Chapter 12 Genomics Fig 12 1 Genomics the












- Slides: 41

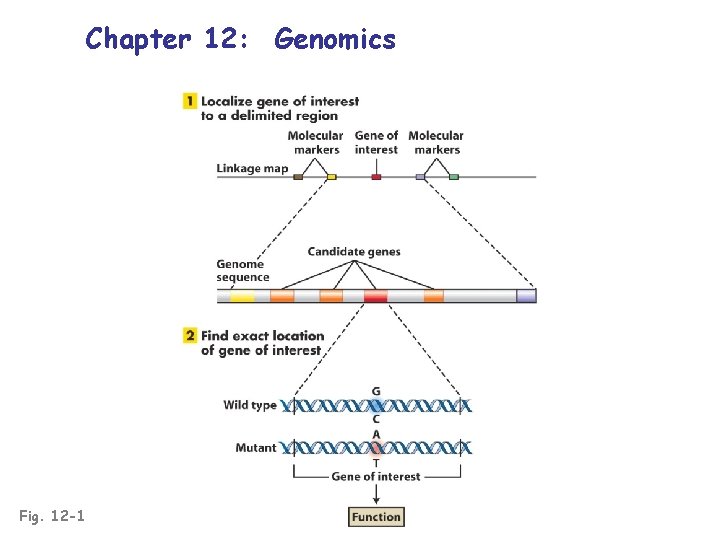
Chapter 12: Genomics Fig. 12 -1

Genomics: the study of whole-genome structure, organization, and function Structural genomics: the physical genome; whole genome mapping Functional genomics: the proteome, expression patterns, networks

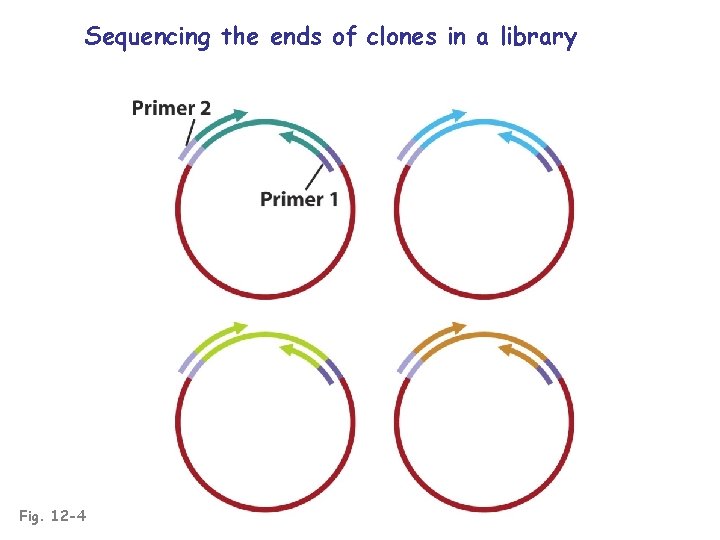
Creating a physical map of the genome • Create a comprehensive genomic library (use a vector that incorporates huge fragments) • Order the clones by identifying overlapping groups (e. g. , sequencing ends to determine “contigs”) • Sequence each contig • Identify genes and chromosomal rearrangements within each contig (correlates the genetic and physical maps)

