0 Managing your NGS Data with em BASE

0 Managing your NGS Data with em. BASE • Sample Annotation • NGS Assays • Data sets grouping in experiments and projects • Programmatic access • Adding, Deleting and Archiving files
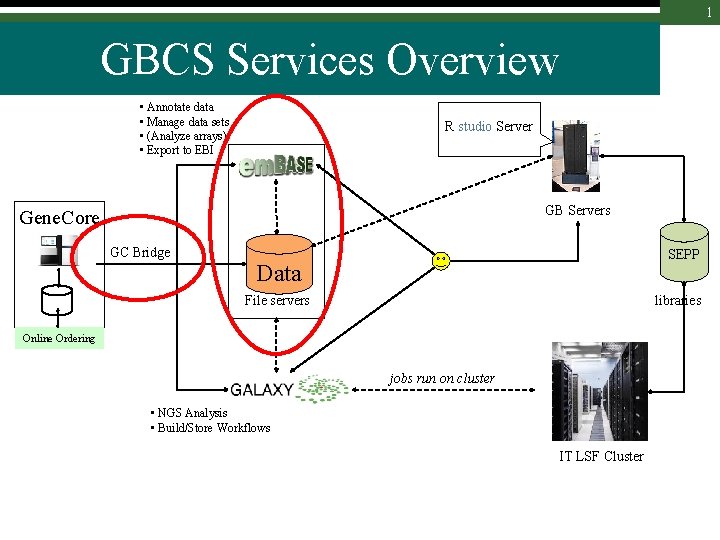
1 GBCS Services Overview • Annotate data • Manage data sets • (Analyze arrays) • Export to EBI R studio Server GB Servers Gene. Core GC Bridge SEPP Data File servers libraries Online Ordering jobs run on cluster • NGS Analysis • Build/Store Workflows IT LSF Cluster

2 NGS data Library 1. em. BASE is a database, with a web front-end, storing all metadata about your data files (e. g. fastq) 2. Your data files remains on your group fileserver in your “NGS data library” and are accessible directly Data web app / My. SQL File Server NGS Data @ GB : : Data Management : : em. BASE
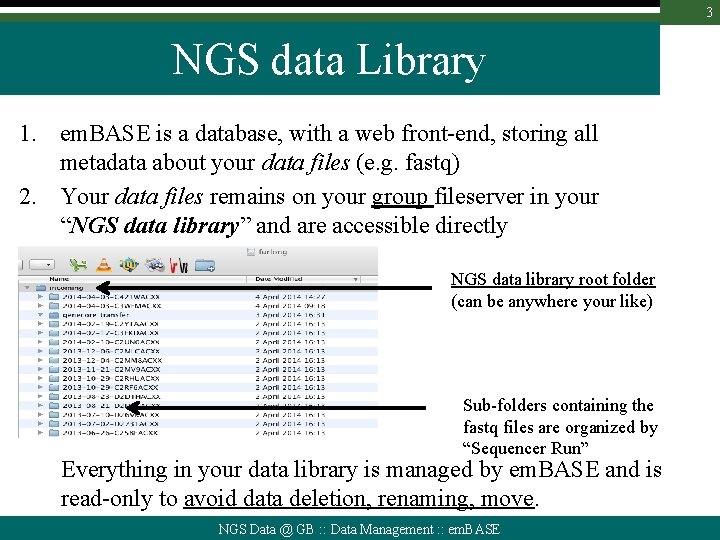
3 NGS data Library 1. em. BASE is a database, with a web front-end, storing all metadata about your data files (e. g. fastq) 2. Your data files remains on your group fileserver in your “NGS data library” and are accessible directly NGS data library root folder (can be anywhere your like) Sub-folders containing the fastq files are organized by “Sequencer Run” Everything in your data library is managed by em. BASE and is read-only to avoid data deletion, renaming, move. NGS Data @ GB : : Data Management : : em. BASE

4 Part I : Fundamental concepts How is the data stored or represented in em. BASE ?
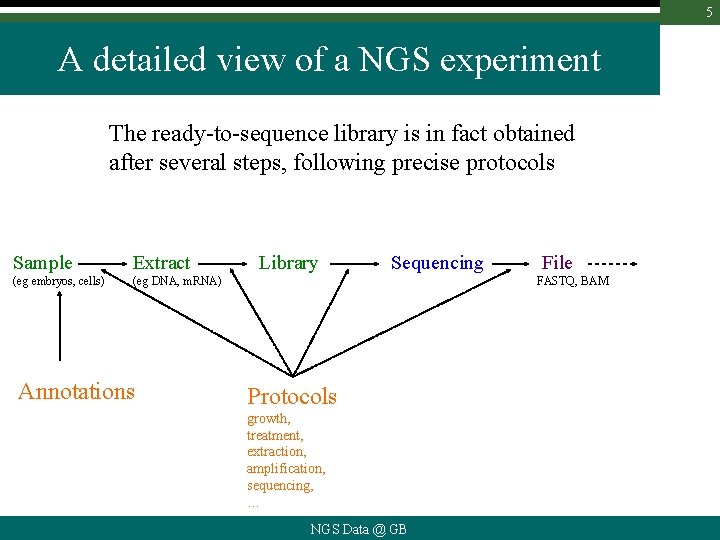
5 A detailed view of a NGS experiment The ready-to-sequence library is in fact obtained after several steps, following precise protocols Sample Extract (eg embryos, cells) (eg DNA, m. RNA) Annotations Library Sequencing File FASTQ, BAM Protocols growth, treatment, extraction, amplification, sequencing, … NGS Data @ GB
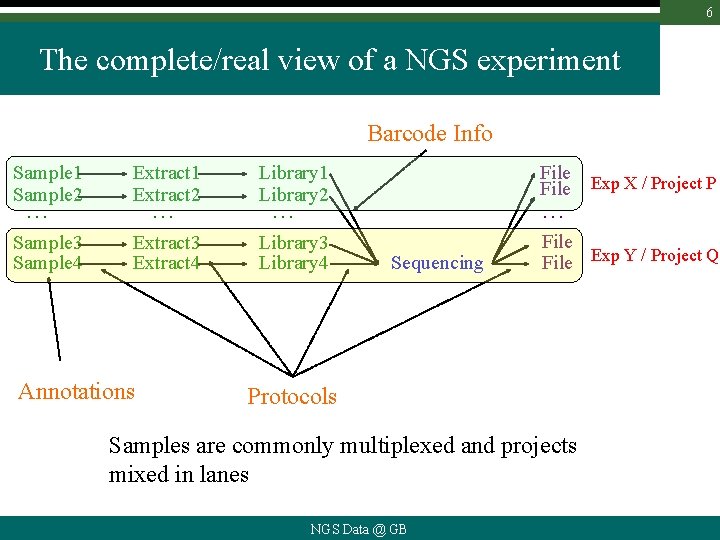
6 The complete/real view of a NGS experiment Barcode Info Sample 1 Sample 2 Extract 1 Extract 2 Library 1 Library 2 File Exp X / Project P Sample 3 Sample 4 Extract 3 Extract 4 Library 3 Library 4 File Exp Y / Project Q … … Annotations … … Sequencing Protocols Samples are commonly multiplexed and projects mixed in lanes NGS Data @ GB

