Intro to NGS analysis Proficio course 2020 NGS











- Slides: 42

Intro to NGS analysis Proficio course 2020 NGS applications and data analysis Vladimir Teif

NGS techniques vs. NGS applications NGS techniques: how to sequence DNA (or RNA) (covered in lecture 1; funny recap in this video https: //www. youtube. com/watch? v=-7 GK 1 HXw. Ct. E) NGS applications: how to design experiments in order to answer a specific biological question

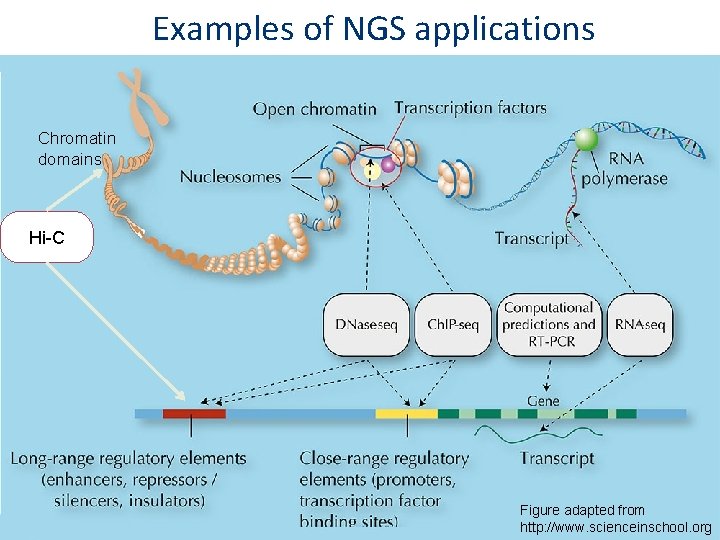
Examples of NGS applications Chromatin domains Hi-C Figure adapted from http: //www. scienceinschool. org

Types of NGS applications q RNA-seq, GRO-seq, CAGE, SAGE, CLIP-seq, Drop-seq Ø gene expression; non-coding RNA q Ch. IP-seq, MNase-seq, DNase-seq, ATAC-se, etc Ø protein binding; histone modifications Ø chromatin accessibility; nucleosome positioning q Bisulfite sequencing (DNA methylation) q Hi-C, 3 C, 4 C, Ch. IA-PET, etc (Chromatin loops) q Amplicon sequencing Ø targeted regions; philogenomics; metagenomics q Whole Genome Sequencing (WGS) Ø de-novo assembly (new species or new analyses) Curated bibliography of *seq methods (~100 methods) can be found at https: //liorpachter. wordpress. com/seq/

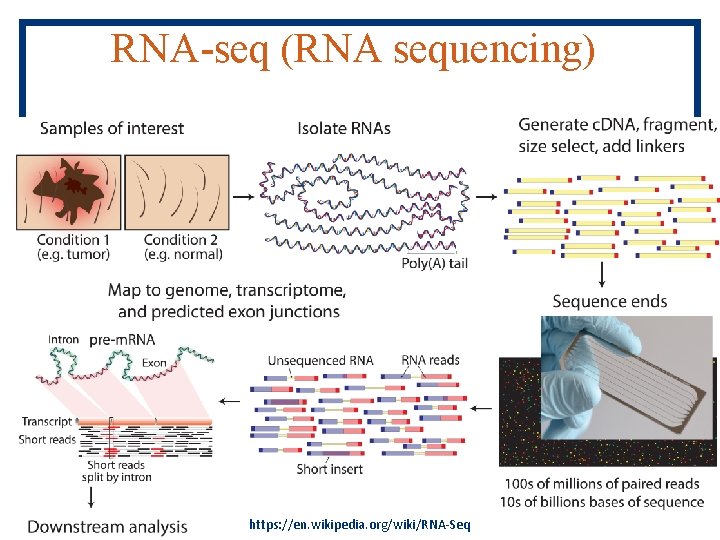
RNA-seq (RNA sequencing) https: //en. wikipedia. org/wiki/RNA-Seq

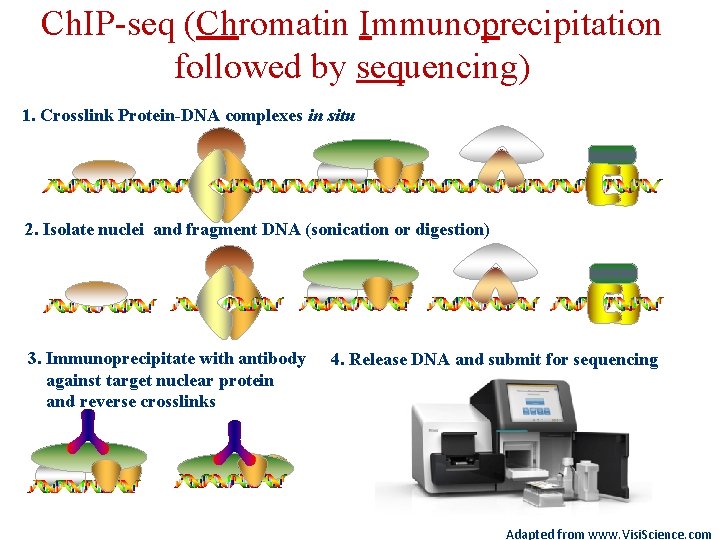
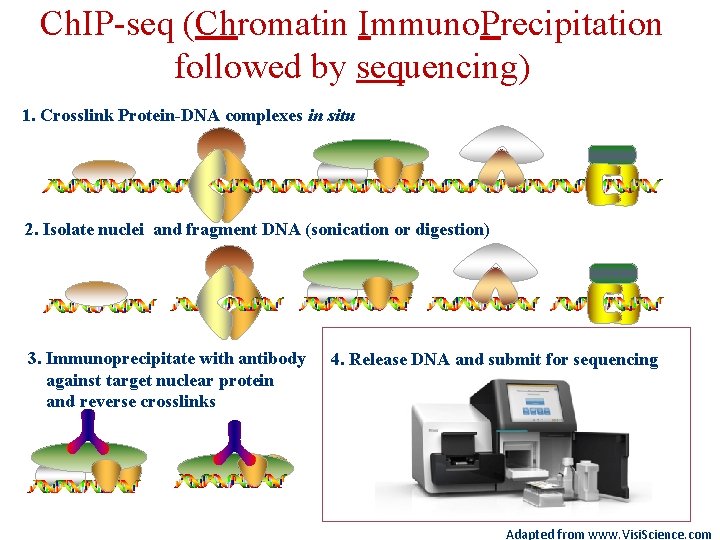
Ch. IP-seq (Chromatin Immunoprecipitation followed by sequencing) 1. Crosslink Protein-DNA complexes in situ 2. Isolate nuclei and fragment DNA (sonication or digestion) 3. Immunoprecipitate with antibody against target nuclear protein and reverse crosslinks 4. Release DNA and submit for sequencing Adapted from www. Visi. Science. com

