Introduction to Bioinformatics Swiss Institute of Bioinformatics Institut

Introduction to Bioinformatics Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

SIB and EMBnet Bioinformatics resources for biomedical scientists Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

The Swiss Institute of Bioinformatics n n n Founded in March 1998 Collaborative structure Lausanne - Geneva - Basel Groups at ISREC, Ludwig Institute, Unil, HUG, Uni. Ge, recently Uni. Bas and soon EPFL. Several roles: teaching, services, research Currently: ~ 130 employees Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

Projects at SIB n Databases n n n Softwares n n Melanie, Deep View, proteomic tools, ESTScan, pftools, Java applets Services n n n SWISS-PROT, PROSITE, EPD, World-2 DPAGE, SWISS-MODEL Tr. EST, Tr. GEN (predicted proteins), tromer (transcriptome) Web servers Ex. PASy, EMBnet Teaching and helpdesk Research n Mostly sequence and expression analysis, 3 D structure, and proteomic Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

Teaching n n n DEA (master degree) in Bioinformatics: 1 year full time, first diploma common to Unige and Unil. EMBnet courses: 2 x 1 week per year in Lausanne, plus 1 week in Basel starting in 2003 (this course!). Pregrade courses in Geneva, Fribourg and Lausanne Universities Other courses at CHUV and EPFL Courses in other countries: Colombia, Cambodia, Peru, … Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

Research n n n New algorithms (faster alignments…) New technology (GRID or cluster computing) New tools (protein analysis, microarrays, confocal microscopy) New databases (microarrays, transcriptome, proteome) Collaborations with lab researchers! Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

Three levels of services n Simple web access to softwares and databases n n n Command-line access with a local Unix account n n n More powerful (automation) and secure Requires to understand Unix system and frequent practice Collaboration with SIB n n n Easy to use for basic occasional research with few sequences Potentially insecure Access to experts in the field (help desk) For projects requiring huge programming or special hardware resources Help desk n helpdesk@mail. ch. embnet. org or http: //www. expasy. org/contact. html Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

SIB’s important sites n Home n n Ex. PASy - Expert Protein Analysis System n n hits. isb-sib. ch EMBnet Switzerland n n www. expasy. org Hits database and tools n n www. isb-sib. ch www. ch. embnet. org Geneva Bioinformatics n www. genebio. ch Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

SIB home Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09
Expert Protein Analysis System Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09
Swiss node http: //www. ch. embnet. org Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

EMBnet organisation n European in 1988, now world-wide spread n n Role n n n 32 country nodes, 8 special nodes. Training, education (EMBER) Software development (EMBOSS, SRS) Computing resources (databases, websites, services) Helpdesk and technical support Publications (EMBnet. news, Briefings in Bioinformatics) Access: www. embnet. org n Each node with “www. xx. embnet. org” where xx is the country code (e. g. , ch for Switzerland) Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

EMBnet home Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

European Molecular Biology Open Software Suite n n Free Open Source (for most Unix plateforms) GCG successor (compatible with GCG file format) More than 150 programs (ver. 2. 7. 1) Easy to install locally n n n Interfaces n n but no interface, requires local databases Unix command-line only Jemboss, www 2 gcg, w 2 h, wemboss… (with account) Pise, EMBOSS-GUI, SRSWWW (no account) Staden, Kaptain, Co. Li. Mate, Jemboss (local) Access: www. emboss. org Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

Other important sites n Ex. PASy - Expert Protein Analysis System n n EBI - European Bioinformatics Institute n n www. ebi. ac. uk NCBI - National Center for Biotechnology Information n n www. expasy. org www. ncbi. nlm. nih. gov Sanger - The Sanger Institute n www. sanger. ac. uk Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

Bioinformatics: definition n Every application of computer science to biology n n Sequence analysis, images analysis, sample management, population modelling, … Analysis of data coming from large-scale biological projects n Genomes, transcriptomes, proteomes, metabolomes, etc… Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

The new biology n Traditional biology n n n Small team working on a specialized topic Well defined experiment to answer precise questions New « high-throughput » biology n n Large international teams using cutting edge technology defining the project Results are given raw to the scientific community without any underlying hypothesis Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

Example of « high-throughput » n n n n n Complete genome sequencing Large-scale sampling of the transcriptome (EST) Simultaneous expression analysis of thousands of genes (DNA microarrays, SAGE) Large-scale sampling of the proteome Protein-protein analysis large-scale 2 -hybrid (yeast, worm) Large-scale 3 D structure production (yeast) Metabolism modelling Simulations Biodiversity Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

