From Datamining to Bioinformatics Limsoon Wong Laboratories for

From Datamining to Bioinformatics Limsoon Wong Laboratories for Information Technology Singapore

What is Bioinformatics?

Themes of Bioinformatics = Data Mgmt + Knowledge Discovery Data Mgmt = Integration + Transformation + Cleansing Knowledge Discovery = Statistics + Algorithms + Databases

Benefits of Bioinformatics To the patient: Better drug, better treatment To the pharma: Save time, save cost, make more $ To the scientist: Better science
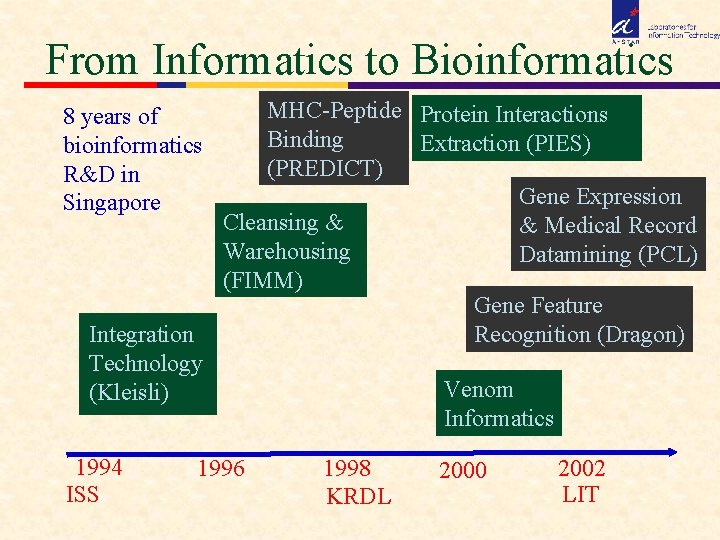
From Informatics to Bioinformatics 8 years of bioinformatics R&D in Singapore Integration Technology (Kleisli) 1994 ISS MHC-Peptide Protein Interactions Binding Extraction (PIES) (PREDICT) Gene Expression Cleansing & & Medical Record Warehousing Datamining (PCL) (FIMM) Gene Feature Recognition (Dragon) 1996 Venom Informatics 1998 KRDL 2000 2002 LIT

Quick Samplings

Epitope Prediction TRAP-559 AA MNHLGNVKYLVIVFLIFFDLFLVNGRDVQNNIVDEIKYSE EVCNDQVDLYLLMDCSGSIRRHNWVNHAVPLAMKLIQQLN LNDNAIHLYVNVFSNNAKEIIRLHSDASKNKEKALIIIRS LLSTNLPYGRTNLTDALLQVRKHLNDRINRENANQLVVIL TDGIPDSIQDSLKESRKLSDRGVKIAVFGIGQGINVAFNR FLVGCHPSDGKCNLYADSAWENVKNVIGPFMKAVCVEVEK TASCGVWDEWSPCSVTCGKGTRSRKREILHEGCTSEIQEQ CEEERCPPKWEPLDVPDEPEDDQPRPRGDNSSVQKPEENI IDNNPQEPSPNPEEGKDENPNGFDLDENPENPPNPDIPEQ KPNIPEDSEKEVPSDVPKNPEDDREENFDIPKKPENKHDN QNNLPNDKSDRNIPYSPLPPKVLDNERKQSDPQSQDNNGN RHVPNSEDRETRPHGRNNENRSYNRKYNDTPKHPEREEHE KPDNNKKKGESDNKYKIAGGLALLACAGLAYKFVVP GAATPYAGEPAPFDETLGEEDKDLDEPEQFRLPEENEWN

