Bioinformatics Ahmet Sacan 06 Protein secondary structure prediction

Bioinformatics Ahmet Sacan 06. Protein (secondary) structure prediction Images are from "Understanding Bioinformatics" and Wikipedia.
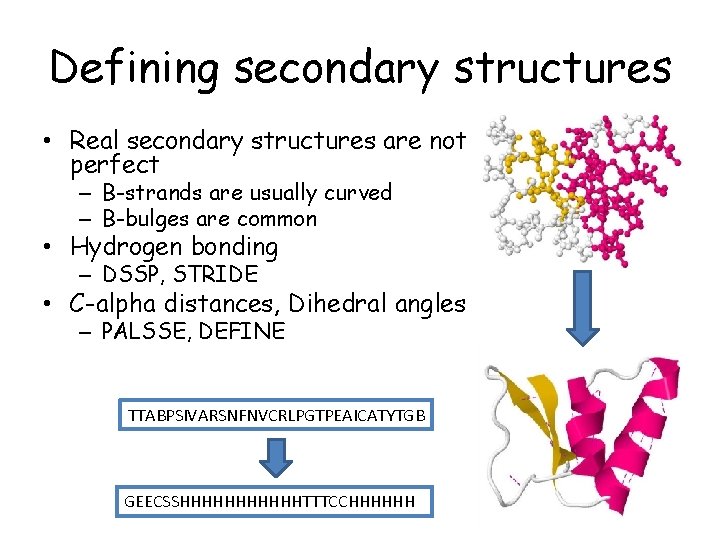
Defining secondary structures • Real secondary structures are not perfect – B-strands are usually curved – B-bulges are common • Hydrogen bonding – DSSP, STRIDE • C-alpha distances, Dihedral angles – PALSSE, DEFINE TTABPSIVARSNFNVCRLPGTPEAICATYTGB GEECSSHHHHHHTTTCCHHHHHH
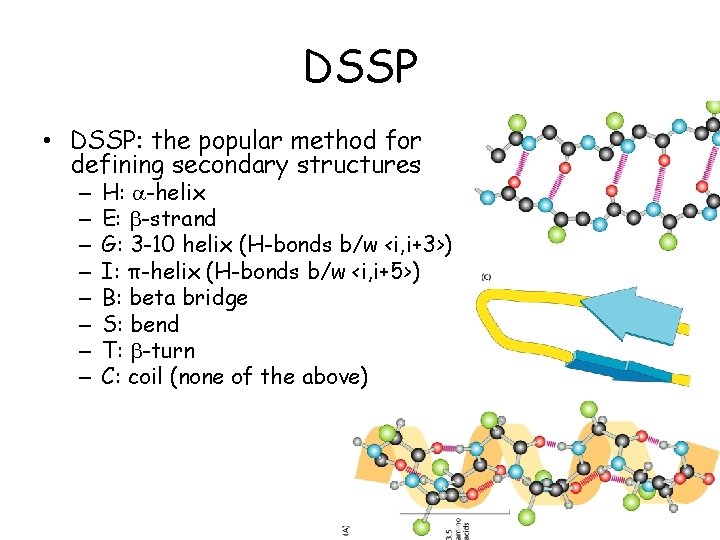
DSSP • DSSP: the popular method for defining secondary structures – – – – H: -helix E: -strand G: 3 -10 helix (H-bonds b/w <i, i+3>) I: π-helix (H-bonds b/w <i, i+5>) B: beta bridge S: bend T: -turn C: coil (none of the above)
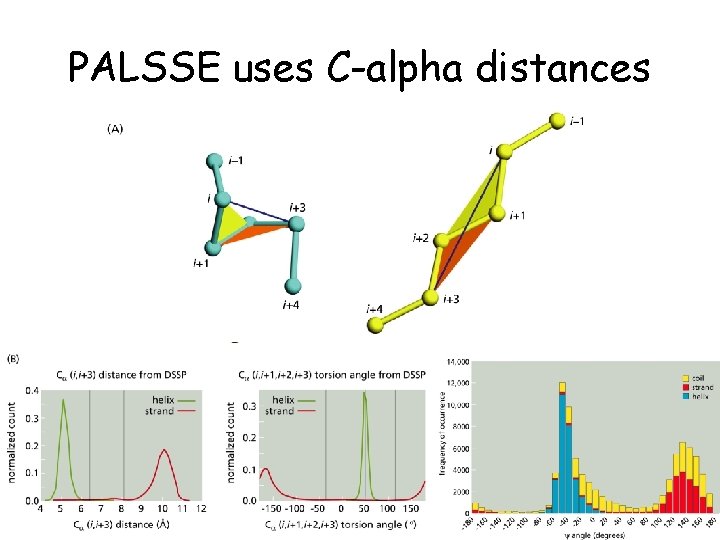
PALSSE uses C-alpha distances
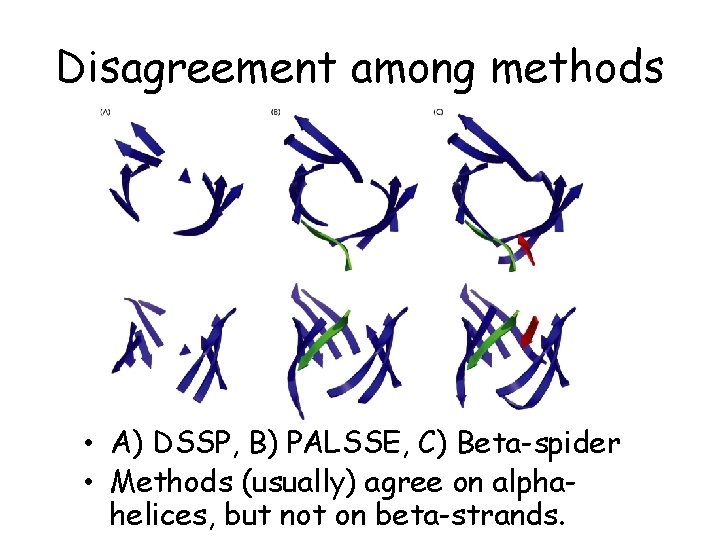
Disagreement among methods • A) DSSP, B) PALSSE, C) Beta-spider • Methods (usually) agree on alphahelices, but not on beta-strands.
