The Web frame for NGS output NGS sequencing

The Web frame for NGS output
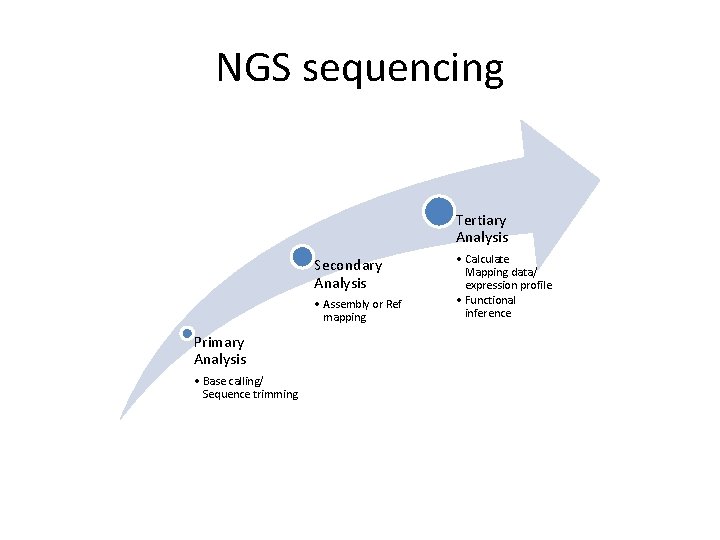
NGS sequencing Tertiary Analysis Secondary Analysis • Assembly or Ref mapping Primary Analysis • Base calling/ Sequence trimming • Calculate Mapping data/ expression profile • Functional inference

No-model Organism Tentative Procedure for RNA –Seq Analysis QC • Discard the low –confident sequences for 3 groups (three time points) • Program: Solexa. QA (http: //solexaqa. sourceforge. net/) Assembly • Merge all reads from 3 Groups for assembly to form Contigs • Program: Trinity (http: //trinityrnaseq. sourceforge. net/), 100 GB RAM requested Mapping • Map pair-end reads from each group on Contigs/ Annotate Contigs • Program: LAST (http: //last. cbrc. jp/), BLASTx, Interpro. Scan Expression • Estimate the expression value for each contig in each group (FPKM) • Program: Cumme. Rbund, an R/Bioconductor package (http: //cufflinks. cbcb. umd. edu/) Functional inference • Functional enrichment analysis in GO and KEGG • Program: Due to no-model organism, we may have to create the mapping identifier in KEGG and GO

No-model Organism for Eel transcriptomics Tentative Procedure for RNA –Seq Analysis QC • Discard the low –confident sequences generated from each library in Hi-seq 200, RNA-seq data, Pairend • Program: Solexa. QA (http: //solexaqa. sourceforge. net/) Assembly • Merge all reads from various libraries for assembly to form Contigs • Program: Trinity (http: //trinityrnaseq. sourceforge. net/), 100 GB RAM requested Mapping • Map pair-end reads from each group on Contigs/ Annotate Contigs • Program: LAST (http: //last. cbrc. jp/), BLASTx, Interpro. Scan Expression Profiling Functional inference • Estimate the expression value for each contig in each group (FPKM) • Program: Cumme. Rbund, an R/Bioconductor package (http: //cufflinks. cbcb. umd. edu/) • Functional enrichment analysis in GO and KEGG • Program: Due to no-model organism, we may have to create the mapping identifier in KEGG and GO

