CS 5263 Bioinformatics RNA Secondary Structure Prediction Outline

CS 5263 Bioinformatics RNA Secondary Structure Prediction

Outline • Biological roles for RNA • RNA secondary structure – What’s “secondary structure”? – How is it represented? – Why is it important? • How to predict?
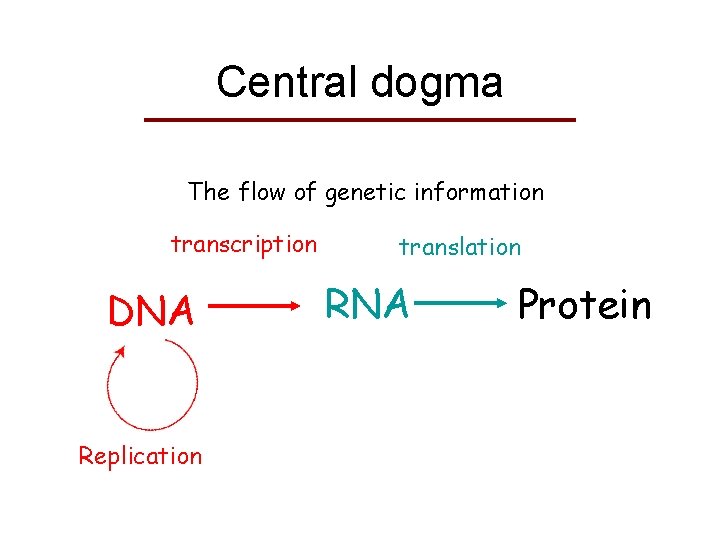
Central dogma The flow of genetic information transcription DNA Replication translation RNA Protein
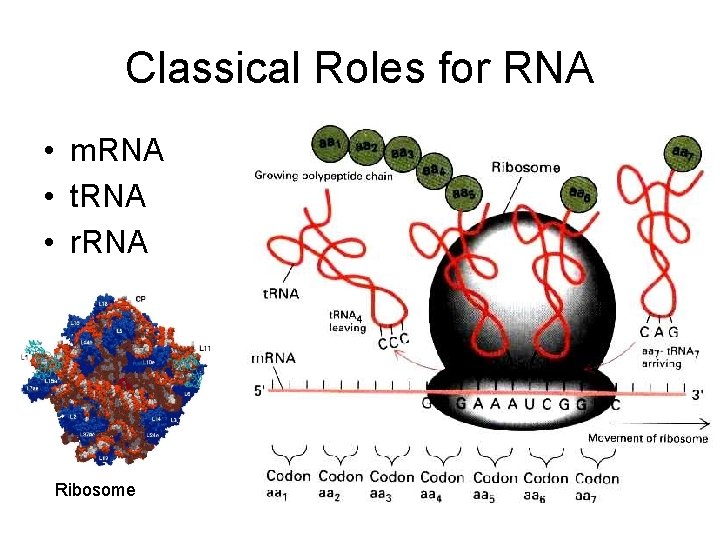
Classical Roles for RNA • m. RNA • t. RNA • r. RNA Ribosome

“Semi-classical” RNA • sn. RNA - small nuclear RNA (60 -300 nt), involved in splicing (removing introns), etc. • RNase. P - t. RNA processing (~300 nt) • SRP RNA - signal recognition particle RNA: membrane targeting (~100 -300 nt) • tm. RNA - resetting stalled ribosomes, destroy aberrant m. RNA • Telomerase - (200 -400 nt) • sno. RNA - small nucleolar RNA (many varieties; 80 -200 nt)

Non-coding RNAs Dramatic discoveries in last 10 years • 100 s of new families • Many roles: regulation, transport, stability, catalysis, … • si. RNA: Small interfering RNA (Nobel prize 2006) and mi. RNAs: both are ~21 -23 nt – Regulating gene expression – Evidence of diseaseassociation • 1% of DNA codes for protein, but 30% of it is copied into RNA, i. e. nc. RNA >> m. RNA

Take-home message • RNAs play many important roles in the cell beyond the classical roles – Many of which yet to be discovered • RNA functions are determined by structures
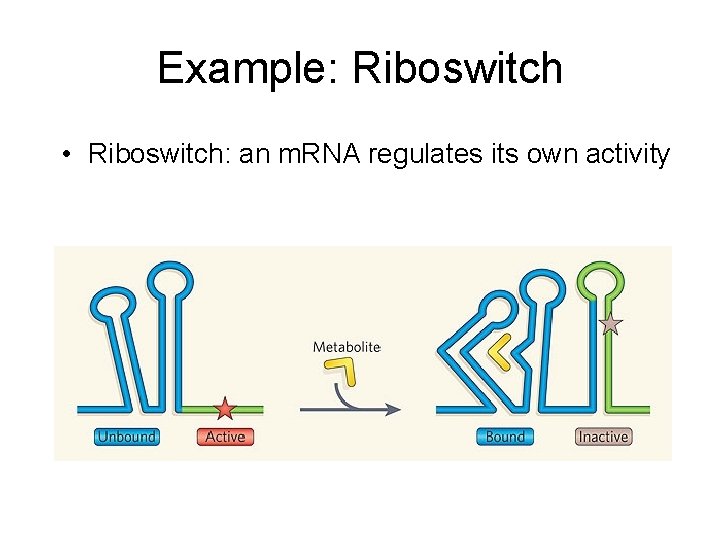
Example: Riboswitch • Riboswitch: an m. RNA regulates its own activity