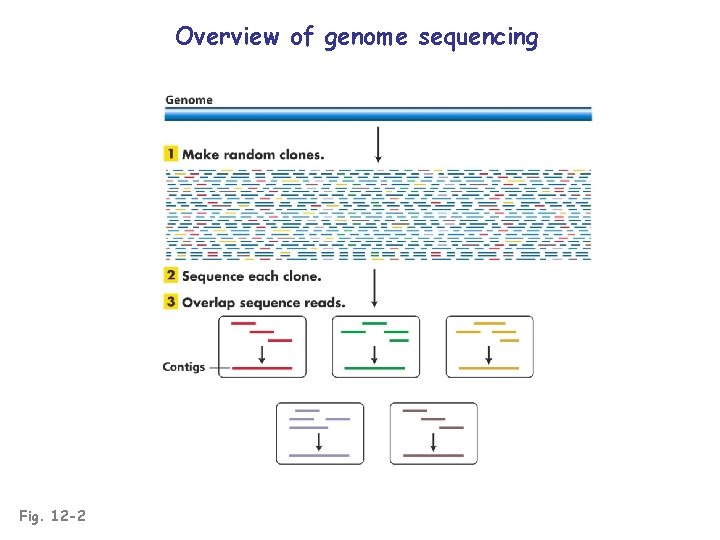
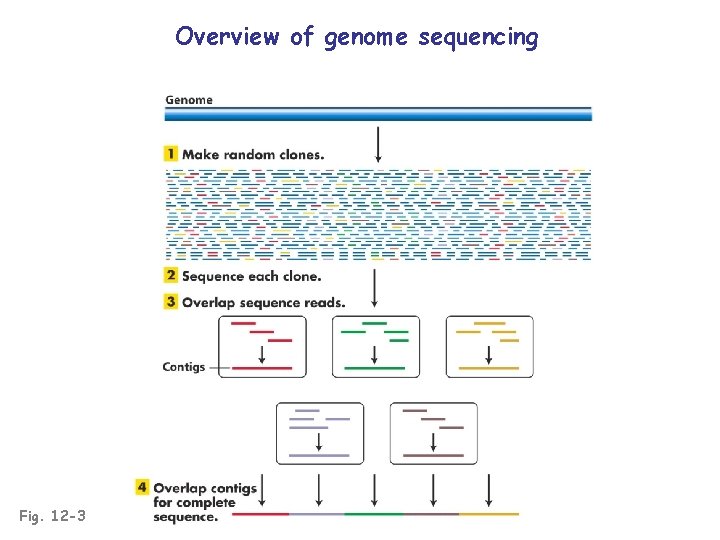
Overview of genome sequencing Fig. 12 -2