No-model Organism Tentative Procedure for RNA –Seq Analysis QC • 去除品質較差的定序結果 • Program: Solexa. QA (http: //solexaqa. sourceforge. net/), Seq. Trim Assembly • 由短序列基因定序結果中,組合出可能的基因表現模組(Merge all reads from 3 Groups for assembly to form Contigs) • Program: Trinity, MIRA, Valvet, etc, multiple CPUs with over 100 GB RAM requested Mapping • 以組合出來的長序列基因片段為主體,將短序列歸位到基因主體上(Map pair-end reads from each group on Contigs) • Program: Bowtie, LAST (http: //last. cbrc. jp/) Expression • 計算與統計不同樣品間同一段基因表現的概況,鑑別出有差異表現基因群(Estimate the expression value for each contig in each group (FPKM)) • Program: Cumme. Rbund, an R/Bioconductor package ( http: //cufflinks. cbcb. umd. edu/), rseq. C (http: //code. google. com/p/rseqc/) Functional inference • 將找出的基因群進行功能性分析,找出在不同時間與組織下,與再生機制相關之調控途徑(Functional enrichment analysis in GO and KEGG) • Program: Due to no-model organism, we may have to create the mapping identifier in KEGG and GO Validation • 以Q-PCR來確認與再生相關之基因群表現概況 • 設計新的實驗來促進或是干擾再生機制,再透過NGS來找出更為精細的調控細節