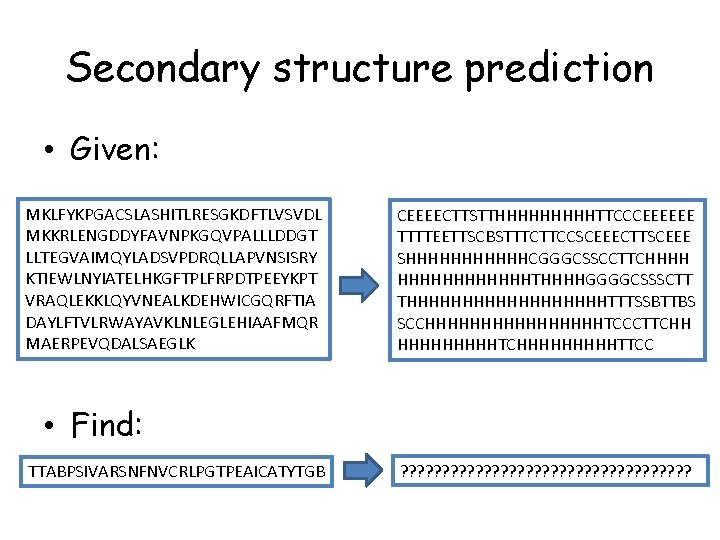
Secondary structure prediction • Given: MKLFYKPGACSLASHITLRESGKDFTLVSVDL MKKRLENGDDYFAVNPKGQVPALLLDDGT LLTEGVAIMQYLADSVPDRQLLAPVNSISRY KTIEWLNYIATELHKGFTPLFRPDTPEEYKPT VRAQLEKKLQYVNEALKDEHWICGQRFTIA DAYLFTVLRWAYAVKLNLEGLEHIAAFMQR MAERPEVQDALSAEGLK CEEEECTTSTTHHHHHTTCCCEEEEEE TTTTEETTSCBSTTTCTTCCSCEEECTTSCEEE SHHHHHHCGGGCSSCCTTCHHHHHHTHHHHGGGGCSSSCTT THHHHHHHHHTTTSSBTTBS SCCHHHHHHHHTCCCTTCHH HHHHHTCHHHHHTTCC • Find: TTABPSIVARSNFNVCRLPGTPEAICATYTGB ? ? ? ? ? ? ? ? ?
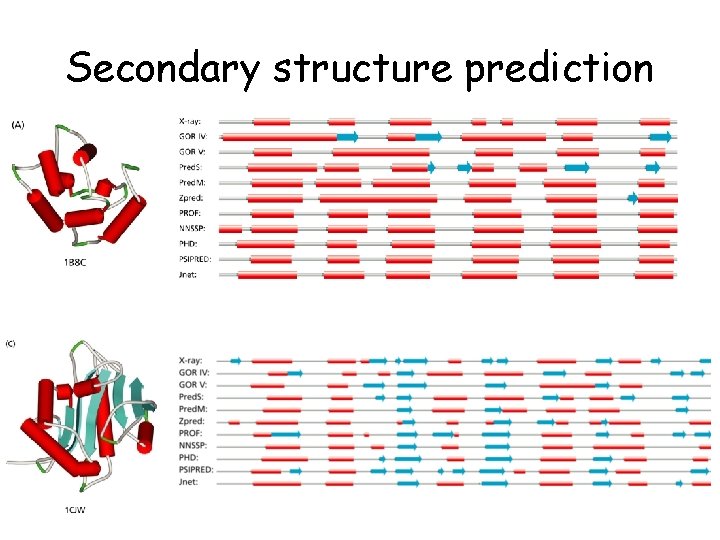
Secondary structure prediction

Determining success •

SOV is more useful for foldrecognition applications SOV Q 3 X-Ray CHHHHHC 100 Prediction 1 CHCHCHCC 12 58 Prediction 2 CCCHHHHHCCCC 63 58 Prediction 3 CHHHCHHC 41 83 Prediction 4 CHHCCHHHHHCC 52 75 Prediction 5 CCCHHHHHHCCC 80 67 • Predictions 1 and 2 have same composition, but 1 is unrealistic. • SOV gives higher scores when only one helix is predicted (2, 5)

Specialized scores for transmembrane helix prediction --> the method misses the first 2 -5 residues of the helix.

