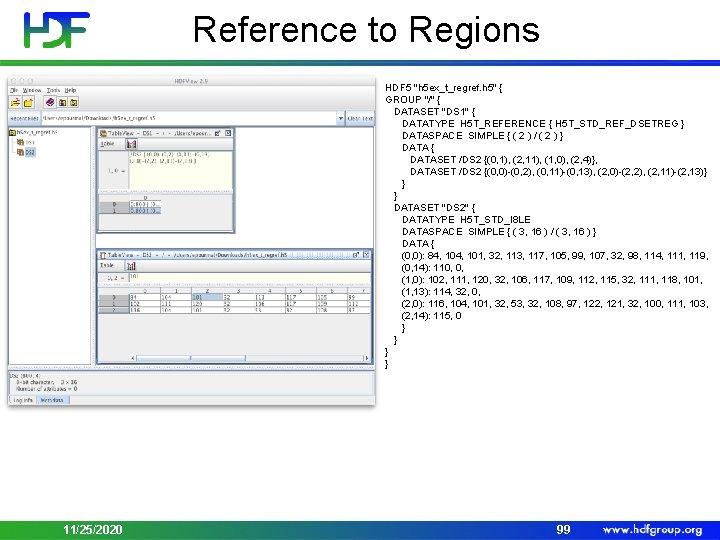
Reference to Regions HDF 5 "h 5 ex_t_regref. h 5" { GROUP "/" { DATASET "DS 1" { DATATYPE H 5 T_REFERENCE { H 5 T_STD_REF_DSETREG } DATASPACE SIMPLE { ( 2 ) / ( 2 ) } DATA { DATASET /DS 2 {(0, 1), (2, 11), (1, 0), (2, 4)}, DATASET /DS 2 {(0, 0)-(0, 2), (0, 11)-(0, 13), (2, 0)-(2, 2), (2, 11)-(2, 13)} } } DATASET "DS 2" { DATATYPE H 5 T_STD_I 8 LE DATASPACE SIMPLE { ( 3, 16 ) / ( 3, 16 ) } DATA { (0, 0): 84, 101, 32, 113, 117, 105, 99, 107, 32, 98, 114, 111, 119, (0, 14): 110, 0, (1, 0): 102, 111, 120, 32, 106, 117, 109, 112, 115, 32, 111, 118, 101, (1, 13): 114, 32, 0, (2, 0): 116, 104, 101, 32, 53, 32, 108, 97, 122, 121, 32, 100, 111, 103, (2, 14): 115, 0 } } } } 11/25/2020 99

Example: h 5 ex_t_objref. c HDF 5 "h 5 ex_t_regref. h 5" { GROUP "/" { DATASET "DS 1" { DATATYPE H 5 T_REFERENCE { H 5 T_STD_REF_DSETREG } DATASPACE SIMPLE { ( 2 ) / ( 2 ) } DATA { DATASET /DS 2 {(0, 1), (2, 11), (1, 0), (2, 4)}, DATASET /DS 2 {(0, 0)-(0, 2), (0, 11)-(0, 13), (2, 0)-(2, 2), (2, 11)-(2, 13)} } } DATASET "DS 2" { DATATYPE H 5 T_STD_I 8 LE DATASPACE SIMPLE { ( 3, 16 ) / ( 3, 16 ) } DATA { (0, 0): 84, 101, 32, 113, 117, 105, 99, 107, 32, 98, 114, 111, 119, 110, 0, (1, 0): 102, 111, 120, 32, 106, 117, 109, 112, 115, 32, 111, 118, 101, 114, 32, 0, (2, 0): 116, 104, 101, 32, 53, 32, 108, 97, 122, 121, 32, 100, 111, 103, 115, 0 } } 11/25/2020 100
Demo • HDView example with SVM 16_npp_d 20120424_t 0302382_e 0304023_b 02536_c 20120502162515786022_noaa_ops. h 5 11/25/2020 101

Storing Records with HDF 5 11/25/2020 102

HDF 5 Compound Datatypes • Compound types • Comparable to C structs • Members can be any datatype • Can write/read by a single field or a set of fields • Not all data filters can be applied (shuffling, SZIP) 11/25/2020 103