Sequencing the ends of clones in a library Fig. 12 -4

Overview of genome sequencing Fig. 12 -2

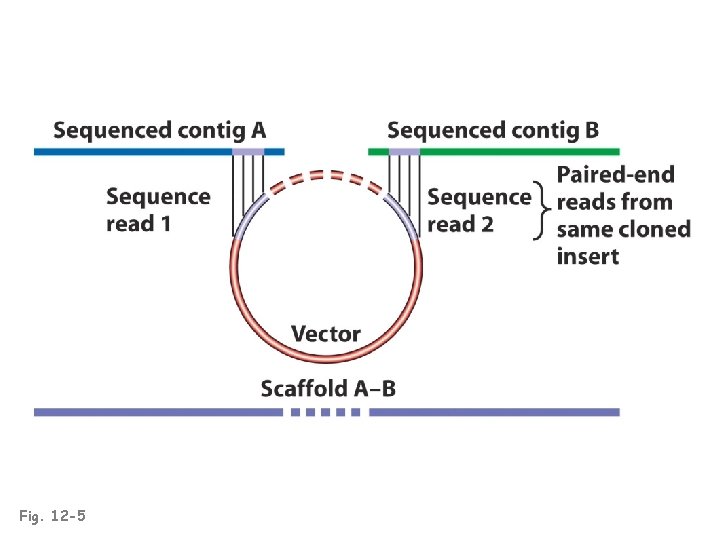
Fig. 12 -5

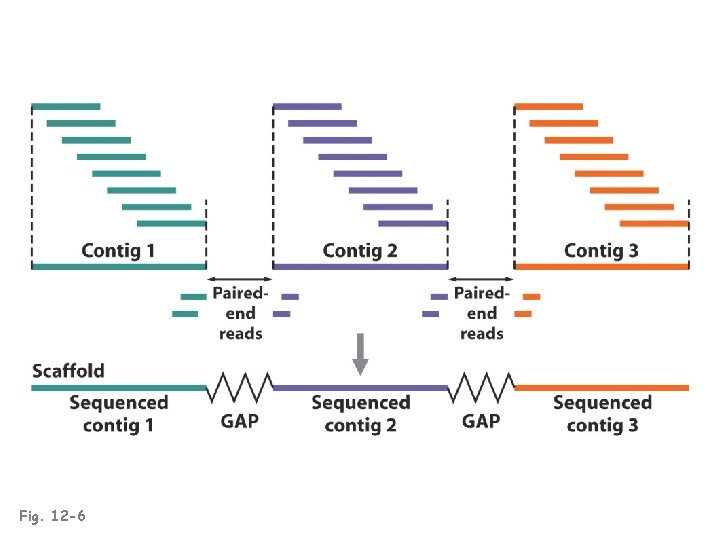
Fig. 12 -6

Overview of genome sequencing Fig. 12 -3

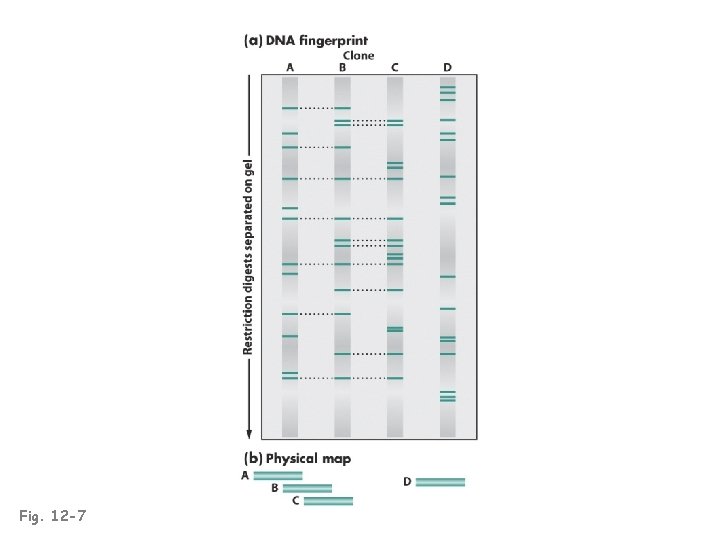
Fig. 12 -7

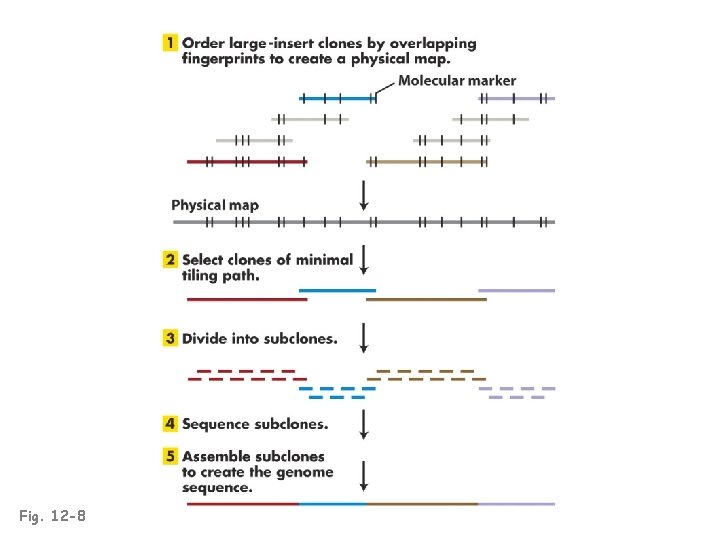
Fig. 12 -8

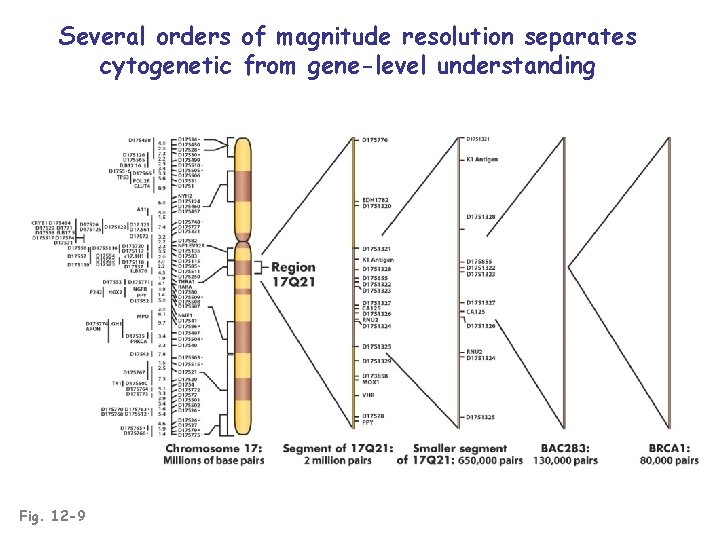
Several orders of magnitude resolution separates cytogenetic from gene-level understanding Fig. 12 -9

Creating a high-resolution genetic map of the genome requires many “markers” • Classic mutations and allelic variations (too few) • Molecular polymorphisms; selectively neutral DNA sequence variations are common in genomes Example: Restriction Fragment Length Polymorphisms (RFLP markers)