Training/Test datasets • Include a range of different and unrelated sequences and structures – Avoid compositional bias (e. g. , more alpha helices and random coils than B-strands) • No protein in test set should be homologous to training set proteins. – Use "non-redundant", "representative" datasets (PDB_SELECT, ASTRAL, etc. )
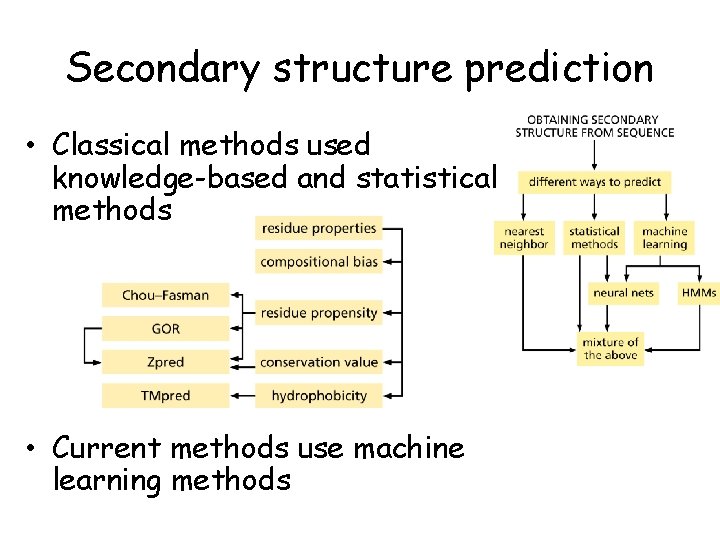
Secondary structure prediction • Classical methods used knowledge-based and statistical methods • Current methods use machine learning methods
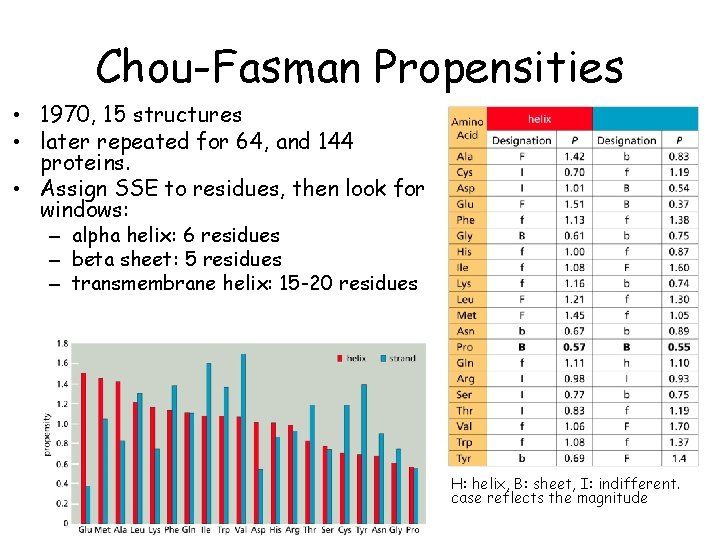
Chou-Fasman Propensities • 1970, 15 structures • later repeated for 64, and 144 proteins. • Assign SSE to residues, then look for windows: – alpha helix: 6 residues – beta sheet: 5 residues – transmembrane helix: 15 -20 residues H: helix, B: sheet, I: indifferent. case reflects the magnitude
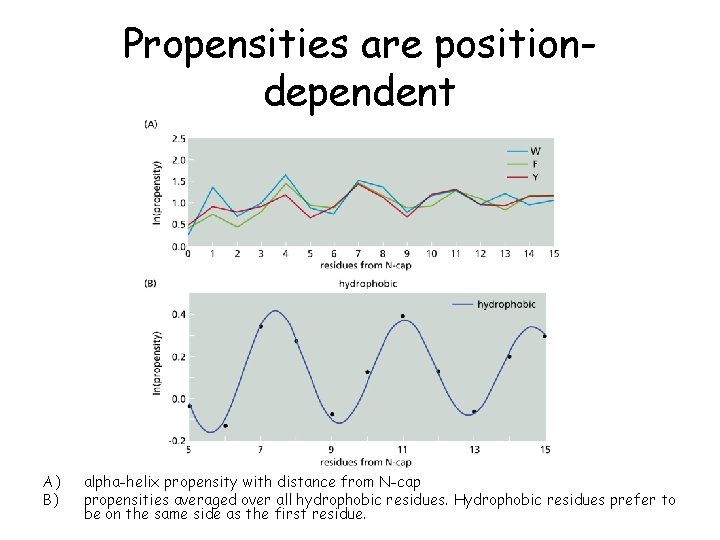
Propensities are positiondependent A) B) alpha-helix propensity with distance from N-cap propensities averaged over all hydrophobic residues. Hydrophobic residues prefer to be on the same side as the first residue.