7 Publication of your NGS experiment requires all this information Barcode Info Sample 1 Sample 2 Extract 1 Extract 2 Library 1 Library 2 File Sample 3 Sample 4 Extract 3 Extract 4 Library 3 Library 4 File (eg embryos, cells) (eg DNA, m. RNA) … … Annotations … … Sequencing FASTQ, BAM Protocols Publish e. g. EBI Specific format eg MAGE-TAB Data management, annotations and publication is the reason of em. BASE models all these different “items” NGS Data @ GB
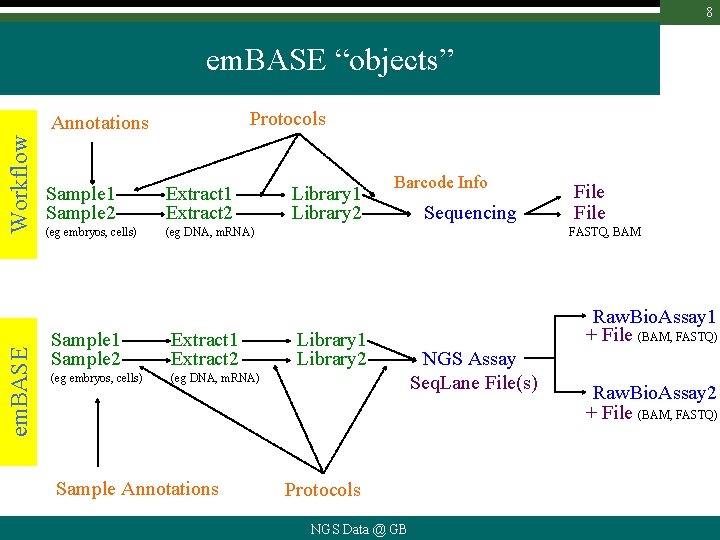
8 em. BASE “objects” Protocols em. BASE Workflow Annotations Sample 1 Sample 2 Extract 1 Extract 2 (eg embryos, cells) (eg DNA, m. RNA) Sample Annotations Library 1 Library 2 Barcode Info Sequencing File FASTQ, BAM Library 1 Library 2 Protocols NGS Data @ GB Raw. Bio. Assay 1 + File (BAM, FASTQ) NGS Assay Seq. Lane File(s) Raw. Bio. Assay 2 + File (BAM, FASTQ)
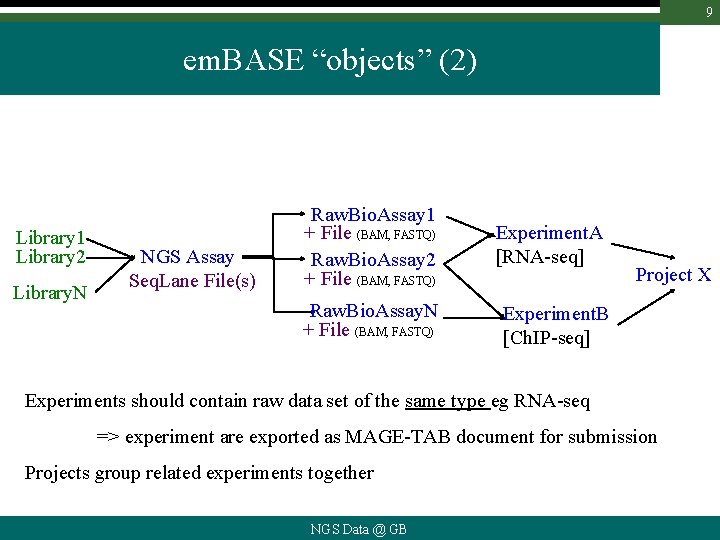
9 em. BASE “objects” (2) Library 1 Library 2 Library. N NGS Assay Seq. Lane File(s) Raw. Bio. Assay 1 + File (BAM, FASTQ) Raw. Bio. Assay 2 + File (BAM, FASTQ) Raw. Bio. Assay. N + File (BAM, FASTQ) Experiment. A [RNA-seq] Project X Experiment. B [Ch. IP-seq] Experiments should contain raw data set of the same type eg RNA-seq => experiment are exported as MAGE-TAB document for submission Projects group related experiments together NGS Data @ GB
10 GBCS web site : primary info. source All about em. BASE All online tutorials and documents

11 All Tutorials and more ~ Whole today tutorial is available in here

12 First steps in em. BASE In this section 1. 2. 3. 4. Login Menus : basic / expert mode Change your defaults, reset pwd Adapt GUI displays Let’s see this for real

13 The different em. BASE Sections => Go to http: //gbcs. embl. de/base and log in Sample, Extract, NGS Library Protocols Annotations NGS Assays (Lane) Raw data sets <=> Experiment linking Experiments, Projects Experiment export (EBI submission) Archiving Account and Default Settings NGS Data @ GB
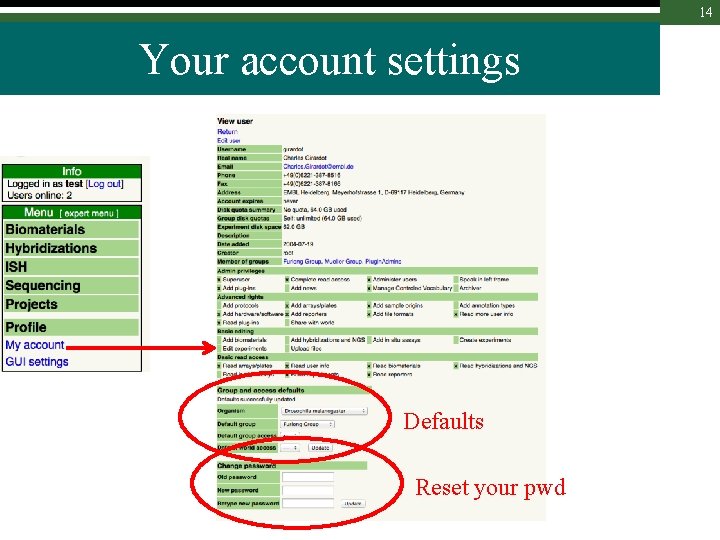
14 Your account settings Defaults Reset your pwd
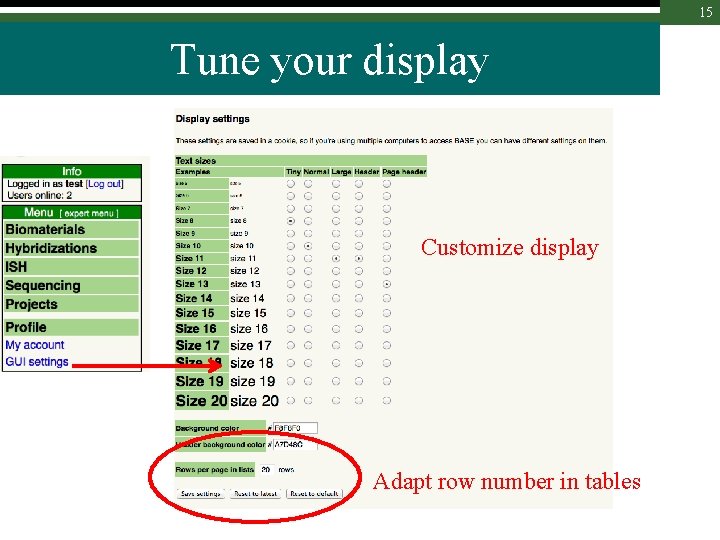
15 Tune your display Customize display Adapt row number in tables