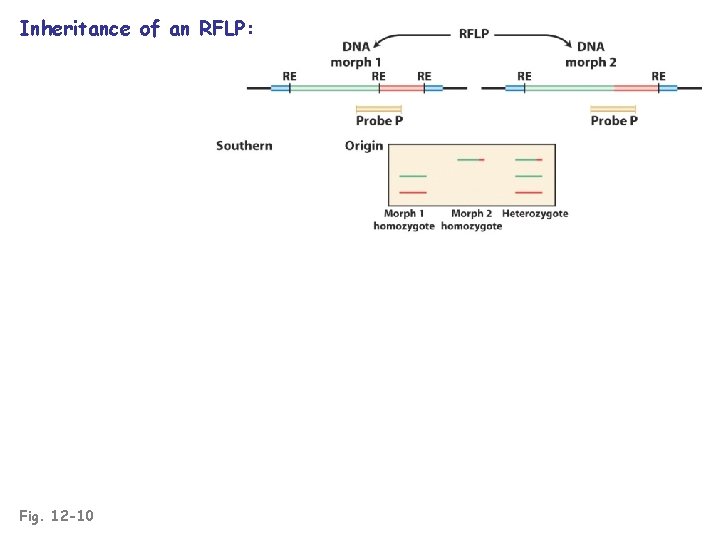
Inheritance of an RFLP: Fig. 12 -10

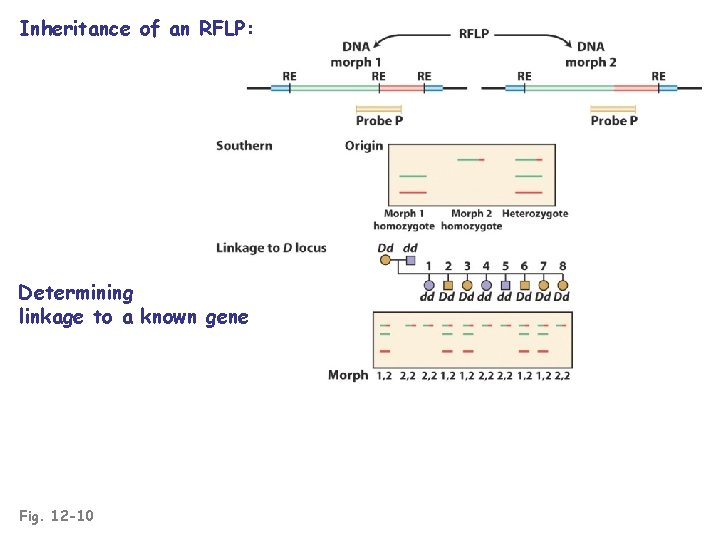
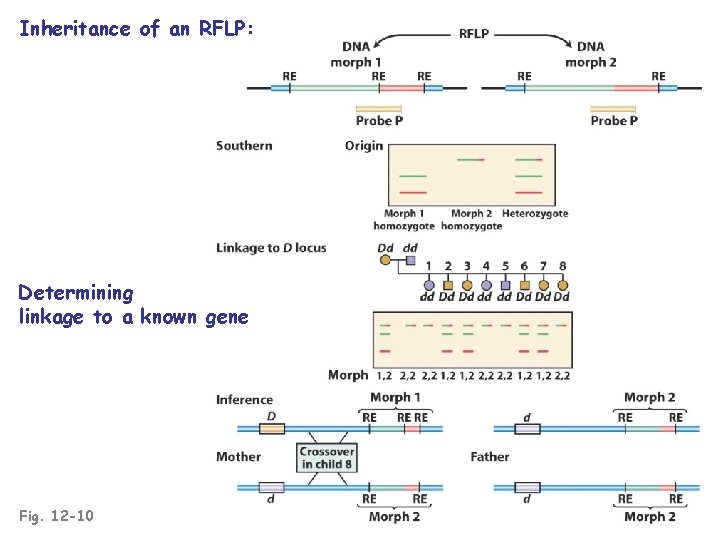
Inheritance of an RFLP: Determining linkage to a known gene Fig. 12 -10

Inheritance of an RFLP: Determining linkage to a known gene Fig. 12 -10

Linkage analysis of a gene and VNTR markers Fig. 12 -11

Creating a high-resolution genetic map of the genome requires many “markers” • Classic mutations and allelic variations • Molecular polymorphisms; selectively neutral DNA sequence variations are common in genomes Example: Restriction Fragment Length Polymorphisms (RFLP markers) Example: Simple Sequence Length Polymorphisms (SSLP markers)

SSLP: Simple sequence length polymorphism • VNTR repeat clusters (minisatellite markers) • dinucleotide repeats (microsatellite markers) VNTRs can be detected by restriction/Southern blot analysis; both detected by PCR using primers for each end of the repeat tract

Variable number tandem repeats (VNTRs) • “minisatellite” DNA • 15 -100 bp units; repeated in 1 -5 kb blocks • expansion/contraction of the block due to meiotic unequal crossingover • crossingover so frequent that each individual has unique pattern (revealed by genomic Southern blot/hybridization analysis)

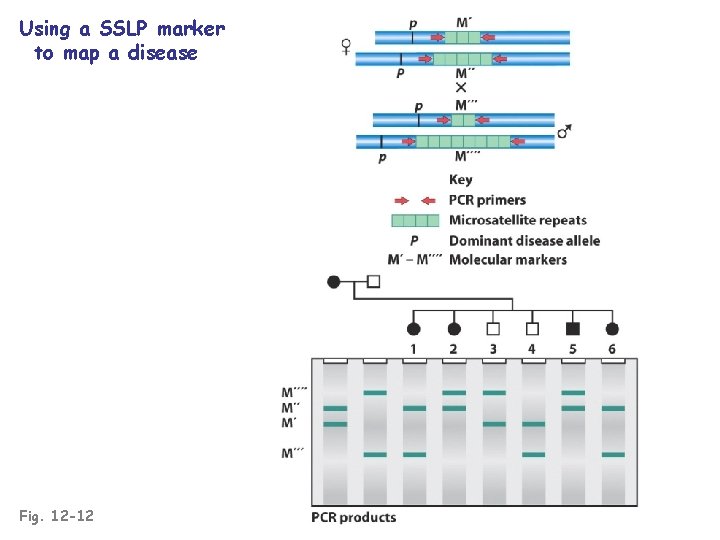
Using a SSLP marker to map a disease Fig. 12 -12