Role of bioinformatics n n Control and management of the data Analysis of primary data e. g. n n n Base calling from chromatograms Mass spectra analysis DNA microarrays images analysis Statistics Database storage and access Results analysis in a biological context Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

First information: a sequence ? n Nucleotide n n n RNA (or c. DNA) Genomic (intron-exon) Complete or incomplete? n n n m. RNA with 5’ and 3’ UTR regions Entire chromosome Protein n n Pre/Pro or functional protein? Function prediction Post-translational modifications? Holy Grail: 3 D structure? Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

Genomes in numbers n Sizes: n n n virus: 103 to 105 nt bacteria: 105 to 107 nt yeast: 1. 35 x 107 nt mammals: 108 to 1010 nt plants: 1010 to 1011 nt n Gene number: n n n virus: 3 to 100 bacteria: ~ 1000 yeast: ~ 7000 mammals: ~ 30’ 000 Plants: 30’ 000 -50’ 000? Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

Sequencing projects n « small » genomes (<107): bacteria, virus n n « large » genomes (107 -1010) eucaryotes n n n Many already sequenced (industry excluded) More than 100 microbial genomes already in the public domain More to come! (one new every two weeks…) 15 finished (S. cerevisiae, S. Pombe, E. cuniculi, G. theta, C. elegans, D. melanogaster, A. gambiae, P. falciparum, P. yoelii, D. rerio, F. rubripes, A. thaliana, O. sativa (2 x), M. musculus, Homo sapiens) Many more to come: rat, pig, cow, maize (and other plants), insects, fishes, many pathogenic parasites (Leishmania…) EST sequencing n Partial m. RNA sequences ~15 x 106 sequences in the public domain Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09
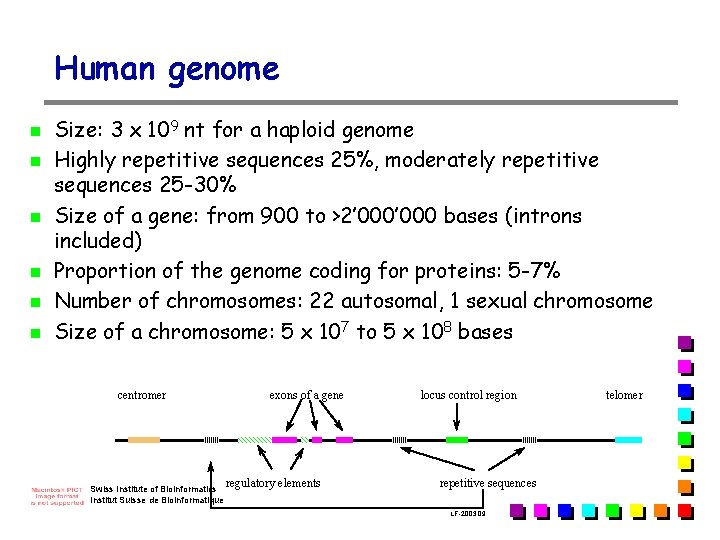
Human genome n n n Size: 3 x 109 nt for a haploid genome Highly repetitive sequences 25%, moderately repetitive sequences 25 -30% Size of a gene: from 900 to >2’ 000 bases (introns included) Proportion of the genome coding for proteins: 5 -7% Number of chromosomes: 22 autosomal, 1 sexual chromosome Size of a chromosome: 5 x 107 to 5 x 108 bases centromer Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique exons of a gene regulatory elements locus control region repetitive sequences LF-2003. 09 telomer

How to sequence the human genome? n Consortium « international » approach: n n Generate genetic maps (meiotic recombination) and pseudogenetic maps (chromosome hybrids) for indicator sequences Generate a physical map based on large clones (BAC or PAC) Sequence enough large clones to cover the genome « commercial » approach (Celera): n n n Generate random libraries of fixed length genomic clones (2 kb and 10 kb) Sequence both ends of enough clones to obtain a 10 x coverage Use computer techniques to reconstitute the chromosomal sequences, check with the public project physical map Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09
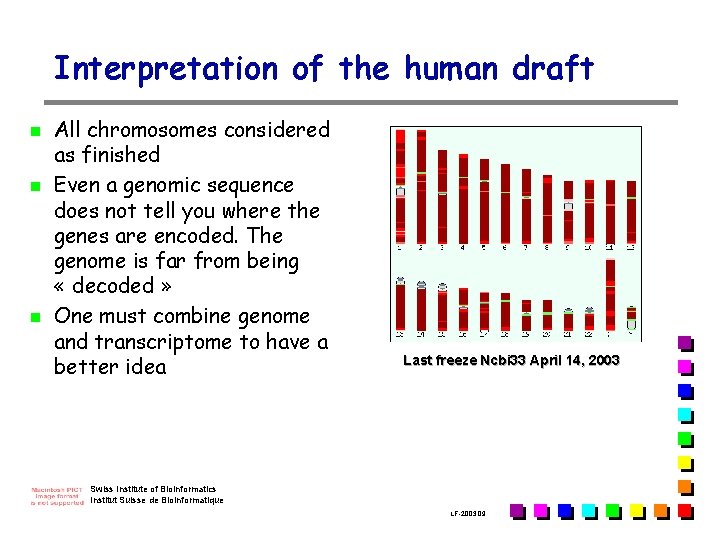
Interpretation of the human draft n n n All chromosomes considered as finished Even a genomic sequence does not tell you where the genes are encoded. The genome is far from being « decoded » One must combine genome and transcriptome to have a better idea Last freeze Ncbi 33 April 14, 2003 Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

