CSE 182 L 6 Protein structure basics Protein

CSE 182 -L 6 Protein structure basics Protein sequencing Fa 05 CSE 182

Announcements • Midterm 1: Nov 1, in class. • Assignment 2: Online, due October 20. Fa 05 CSE 182

Distinguishing between families Fa 05 CSE 182
Distinguishing between families Assignment 2 Fa 05 CSE 182
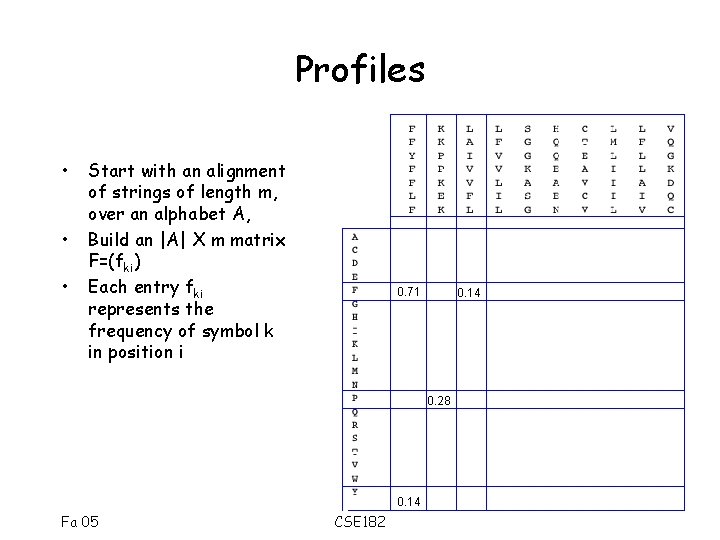
Profiles • • • Start with an alignment of strings of length m, over an alphabet A, Build an |A| X m matrix F=(fki) Each entry fki represents the frequency of symbol k in position i 0. 71 0. 14 0. 28 0. 14 Fa 05 CSE 182
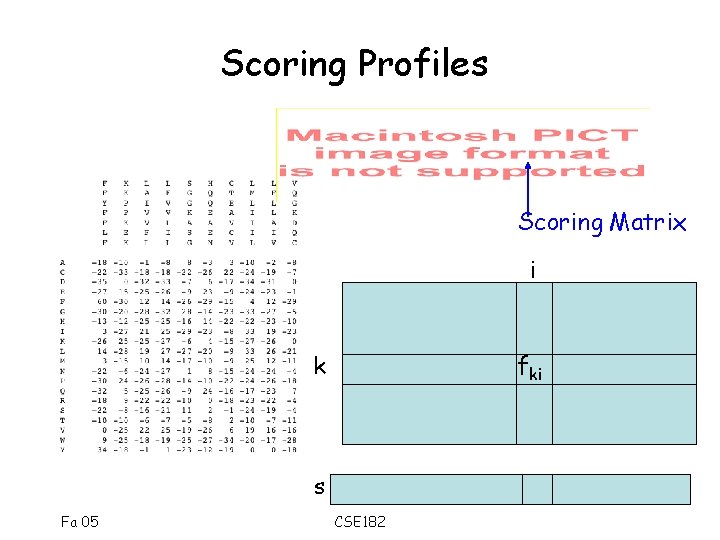
Scoring Profiles Scoring Matrix i k fki s Fa 05 CSE 182

Psi-BLAST idea • Multiple alignments are important for capturing remote homology. • Profile based scores are a natural way to handle this. • Q: What if the query is a single sequence. • A: Iterate: – Find homologs using Blast on query – Discard very similar homologs – Align, make a profile, search with profile. Fa 05 CSE 182

