From Informatics to Bioinformatics Limsoon Wong Institute for

From Informatics to Bioinformatics Limsoon Wong Institute for Infocomm Research Singapore
What is Bioinformatics?

Themes of Bioinformatics = Data Mgmt + Knowledge Discovery Data Mgmt = Integration + Transformation + Cleansing Knowledge Discovery = Statistics + Algorithms + Databases

Benefits of Bioinformatics To the patient: Better drug, better treatment To the pharma: Save time, save cost, make more $ To the scientist: Better science
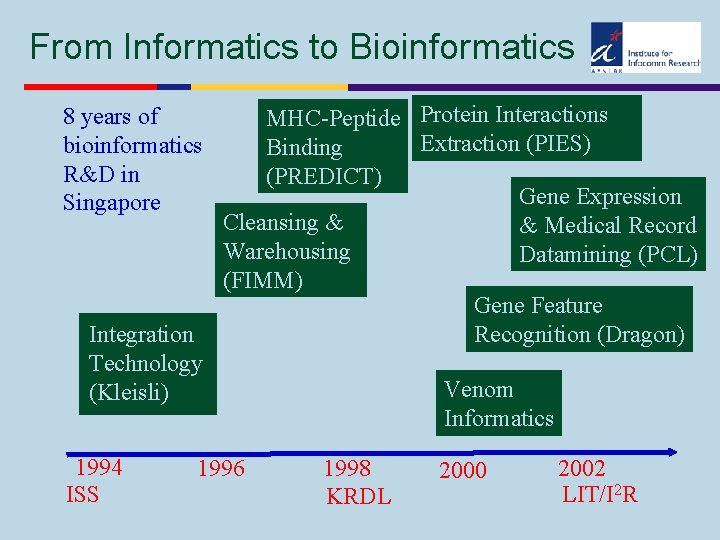
From Informatics to Bioinformatics 8 years of bioinformatics R&D in Singapore Integration Technology (Kleisli) 1994 ISS MHC-Peptide Protein Interactions Extraction (PIES) Binding (PREDICT) Gene Expression Cleansing & & Medical Record Warehousing Datamining (PCL) (FIMM) Gene Feature Recognition (Dragon) 1996 Venom Informatics 1998 KRDL 2000 2002 LIT/I 2 R

Data Integration A DOE “impossible query”: For each gene on a given cytogenetic band, find its non-human homologs.

Data Integration Results • Using Kleisli: • Clear • Succinct • Efficient sybase-add (#name: ”GDB", . . . ); create view L from locus_cyto_location using GDB; create view E from object_genbank_eref using GDB; select #accn: g. #genbank_ref, #nonhuman-homologs: H from L as c, E as g, • Handles • heterogeneity • complexity {select u from g. #genbank_ref. na-get-homolog-summary as u where not(u. #title string-islike "%Human%") andalso not(u. #title string-islike "%H. sapien%")} as H where c. #chrom_num = "22” andalso g. #object_id = c. #locus_id andalso not (H = { });

Data Warehousing Motivation efficiency availabilty “denial of service” data cleansing Requirements efficient to query easy to update. model data naturally {(#uid: 6138971, #title: "Homo sapiens adrenergic. . . ", #accession: "NM_001619", #organism: "Homo sapiens", #taxon: 9606, #lineage: ["Eukaryota", "Metazoa", …], #seq: "CTCGGCCTCGGGCGCGGC. . . ", #feature: { (#name: "source", #continuous: true, #position: [ (#accn: "NM_001619", #start: 0, #end: 3602, #negative: false)], #anno: [ (#anno_name: "organism", #descr: "Homo sapiens"), …] ), …)}

Data Warehousing Results Relational DBMS is insufficient because it forces us to fragment data into 3 NF. Kleisli turns flat relational DBMS into nested relational DBMS. It can use flat relational DBMS such as Sybase, Oracle, My. SQL, etc. to be its update-able complex object store. ! Log in oracle-cplobj-add (#name: "db", . . . ); ! Define table create table GP (#uid: "NUMBER", #detail: "LONG") using db; ! Populate table with Gen. Pept reports select #uid: x. #uid, #detail: x into GP from aa-get-seqfeat-general "PTP” as x using db; ! Map GP to that table create view GP from GP using db; ! Run a queryto get title of 131470 select x. #detail. #title from GP as x where x. #uid = 131470;