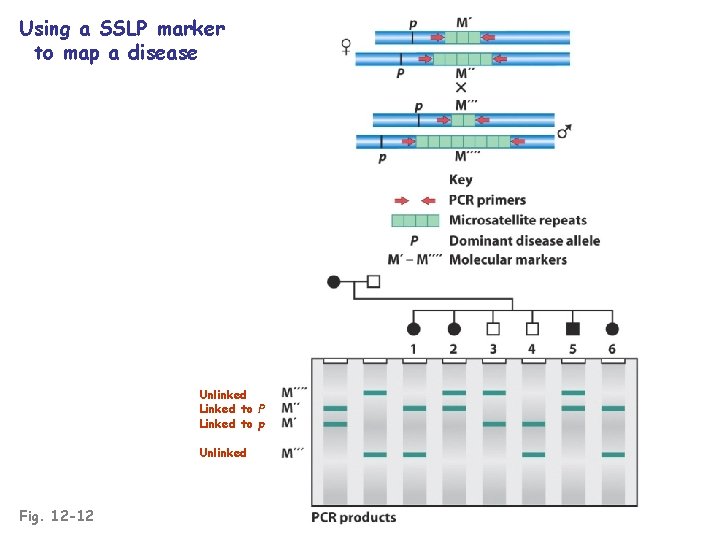
Using a SSLP marker to map a disease Unlinked Linked to P Linked to p Unlinked Fig. 12 -12

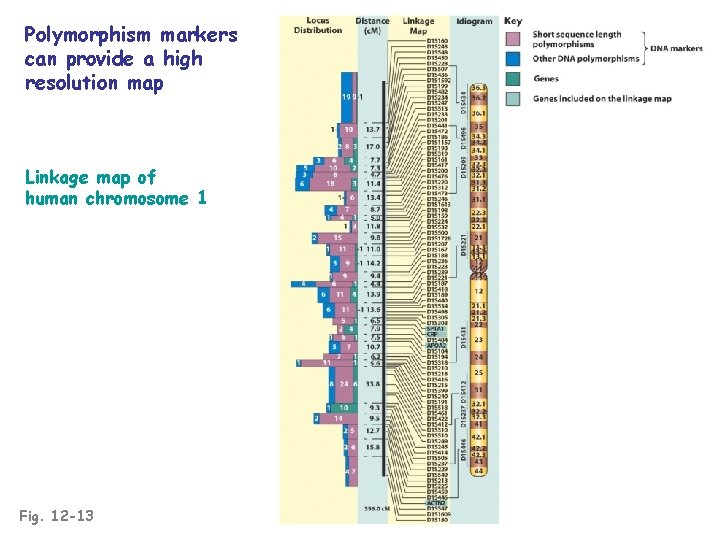
Polymorphism markers can provide a high resolution map Linkage map of human chromosome 1 Fig. 12 -13

High-resolution cytogenetic mapping is based on: • In situ hybridization: hybridization of known sequences directly to chromosome preparations • Rearrangement break mapping • Radiation hybrid mapping

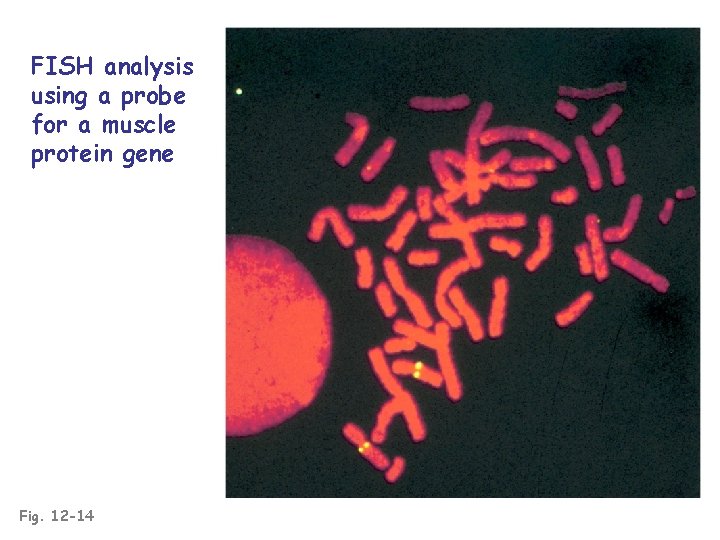
FISH analysis using a probe for a muscle protein gene Fig. 12 -14