RNA structure • Primary: sequence • Secondary: base-pairing • Tertiary: 3 D shape
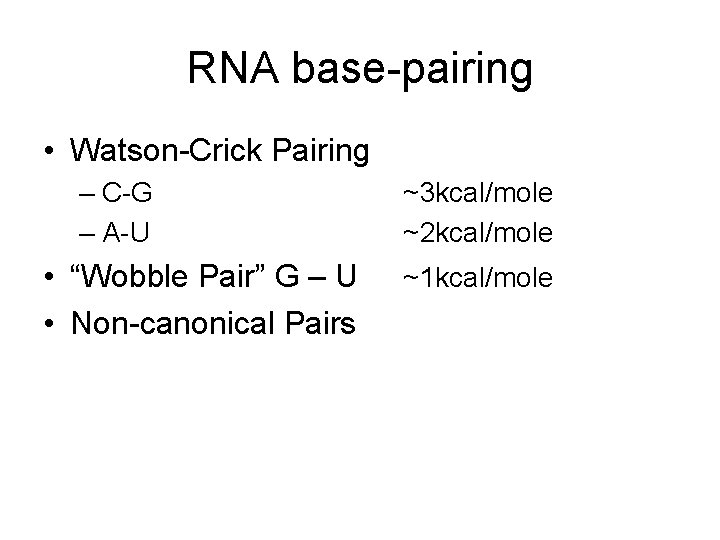
RNA base-pairing • Watson-Crick Pairing – C-G – A-U • “Wobble Pair” G – U • Non-canonical Pairs ~3 kcal/mole ~2 kcal/mole ~1 kcal/mole
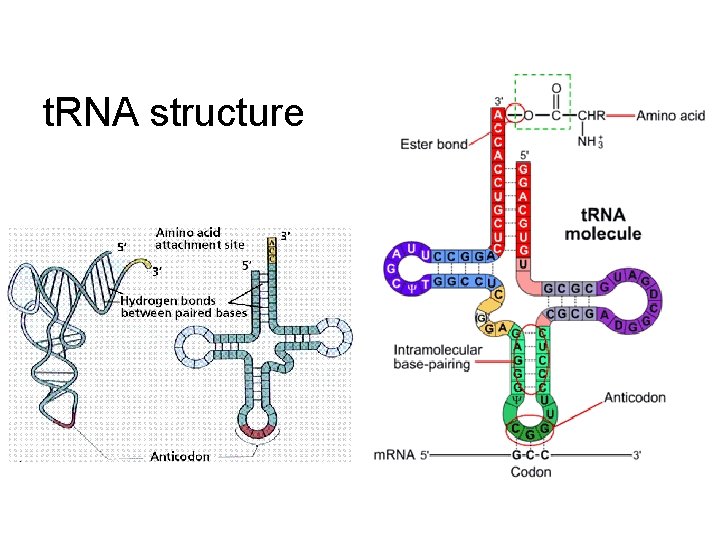
t. RNA structure
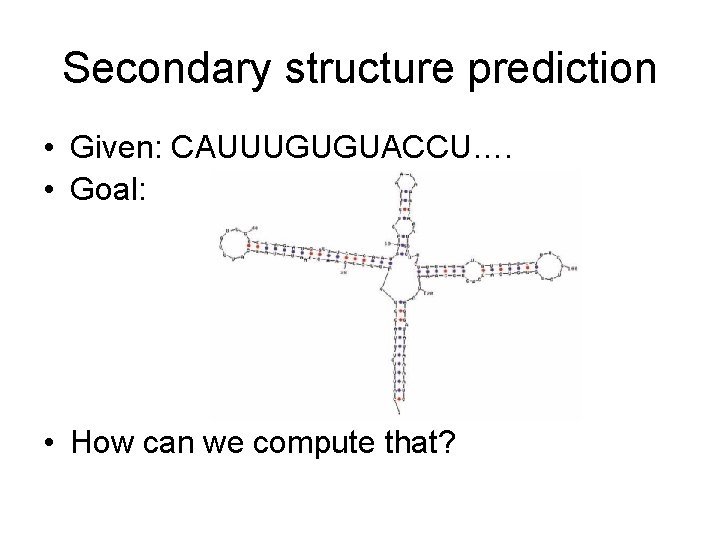
Secondary structure prediction • Given: CAUUUGUGUACCU…. • Goal: • How can we compute that?
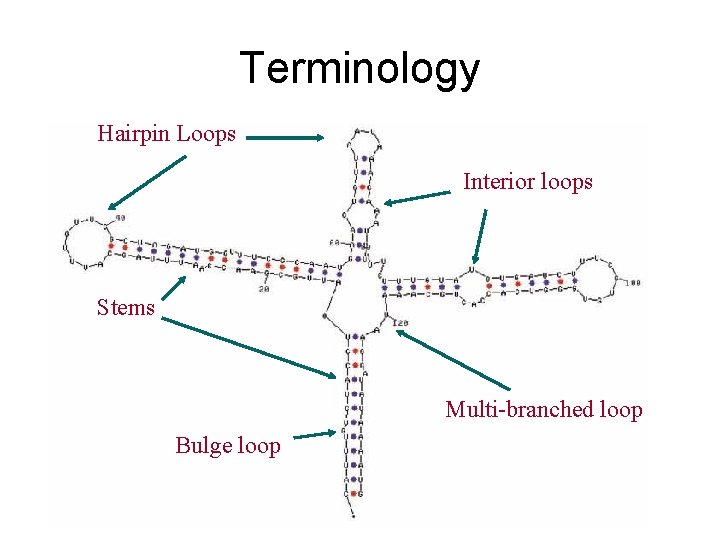
Terminology Hairpin Loops Interior loops Stems Multi-branched loop Bulge loop
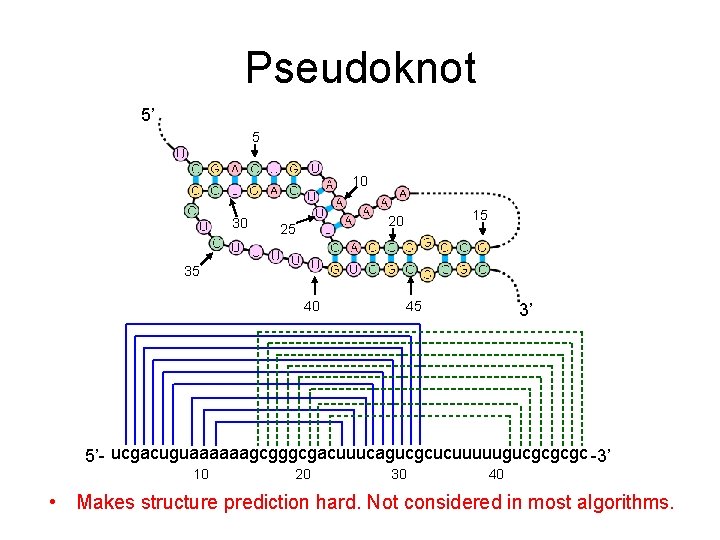
Pseudoknot 5’ 5 10 30 15 20 25 35 40 45 3’ 5’- ucgacuguaaaaaagcgggcgacuuucagucgcucuuuuugucgcgcgc -3’ 10 20 30 40 • Makes structure prediction hard. Not considered in most algorithms.