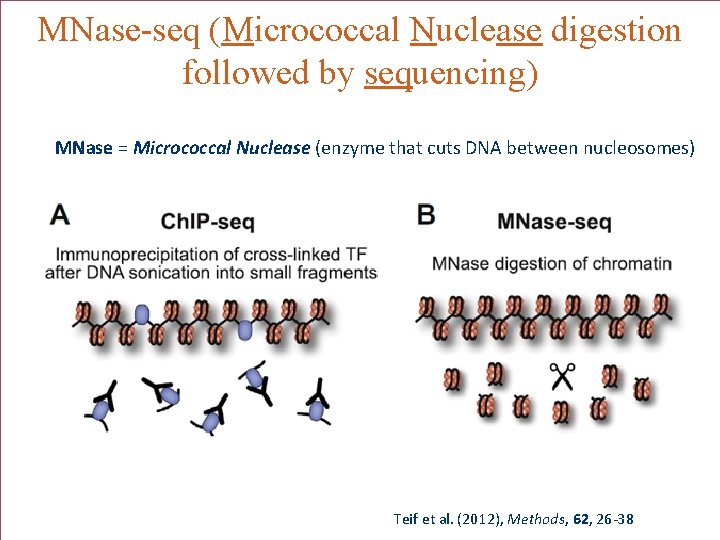
MNase-seq (Micrococcal Nuclease digestion followed by sequencing) MNase = Micrococcal Nuclease (enzyme that cuts DNA between nucleosomes) MM Teif et al. (2012), Methods, 62, 26 -38

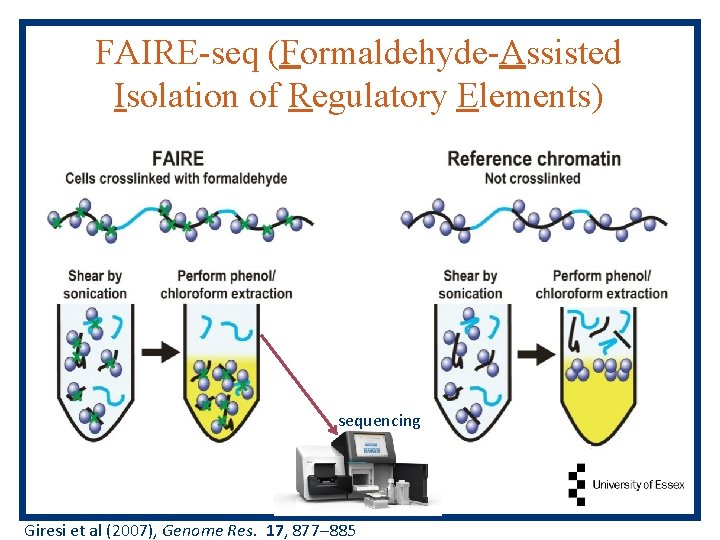
FAIRE-seq (Formaldehyde-Assisted Isolation of Regulatory Elements) sequencing Giresi et al (2007), Genome Res. 17, 877– 885

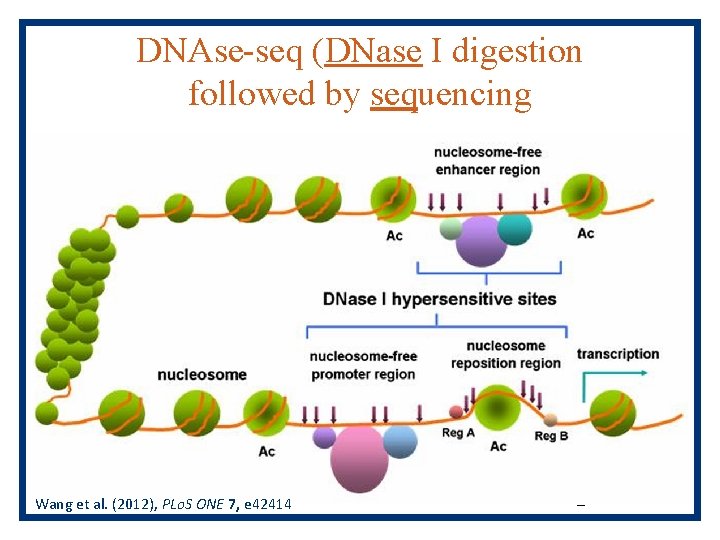
DNAse-seq (DNase I digestion followed by sequencing Wang et al. (2012), PLo. S ONE 7, e 42414