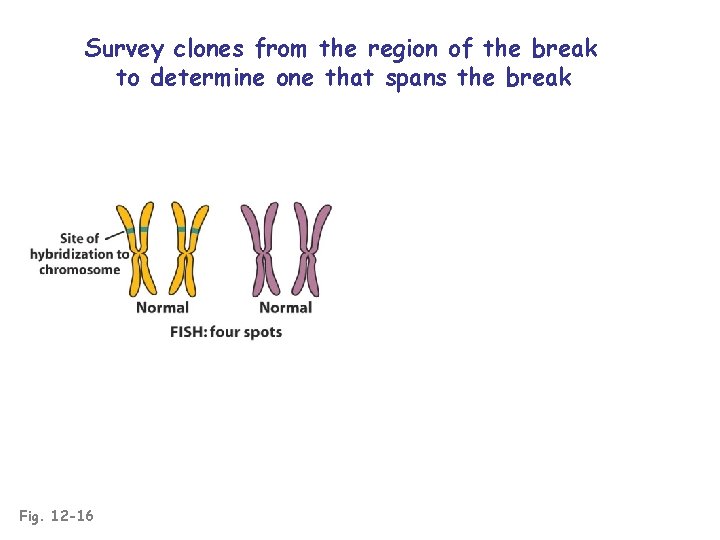
Survey clones from the region of the break to determine one that spans the break Fig. 12 -16

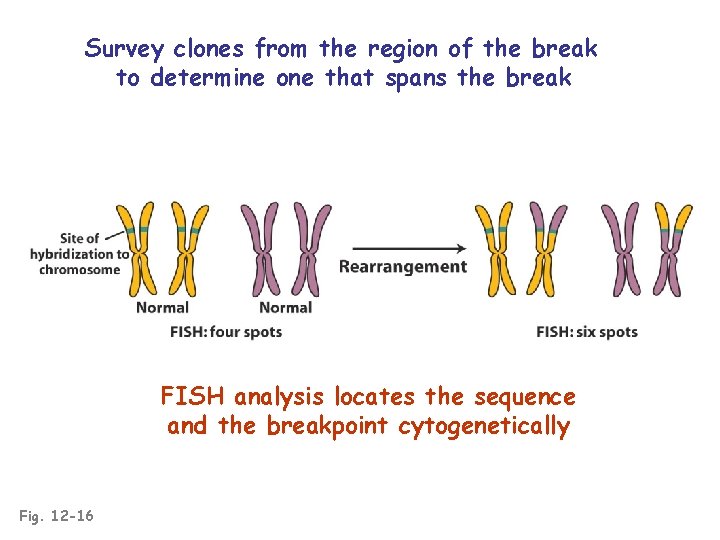
Survey clones from the region of the break to determine one that spans the break FISH analysis locates the sequence and the breakpoint cytogenetically Fig. 12 -16