The Nussinov algorithm • Goal: maximizing the number of basepairs • Idea: Dynamic programming – Loop matching – Nussinov, Pieczenik, Griggs, Kleitman ’ 78 • Too simple for accurate prediction, but stepping-stone for later algorithms
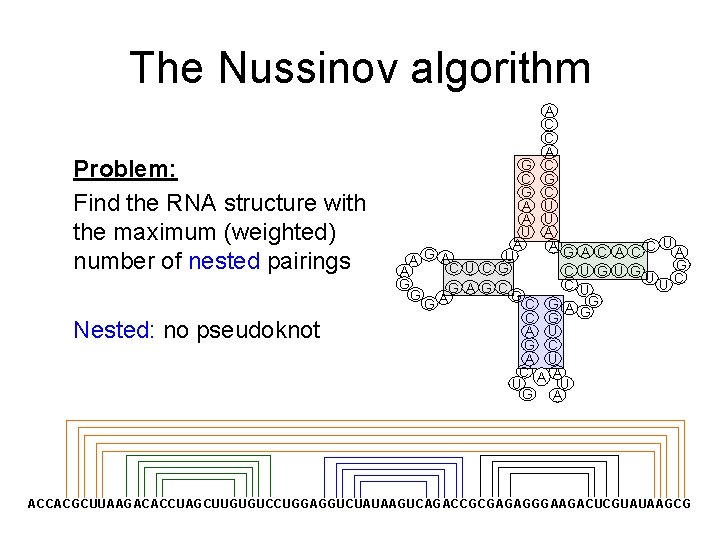
The Nussinov algorithm Problem: Find the RNA structure with the maximum (weighted) number of nested pairings Nested: no pseudoknot A C C A G C C G G C A U U A A AG G U A A C U C G C A G C G A G G C G A U G C A U A U G A U A CC A G U G UUC U G G ACCACGCUUAAGACACCUAGCUUGUGUCCUGGAGGUCUAUAAGUCAGACCGCGAGAGGGAAGACUCGUAUAAGCG
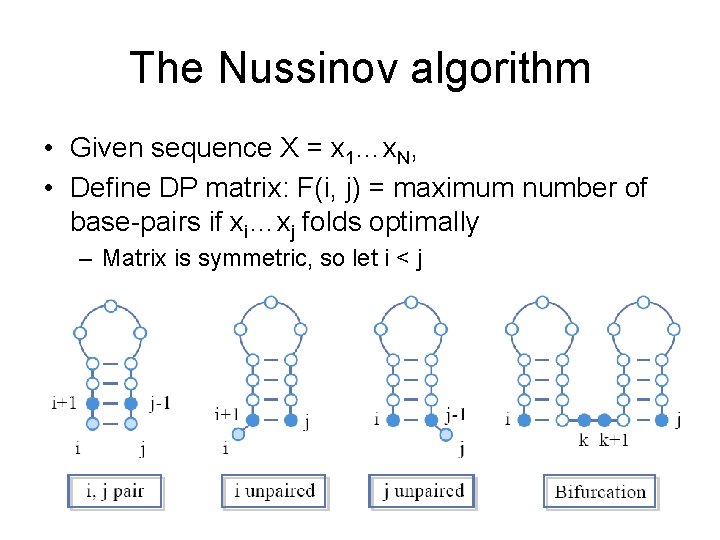
The Nussinov algorithm • Given sequence X = x 1…x. N, • Define DP matrix: F(i, j) = maximum number of base-pairs if xi…xj folds optimally – Matrix is symmetric, so let i < j
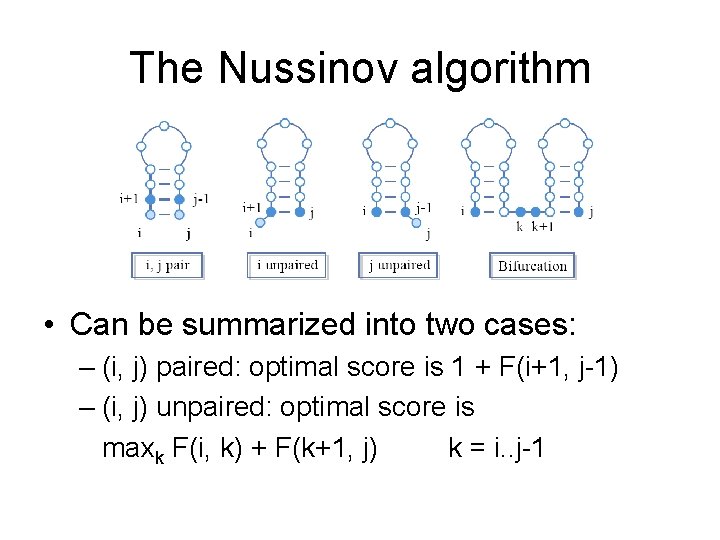
The Nussinov algorithm • Can be summarized into two cases: – (i, j) paired: optimal score is 1 + F(i+1, j-1) – (i, j) unpaired: optimal score is maxk F(i, k) + F(k+1, j) k = i. . j-1