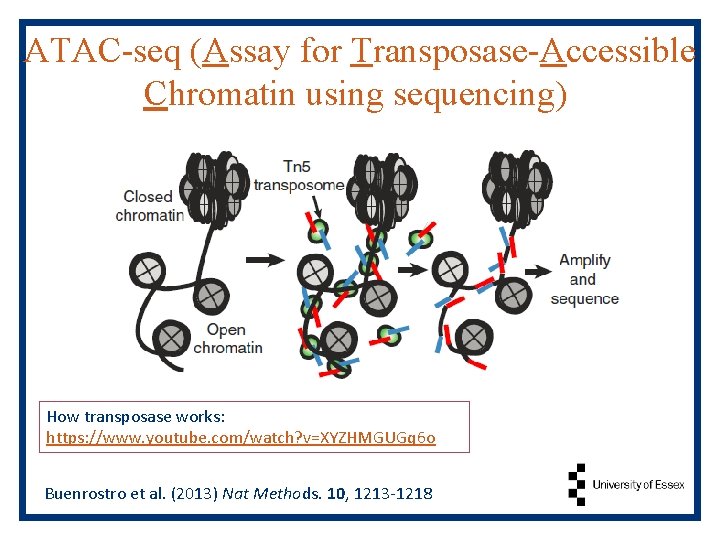
ATAC-seq (Assay for Transposase-Accessible Chromatin using sequencing) How transposase works: https: //www. youtube. com/watch? v=XYZHMGUGq 6 o Buenrostro et al. (2013) Nat Methods. 10, 1213 -1218

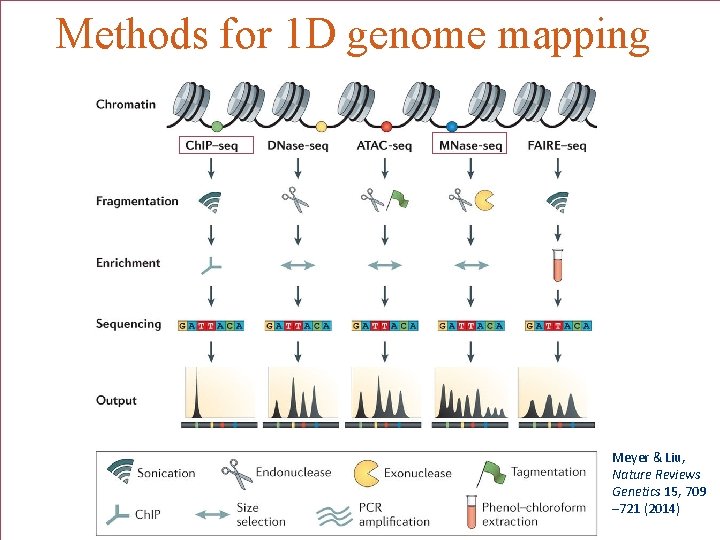
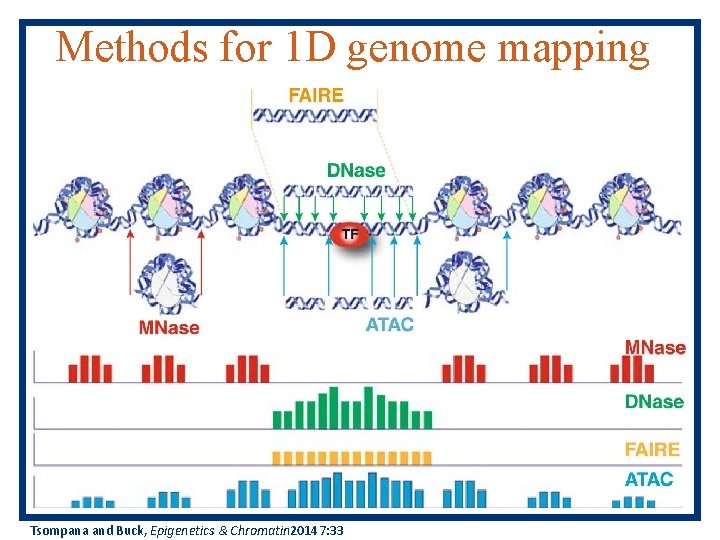
Methods for 1 D genome mapping MM Meyer & Liu, Nature Reviews Genetics 15, 709 – 721 (2014)

Methods for 1 D genome mapping Tsompana and Buck, Epigenetics & Chromatin 20147: 33

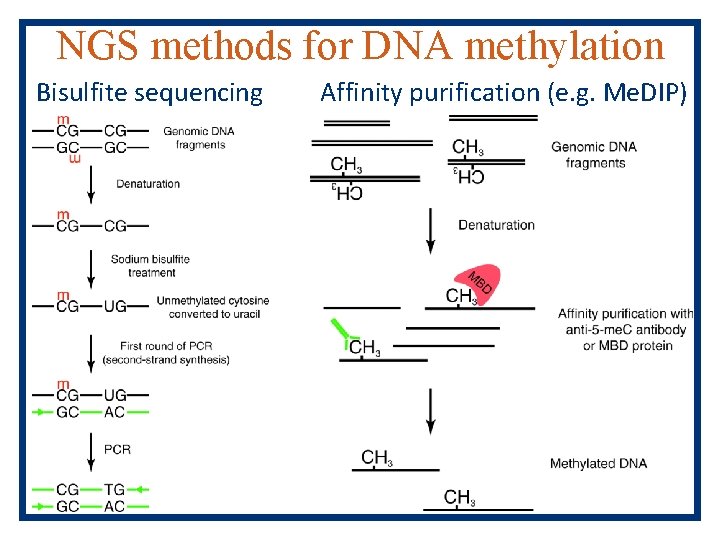
NGS methods for DNA methylation Bisulfite sequencing Affinity purification (e. g. Me. DIP)

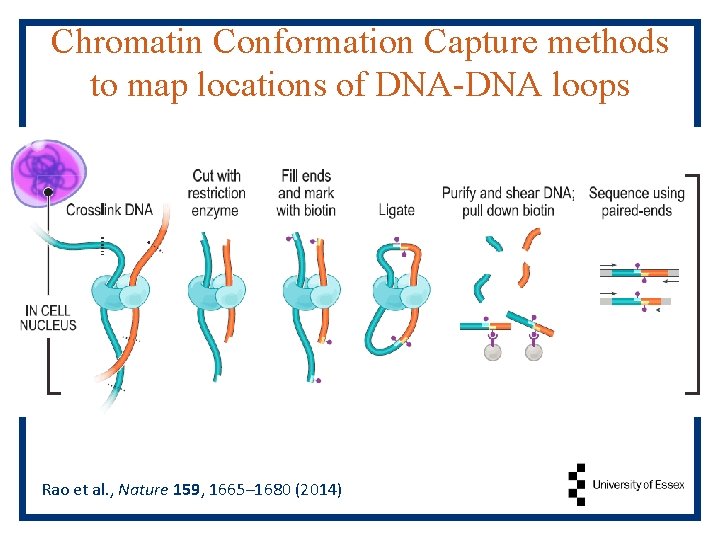
Chromatin Conformation Capture methods to map locations of DNA-DNA loops Rao et al. , Nature 159, 1665– 1680 (2014)