Epitope Prediction TRAP-559 AA MNHLGNVKYLVIVFLIFFDLFLVNGRDVQNNIVDEIKYSE EVCNDQVDLYLLMDCSGSIRRHNWVNHAVPLAMKLIQQLN LNDNAIHLYVNVFSNNAKEIIRLHSDASKNKEKALIIIRS LLSTNLPYGRTNLTDALLQVRKHLNDRINRENANQLVVIL TDGIPDSIQDSLKESRKLSDRGVKIAVFGIGQGINVAFNR FLVGCHPSDGKCNLYADSAWENVKNVIGPFMKAVCVEVEK TASCGVWDEWSPCSVTCGKGTRSRKREILHEGCTSEIQEQ CEEERCPPKWEPLDVPDEPEDDQPRPRGDNSSVQKPEENI IDNNPQEPSPNPEEGKDENPNGFDLDENPENPPNPDIPEQ KPNIPEDSEKEVPSDVPKNPEDDREENFDIPKKPENKHDN QNNLPNDKSDRNIPYSPLPPKVLDNERKQSDPQSQDNNGN RHVPNSEDRETRPHGRNNENRSYNRKYNDTPKHPEREEHE KPDNNKKKGESDNKYKIAGGLALLACAGLAYKFVVP GAATPYAGEPAPFDETLGEEDKDLDEPEQFRLPEENEWN