GOR • GOR method uses information theory – Self: information contained by residue type – Directional: information contained by types of other residues (without using self-type) – Pair: same as directional, but including the self residue type. The effect of Pro vs. Met at position j+5 on alpha helix probability
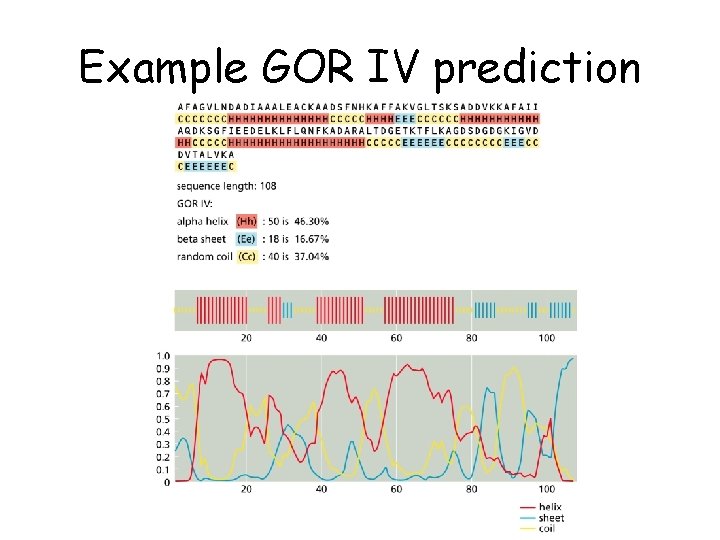
Example GOR IV prediction
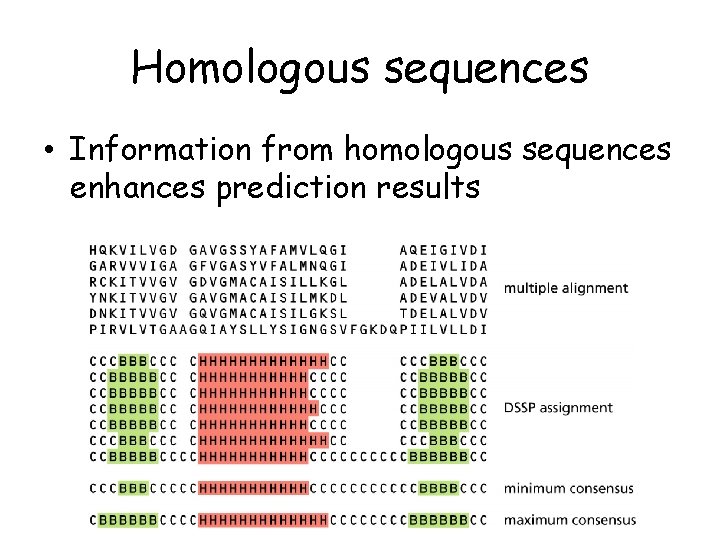
Homologous sequences • Information from homologous sequences enhances prediction results
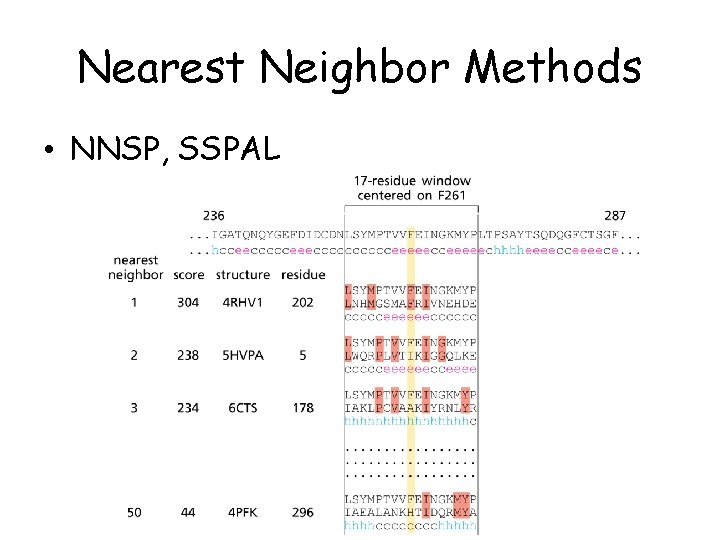
Nearest Neighbor Methods • NNSP, SSPAL
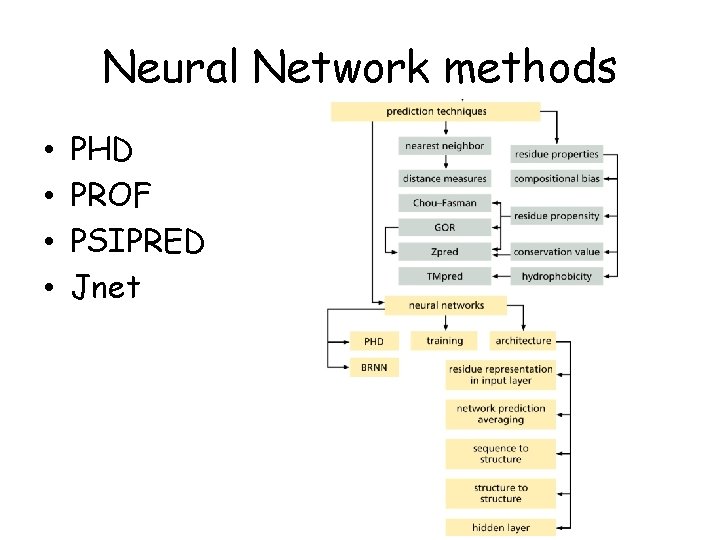
Neural Network methods • • PHD PROF PSIPRED Jnet