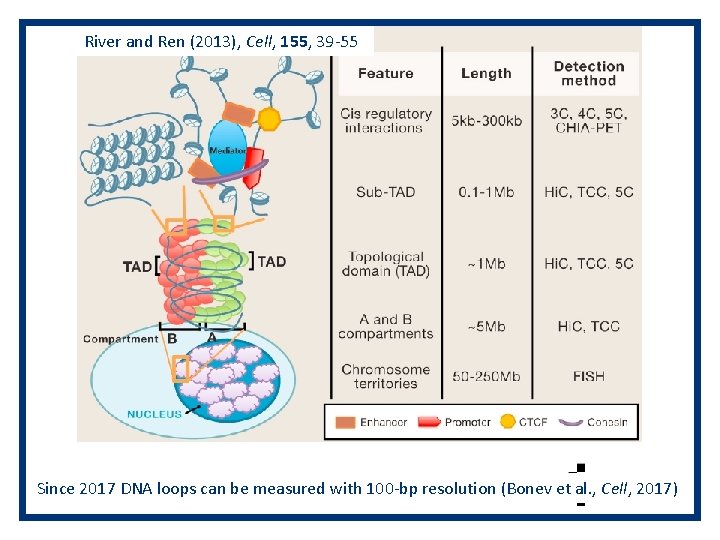
River and Ren (2013), Cell, 155, 39 -55 Since 2017 DNA loops can be measured with 100 -bp resolution (Bonev et al. , Cell, 2017)

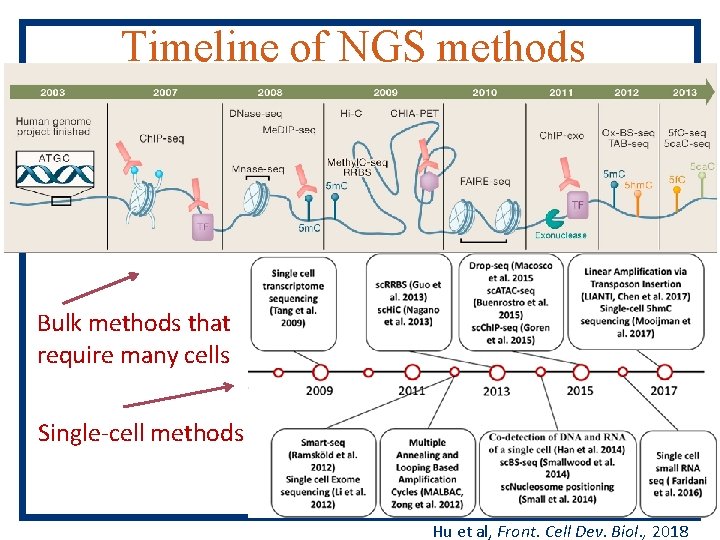
Timeline of NGS methods River and Ren (2013), Cell, 155, 39 -55 Bulk methods that require many cells Single-cell methods Hu et al, Front. Cell Dev. Biol. , 2018

Where to get NGS data? Ø Do your own experiment Ø Gene Expression Omnibus (GEO) https: //www. ncbi. nlm. nih. gov/geo Ø Sequence read archive (SRA) https: //www. ncbi. nlm. nih. gov/sra Ø European Nucleotide Archive https: //www. ebi. ac. uk/ena Ø The Cancer Genome Atlas (TCGA) https: //tcga-data. nci. nih. gov/tcga ØExome Aggregation Consortium (Ex. AC) http: //exac. broadinstitute. org/ You also have to upload your data!

Next generation sequencing analysis

How to analyze NGS data? q Ask a bioinformatician Ø you need to explain what do you want, and for that you need to understand what/how can be done q Do it yourself Ø Command line –> become a bioinformatician Ø Online wrappers –> simpler, but file size limits Example of a convenient online tool: Galaxy http: //galaxy. essex. ac. uk/

Ch. IP-seq (Chromatin Immuno. Precipitation followed by sequencing) 1. Crosslink Protein-DNA complexes in situ 2. Isolate nuclei and fragment DNA (sonication or digestion) 3. Immunoprecipitate with antibody against target nuclear protein and reverse crosslinks 4. Release DNA and submit for sequencing Adapted from www. Visi. Science. com

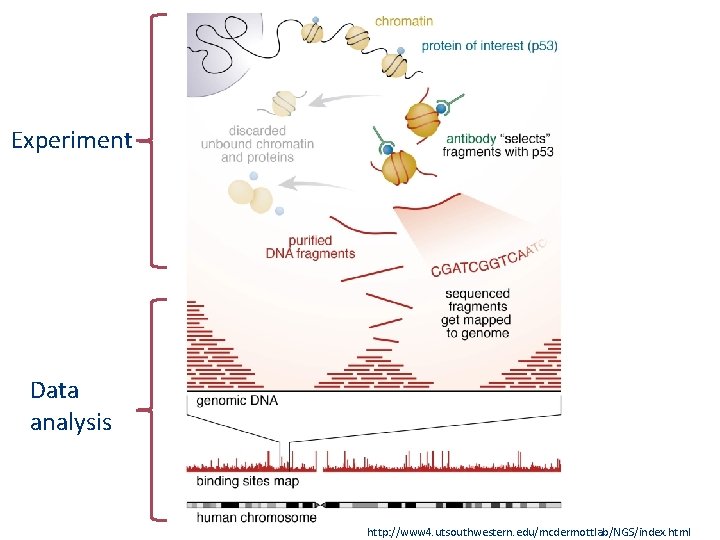
Experiment Data analysis http: //www 4. utsouthwestern. edu/mcdermottlab/NGS/index. html