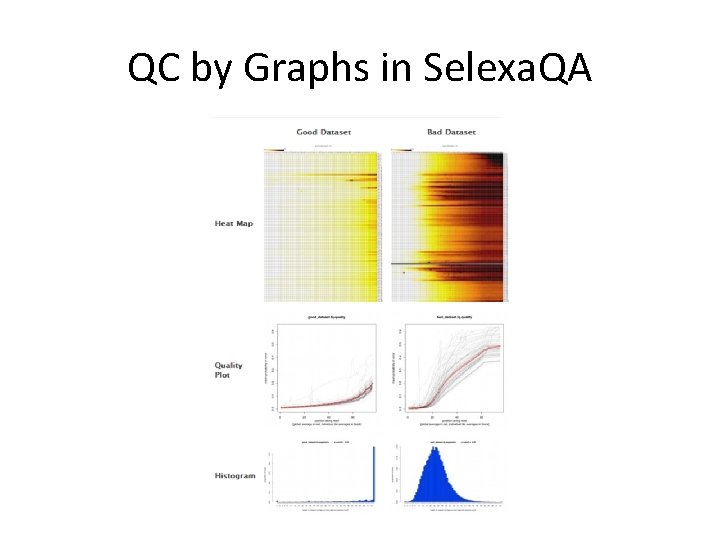
QC by Graphs in Selexa. QA

Annotations for each Contig • Contig in FASTA (N. A) Translated sequence (AA) in longest ORF • Then perform Sequence Search (BLASTp) on NR, KEGG, GO, p. Fam (Interpro)
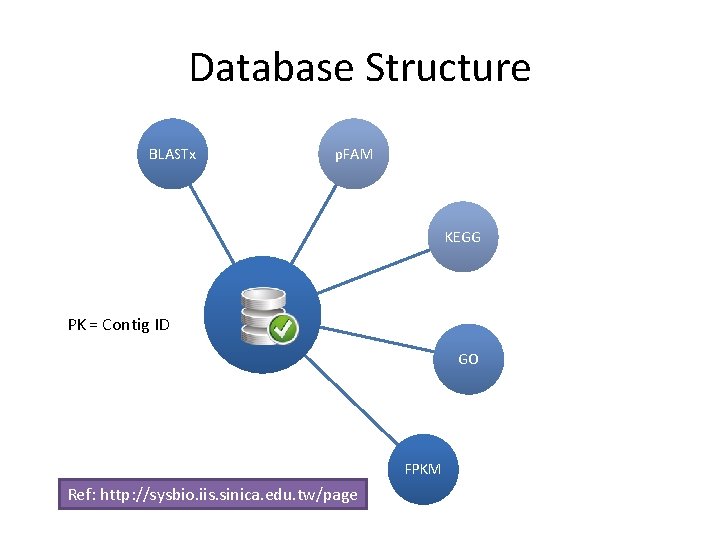
Database Structure BLASTx p. FAM KEGG PK = Contig ID GO FPKM Ref: http: //sysbio. iis. sinica. edu. tw/page
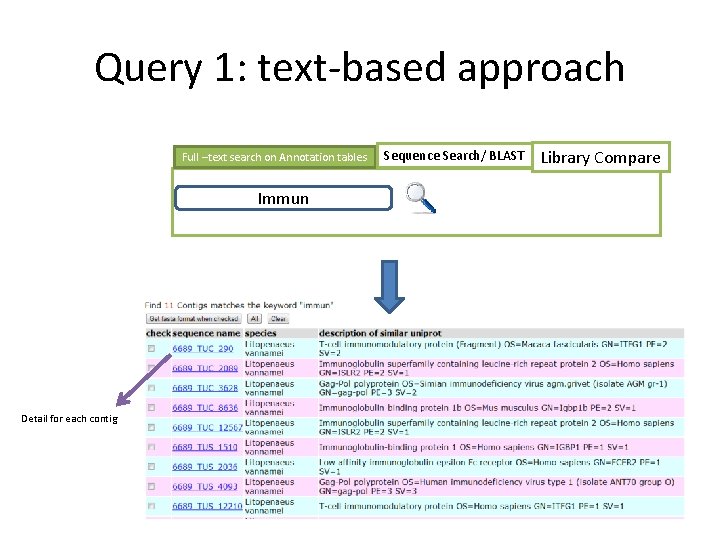
Query 1: text-based approach Full –text search on Annotation tables Immun Detail for each contig Sequence Search/ BLAST Library Compare
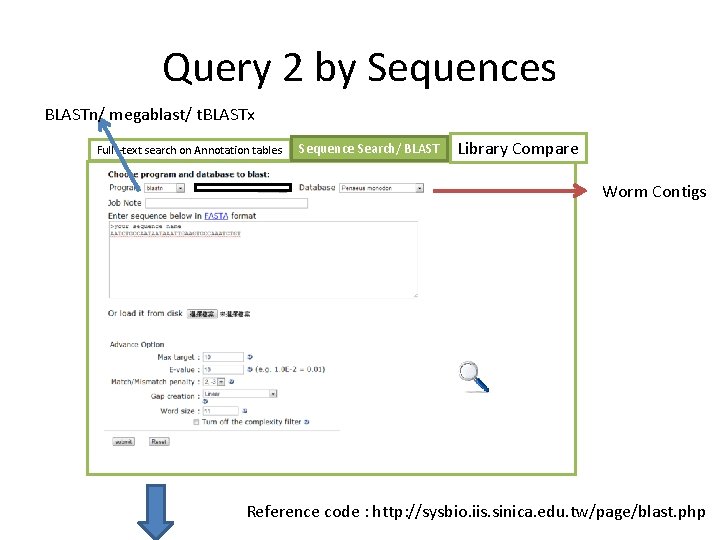
Query 2 by Sequences BLASTn/ megablast/ t. BLASTx Full –text search on Annotation tables Sequence Search/ BLAST Library Compare Worm Contigs Reference code : http: //sysbio. iis. sinica. edu. tw/page/blast. php