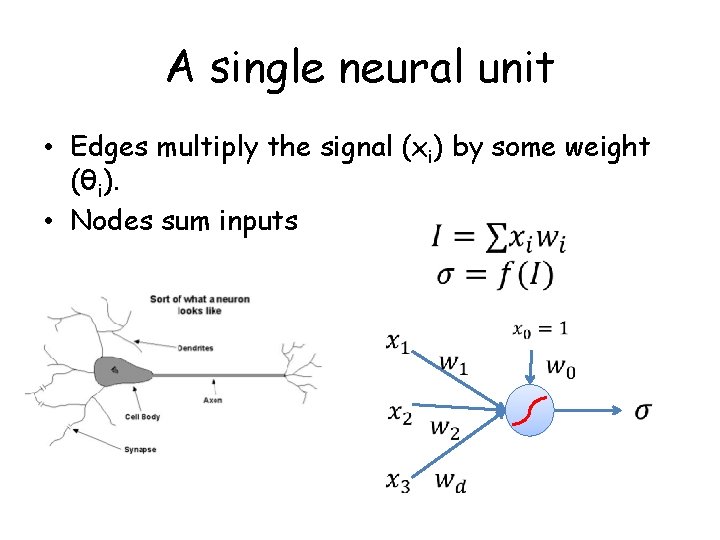
A single neural unit • Edges multiply the signal (xi) by some weight (θi). • Nodes sum inputs
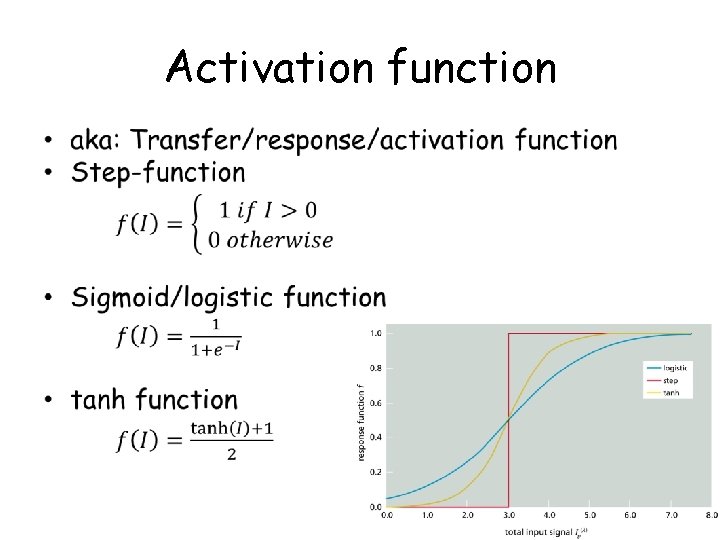
Activation function •
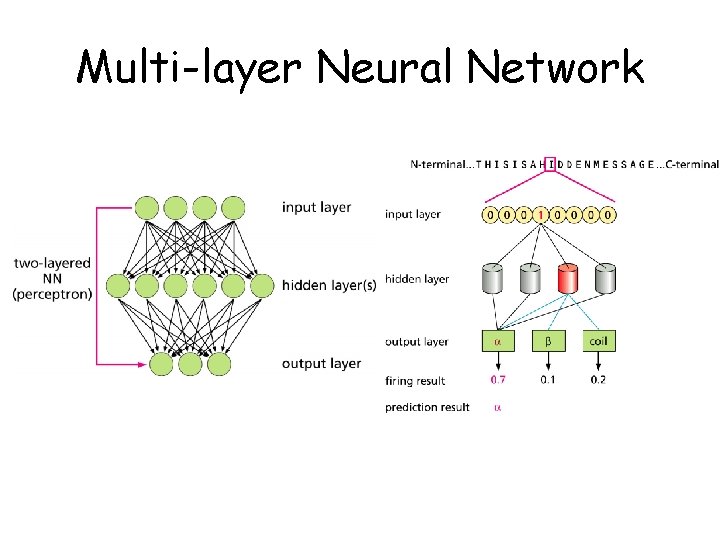
Multi-layer Neural Network
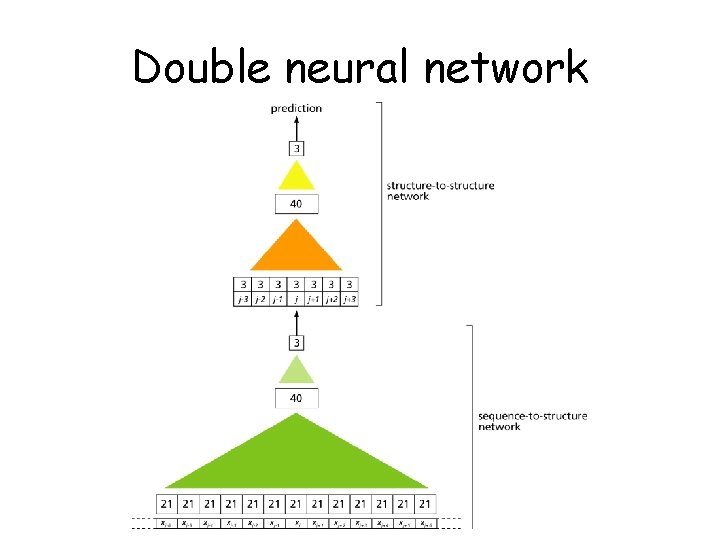
Double neural network
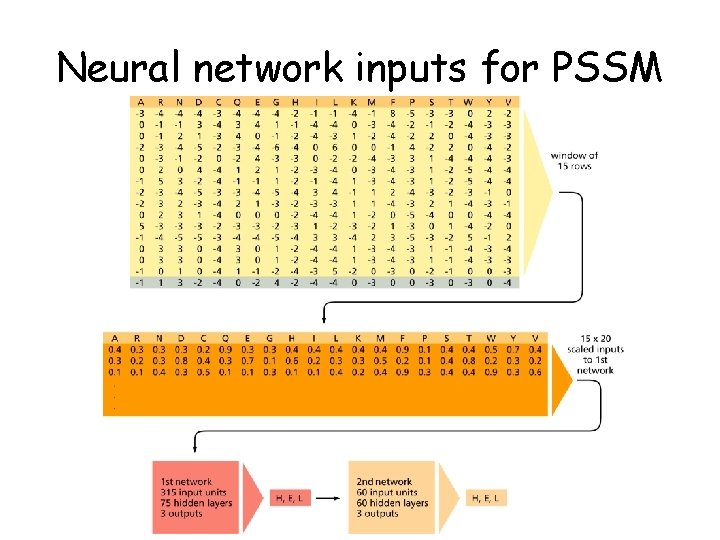
Neural network inputs for PSSM

Training Neural Network • Given: – sets of input x and correct output y – adjust weights, so network output is similar to y. • Gradient Descent: – Repeatedly move in the best direction that minimizes error. • Back-propagation: – Back-propagate the error into previous layers, so their weights can also be adjusted.