Creating and Writing Compound Dataset h 5_compound. c example typedef struct s 1_t { int a; float b; double c; } s 1_t; s 1_t 11/25/2020 s 1[LENGTH]; 104

Creating and Writing Compound Dataset /* Create datatype in memory. */ s 1_tid = H 5 Tcreate(H 5 T_COMPOUND, sizeof(s 1_t)); H 5 Tinsert(s 1_tid, "a_name", HOFFSET(s 1_t, a), H 5 T_NATIVE_INT); H 5 Tinsert(s 1_tid, "c_name", HOFFSET(s 1_t, c), H 5 T_NATIVE_DOUBLE); H 5 Tinsert(s 1_tid, "b_name", HOFFSET(s 1_t, b), H 5 T_NATIVE_FLOAT); Note: • Use HOFFSET macro instead of calculating offset by hand. • Order of H 5 Tinsert calls is not important if HOFFSET is used. 11/25/2020 105

Creating and Writing Compound Dataset /* Create dataset and write data */ dataset = H 5 Dcreate(file, DATASETNAME, s 1_tid, space, H 5 P_DEFAULT); status = H 5 Dwrite(dataset, s 1_tid, H 5 S_ALL, H 5 P_DEFAULT, s 1); Note: • In this example memory and file datatypes are the same. • Type is not packed (i. e. , it still has internal padding). • Use H 5 Tpack to save space in the file. status = H 5 Tpack(s 1_tid); status = H 5 Dcreate(file, DATASETNAME, s 1_tid, space, H 5 P_DEFAULT); 11/25/2020 106

Reading Compound Dataset /* Create datatype in memory and read data. */ dataset = H 5 Dopen(file, DATASETNAME, H 5 P_DEFAULT); s 2_tid = H 5 Dget_type(dataset); mem_tid = H 5 Tget_native_type(s 2_tid); buf = malloc(H 5 Tget_size(mem_tid)*number_of_elements); status = H 5 Dread(dataset, mem_tid, H 5 S_ALL, H 5 P_DEFAULT, buf); Note: • We could construct memory type as we did in writing example. • For general applications we need to discover the type in the file, find out corresponding memory type, allocate space and do read. 11/25/2020 107

Reading Compound Dataset by Fields typedef struct s 2_t { double c; int a; } s 2_t; s 2_t s 2[LENGTH]; … s 2_tid = H 5 Tcreate (H 5 T_COMPOUND, sizeof(s 2_t)); H 5 Tinsert(s 2_tid, "c_name", HOFFSET(s 2_t, c), H 5 T_NATIVE_DOUBLE); H 5 Tinsert(s 2_tid, "a_name", HOFFSET(s 2_t, a), H 5 T_NATIVE_INT); … status = H 5 Dread(dataset, s 2_tid, H 5 S_ALL, H 5 P_DEFAULT, s 2); 11/25/2020 108
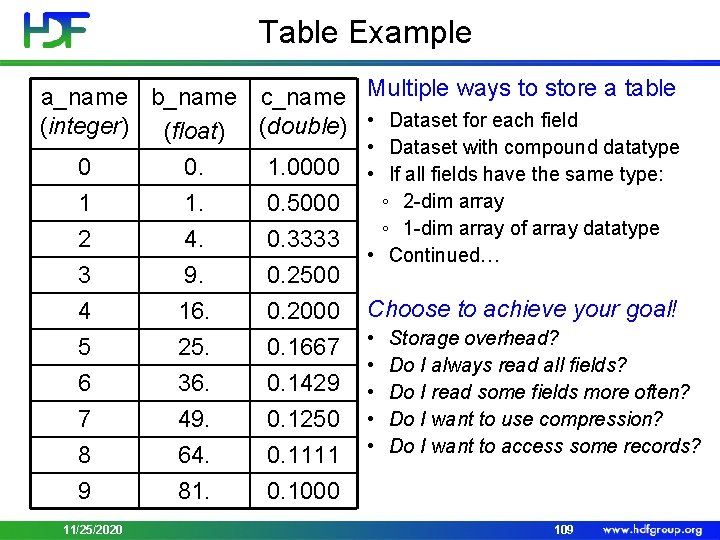
Table Example a_name b_name c_name (integer) (float) (double) 0 0. 1. 0000 1 1. 0. 5000 2 4. 0. 3333 3 4 5 6 7 8 9 9. 16. 25. 36. 49. 64. 81. 0. 2500 0. 2000 0. 1667 0. 1429 0. 1250 0. 1111 0. 1000 11/25/2020 Multiple ways to store a table • Dataset for each field • Dataset with compound datatype • If all fields have the same type: ◦ 2 -dim array ◦ 1 -dim array of array datatype • Continued… Choose to achieve your goal! • • • Storage overhead? Do I always read all fields? Do I read some fields more often? Do I want to use compression? Do I want to access some records? 109