16 Managing your “Biomaterial” In this section 1. 2. 3. 4. Using the search interface to narrow down interesting samples Customizing list pages (item number, columns) Change samples property in batch Sample Annotation : • individually • in batch online • using a file 5. Add protocols Let’s see this for real
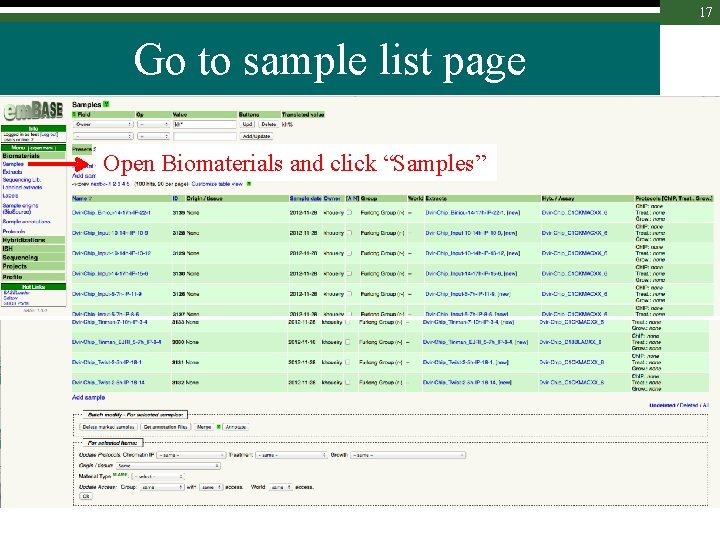
17 Go to sample list page Open Biomaterials and click “Samples”
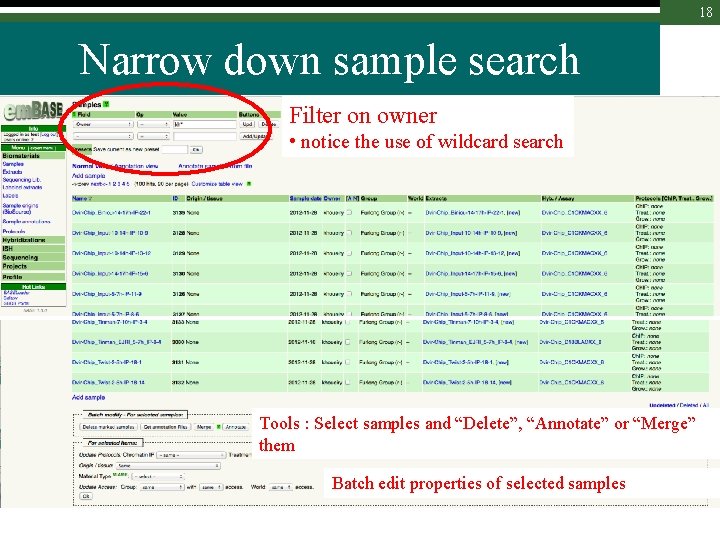
18 Narrow down sample search Filter on owner • notice the use of wildcard search Tools : Select samples and “Delete”, “Annotate” or “Merge” them Batch edit properties of selected samples
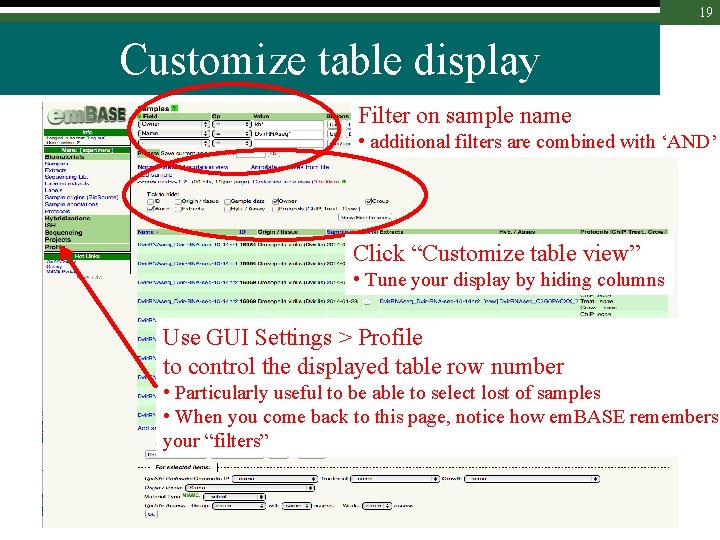
19 Customize table display Filter on sample name • additional filters are combined with ‘AND’ Click “Customize table view” • Tune your display by hiding columns Use GUI Settings > Profile to control the displayed table row number • Particularly useful to be able to select lost of samples • When you come back to this page, notice how em. BASE remembers your “filters”
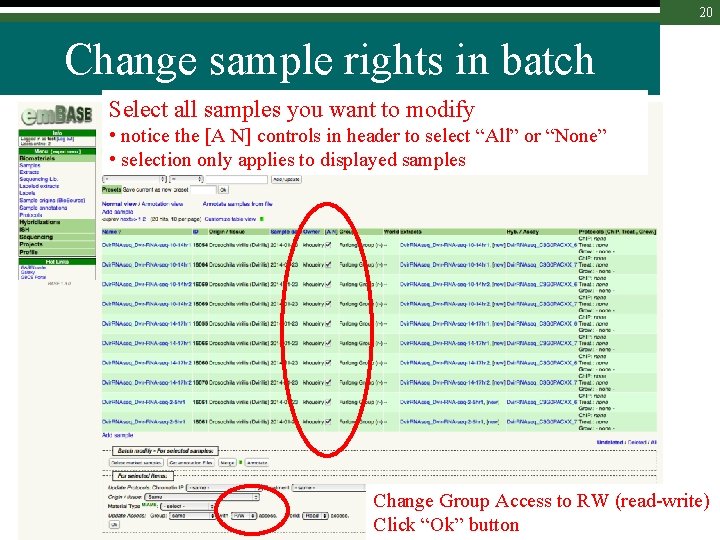
20 Change sample rights in batch Select all samples you want to modify • notice the [A N] controls in header to select “All” or “None” • selection only applies to displayed samples Change Group Access to RW (read-write) Click “Ok” button
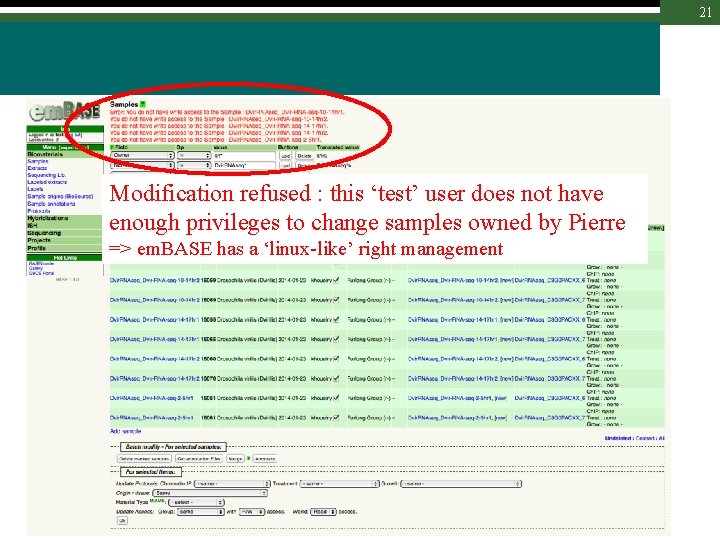
21 Modification refused : this ‘test’ user does not have enough privileges to change samples owned by Pierre => em. BASE has a ‘linux-like’ right management