Epitope Prediction Results w Prediction by our ANN model for HLA-A 11 w w w 29 predictions 22 epitopes 76% specificity w Prediction by BIMAS matrix for HLA-A*1101 Number of experimental binders 19 (52. 8%) 5 (13. 9%) 12 (33. 3%) 1 66 100 Rank by BIMAS
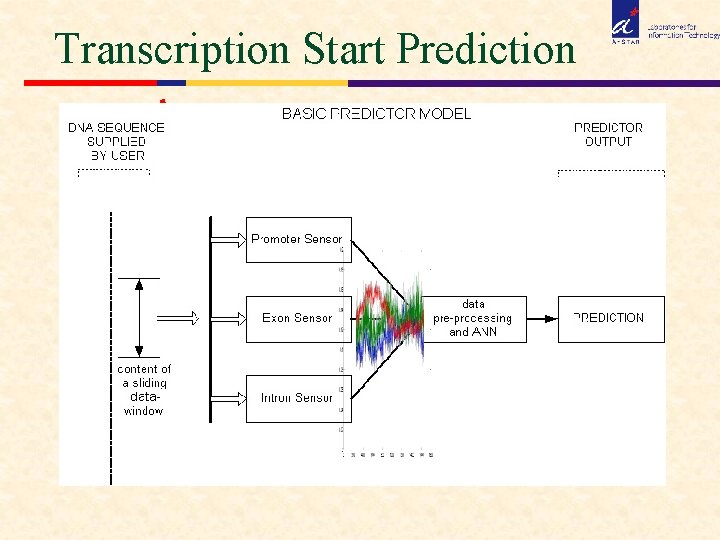
Transcription Start Prediction
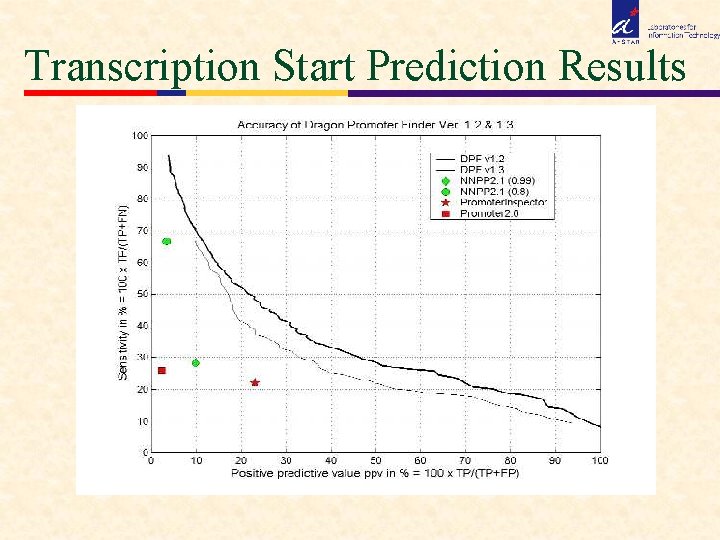
Transcription Start Prediction Results

Medical Record Analysis w Looking for patterns that are w w valid novel useful understandable
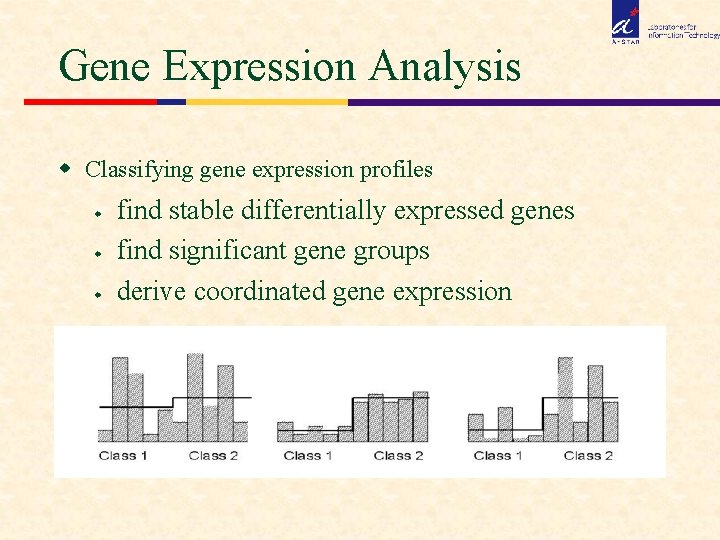
Gene Expression Analysis w Classifying gene expression profiles w w w find stable differentially expressed genes find significant gene groups derive coordinated gene expression
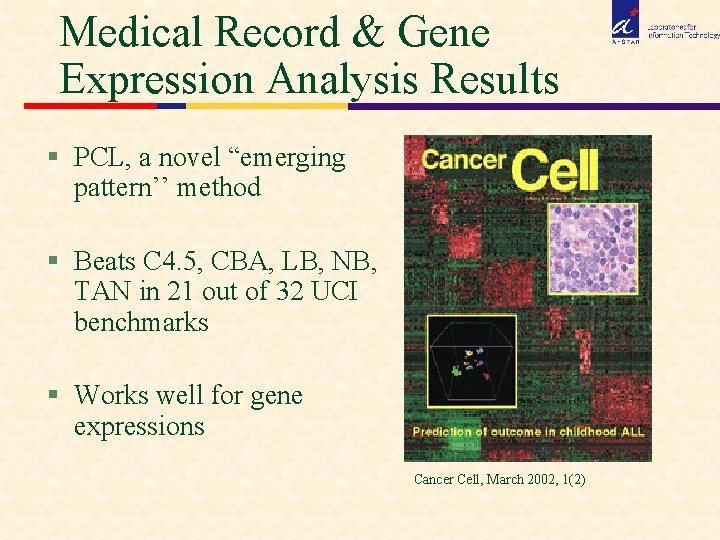
Medical Record & Gene Expression Analysis Results § PCL, a novel “emerging pattern’’ method § Beats C 4. 5, CBA, LB, NB, TAN in 21 out of 32 UCI benchmarks § Works well for gene expressions Cancer Cell, March 2002, 1(2)