Demo • h 5 ex_t_cmpd. c 11/25/2020 110

STORING VARIABLE-LENGTH DATA IN HDF 5 11/25/2020 111

HDF 5 Fixed and Variable Length Array Storage • Data Time • Data Time • Data 11/25/2020 112

Storing Variable Length Data in HDF 5 • Each element is represented by C structure typedef struct { size_t length; void *p; } hvl_t; • Base type can be any HDF 5 type H 5 Tvlen_create(base_type) 11/25/2020 113
![Example hvl_t data[LENGTH]; for(i=0; i<LENGTH; i++) { data[i]. p = malloc((i+1)*sizeof(unsigned int)); data[i]. len Example hvl_t data[LENGTH]; for(i=0; i<LENGTH; i++) { data[i]. p = malloc((i+1)*sizeof(unsigned int)); data[i]. len](http://slidetodoc.com/presentation_image_h/99bd80d275a8699c0c8ea57042376157/image-112.jpg)
Example hvl_t data[LENGTH]; for(i=0; i<LENGTH; i++) { data[i]. p = malloc((i+1)*sizeof(unsigned int)); data[i]. len = i+1; } tvl = H 5 Tvlen_create (H 5 T_NATIVE_UINT); data[0]. p • Data data[4]. len 11/25/2020 • Data 114

Reading HDF 5 Variable Length Array • HDF 5 library allocates memory to read data in • Application only needs to allocate array of hvl_t elements (pointers and lengths) • Application must reclaim memory for data read in hvl_t rdata[LENGTH]; /* Create the memory vlen type */ tvl = H 5 Tvlen_create(H 5 T_NATIVE_INT); ret = H 5 Dread(dataset, tvl, H 5 S_ALL, H 5 P_DEFAULT, rdata); /* Reclaim the read VL data */ H 5 Dvlen_reclaim(tvl, H 5 S_ALL, H 5 P_DEFAULT, rdata); 11/25/2020 115

Variable Length vs. Array • Pros of variable length datatypes vs. arrays: • Uses less space if compression unavailable • Automatically stores length of data • No maximum size • Size of an array is its effective maximum size • Cons of variable length datatypes vs. arrays: • Substantial performance overhead • Each element a “pointer” to piece of metadata • Variable length data cannot be compressed • Unused space in arrays can be “compressed away” • Must be 1 -dimensional 11/25/2020 116

Demo • h 5 ex_t_vlen. c 11/25/2020 117

OVERVIEW OF HDF 5 FILTERS 11/25/2020 118

What is an HDF 5 filter? • Data transformation performed by the HDF 5 library during I/O operations • HDF 5 filters (or built-in filters) • Supported by The HDF Group • Come with the HDF 5 library source code • User-defined filters • Filters written by HDF 5 users and/or available with some applications (h 5 py, Py. Tables) • May be or may not be registered with The HDF Group 11/25/2020 119