Epitope Prediction Results w Prediction by our ANN model for HLA-A 11 w w w 29 predictions 22 epitopes 76% specificity w Prediction by BIMAS matrix for HLA-A*1101 Number of experimental binders 19 (52. 8%) 5 (13. 9%) 12 (33. 3%) 1 66 100 Rank by BIMAS
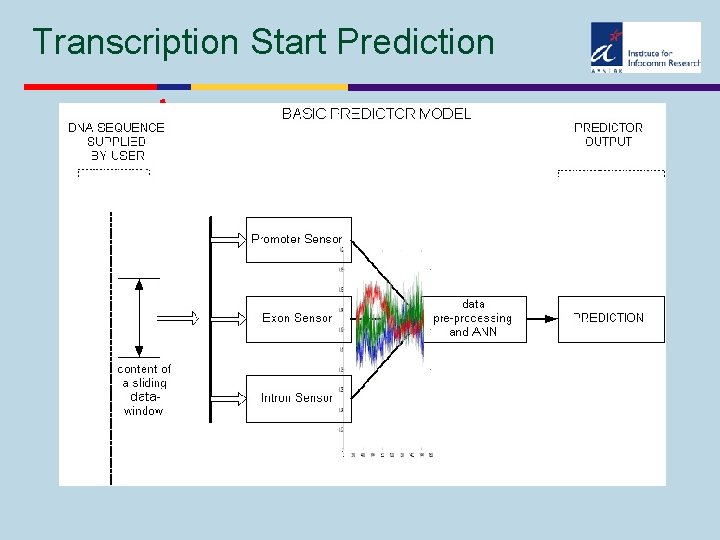
Transcription Start Prediction
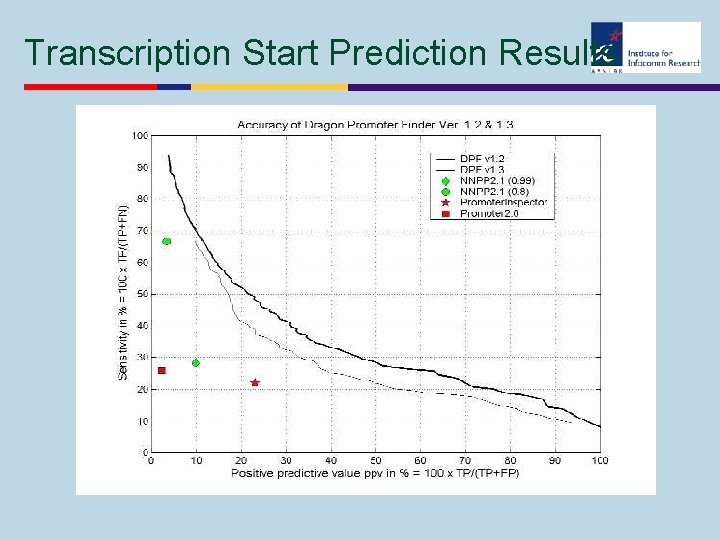
Transcription Start Prediction Results
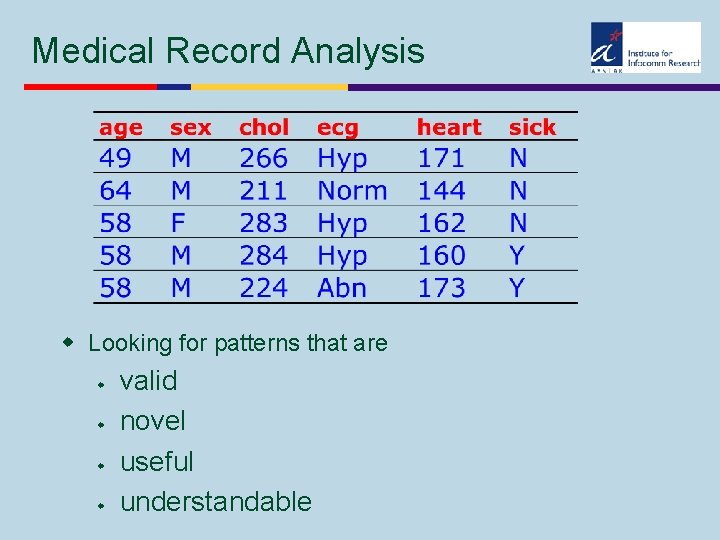
Medical Record Analysis w Looking for patterns that are w w valid novel useful understandable
Gene Expression Analysis w Classifying gene expression profiles w w w find stable differentially expressed genes find significant gene groups derive coordinated gene expression
Medical Record & Gene Expression Analysis Results § PCL, a novel “emerging pattern’’ method § Beats C 4. 5, CBA, LB, NB, TAN in 21 out of 32 UCI benchmarks § Works well for gene expressions Cancer Cell, March 2002, 1(2)
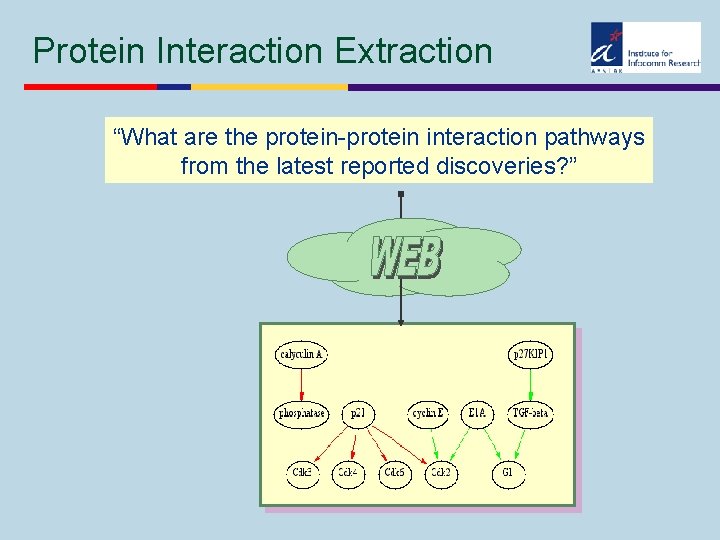
Protein Interaction Extraction “What are the protein-protein interaction pathways from the latest reported discoveries? ”
Protein Interaction Extraction Results w Rule-based system for processing free texts in scientific abstracts w Specialized in w extracting protein names w extracting protein interactions

Behind the Scene w Vladimir Bajic w Vladimir Brusic w Jinyan Li w See-Kiong Ng w Limsoon Wong w Louxin Zhang w Allen Chong w Judice Koh w SPT Krishnan w Huiqing Liu w Seng Hong Seah w Soon Heng Tan w Guanglan Zhang w Zhuo Zhang and many more: students, folks from genetic. Xchange, Molecular. Connections, and other collaborators….