The transcriptome n n n The set of all functional RNAs (t. RNA, r. RNA, m. RNA etc…) that can potentially be transcribed from the genome The documentation of the localization (cell type) and conditions under which these RNAs are expressed The documentation of the biological function(s) of each RNA species Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

Public draft transcriptome n Information about the expression specificity and the function of m. RNAs n n n « full » c. DNA sequences of know function « full » c. DNA sequences, but « anonymous » (e. g. KIAA or DKFZ collections) EST sequences n n n c. DNA libraries derived from many different tissues Rapid random sequencing of the ends of all clones ORESTES sequences Growing set of expression data (microarrays, SAGE etc…) Increasing evidences for multiple alternative splicing and polyadenylation Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09
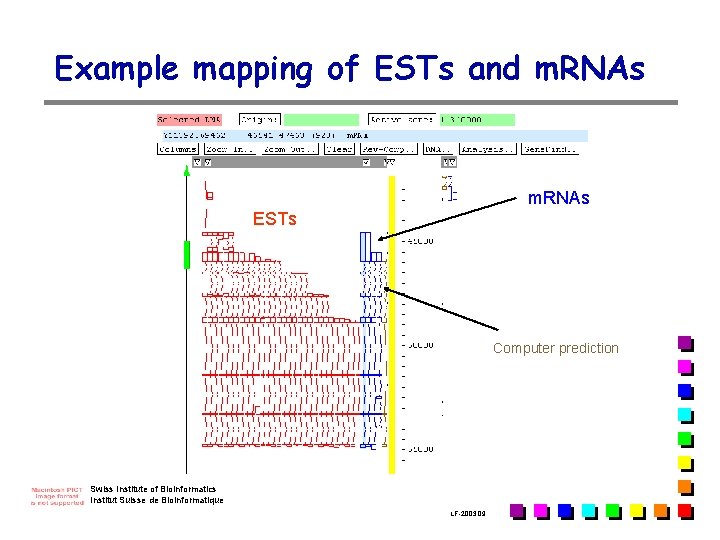
Example mapping of ESTs and m. RNAs ESTs Computer prediction Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

The proteome n n n Set of proteins present in a particular cell type under particular conditions Set of proteins potentially expressed from the genome Information about the specific expression and function of the proteins Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

Information on the proteome n Separation of a complex mixture of proteins n n n Individual characterisation of proteins n n n 2 D PAGE (IEF + SDS PAGE) Capillary chromatography Tryptic peptides signature (MS) Sequencing by chemistry or MS/MS All post-translational modifications (PTMs) ! Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

Tridimentional structures n Methods to determine structures n n n Data format n n X-ray cristallography NMR Atoms coordinates (except H) in a cartesian space Databases n n For proteins and nucleic acids (RSCB, was PDB) Independent databases for sugars and small organic molecules Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

Visualisation of the structures n Secondary structure elements n n Alpha helices, beta sheets, other Softwares n n Various representations (atoms, bonds, secondary…) Big choice of commercial and free software (e. g. , Deep. View) Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

Sequence information, and so what ? n How to store and organise ? n n Databases (next lecture) How to access, search, compare ? n n n n Pairwise alignments, dot plots (Tuesday) BLAST searches in db (Tuesday) Patterns, PSI-BLAST, Profiles and HMMs (Wednesday) Gene prediction (Wednesday) EST clustering (Thursday) Multiple Alignments (Thursday) Protein function prediction (Friday) Users problems (Friday) Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09

Thanks you Swiss Institute of Bioinformatics Institut Suisse de Bioinformatique LF-2003. 09
- Slides: 35