Psi-BLAST speed • Two time consuming steps. 1. Multiple alignment of homologs 2. Searching with Profiles. 1. Does the keyword search idea work? • • Multiple alignment: – Use ungapped multiple alignments only Pigeonhole principle again: – – Fa 05 CSE 182 If profile of length m must score >= T Then, a sub-profile of length l must score >= l. T|/m Generate all l-mers that score at least l. T|/M Search using an automaton
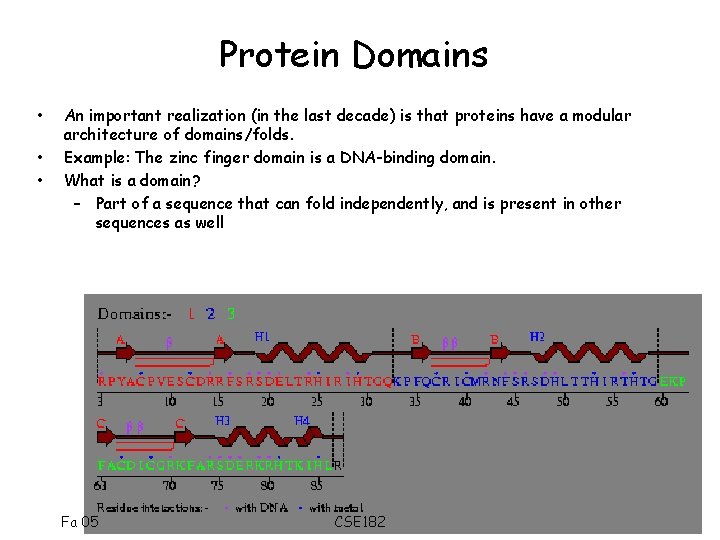
Protein Domains • • • An important realization (in the last decade) is that proteins have a modular architecture of domains/folds. Example: The zinc finger domain is a DNA-binding domain. What is a domain? – Part of a sequence that can fold independently, and is present in other sequences as well Fa 05 CSE 182

Domain review • What is a domain? • How are domains expressed – – Fa 05 Motifs (Regular expression & others) Multiple alignments Profile HMMs CSE 182

Domain databases Can you speed up HMM search? Fa 05 CSE 182

A structural view of proteins Fa 05 CSE 182

CS view of a protein • >sp|P 00974|BPT 1_BOVIN Pancreatic trypsin inhibitor precursor (Basic protease inhibitor) (BPI) (BPTI) (Aprotinin) - Bos taurus (Bovine). • MKMSRLCLSVALLVLLGTLAASTPGCDTSNQAKAQ RPDFCLEPPYTGPCKARIIRYFYNAKAGLCQTFVYGG CRAKRNNFKSAEDCMRTCGGAIGPWENL Fa 05 CSE 182
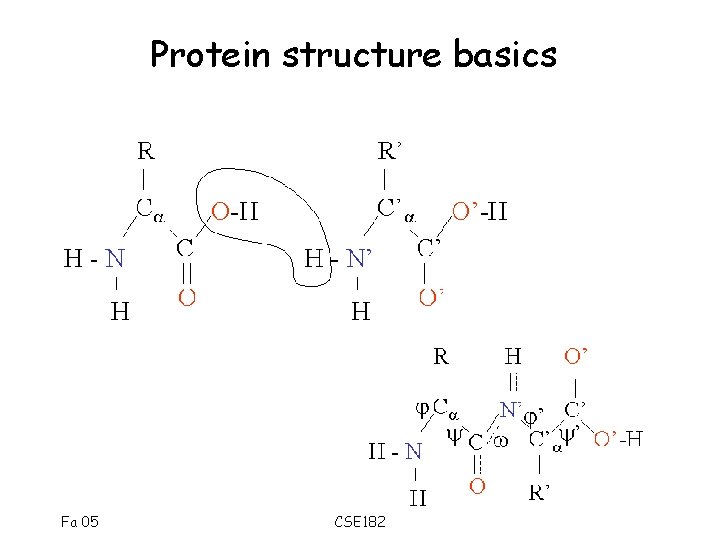
Protein structure basics Fa 05 CSE 182

Side chains determine amino-acid type • The residues may have different properties. • Aspartic acid (D), and Glutamic Acid (E) are acidic residues Fa 05 CSE 182
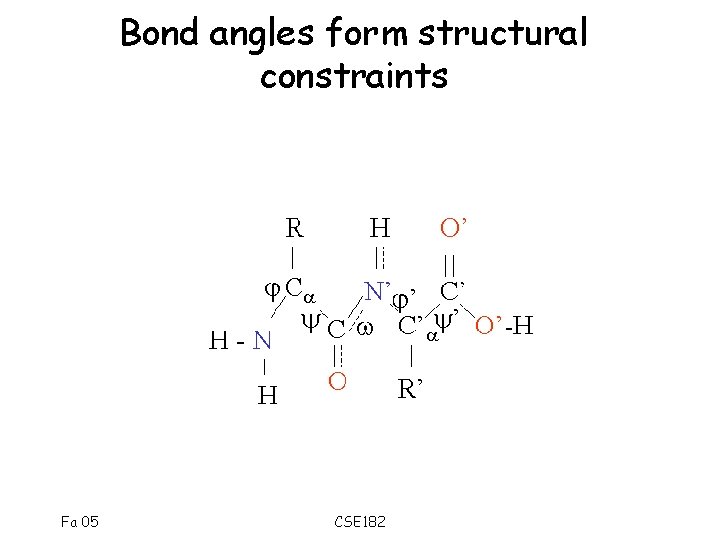
Bond angles form structural constraints Fa 05 CSE 182