The Nussinov algorithm • F(i, i) = 0 F(i+1, j-1) + S(xi, xj) • F(i, j) = maxk=i. . j-1 F(i, k) + F(k+1, j) • S(xi, xj) = 1 if xi, xj can form a base-pair, and 0 otherwise – Generalize: S(A, U) = 2, S(C, G) = 3, S(G, U) = 1 – Or other types of scores (later) • F(1, N) gives the optimal score for the whole seq
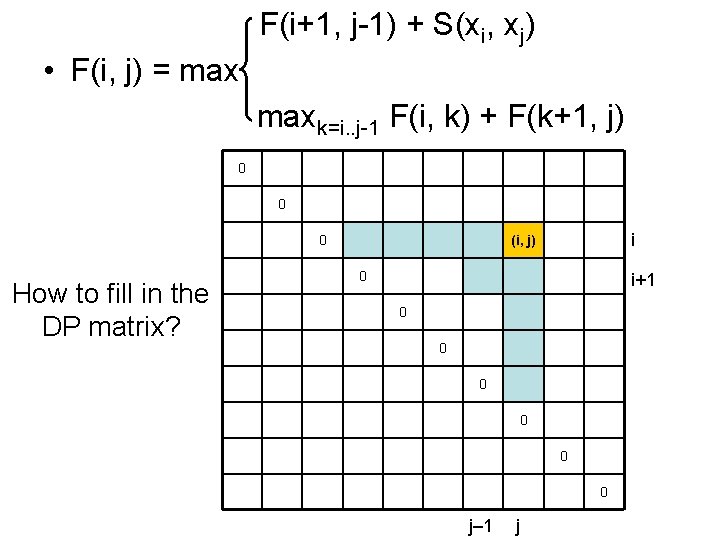
F(i+1, j-1) + S(xi, xj) • F(i, j) = maxk=i. . j-1 F(i, k) + F(k+1, j) 0 0 0 How to fill in the DP matrix? i (i, j) 0 i+1 0 0 0 j– 1 j
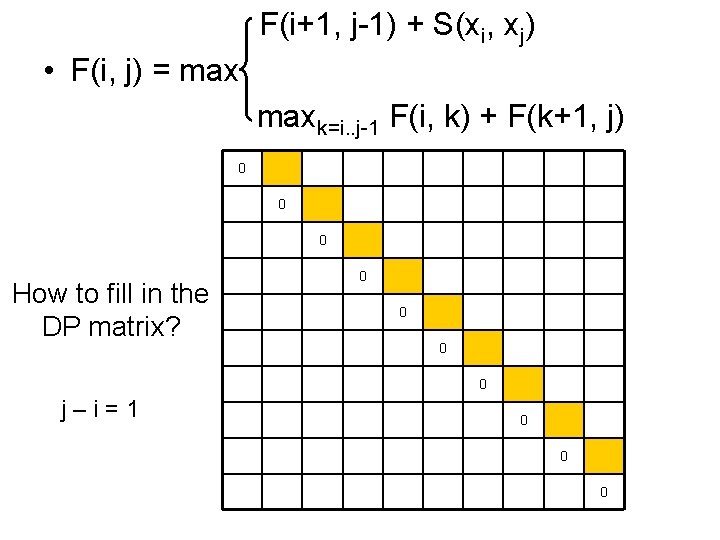
F(i+1, j-1) + S(xi, xj) • F(i, j) = maxk=i. . j-1 F(i, k) + F(k+1, j) 0 0 0 How to fill in the DP matrix? 0 0 j–i=1 0 0 0
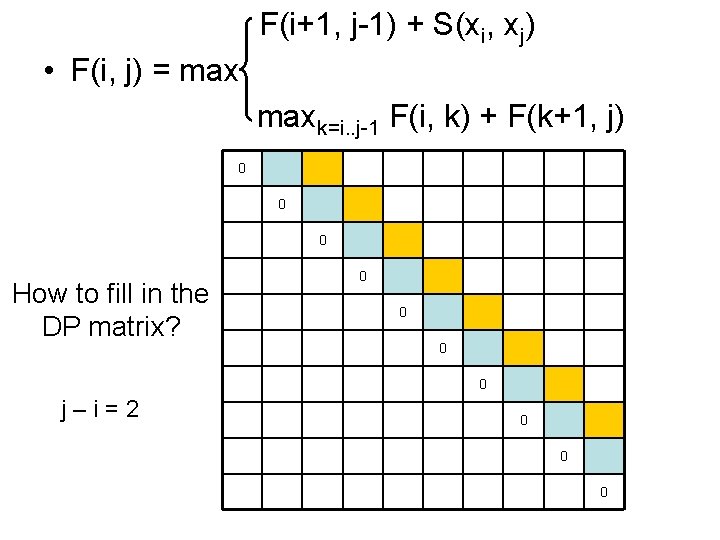
F(i+1, j-1) + S(xi, xj) • F(i, j) = maxk=i. . j-1 F(i, k) + F(k+1, j) 0 0 0 How to fill in the DP matrix? 0 0 j–i=2 0 0 0
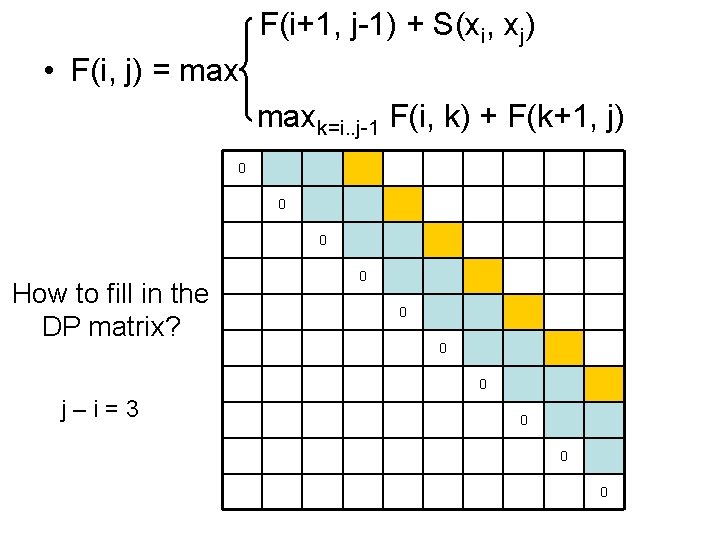
F(i+1, j-1) + S(xi, xj) • F(i, j) = maxk=i. . j-1 F(i, k) + F(k+1, j) 0 0 0 How to fill in the DP matrix? 0 0 j–i=3 0 0 0
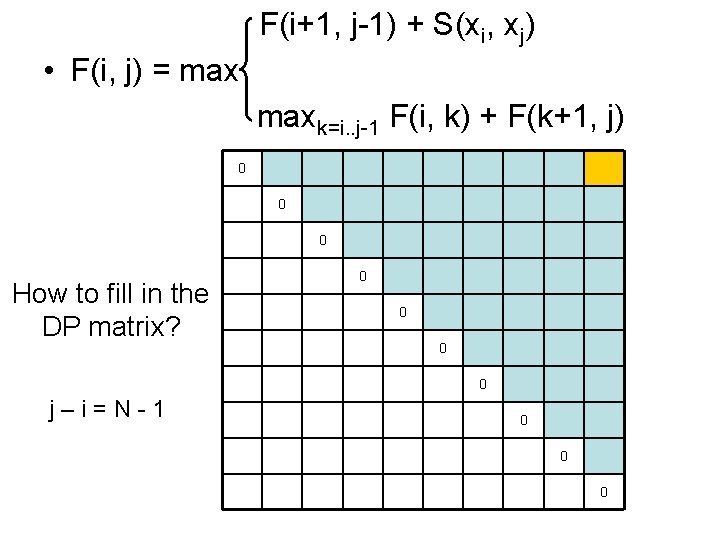
F(i+1, j-1) + S(xi, xj) • F(i, j) = maxk=i. . j-1 F(i, k) + F(k+1, j) 0 0 0 How to fill in the DP matrix? 0 0 j–i=N-1 0 0 0
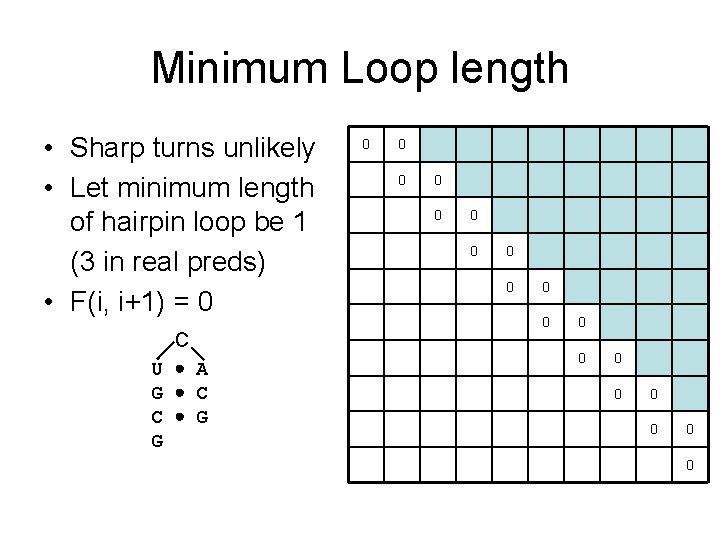
Minimum Loop length • Sharp turns unlikely • Let minimum length of hairpin loop be 1 (3 in real preds) • F(i, i+1) = 0 C U A G C C G G 0 0 0 0 0