Using Feature Generation & Feature Selection for Accurate Prediction of Translation Initiation Sites A more detailed example of post-genome knowledge discovery
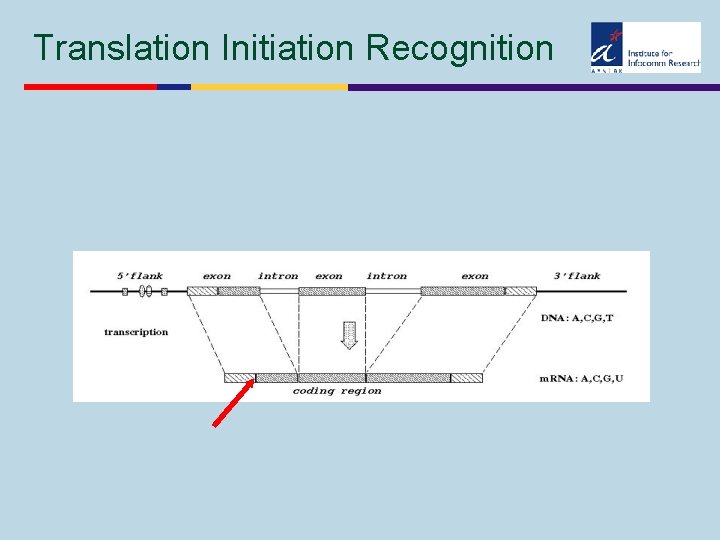
Translation Initiation Recognition

A Sample c. DNA 299 HSU 27655. 1 CAT U 27655 Homo sapiens CGTGTGTGCAGCAGCCTGCAGCTGCCCCAAGCCATGGCTGAACACTGACTCCCAGCTGTG CCCAGGGCTTCAAAGACTTCTCAGCTTCGAGCATGGCTTTTGGCTGTCAGGGCAGCTGTA GGAGGCAGATGAGAAGAGGGAGATGGCCTTGGAGGAAGGGGCCTGGTGCCGAGGA CCTCTCCTGGCCAGGAGCTTCCTCCAGGACAAGACCTTCCACCCAACAAGGACTCCCCT. . . . . . i. EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE What makes the second ATG the translation initiation site? 80 160 240

Approach • Training data gathering • Signal generation § k-grams, distance, domain know-how, . . . • Signal selection § Entropy, 2, CFS, t-test, domain knowhow. . . • Signal integration § SVM, ANN, PCL, CART, C 4. 5, k. NN, . . .
![Training & Testing Data • Vertebrate dataset of Pedersen & Nielsen [ISMB’ 97] • Training & Testing Data • Vertebrate dataset of Pedersen & Nielsen [ISMB’ 97] •](http://slidetodoc.com/presentation_image/654466690ccd1c8962c1331731a931f5/image-24.jpg)
Training & Testing Data • Vertebrate dataset of Pedersen & Nielsen [ISMB’ 97] • • • 3312 sequences 13503 ATG sites 3312 (24. 5%) are TIS 10191 (75. 5%) are non-TIS Use for 3 -fold x-validation expts

Signal Generation • K-grams (ie. , k consecutive letters) • • K = 1, 2, 3, 4, 5, … Window size vs. fixed position Up-stream, downstream vs. any where in window In-frame vs. any frame

Too Many Signals • For each value of k, there are 4 k * 3 * 2 k-grams • If we use k = 1, 2, 3, 4, 5, we have 4 + 24 + 96 + 384 + 1536 + 6144 = 8188 features! • This is too many for most machine learning algorithms

Signal Selection (Basic Idea) • Choose a signal w/ low intra-class distance • Choose a signal w/ high inter-class distance • Which of the following 3 signals is good?