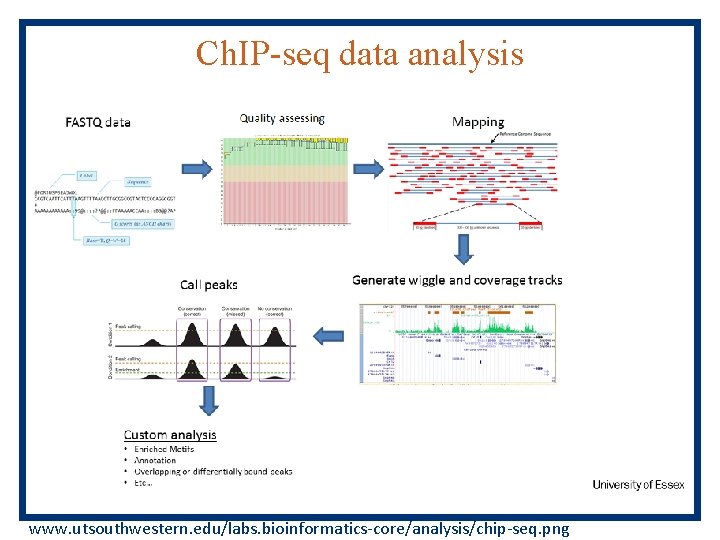
Ch. IP-seq data analysis www. utsouthwestern. edu/labs. bioinformatics-core/analysis/chip-seq. png

Unmapped sequenced reads (this is “raw”, primary data):

Mapped reads are characterised by their locations in the genome Bowtie, BWA, ELAND, Novoalign, BLAST, Clustal. W Top. Hat (for RNA-seq)

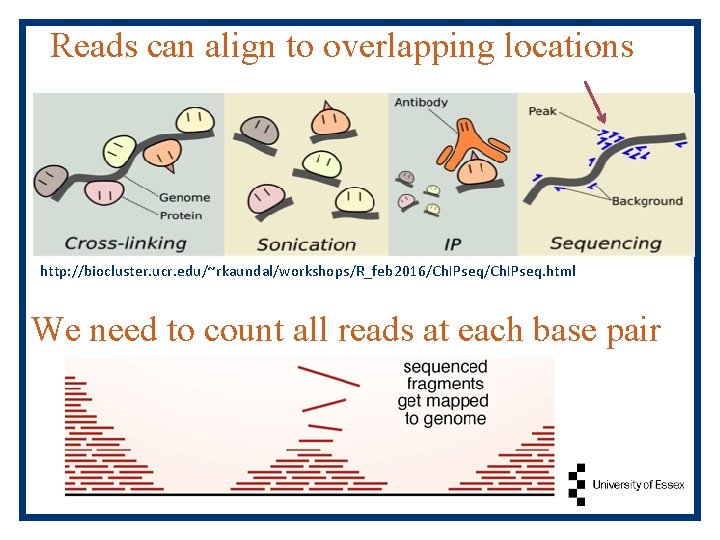
Reads can align to overlapping locations http: //biocluster. ucr. edu/~rkaundal/workshops/R_feb 2016/Ch. IPseq. html We need to count all reads at each base pair

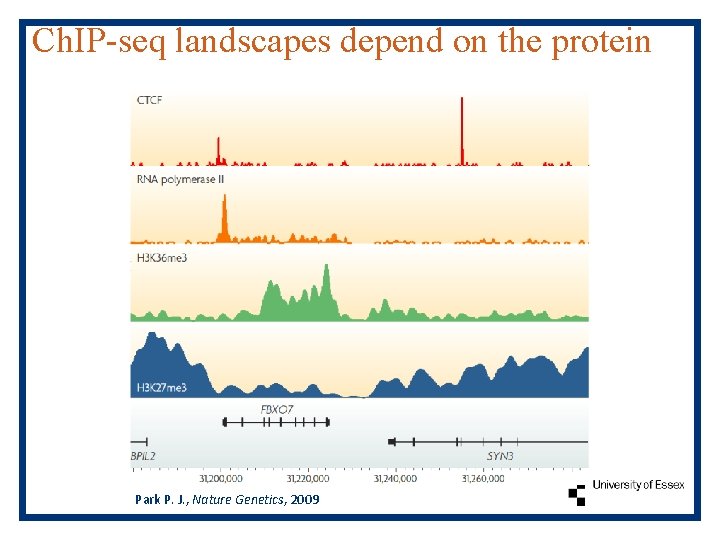
Ch. IP-seq landscapes depend on the protein Park P. J. , Nature Genetics, 2009

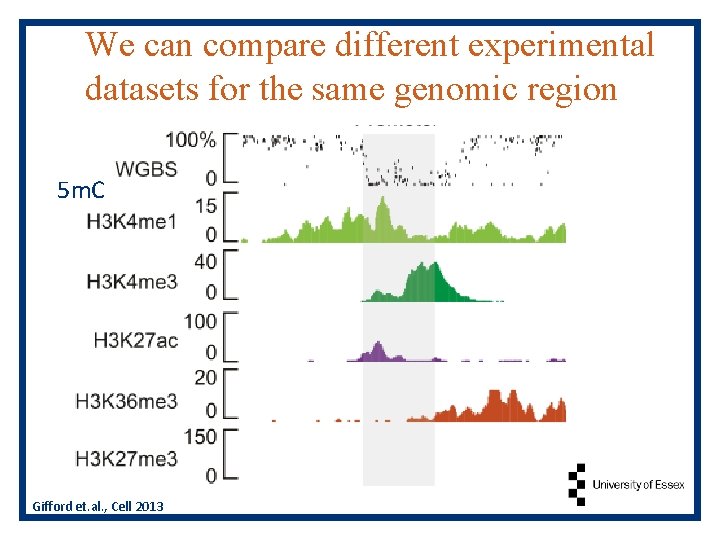
We can compare different experimental datasets for the same genomic region 5 m. C Gifford et. al. , Cell 2013