Algorithm Initialization: F(i, i) = 0; F(i, i+1) = 0; for i = 1 to N-1 Iteration: For L = 1 to N-1 For i = 1 to N – l j = min(i + L, N) F(i, j) = max F(i+1, j -1) + s(xi, xj) max{ i k < j } F(i, k) + F(k+1, j) Termination: Best score is given by F(1, N) (For trace back, refer to the Durbin book)

Complexity For L = 1 to N-1 For i = 1 to N – l j = min(i + L, N) F(i+1, j -1) + s(xi, xj) F(i, j) = max{ i k < j } F(i, k) + F(k+1, j) • Time complexity: O(N 3) • Memory: O(N 2)

Example • RNA sequence: GGGAAAUCC • Only count # of base-pairs – A-U = 1 – G-C = 1 – G-U = 1 • Minimum hairpin loop length = 1
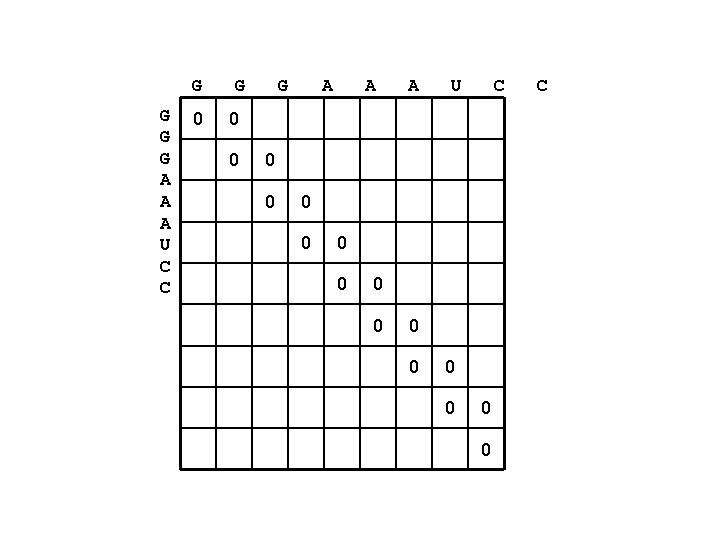
G G A A A U C C 0 G G A A A U C 0 0 0 0 C
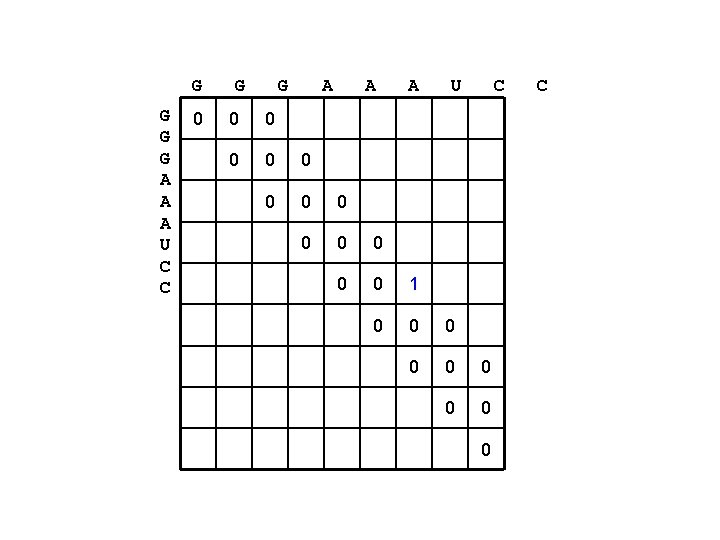
G G A A A U C C 0 G G A A A U C 0 0 0 0 1 0 0 0 0 0 C