HDF 5 filters • Filters are arranged in a pipeline so the output of one filter becomes the input of the next filter • The filter pipeline can be only applied to - Chunked dataset - HDF 5 library passes each chunk through the filter pipeline on the way to or from disk - Group - Link names are stored in a local heap, which may be compressed with a filter pipeline • Filter pipeline is permanent for dataset or a group 11/25/2020 120
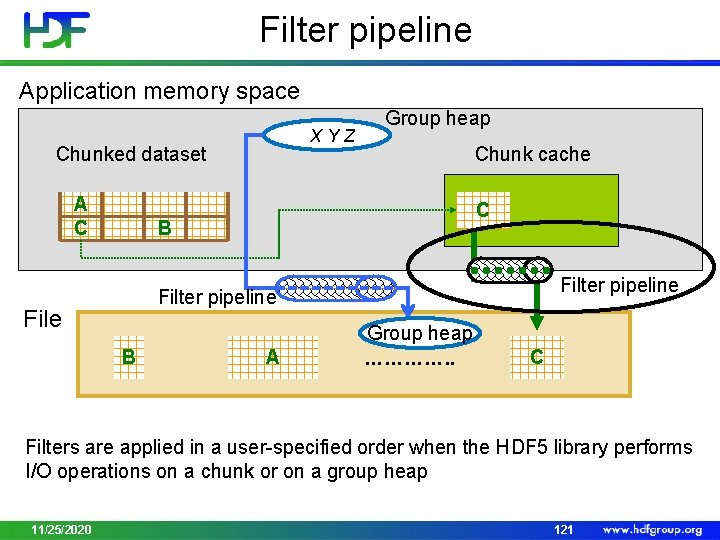
Filter pipeline Application memory space XYZ Chunked dataset A C Group heap Chunk cache C B Filter pipeline File B A Group heap …………. . C Filters are applied in a user-specified order when the HDF 5 library performs I/O operations on a chunk or on a group heap 11/25/2020 121

Filter pipeline programming model • Operations on the HDF 5 filter pipeline • Query - Number of filters in a pipeline - H 5 Pget_nfilters - Information about a filter using filter identifier - H 5 Pget_filter_by_id - Check if a filter is available in the library - H 5 Zfilter_avail • Modify - Change properties of existing filter - H 5 Pmodify_filter - Remove filter from pipeline - H 5 Premove_filter 11/25/2020 122

Filter pipeline programming model • Filter pipeline is permanent for dataset or a group • Filters are part of an HDF 5 object (group or dataset) creation property • The object’s filter pipeline cannot be modified after the object has been created 11/25/2020 123
![Applying filters to a dataset dcpl_id = H 5 Pcreate(H 5 P_DATASET_CREATE); cdims[0] = Applying filters to a dataset dcpl_id = H 5 Pcreate(H 5 P_DATASET_CREATE); cdims[0] =](http://slidetodoc.com/presentation_image_h/99bd80d275a8699c0c8ea57042376157/image-122.jpg)
Applying filters to a dataset dcpl_id = H 5 Pcreate(H 5 P_DATASET_CREATE); cdims[0] = 100; cdims[1] = 100; H 5 Pset_chunk(dcpl_id, 2, cdims); H 5 Pset_shuffle(dcpl); H 5 Pset_deflate(dcpl_id, 9); dset_id = H 5 Dcreate (…, dcpl_id); H 5 Pclose(dcpl_id); 11/25/2020 124
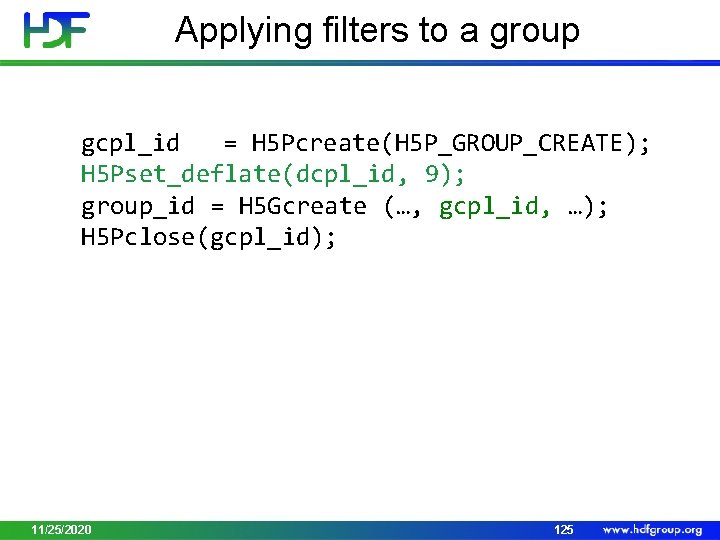
Applying filters to a group gcpl_id = H 5 Pcreate(H 5 P_GROUP_CREATE); H 5 Pset_deflate(dcpl_id, 9); group_id = H 5 Gcreate (…, gcpl_id, …); H 5 Pclose(gcpl_id); 11/25/2020 125