Behind the Scene w Vladimir Bajic w Vladimir Brusic w Jinyan Li w See-Kiong Ng w Limsoon Wong w Louxin Zhang w Allen Chong w Judice Koh w SPT Krishnan w Huiqing Liu w Seng Hong Seah w Soon Heng Tan w Guanglan Zhang w Zhuo Zhang and many more: students, folks from genetic. Xchange, Molecular. Connections, and other collaborators….

Questions?

A More Detailed Account
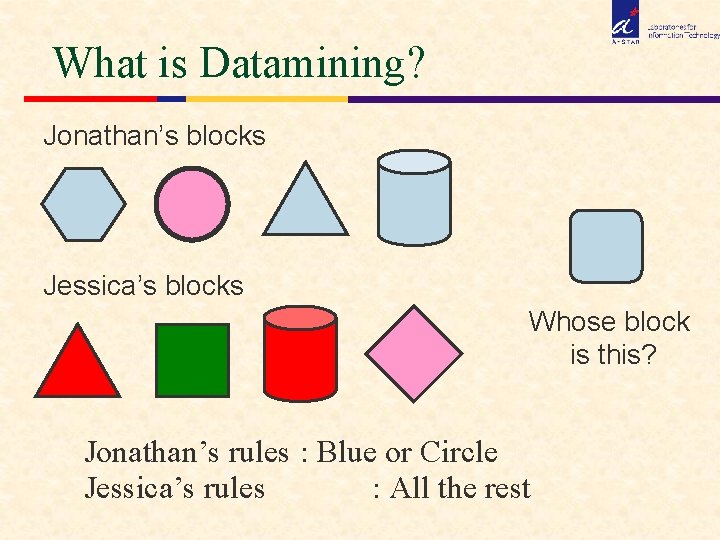
What is Datamining? Jonathan’s blocks Jessica’s blocks Whose block is this? Jonathan’s rules : Blue or Circle Jessica’s rules : All the rest

What is Datamining? Question: Can you explain how?

The Steps of Data Mining § Training data gathering § Signal generation § k-grams, colour, texture, domain know-how, . . . § Signal selection § Entropy, 2, CFS, t-test, domain know-how. . . § Signal integration § SVM, ANN, PCL, CART, C 4. 5, k. NN, . . .
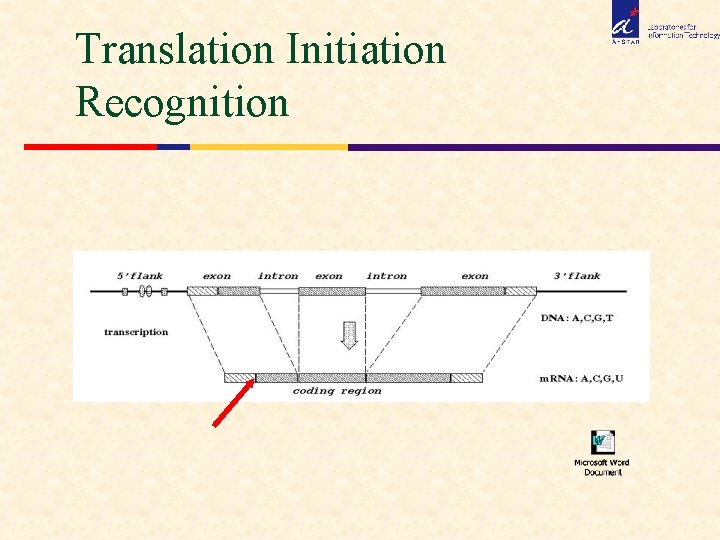
Translation Initiation Recognition

A Sample c. DNA 299 HSU 27655. 1 CAT U 27655 Homo sapiens CGTGTGTGCAGCAGCCTGCAGCTGCCCCAAGCCATGGCTGAACACTGACTCCCAGCTGTG CCCAGGGCTTCAAAGACTTCTCAGCTTCGAGCATGGCTTTTGGCTGTCAGGGCAGCTGTA GGAGGCAGATGAGAAGAGGGAGATGGCCTTGGAGGAAGGGGCCTGGTGCCGAGGA CCTCTCCTGGCCAGGAGCTTCCTCCAGGACAAGACCTTCCACCCAACAAGGACTCCCCT. . . . . . i. EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE What makes the second ATG the translation initiation site? 80 160 240