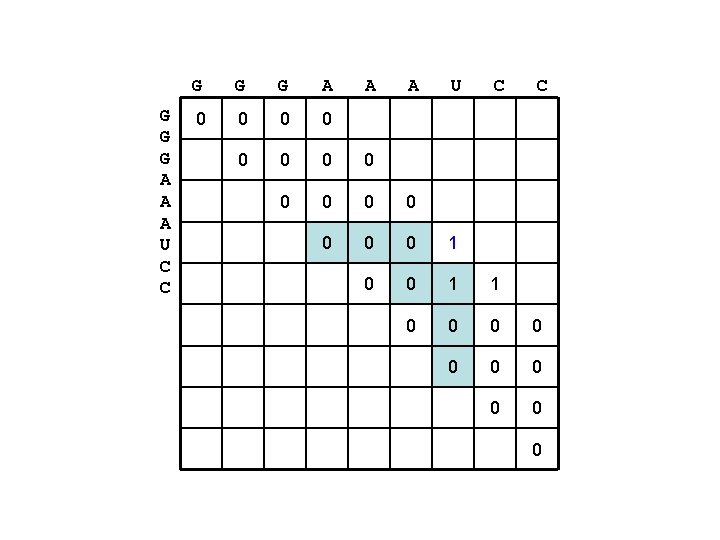
G G G A A A U C C 0 0 0 0 1 0 0 1 1 0 0 0 0 0
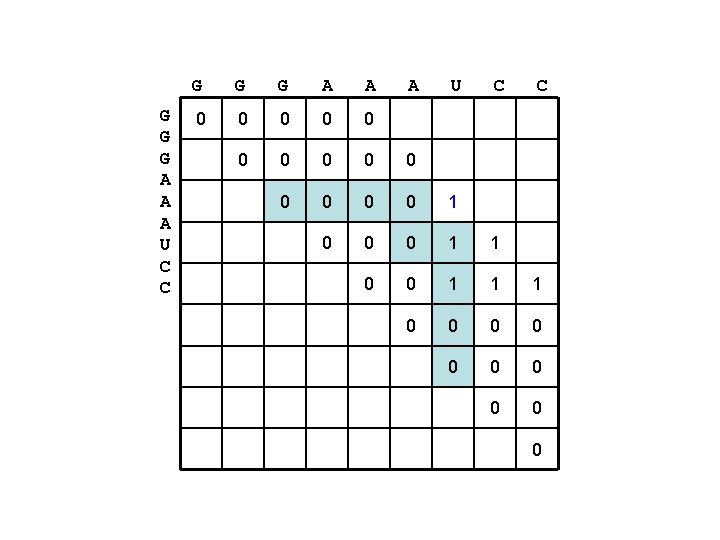
G G G A A A U C C 0 0 0 0 1 0 0 0 1 1 1 0 0 0 0 0
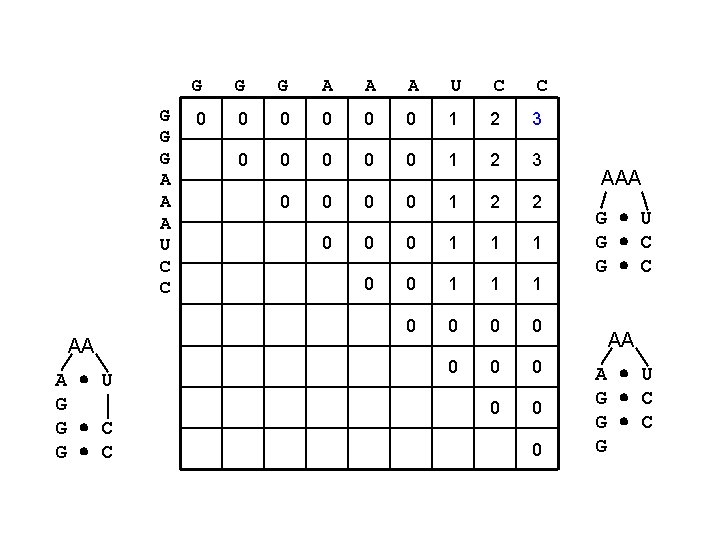
G G G A A A U C C AA A U G G C G G G A A A U C C 0 0 0 1 2 3 0 0 1 2 2 0 0 0 1 1 1 0 0 0 0 0 AAA G U G C AA A U G C G
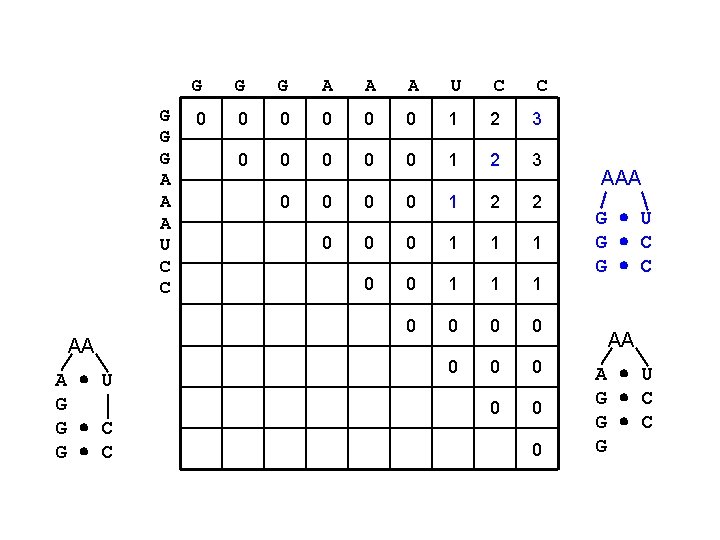
G G G A A A U C C AA A U G G C G G G A A A U C C 0 0 0 1 2 3 0 0 1 2 2 0 0 0 1 1 1 0 0 0 0 0 AAA G U G C AA A U G C G
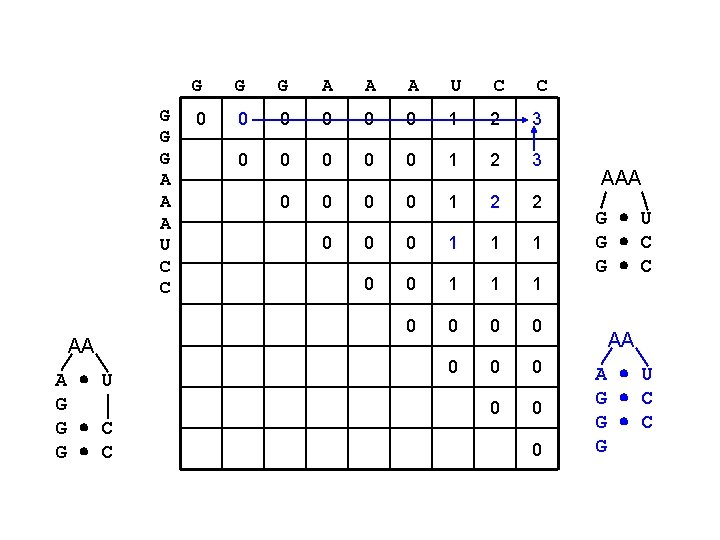
G G G A A A U C C AA A U G G C G G G A A A U C C 0 0 0 1 2 3 0 0 1 2 2 0 0 0 1 1 1 0 0 0 0 0 AAA G U G C AA A U G C G
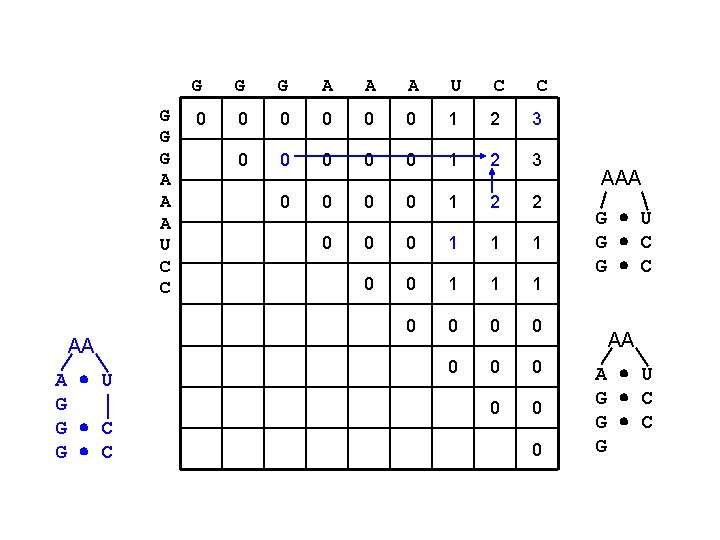
G G G A A A U C C AA A U G G C G G G A A A U C C 0 0 0 1 2 3 0 0 1 2 2 0 0 0 1 1 1 0 0 0 0 0 AAA G U G C AA A U G C G