HDF 5 DEAFAULT FILTERS 11/25/2020 126

External HDF 5 Filters • External HDF 5 filters rely on the third-party libraries installed on the system • GZIP • By default HDF 5 configure uses ZLIB installed on the system • Configure will proceed if ZLIB is not found on the system • SZIP (added by NASA request) • Optional; have to be configured in using –withszlib=/path…. • Configure will proceed if SZIP is not found • Comes with a license http: //www. hdfgroup. org/doc_resource/SZIP/Commercial _szip. html • Decoder is free; for encoder see the license terms 11/25/2020 127

Internal HDF 5 Filters • Internal filters are implemented by The HDF Group and come with the library • HDF 5 internal filters • • 11/25/2020 FLETCHER 32 SHUFFLE SCALEOFFSET NBIT 128
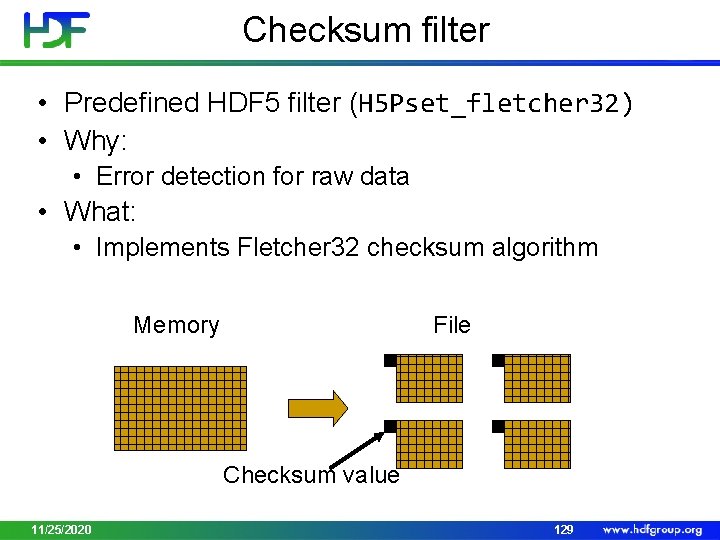
Checksum filter • Predefined HDF 5 filter (H 5 Pset_fletcher 32) • Why: • Error detection for raw data • What: • Implements Fletcher 32 checksum algorithm Memory File Checksum value 11/25/2020 129

Shuffling filter • Predefined HDF 5 filter (H 5 Pset_shuffle) • Why: • Better compression of unused bytes • What: • Changes byte order in a stream of data 00 00 00 01 00 00 00 17 00 00 00 2 B 00 00 00 01 17 2 B 11/25/2020 130
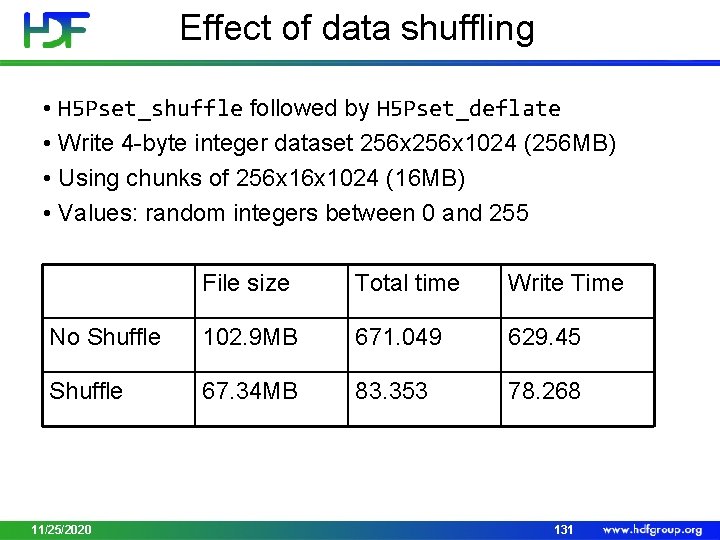
Effect of data shuffling • H 5 Pset_shuffle followed by H 5 Pset_deflate • Write 4 -byte integer dataset 256 x 1024 (256 MB) • Using chunks of 256 x 1024 (16 MB) • Values: random integers between 0 and 255 File size Total time Write Time No Shuffle 102. 9 MB 671. 049 629. 45 Shuffle 67. 34 MB 83. 353 78. 268 11/25/2020 131