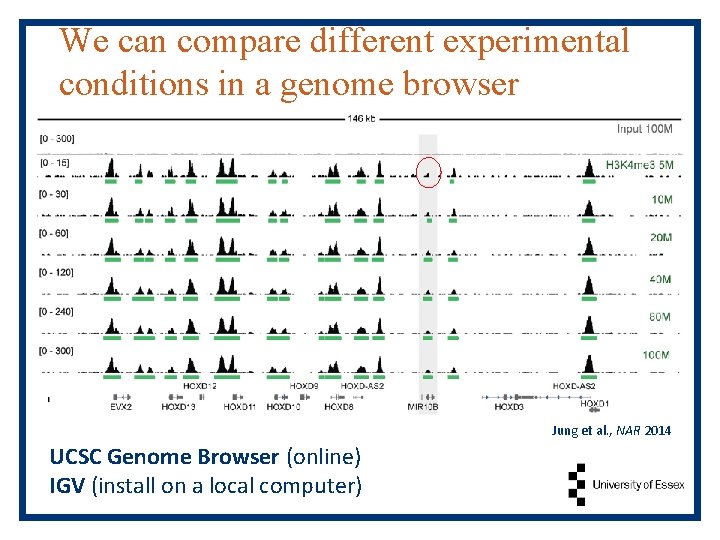
We can compare different experimental conditions in a genome browser Jung et al. , NAR 2014 UCSC Genome Browser (online) IGV (install on a local computer)

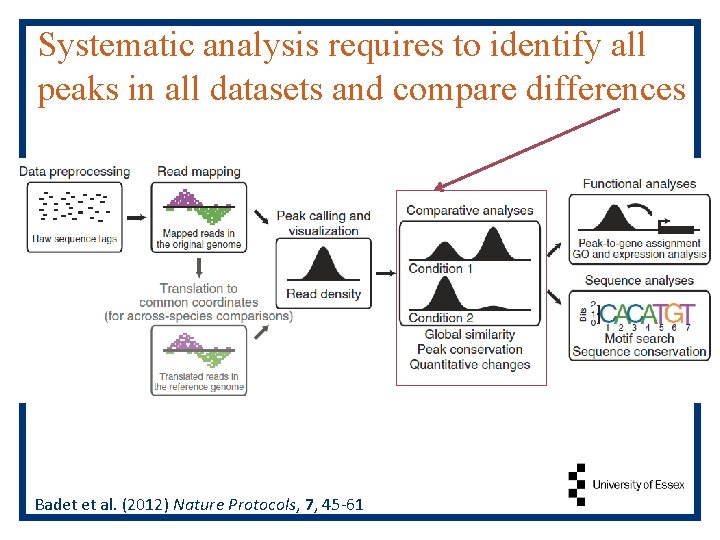
Systematic analysis requires to identify all peaks in all datasets and compare differences Badet et al. (2012) Nature Protocols, 7, 45 -61

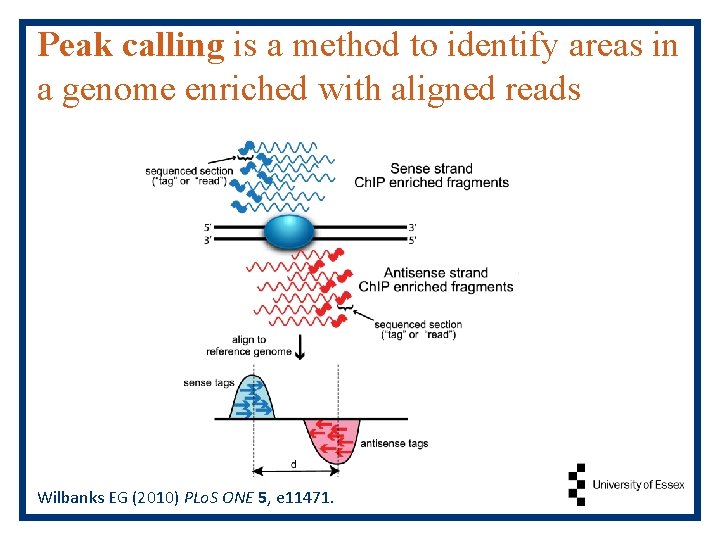
Peak calling is a method to identify areas in a genome enriched with aligned reads Wilbanks EG (2010) PLo. S ONE 5, e 11471.

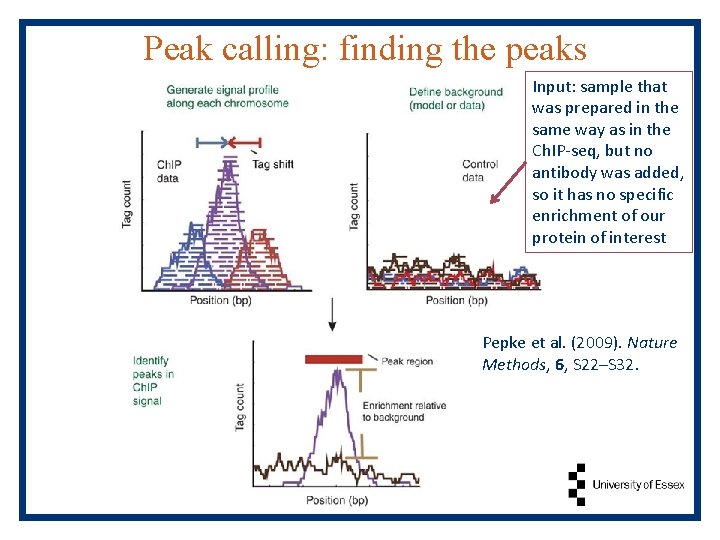
Peak calling: finding the peaks Input: sample that was prepared in the same way as in the Ch. IP-seq, but no antibody was added, so it has no specific enrichment of our protein of interest Pepke et al. (2009). Nature Methods, 6, S 22–S 32.