Various constraints determine 3 d structure • Constraints – Structural constraints due to physiochemical properties – Constraints due to bond angles – H-bond formation • Surprisingly, a few conformations are seen over and over again. Fa 05 CSE 182

Alpha-helix • 3. 6 residues per turn • H-bonds between 1 st and 4 th residue stabilize the structure. • First discovered by Linus Pauling Fa 05 CSE 182

Beta-sheet • • • Each strand by itself has 2 residues per turn, and is not stable. Adjacent strands hydrogen-bond to form stable beta-sheets, parallel or anti-parallel. Beta sheets have long range interactions that stabilize the structure, while alpha-helices have local interactions. Fa 05 CSE 182

Domains • The basic structures (helix, strand, loop) combine to form complex 3 D structures. • Certain combinations are popular. Many sequences, but only a few folds Fa 05 CSE 182
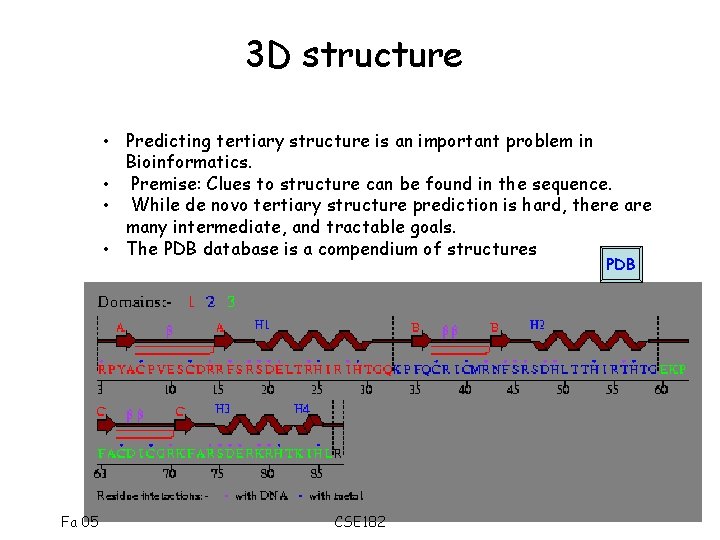
3 D structure • Predicting tertiary structure is an important problem in Bioinformatics. • Premise: Clues to structure can be found in the sequence. • While de novo tertiary structure prediction is hard, there are many intermediate, and tractable goals. • The PDB database is a compendium of structures PDB Fa 05 CSE 182

Searching structure databases • Threading, and other 3 d Alignments can be used to align structures. • Database filtering is possible through geometric hashing. Fa 05 CSE 182

Trivia Quiz • What research won the Nobel prize in Chemistry in 2004? • In 2002? Fa 05 CSE 182