N-bit compression filter • Predefined HDF 5 filter (H 5 Pset_nbit) • Why: Compact storage for user-defined datatypes • What: • When data stored on disk, padding bits chopped off and only significant bits stored • Supports most datatypes • Works with compound datatypes 11/25/2020 132

N-bit compression example • In memory, one value of N-Bit datatype is stored like this: | byte 3 | byte 2 | byte 1 | byte 0 | |? ? ? ? |? ? SPPP|PPPP|PPPP? ? | S-sign bit P-significant bit ? -padding bit • After passing through the N-Bit filter, all padding bits are chopped off, and the bits are stored on disk like this: | 1 st value | 2 nd value | |SPPPPPPPP|. . . • Opposite (decompress) when going from disk to memory 11/25/2020 133

“Scale+offset” filter • Predefined HDF 5 filter (H 5 Pset_scaleoffset) • Why: • Use less storage when less precision needed • What: • Performs scale/offset operation on each value • Truncates result to fewer bits before storing • Currently supports integers and floats 11/25/2020 134

Example with floating-point type • Data: {104. 561, 99. 459, 100. 545, 105. 644} • Choose scaling factor: decimal precision to keep E. g. scale factor D = 2 1. Find minimum value (offset): 99. 459 2. Subtract minimum value from each element Result: {5. 102, 0, 1. 086, 6. 185} 3. Scale data by multiplying 10 D = 100 Result: {510. 2, 0, 108. 6, 618. 5} 4. Round the data to integer Result: {510 , 0, 109, 619} 5. Pack and store using min number of bits 11/25/2020 135

Checking available HDF 5 Filters • Use API (H 5 Zfilter_avail) • Check libhdf 5. settings file Features: Parallel HDF 5: no ………………………. I/O filters (external): deflate(zlib), szip(encoder) I/O filters (internal): shuffle, fletcher 32, nbit, scaleoffset ………………………. 11/25/2020 136

THIRD PARTY HDF 5 FILTERS 11/25/2020 137

Third-party HDF 5 filters • Compression methods supported by HDF 5 user community http: //www. hdfgroup. org/services/contributions - LZO, BZIP 2, BLOSC (Py. Tables) - LZF (h 5 py) - MAFISC - … - May come as shared libraries • Registration process - 11/25/2020 Helps with filter’s provenance 138

Programming model • Set up search path for filter libraries • Where to search for write and read? • Linux/UNIX “/usr/local/hdf 5/lib/plugin” • Windows "%ALLUSERSPROFILE%/hdf 5/lib/plugin” • 11/25/2020 Default path can be overwritten with the HDF 5_PLUGIN_PATH environment variable 139

HDF 5 Tools • No need to modify “reading tools” such as h 5 dump, h 5 ls, and h 5 copy, and HDFView • Improvement to h 5 dump: replaced “UNKNOWN_FILTER” with “USER_DEFINED_FILTER” • h 5 repack –f UD={ID: k; N: m; CD_VAL: [n 1, …, nm]}…. . • BZIP 2 example h 5 repack –f UD={ID: 307; N: 1; CD_VAL: [9]} file 1. h 5 file 2. h 5 11/25/2020 140

Thank You! Questions? 11/25/2020 141
- Slides: 139