Signal Selection (eg. , t-statistics)

Signal Selection (eg. , CFS) • Instead of scoring individual signals, how about scoring a group of signals as a whole? • CFS • A good group contains signals that are highly correlated with the class, and yet uncorrelated with each other
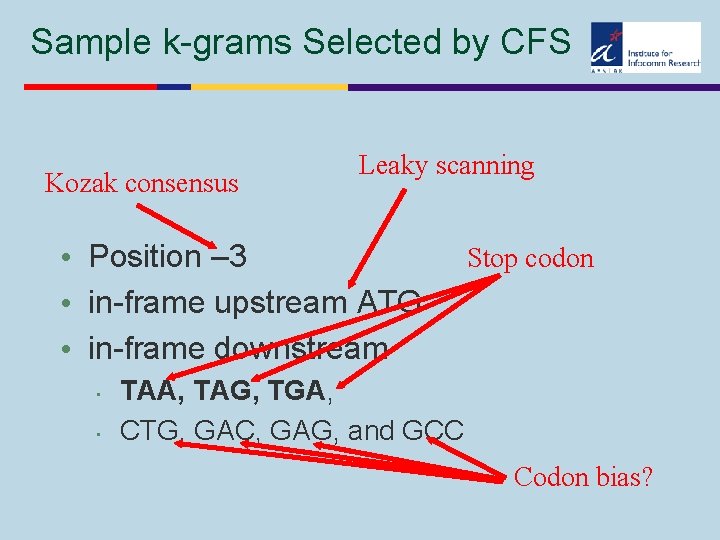
Sample k-grams Selected by CFS Kozak consensus Leaky scanning • Position – 3 • in-frame upstream ATG • in-frame downstream • • Stop codon TAA, TAG, TGA, CTG, GAC, GAG, and GCC Codon bias?