Energy minimization For L = 1 to N-1 For i = 1 to N – l j = min(i + L, N); E(i+1, j -1) + e(xi, xj) E(i, j) = min{ i k < j } E(i, k) + E(k+1, j) e(xi, xj) represents the energy for xi base pair with xj • Energy are negative values. Therefore minimization rather than maximize. • More complex energy rules: energy depends on neighboring bases
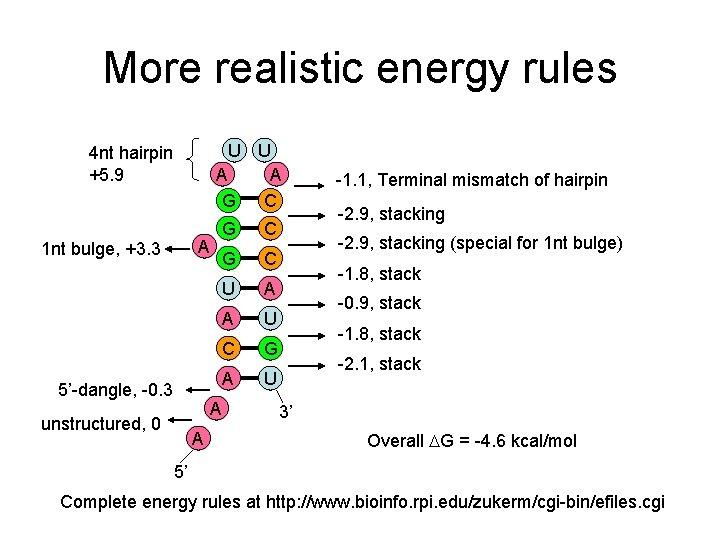
More realistic energy rules U U A A G C 4 nt hairpin +5. 9 A 1 nt bulge, +3. 3 5’-dangle, -0. 3 G C U A A U C G A U A unstructured, 0 A -1. 1, Terminal mismatch of hairpin -2. 9, stacking (special for 1 nt bulge) -1. 8, stack -0. 9, stack -1. 8, stack -2. 1, stack 3’ Overall G = -4. 6 kcal/mol 5’ Complete energy rules at http: //www. bioinfo. rpi. edu/zukerm/cgi-bin/efiles. cgi

The Zuker algorithm – main ideas 1. Instead of base pairs, pairs of base pairs (more accurate) Separate score for bulges Separate score for different-size & composition of loops Separate score for interactions between stem & beginning of loop Use additional matrix to remember current state. e. g, to model stacking energy: 2. 3. 4. 5. • • • W(i, j): energy of the best structure on i, j V(i, j): energy of the best structure on i, j given that i, j are paired Similar to affine-gap alignment.

Two popular implementations • mfold (Zuker) http: //mfold. bioinfo. rpi. edu/ • RNAfold in the Vienna package (Hofacker) http: //www. tbi. univie. ac. at/~ivo/RNA/

Accuracy • 50 -70% for sequences up to 300 nt • Not perfect, but useful • Possible reasons: – Energy rule not perfect: 5 -10% error – Many alternative structures within this error range – Alternative structure do exist – Structure may change in presence of other molecules
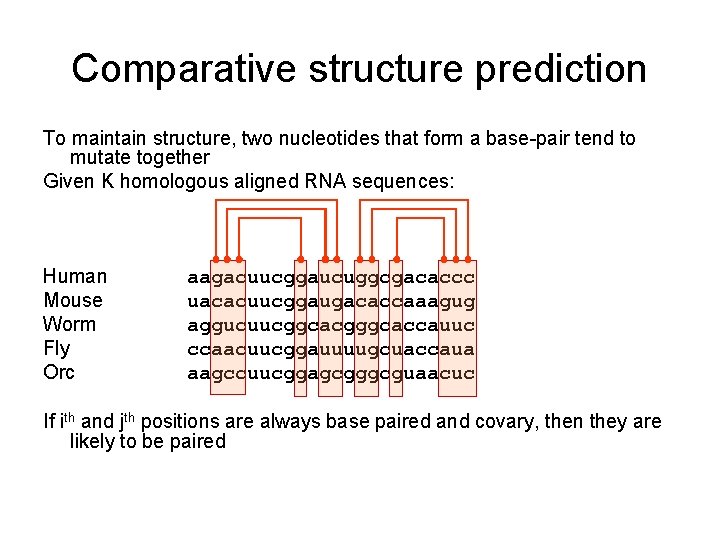
Comparative structure prediction To maintain structure, two nucleotides that form a base-pair tend to mutate together Given K homologous aligned RNA sequences: Human Mouse Worm Fly Orc aagacuucggaucuggcgacaccc uacacuucggaugacaccaaagug aggucuucggcacgggcaccauuc ccaacuucggauuuugcuaccaua aagccuucggagcgggcguaacuc If ith and jth positions are always base paired and covary, then they are likely to be paired