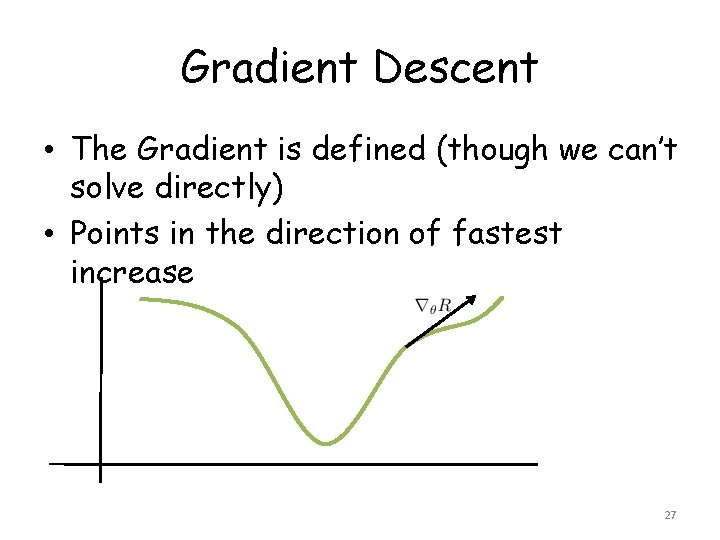
Gradient Descent • The Gradient is defined (though we can’t solve directly) • Points in the direction of fastest increase 27
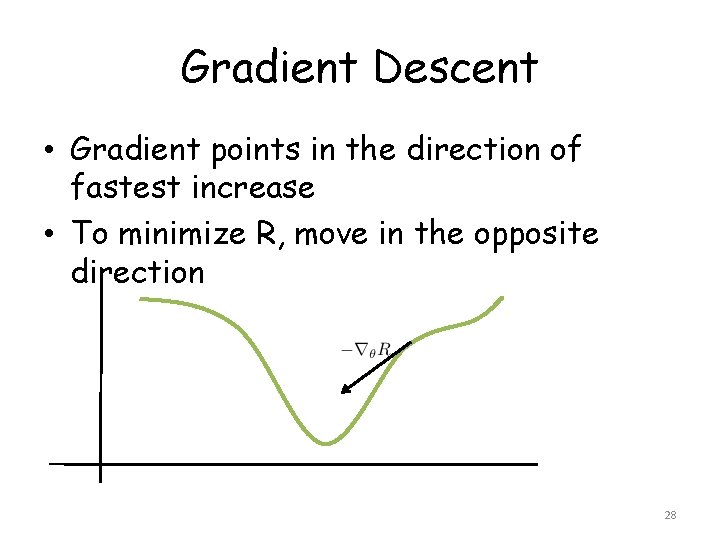
Gradient Descent • Gradient points in the direction of fastest increase • To minimize R, move in the opposite direction 28
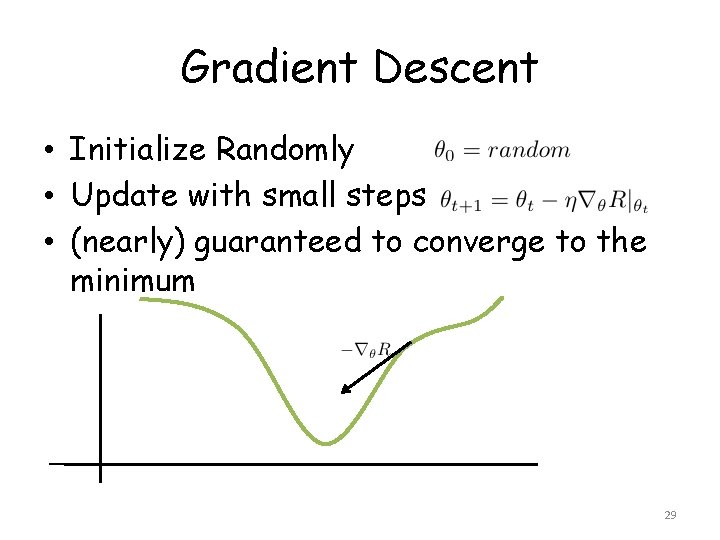
Gradient Descent • Initialize Randomly • Update with small steps • (nearly) guaranteed to converge to the minimum 29
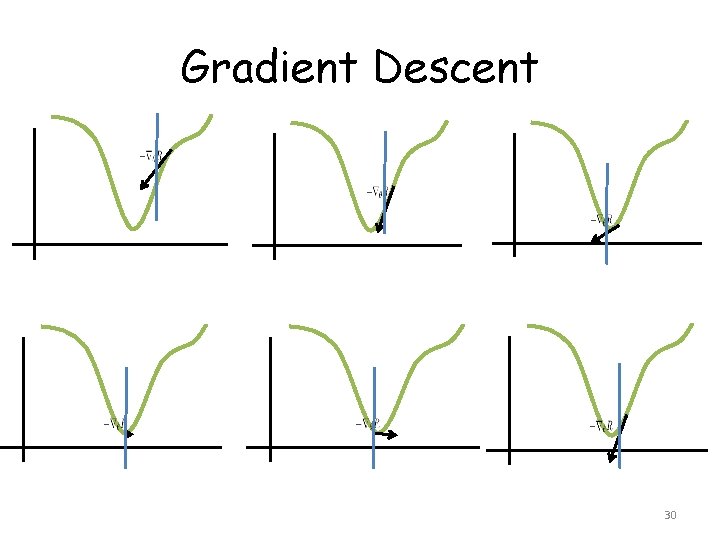
Gradient Descent 30