Signal Integration • k. NN Given a test sample, find the k training samples that are most similar to it. Let the majority class win. • SVM Given a group of training samples from two classes, determine a separating plane that maximises the margin of error. • Naïve Bayes, ANN, C 4. 5, . . .
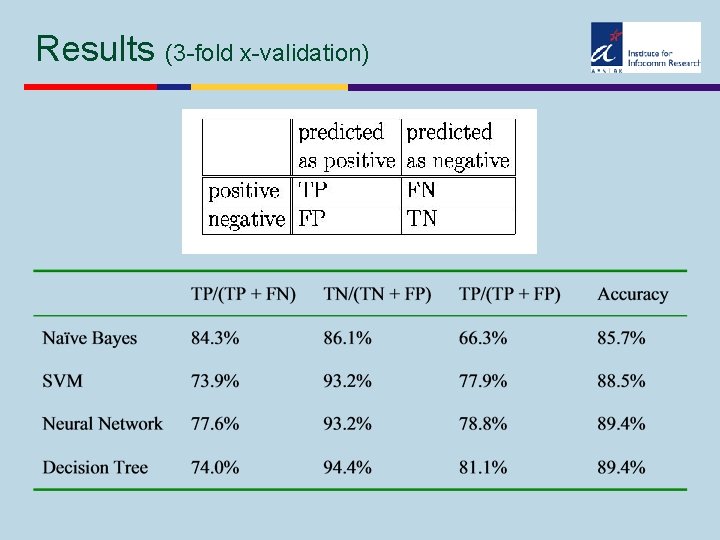
Results (3 -fold x-validation)
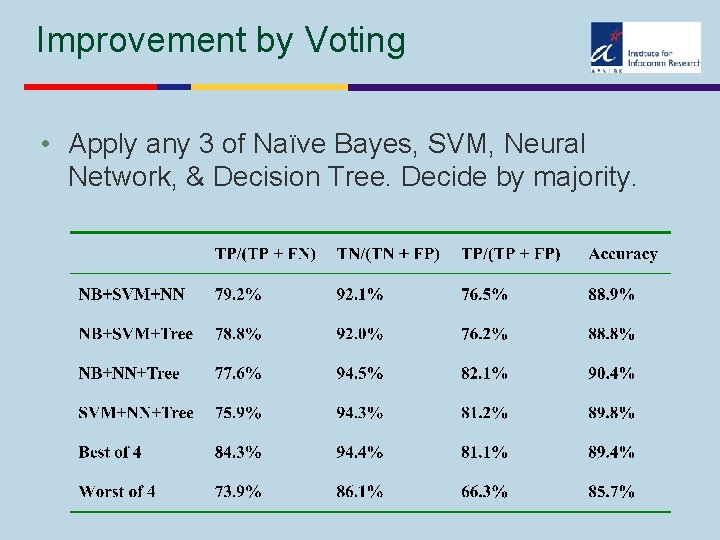
Improvement by Voting • Apply any 3 of Naïve Bayes, SVM, Neural Network, & Decision Tree. Decide by majority.
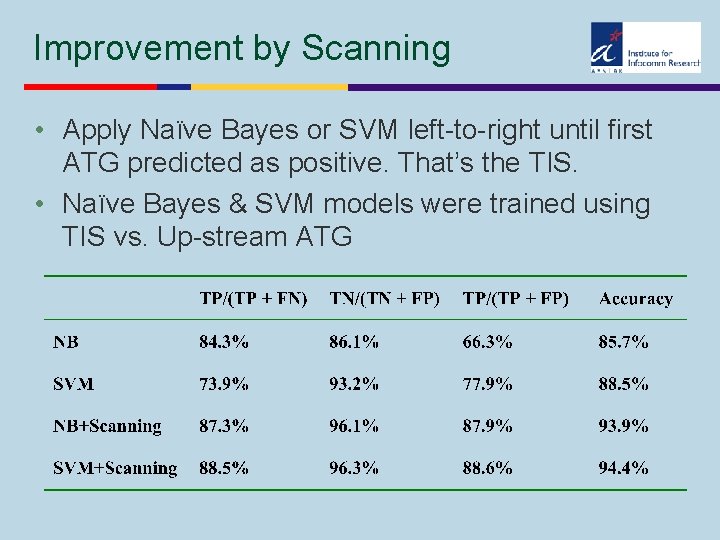
Improvement by Scanning • Apply Naïve Bayes or SVM left-to-right until first ATG predicted as positive. That’s the TIS. • Naïve Bayes & SVM models were trained using TIS vs. Up-stream ATG
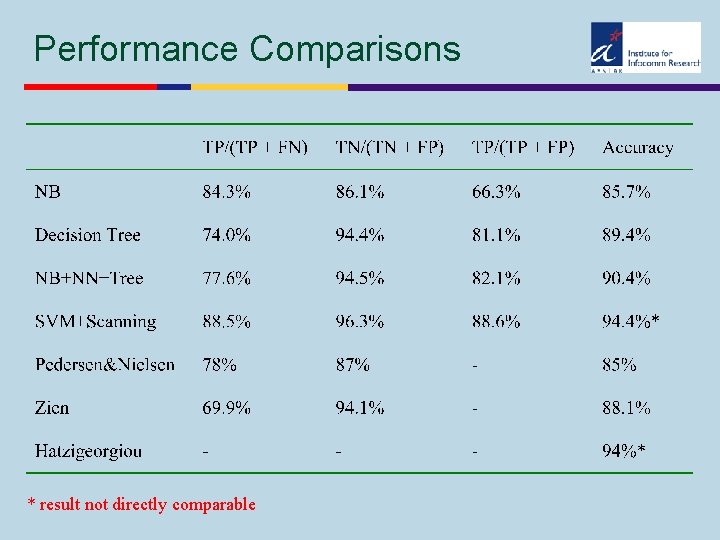
Performance Comparisons * result not directly comparable
![Technique Comparisons • Pedersen&Nielsen [ISMB’ 97] • Our approach • • Neural network No Technique Comparisons • Pedersen&Nielsen [ISMB’ 97] • Our approach • • Neural network No](http://slidetodoc.com/presentation_image/654466690ccd1c8962c1331731a931f5/image-36.jpg)
Technique Comparisons • Pedersen&Nielsen [ISMB’ 97] • Our approach • • Neural network No explicit features • Zien [Bioinformatics’ 00] • SVM+kernel engineering • No explicit features • Hatzigeorgiou [Bioinformatics’ 02] • • • Multiple neural networks Scanning rule No explicit features • • Explicit feature generation Explicit feature selection Use any machine learning method w/o any form of complicated tuning Scanning rule is optional

Acknowledgements • • A. G. Pedersen H. Nielsen Roland Yap Fanfan Zeng
- Slides: 37