Signal Generation § K-grams (ie. , k consecutive letters) l l K = 1, 2, 3, 4, 5, … Window size vs. fixed position Up-stream, downstream vs. any where in window In-frame vs. any frame

Too Many Signals § For each value of k, there are 4 k * 3 * 2 k-grams § If we use k = 1, 2, 3, 4, 5, we have 4 + 24 + 96 + 384 + 1536 + 6144 = 8188 features! § This is too many for most machine learning algorithms

Signal Selection (Basic Idea) § Choose a signal w/ low intra-class distance § Choose a signal w/ high inter-class distance § Which of the following 3 signals is good?

Signal Selection (eg. , t-statistics)

Signal Selection (eg. , MIT-correlation)

Signal Selection (eg. , 2)

Signal Selection (eg. , CFS) § Instead of scoring individual signals, how about scoring a group of signals as a whole? § CFS l A good group contains signals that are highly correlated with the class, and yet uncorrelated with each other § Homework: find a formula that captures the key idea of CFS above

Sample k-grams Selected Kozak consensus Leaky scanning § Position – 3 § in-frame upstream ATG § in-frame downstream l l Stop codon TAA, TAG, TGA, CTG, GAC, GAG, and GCC Codon bias

Signal Integration § k. NN Given a test sample, find the k training samples that are most similar to it. Let the majority class win. § SVM Given a group of training samples from two classes, determine a separating plane that maximises the margin of error. § Naïve Bayes, ANN, C 4. 5, . . .
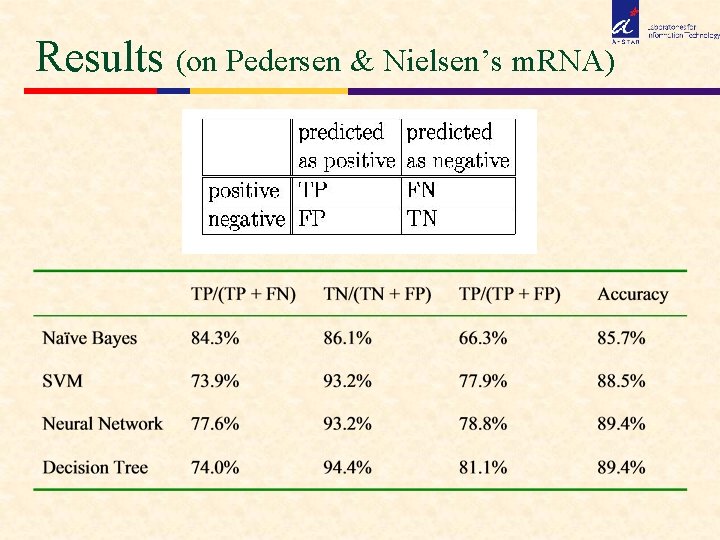
Results (on Pedersen & Nielsen’s m. RNA)

Acknowledgements § § Roland Yap Zeng Fanfan A. G. Pedersen H. Nielsen

Questions?

Common Mistakes

Self-fulfilling Oracle § Consider this scenario l l l Given classes C 1 and C 2 w/ explicit signals Use 2 to C 1 and C 2 to select signals s 1, s 2, s 3 Run 3 -fold x-validation on C 1 and C 2 using s 1, s 2, s 3 and get accuracy of 90% § Is the accuracy really 90%? § What can be wrong with this?

Phil Long’s Experiment § Let there be classes C 1 and C 2 w/ 100000 features having randomly generated values § Use 2 to select 20 features § Run k-fold x-validation on C 1 and C 2 w/ these 20 features § Expect: 50% accuracy § Get: 90% accuracy! § Lesson: choose features at each fold

Apples vs Oranges § Consider this scenario: l l Fanfan reported 89% accuracy on his TIS prediction method Hatzigeorgiou reported 94% accuracy on her TIS prediction method § So Hatzigeorgiou’s method is better § What is wrong with this conclusion?

Apples vs Oranges § Differences in datasets used: l l Fanfan’s expt used Pedersen’s dataset Hatzigeorgiou’s used her own dataset § Differences in counting: l l Fanfan’s expt was on a per ATG basis Hatzigeorgiou’s expt used the scanning rule and thus was on a per c. DNA basis § When Fanfan ran the same dataset and count the same way as Hatzigeorgiou, got 94% also!

Questions?
- Slides: 39