22 In the context of the training I now log in as a privileged user to be the owner of the data we are playing with (e. g. you and your data)
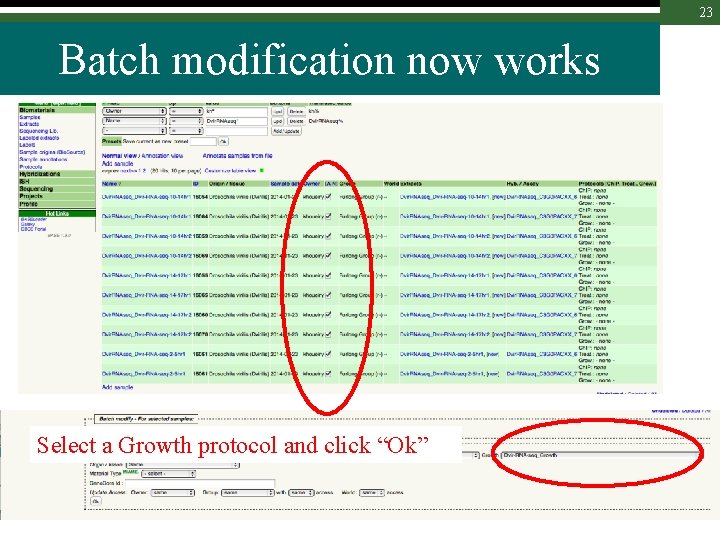
23 Batch modification now works Select a Growth protocol and click “Ok”
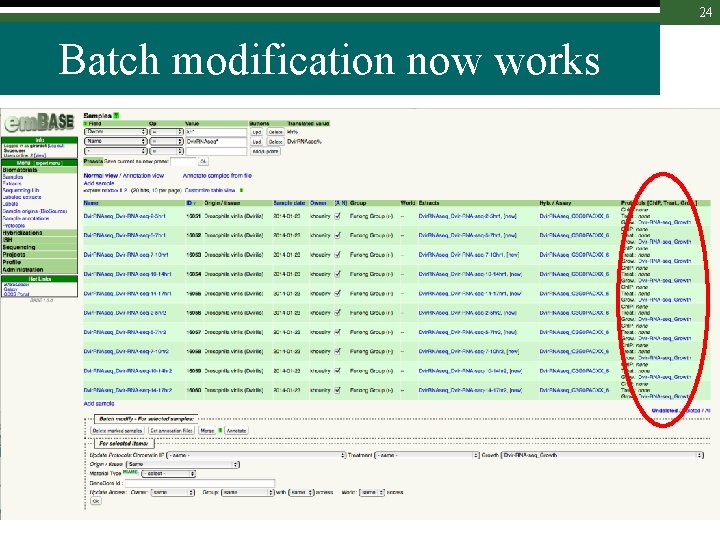
24 Batch modification now works