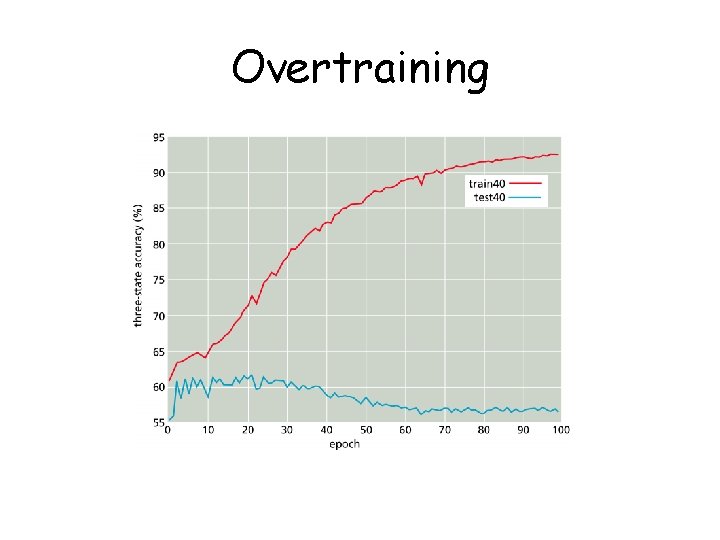
Overtraining
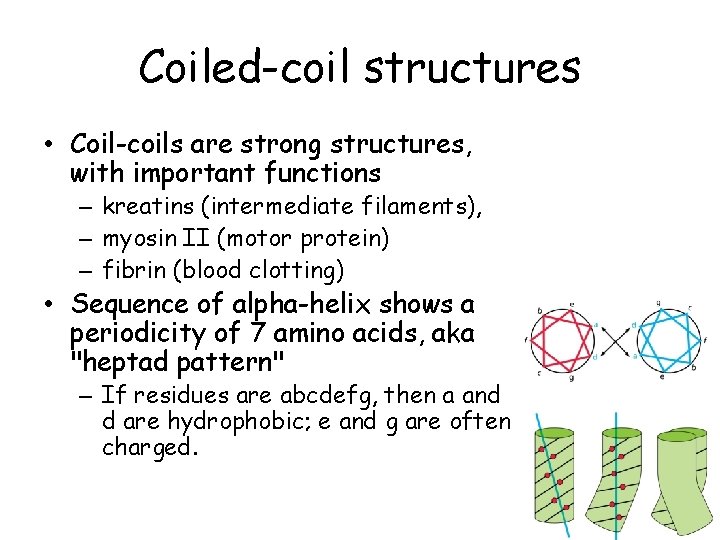
Coiled-coil structures • Coil-coils are strong structures, with important functions – kreatins (intermediate filaments), – myosin II (motor protein) – fibrin (blood clotting) • Sequence of alpha-helix shows a periodicity of 7 amino acids, aka "heptad pattern" – If residues are abcdefg, then a and d are hydrophobic; e and g are often charged.

Coiled-coils • Leucine zippers – The repeating residues a and d are Leu. – Occur in transcription factors.