How are Proteins Sequenced? Mass Spec 101: Fa 05 CSE 182

Nobel Citation 2002 Fa 05 CSE 182

Nobel Citation, 2002 Fa 05 CSE 182
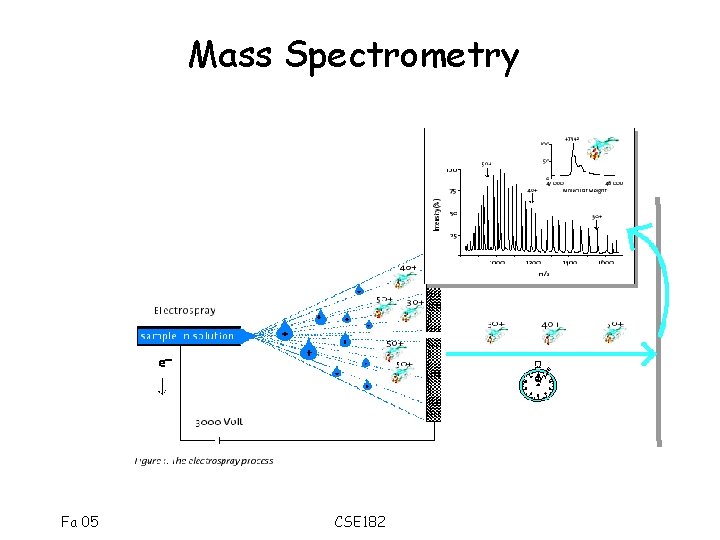
Mass Spectrometry Fa 05 CSE 182
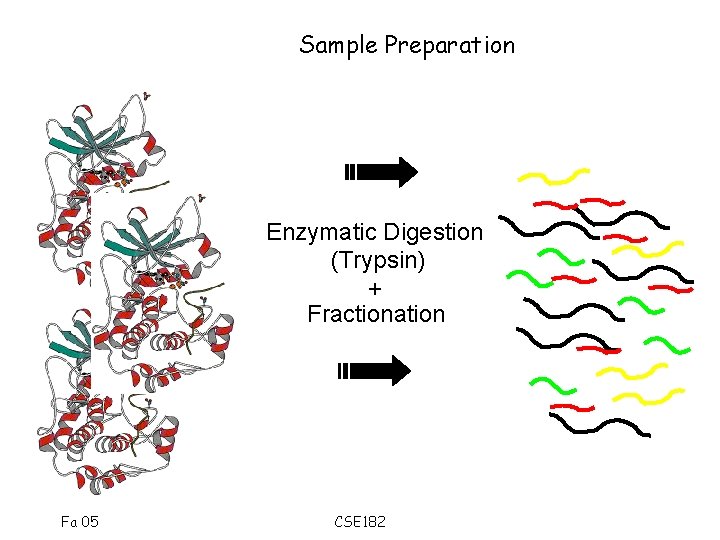
Sample Preparation Enzymatic Digestion (Trypsin) + Fractionation Fa 05 CSE 182
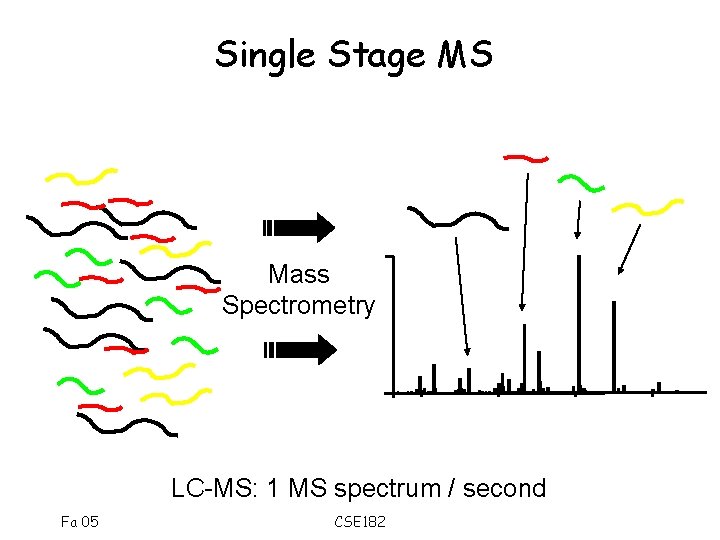
Single Stage MS Mass Spectrometry LC-MS: 1 MS spectrum / second Fa 05 CSE 182
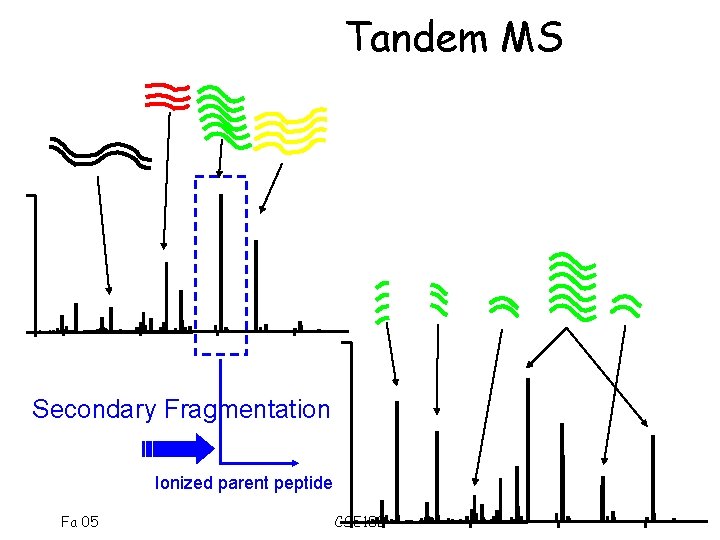
Tandem MS Secondary Fragmentation Ionized parent peptide Fa 05 CSE 182
- Slides: 30