25 Setting Protocols and properties must be done on : • Sample • Extracts • NGS Libraries We won’t demonstrate this ; it is exactly like for samples
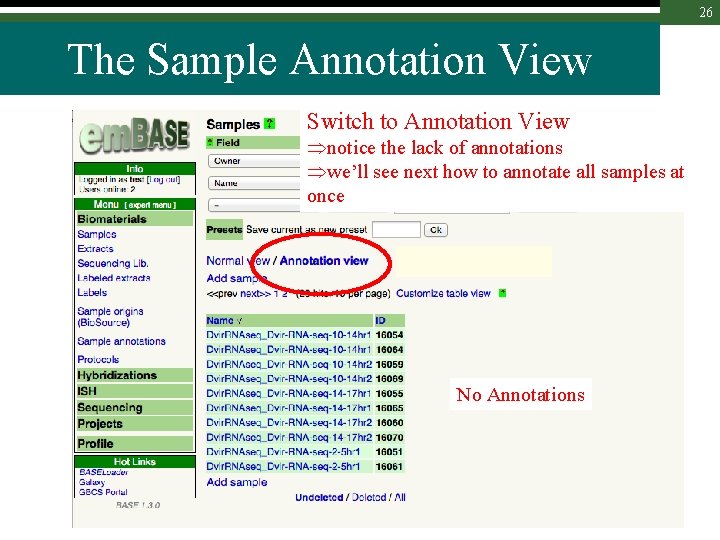
26 The Sample Annotation View Switch to Annotation View notice the lack of annotations we’ll see next how to annotate all samples at once No Annotations
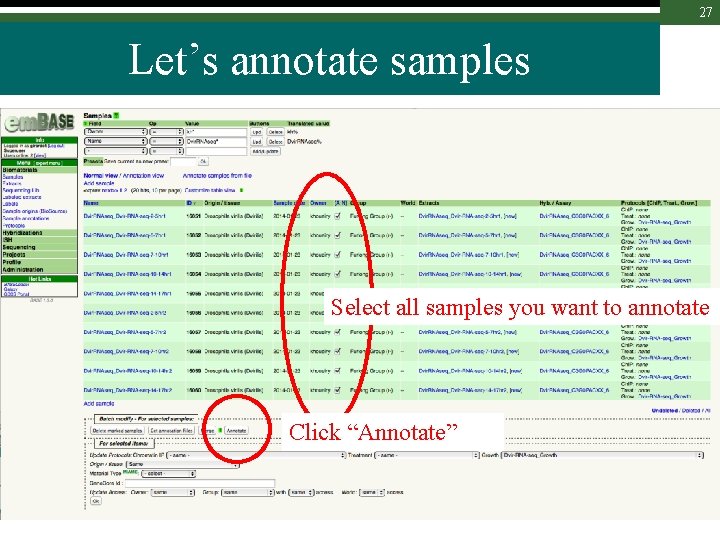
27 Let’s annotate samples Select all samples you want to annotate Click “Annotate”
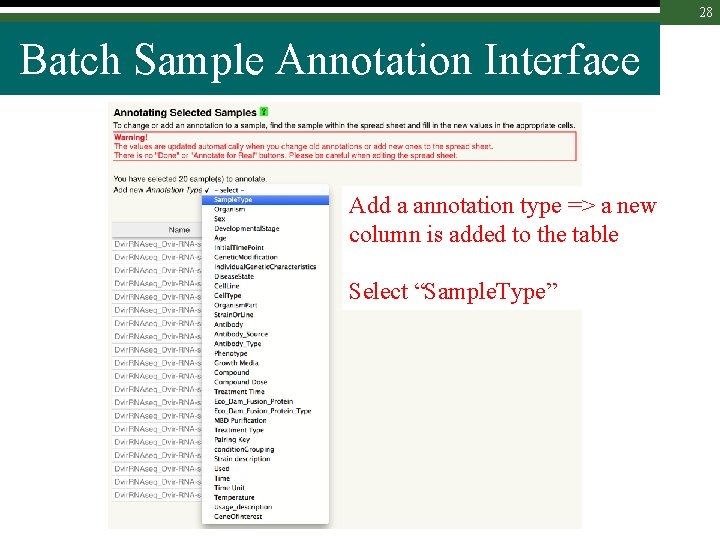
28 Batch Sample Annotation Interface Add a annotation type => a new column is added to the table Select “Sample. Type”
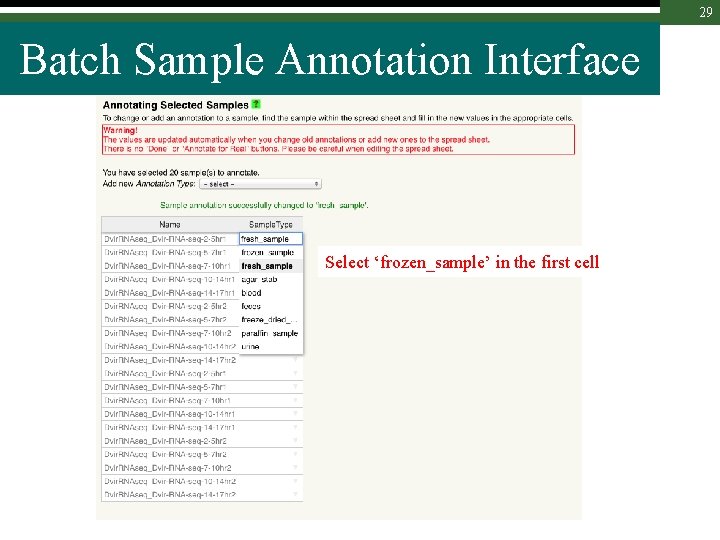
29 Batch Sample Annotation Interface Select ‘frozen_sample’ in the first cell
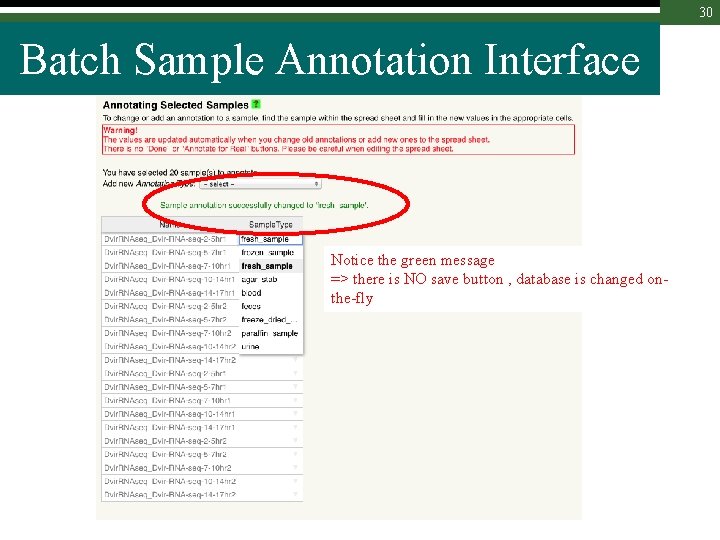
30 Batch Sample Annotation Interface Notice the green message => there is NO save button , database is changed onthe-fly
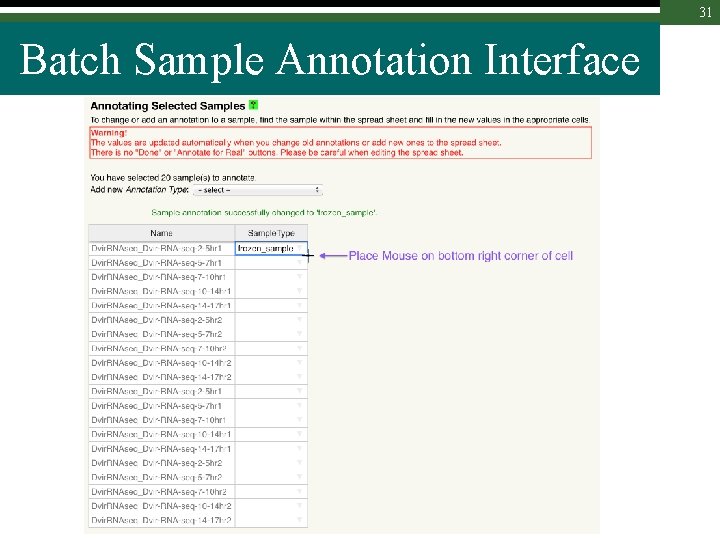
31 Batch Sample Annotation Interface
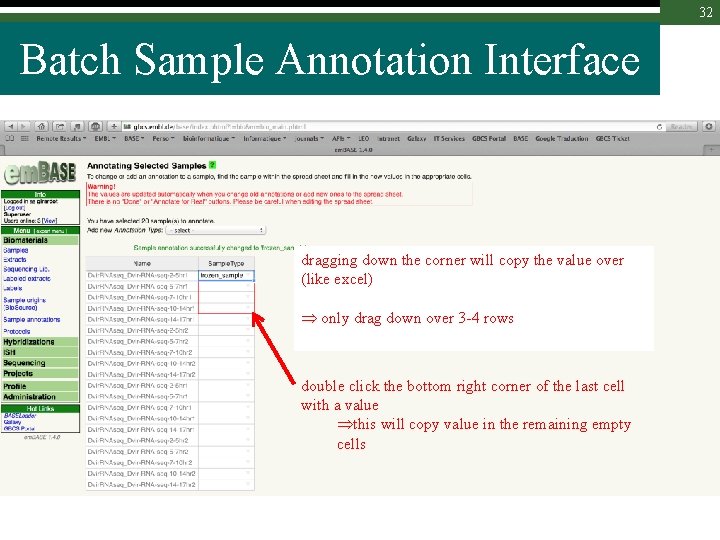
32 Batch Sample Annotation Interface dragging down the corner will copy the value over (like excel) only drag down over 3 -4 rows double click the bottom right corner of the last cell with a value this will copy value in the remaining empty cells
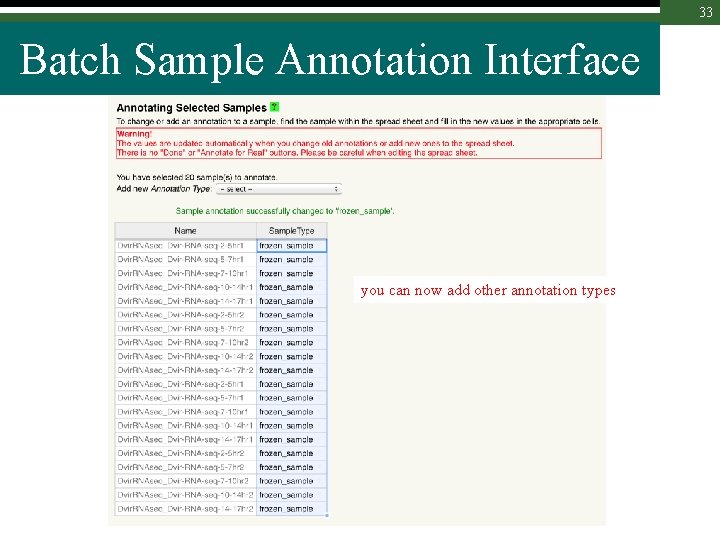
33 Batch Sample Annotation Interface you can now add other annotation types
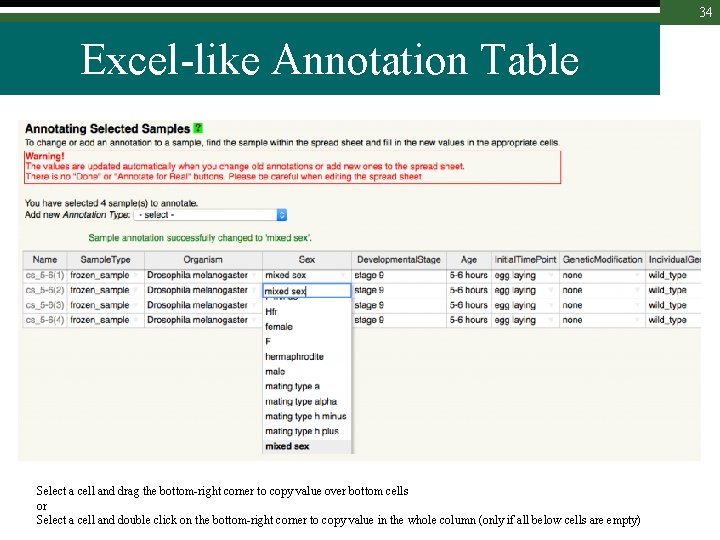
34 Excel-like Annotation Table Select a cell and drag the bottom-right corner to copy value over bottom cells or Select a cell and double click on the bottom-right corner to copy value in the whole column (only if all below cells are empty)