Transmembrane helix prediction • Find just the membrane helices; or additionally find inside/outside helices and loops and topology. • Simplest methods use hydrohobicity – hydropathic profile: average hydrophobicity over a sliding window (e. g. , 15 -residues)
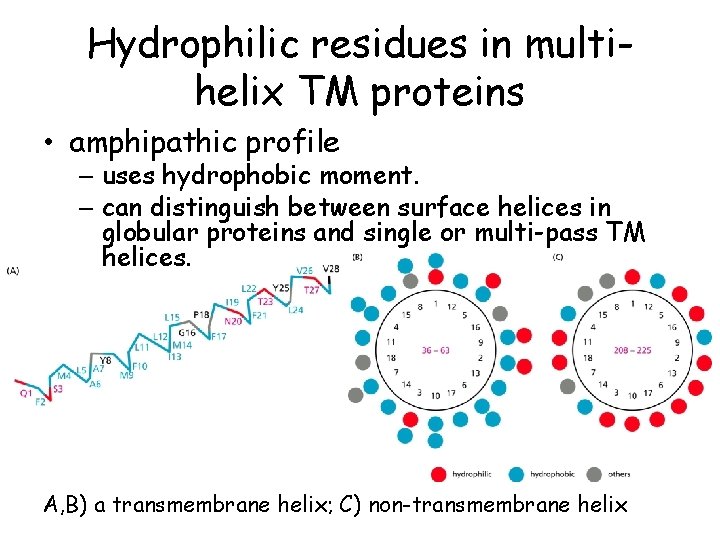
Hydrophilic residues in multihelix TM proteins • amphipathic profile – uses hydrophobic moment. – can distinguish between surface helices in globular proteins and single or multi-pass TM helices. A, B) a transmembrane helix; C) non-transmembrane helix

Orienting the helix • Positive inside rule: Intracellular loops have higher concentration of Arg (R) and Lys (K). • Or, build the inside/outside into the model. • SOSUI assumes at least one hydrophobic helix is present (primary helix), and it uses the following features to identify others: – hydropathy index – ampiphilicity index – index of amino acid charges
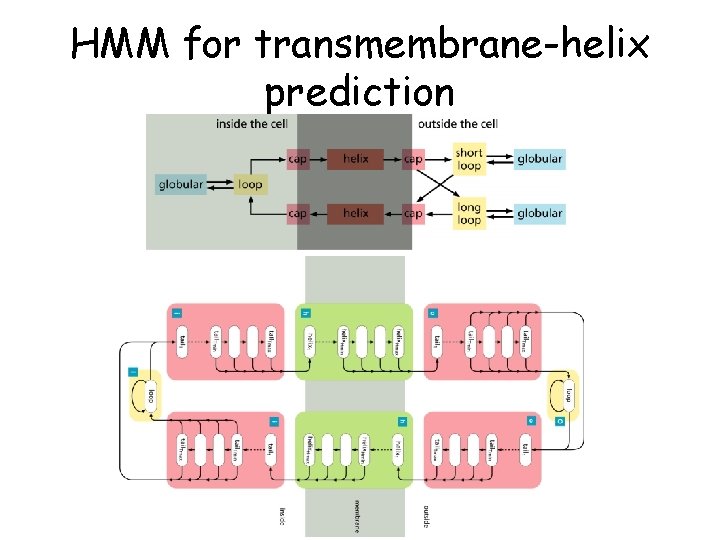
HMM for transmembrane-helix prediction
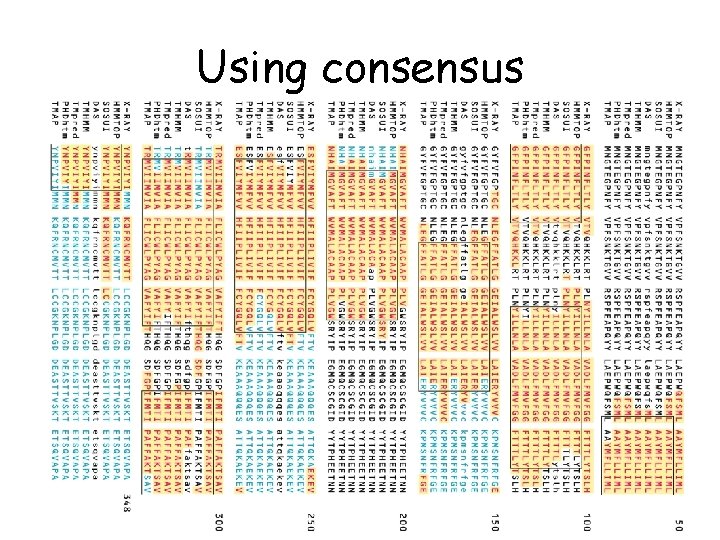
Using consensus
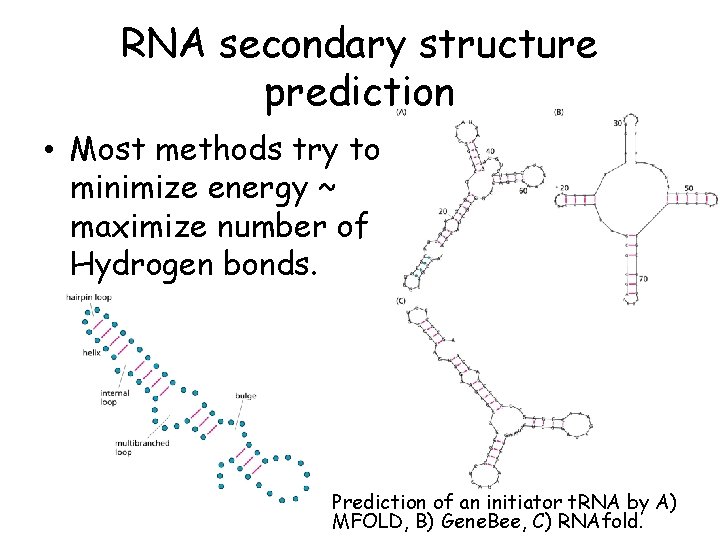
RNA secondary structure prediction • Most methods try to minimize energy ~ maximize number of Hydrogen bonds. Prediction of an initiator t. RNA by A) MFOLD, B) Gene. Bee, C) RNAfold.

- Slides: 40