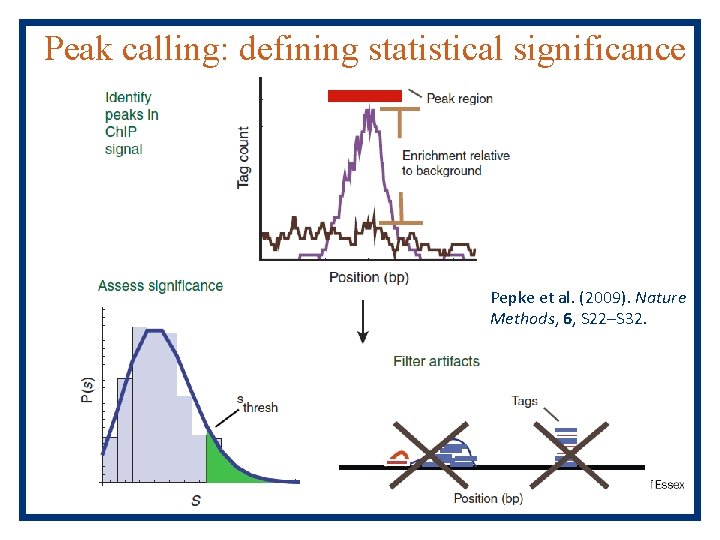
Peak calling: defining statistical significance Pepke et al. (2009). Nature Methods, 6, S 22–S 32.

Peak calling: defining statistical significance Is this peak statistically significant? Park P. J. , Nature Genetics, 2009 MACS (good for TFs) CISER (histones, etc) HOMER (universal) Peak. Seq edge. R Cis. Genome

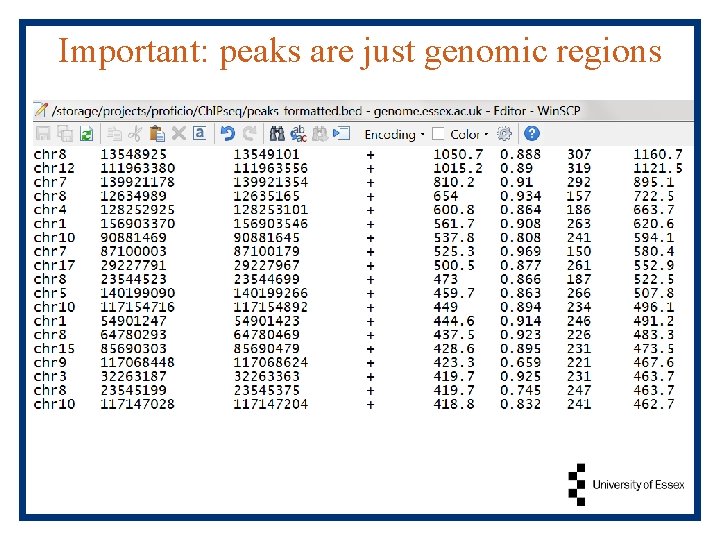
Important: peaks are just genomic regions

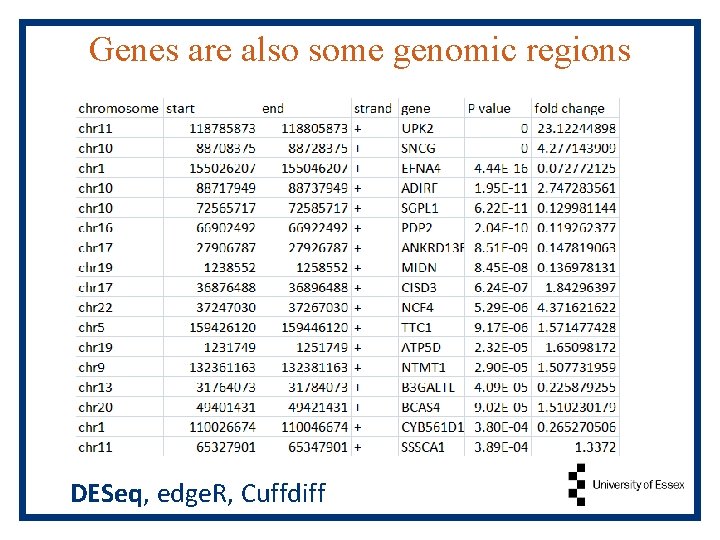
Genes are also some genomic regions DESeq, edge. R, Cuffdiff

DNA methylation: also genomic regions Individual Cp. Gs BISMARK Differentially methylated regions DMRcaller

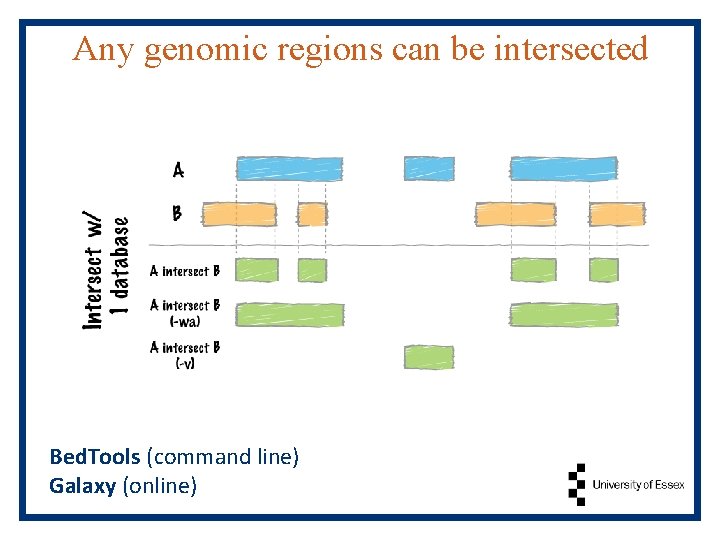
Any genomic regions can be intersected Bed. Tools (command line) Galaxy (online)