35 The next slides show to batch annotate samples using a file you created in excel

36 • COPY SLIDES IN

37 Managing your “NGS Assay” and “RBA” Let’s see this for real 1. Understanding NGS Assay content • multiplexed libraries • QC flag 2. Raw. Bio. Assay (aka RBA) <=> data file relationship • QC flag 3. Understanding how files are stored in your NGS Library 4. File Locking/Unlocking concept 5. Lane and RBA File deletion philosophy 6. Getting and Adding RBA files from the command line em. BASE: : Storing Sequencing Assay
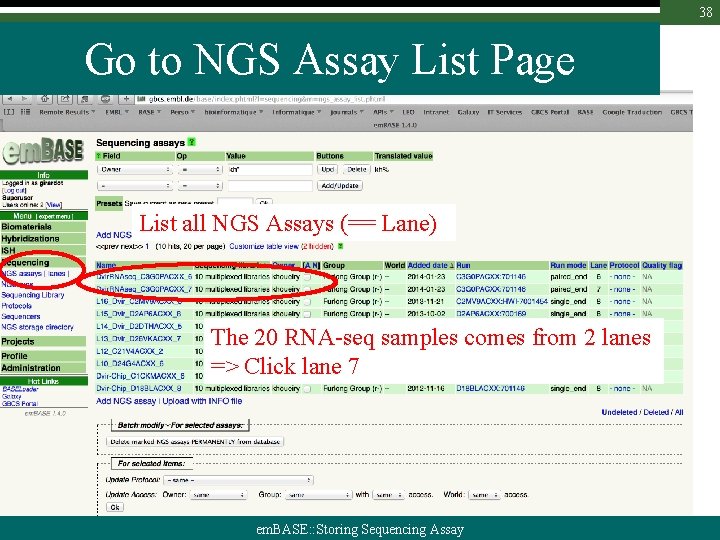
38 Go to NGS Assay List Page List all NGS Assays (== Lane) The 20 RNA-seq samples comes from 2 lanes => Click lane 7 em. BASE: : Storing Sequencing Assay

39 NGS Assay: Example of a multiplexed lane Assay (=Lane) info & rights Sequencing run info (notice link to the “run”) Lane File & Location Individual raw data sets & De-multiplexed Files Let’s zoom in raw data sets
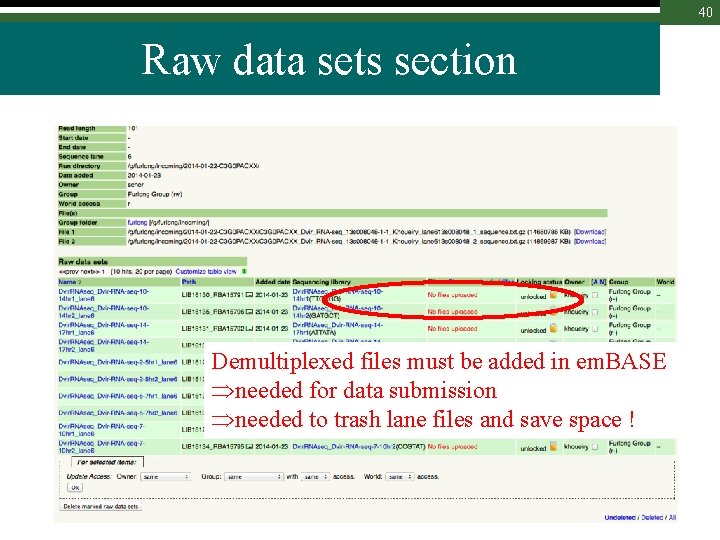
40 Raw data sets section Demultiplexed files must be added in em. BASE needed for data submission needed to trash lane files and save space !
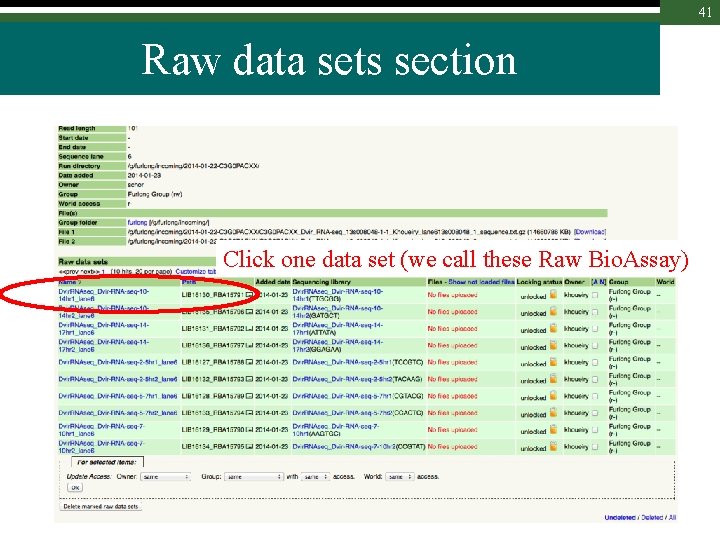
41 Raw data sets section Click one data set (we call these Raw Bio. Assay)
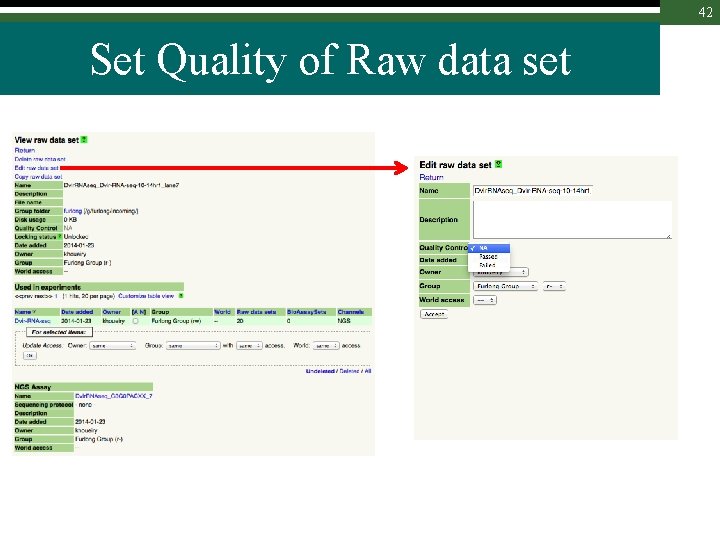
42 Set Quality of Raw data set
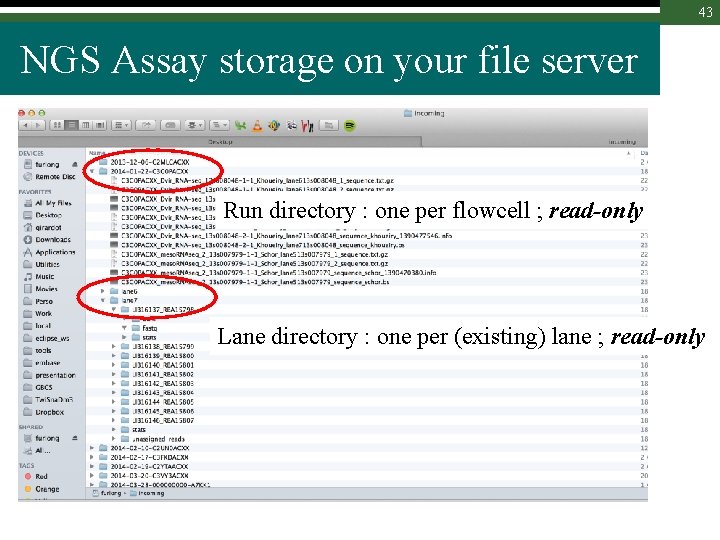
43 NGS Assay storage on your file server Run directory : one per flowcell ; read-only Lane directory : one per (existing) lane ; read-only
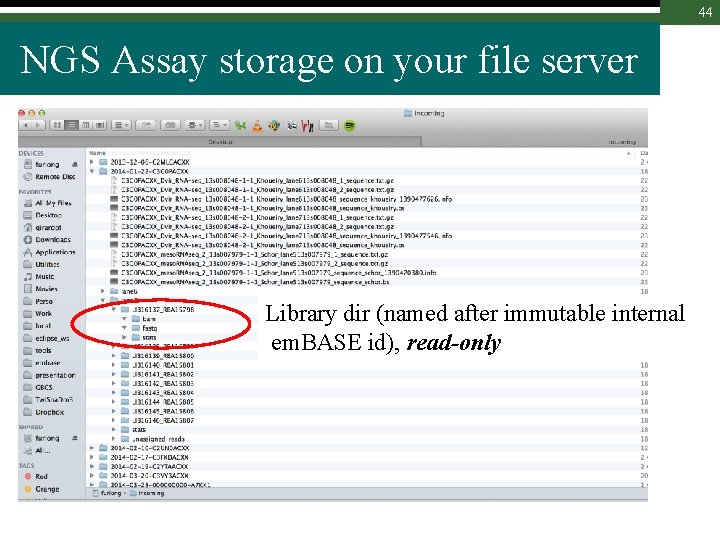
44 NGS Assay storage on your file server Library dir (named after immutable internal em. BASE id), read-only
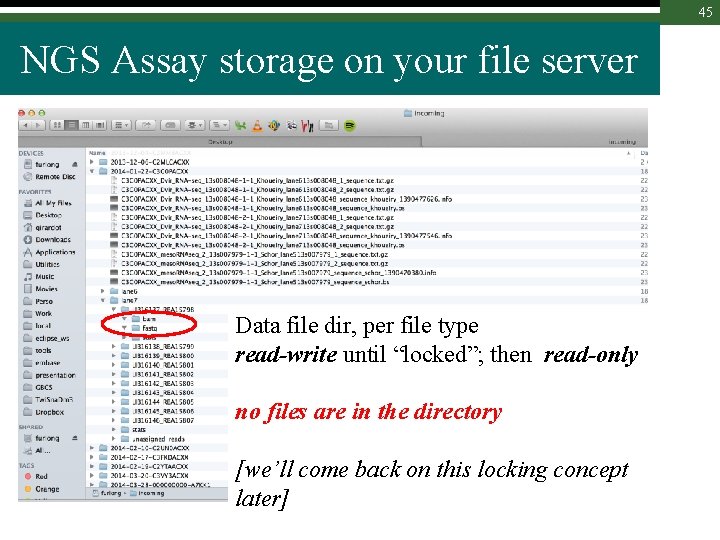
45 NGS Assay storage on your file server Data file dir, per file type read-write until “locked”; then read-only no files are in the directory [we’ll come back on this locking concept later]
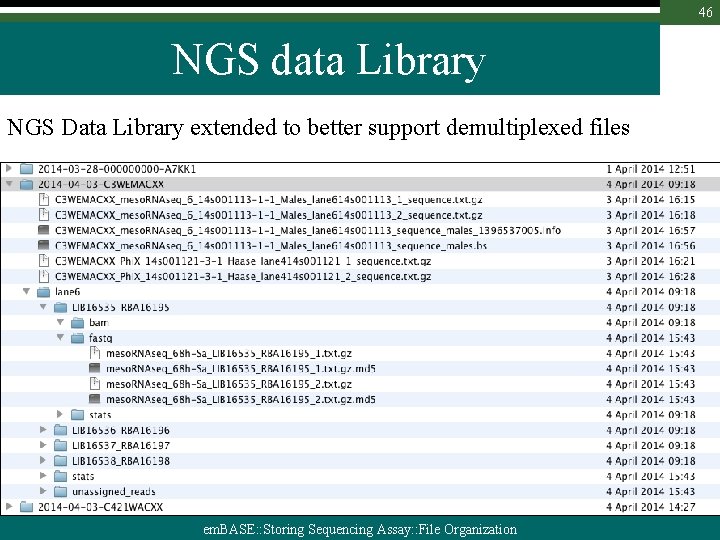
46 NGS data Library NGS Data Library extended to better support demultiplexed files em. BASE: : Storing Sequencing Assay: : File Organization