Mutual information fab(i, j): Prob for a, b to be in positions i, j fa (i): Prob for a to be in positions i aagacuucggaucuggcgacaccc uacacuucggaugacaccaaagug aggucuucggcacgggcaccauuc ccaacuucggauuuugcuaccaua aagccuucggagcgggcguaacuc fgc(3, 13) = 3/5 fcg(3, 13) = 1/5 fau(3, 13) = 1/5 fg(3) = 3/5 fc(3) = 1/5 fa(3) = 1/5 fc(13) = 3/5 fg(13) = 1/5 fu(13) = 1/5

Mutual information • • Also called covariance score M is high if base a in position i always follow by base b in position j – Does not require a to base-pair with b – Advantage: can detect non-canonical base-pairs • However, M = 0 if no mutation at all, even if perfect base-pairs aagacuucggaucuggcgacaccc uacacuucggaugacaccaaagug aggucuucggcacgggcaccauuc ccaacuucggauuuugcuaccaua aagccuucggagcgggcguaacuc One way to get around is to combine covariance and energy scores
Comparative structure prediction • • Given a multiple alignment, can infer structure that maximizes the sum of mutual information, by DP However, alignment is hard, since structure often more important than sequence

Comparative structure prediction In practice: 1. Get multiple alignment 2. Find covarying bases – deduce structure 3. Improve multiple alignment (by hand) 4. Go to 2 A manual EM process!!

Comparative structure prediction • Align then fold • Fold then align • Align and fold
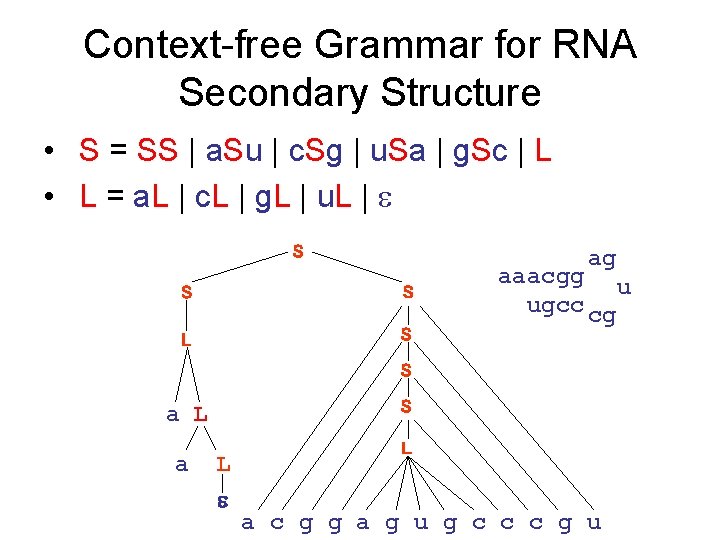
Context-free Grammar for RNA Secondary Structure • S = SS | a. Su | c. Sg | u. Sa | g. Sc | L • L = a. L | c. L | g. L | u. L | S ag S S L S aaacgg u ugcc cg S S a L L a c g g a g u g c c c g u

Stochastic Context-free Grammar (SCFG) • Probabilistic context-free grammar • Probabilities can be converted into weights • CFG vs SCFG is similar to RG vs HMM • • • 0 S =2 SS S =3 a. Su | u. Sa S(i, j) = max S =1 c. Sg | g. Sc L(i, j) = 0 S = u. Sg | g. Su 0 S=L 0 L = a. L | c. L | g. L | u. L | e(xi, xj) + S(i+1, j-1) L(i, j) maxk (S(i, k) + S(k+1, j))

SCFG Decoding • Decoding: given a grammar (SCFG/HMM) and a sequence, find the best parse (highest probability or score) – Cocke-Younger-Kasami (CYK) algorithm (analogous to Viterbi in HMM) – The Nussinov and Zuker algorithms are essentially special cases of CYK – CYK and SCFG are also used in other domains (NLP, Compiler, etc).

SCFG Evaluation • Given a sequence and a SCFG model – Estimate P(seq is generated by model), summing over all possible paths (analogous to forward-algorithm in HMM) • Inside-outside algorithm – Analogous to forward-background – Inside: bottom-up parsing (P(xi. . xj)) – Outside: top-down parsing (P(x 1. . xi-1 xj+1. . x. N)) • Can calculate base-paring probability – Analogous to posterior decoding – Essentially the same idea implemented in the Vienna RNAfold package

SCFG Learning • Covariance model: similar to profile HMMs – Given a set of sequences with common structures, simultaneously learn SCFG parameters and optimally parse sequences into states – EM on SCFG – Inside-outside algorithm – Efficiency is a bottleneck • Have been successfully applied to predict t. RNA genes and structures – t. RNAScan

Summary: SCFG and HMM algorithms GOAL HMM algorithm SCFG algorithm Optimal parse Viterbi CYK Estimation Forward Backward Inside Outside Learning EM: Fw/Bck EM: Ins/Outs Memory Complexity Time Complexity O(N K) O(N K 2) O(N 2 K) O(N 3 K 3) Where K: # of states in the HMM # of nonterminal symbols in the SCFG

Open research problems • nc. RNA gene prediction • nc. RNA regulatory networks • Structure prediction – Secondary, including pseudoknots – Tertiary • Structural comparison tools – Structural alignment • Structure search tools – “RNA-BLAST” • Structural motif finding – “RNA-MEME”
- Slides: 54