Blast Result Detail for each contig

Detail for Each Contig Interpro/ p. FAM
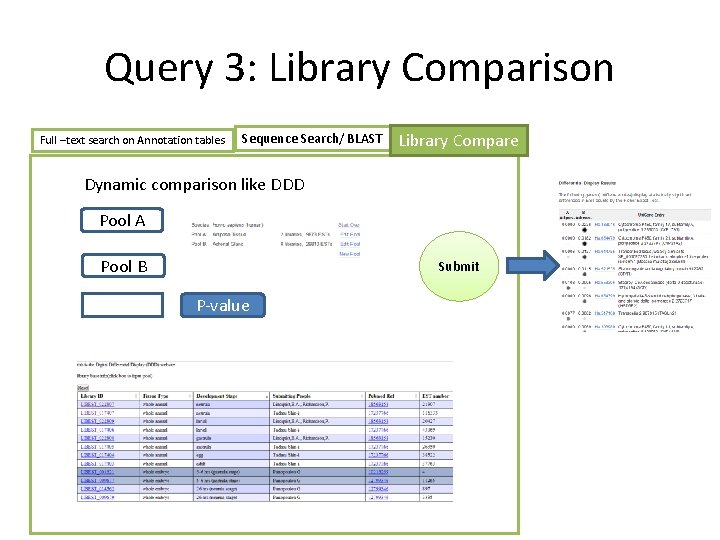
Query 3: Library Comparison Full –text search on Annotation tables Sequence Search/ BLAST Library Compare Dynamic comparison like DDD Pool A Pool B Submit P-value
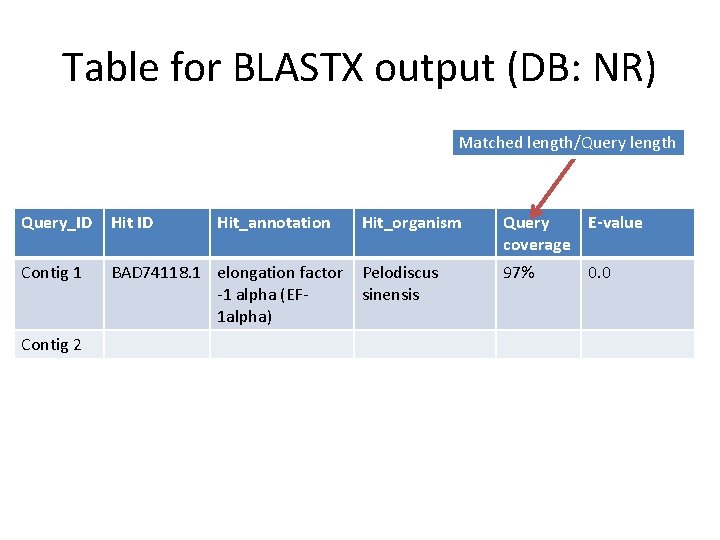
Table for BLASTX output (DB: NR) Matched length/Query length Query_ID Hit ID Contig 1 BAD 74118. 1 elongation factor -1 alpha (EF 1 alpha) Contig 2 Hit_annotation Hit_organism Query E-value coverage Pelodiscus sinensis 97% 0. 0
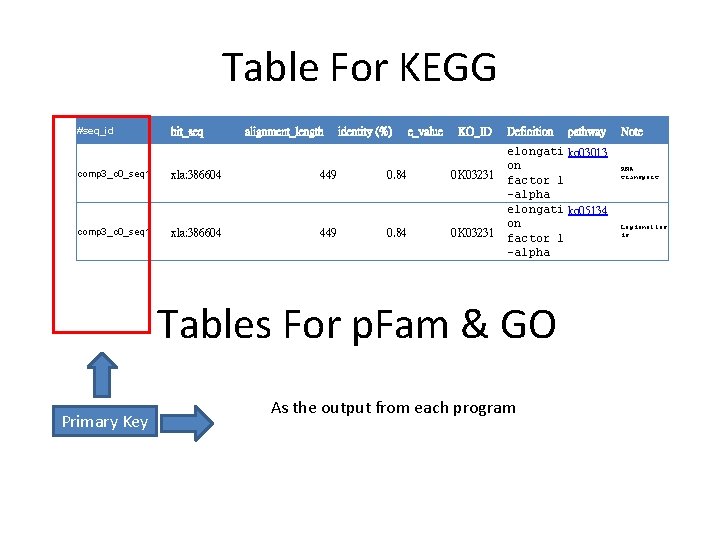
Table For KEGG #seq_id hit_seq alignment_length identity (%) e_value KO_ID comp 3_c 0_seq 1 xla: 386604 449 0. 84 0 K 03231 Definition elongati ko 03013 on factor 1 -alpha elongati ko 05134 on factor 1 -alpha Tables For p. Fam & GO Primary Key pathway As the output from each program Note RNA transport Legionellos is
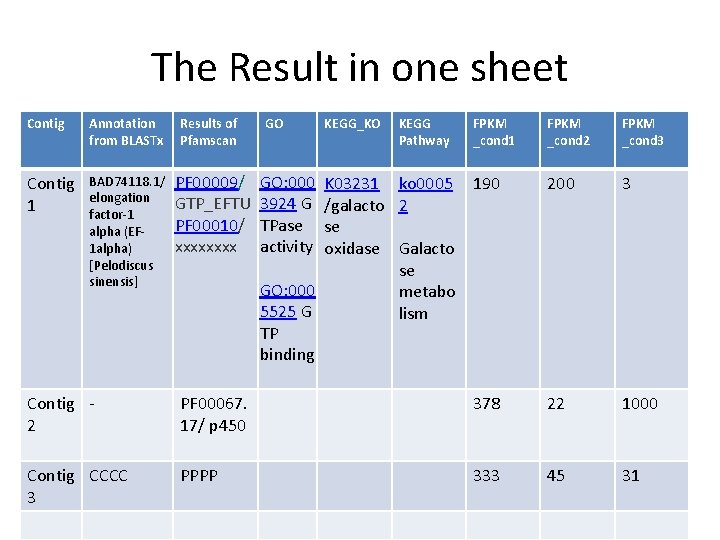
The Result in one sheet Contig Annotation from BLASTx Results of Pfamscan Contig 1 BAD 74118. 1/ elongation factor-1 alpha (EF 1 alpha) [Pelodiscus sinensis] PF 00009/ GTP_EFTU PF 00010/ xxxx GO KEGG_KO KEGG Pathway FPKM _cond 1 GO: 000 3924 G TPase activity K 03231 ko 0005 190 /galacto 2 se oxidase Galacto se GO: 000 metabo 5525 G lism TP binding FPKM _cond 2 FPKM _cond 3 200 3 Contig 2 PF 00067. 17/ p 450 378 22 1000 Contig CCCC 3 PPPP 333 45 31
Library Compare 0 hr 24 hrs 48 hrs

The Way of Redundancy Reduction Input 700 Million reads 1 st Trinity Run 500, 000 genes Abundance Sorting 48, 000 Genes Refinement Final Set Mapping by BOWTIE 2 (LAST? ), pick longest one as reduced set
- Slides: 18