47 Adding demultiplexed files one can of course manually copy files in these directories (fastq or bam dir) or use our command line utilities Time to stop clicking ! log on spinoza [as galaxy] cd /g/furlong/project/21_dvir/fastq/RNA-seq …

48 • SCREENSHOOT OF LOCKING AND LANE FILE DELETION

49 Locking / Unlocking concept 1. Library file sub-directories are unlocked (writable for group) – you can work and replace files as you wish 2. At some point, files are ready and directories can be locked (only readable): 1. em. BASE starts, at this point, to track these files 2. em. BASE will allow lane file deletion when all its multiplexed libraries are locked. 3. Locking is operated via the web interface, on the whole lane or per library (case of shared lanes) em. BASE: : Storing Sequencing Assay: : File Organization

50 Organizing your RBA (files) into Experiments and Projects Let’s see this for real 1. 2. 3. 4. Experiments Adding/Removing Raw. Bio. Assay to/from Experiments Sync’ing with Galaxy Exporting an experiment to MAGE-TAB (for submission) • do not demonstrate, too long 5. Grouping Experiment in Project 6. Archiving Experiments/Project to tape • what happens ? (price, replacement file, duration) • how is the archiving info stored and accessible ? em. BASE: : Working With Data File Sets
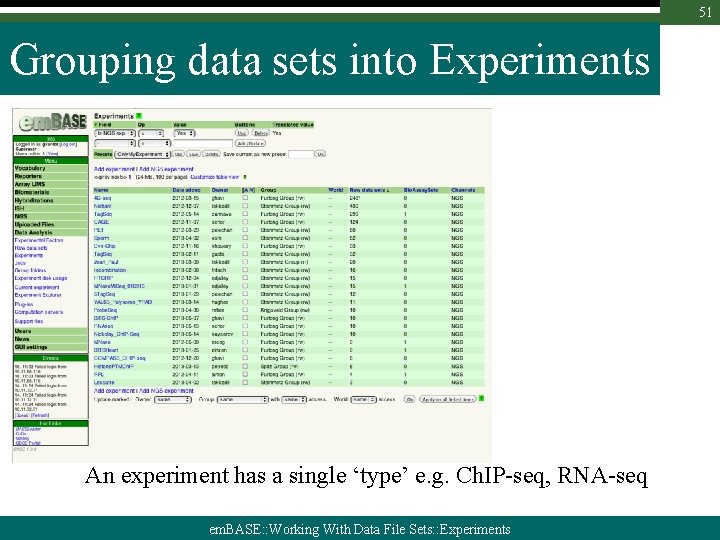
51 Grouping data sets into Experiments An experiment has a single ‘type’ e. g. Ch. IP-seq, RNA-seq em. BASE: : Working With Data File Sets: : Experiments
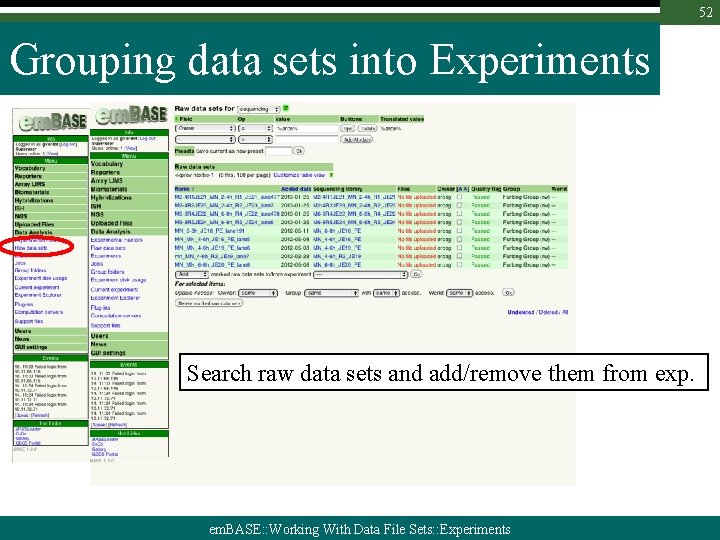
52 Grouping data sets into Experiments Search raw data sets and add/remove them from exp. em. BASE: : Working With Data File Sets: : Experiments