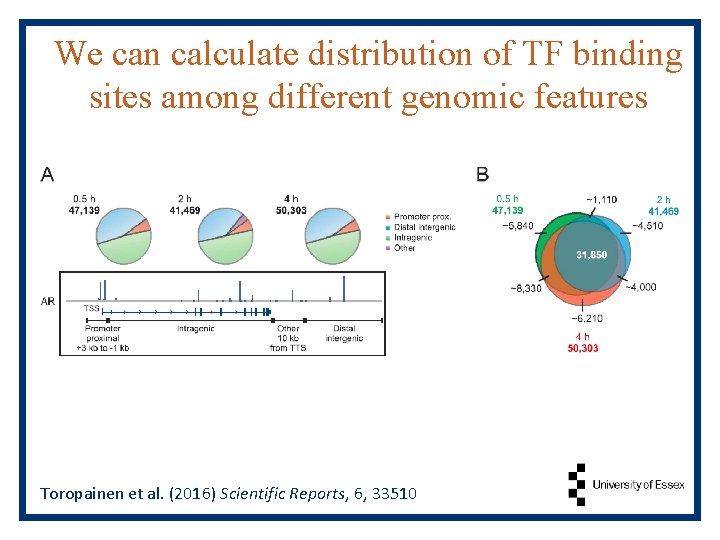
We can calculate distribution of TF binding sites among different genomic features Toropainen et al. (2016) Scientific Reports, 6, 33510

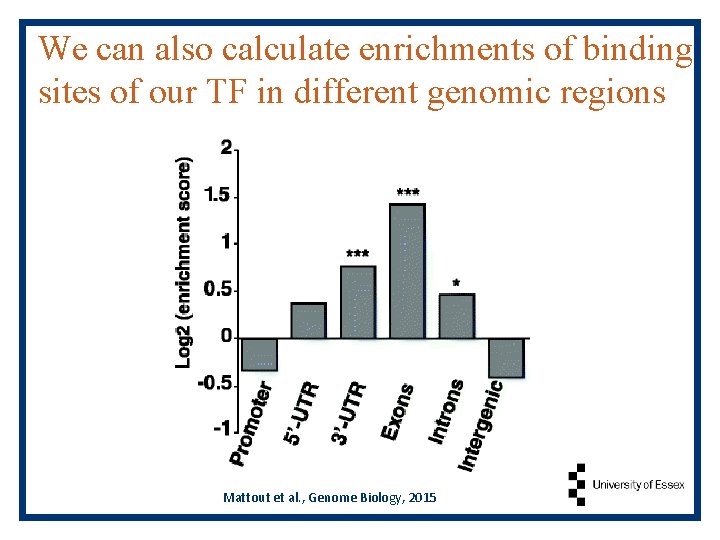
We can also calculate enrichments of binding sites of our TF in different genomic regions Mattout et al. , Genome Biology, 2015

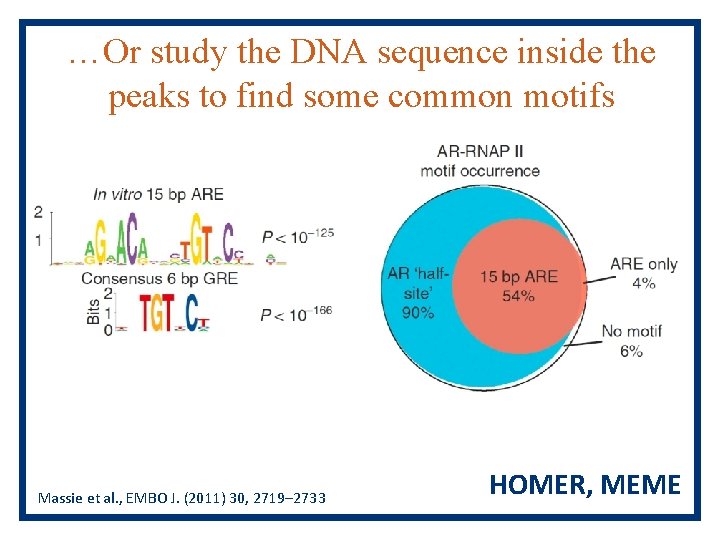
…Or study the DNA sequence inside the peaks to find some common motifs Massie et al. , EMBO J. (2011) 30, 2719– 2733 HOMER, MEME

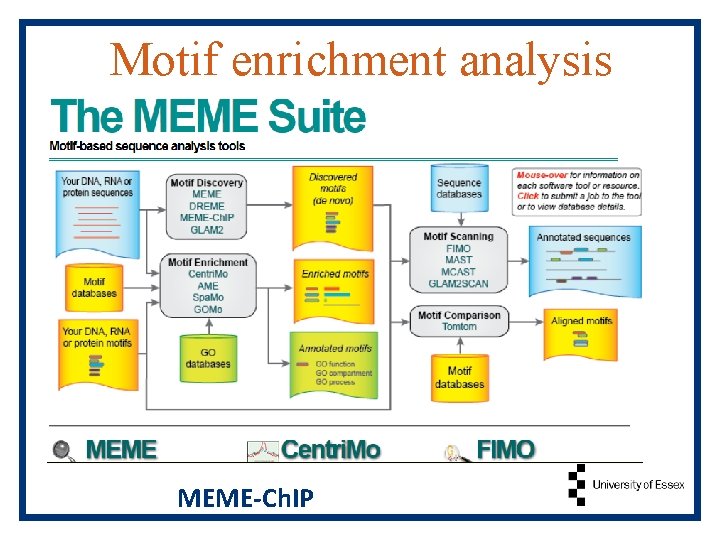
Motif enrichment analysis MEME-Ch. IP

What else can we do with peaks? q Compare two experimental conditions to see which peaks appear/disappear (e. g. protein binding gained/lost); q Compute associations of our protein with different genes (e. g. define which genes are regulated by this protein) q Study the DNA sequence inside the peaks (e. g. to find which other TFs co-bind with our protein of interest) q Look how our peaks are arranged with respect to other peaks (e. g. to check for interactions with other proteins) q etc