53 Other with Experiments • Galaxy Sync • MAGE-TAB Export em. BASE: : Working With Data File Sets: : Experiments

54 Regrouping Experiments in Projects • Show Project page em. BASE: : Working With Data File Sets: : Projects
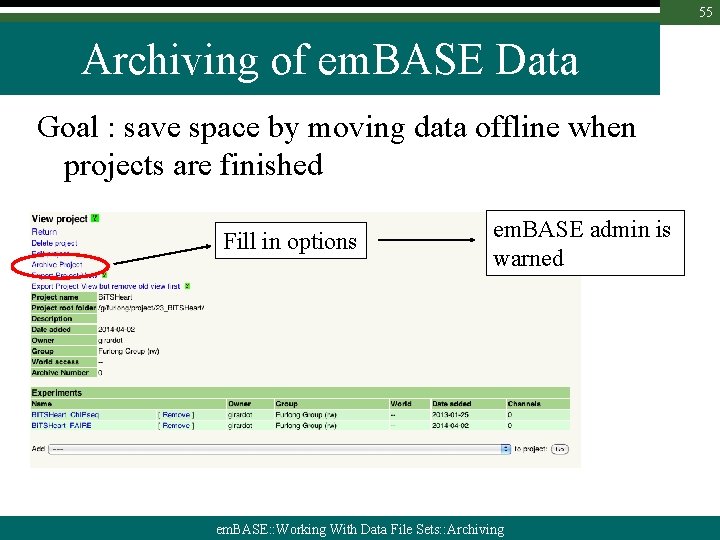
55 Archiving of em. BASE Data Goal : save space by moving data offline when projects are finished Fill in options em. BASE admin is warned em. BASE: : Working With Data File Sets: : Archiving

56 Archiving of em. BASE Data Please see online tutorial at http: //gbcs. embl. de/portal/tiki-index. php? page=archiving. Tutorial em. BASE: : Working With Data File Sets: : Archiving

57 Archiving of em. BASE Data What happens next ? • All data files connected to the experiments are exported • IT performs back up on tape • We delete ‘deletable’ files (concept of active experiment): – em. BASE knows which files can be deleted, which ones have been deleted and how to get them back, if needed – delete files are locally replaced with the a small file containing back up information • You can follow the archiving status in em. BASE This is a couple of clicks on your side but remember that you still pay the bill ! em. BASE: : Working With Data File Sets: : Archiving

58 More for the command line user em. BASE: : Working With Data File Sets: : Jem. BASEAPI

59 Working with this new structure 1. Use the command line em. BASE API to learn where files are or should be placed – These commands extracts all info from em. BASE for a lane, an experiment or a project 2. Use the command line em. BASE API to add RBA files (Fastq, BAM) in em. BASE Storage Documentation at : http: //gbcs. embl. de/tikiwiki/tiki-index. php? page=BASEJava. Cmd. Line. Utilities em. BASE: : Storing Sequencing Assay: : API
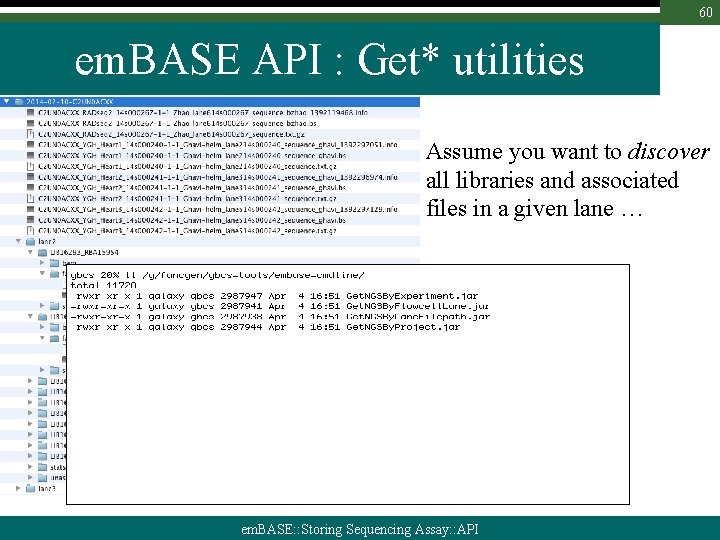
60 em. BASE API : Get* utilities Assume you want to discover all libraries and associated files in a given lane … em. BASE: : Storing Sequencing Assay: : API

61 A Get* Example • Available from anywhere • Logged in user used to authenticate in em. BASE • Rights apply the same way as in em. BASE: : Storing Sequencing Assay: : API
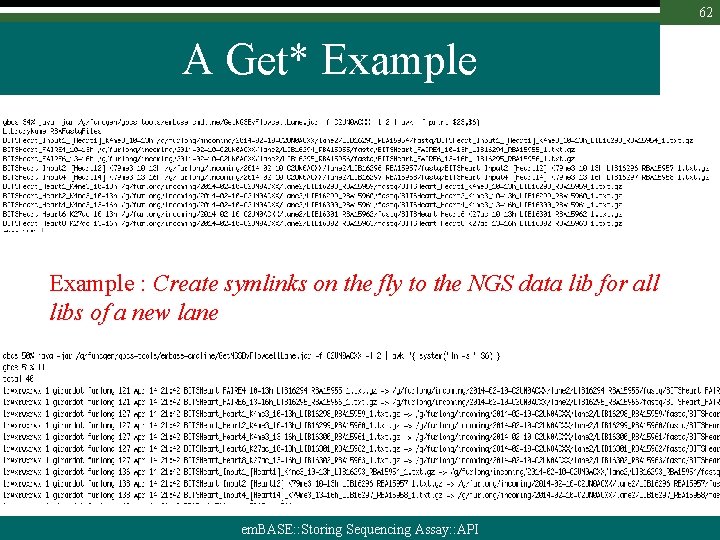
62 A Get* Example : Create symlinks on the fly to the NGS data lib for all libs of a new lane em. BASE: : Storing Sequencing Assay: : API

63 Loading your data with GCBridge Step-by-step tutorial at http: //gbcs. embl. de (then GC Bridge Menu) 1. The 3 common situations : a) single sample lane b) internally multiplexed lane c) demultiplexed lanes using Illumina index 2. Re-using existing samples 3. Handling Lane Mate in situations b) and c) 4. Coping with multiple projects lane i. e. situation c) only 5. Data demultiplexing i. e. situation c) only 6. Handling mistakes GCBridge: : Batch Data Upload

64 Thank you Joscha Sauer Shu-yi Su Aziz Moussa M Chaturvedi Alumni L-A Schmitt Nicolas Delhomme Leila Tlili Arnaud Huaulme IT Services Michael Wahlers Andres Lindau Gene. Core Jonathon Blake Juergen Zimmermann Markus Fritz Vladimir Benes All GB members Julien Gagneur (now LMU) Chenchen Zhu Lin Gen Simon Anders Tobias Rausch Eileen Furlong and Lars Steinmetz
- Slides: 65