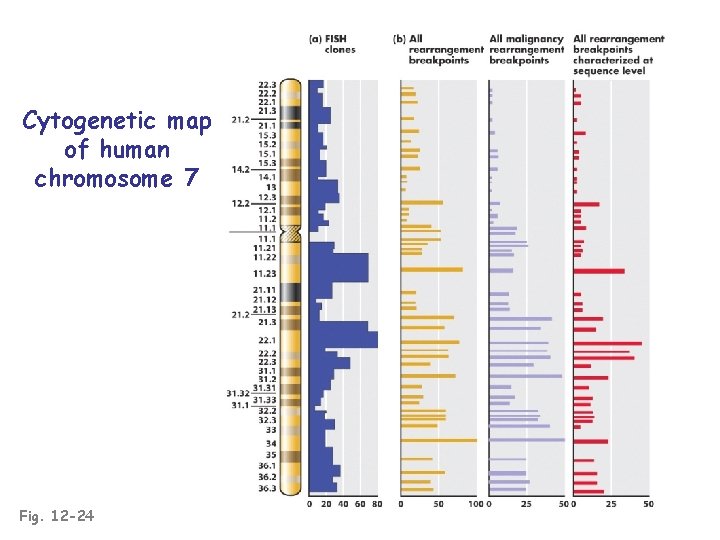
Cytogenetic map of human chromosome 7 Fig. 12 -24

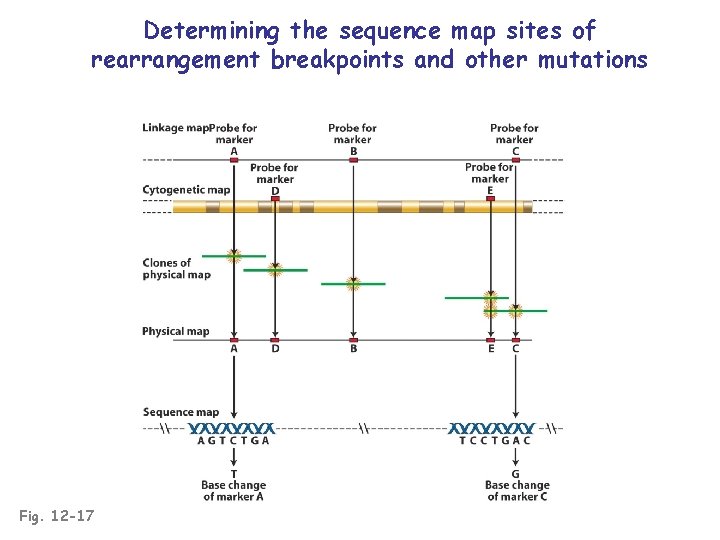
Determining the sequence map sites of rearrangement breakpoints and other mutations Fig. 12 -17

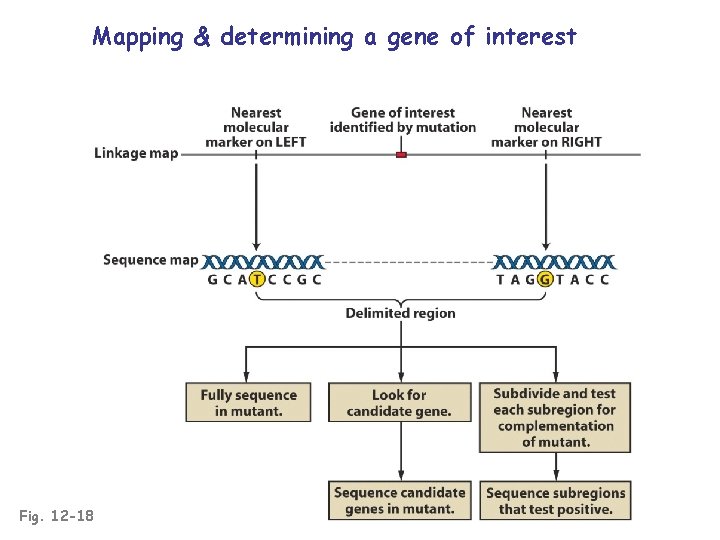
Mapping & determining a gene of interest Fig. 12 -18

Genome sequencing projects • Sequence individual clones and subclones (extensive use of robotics) • Identify overlaps to assemble sequence contigs (extensive use of computer-assisted analysis) • Identify putative genes by identifying open reading frames, consensus sequences and other bioinformatic tools

Once a genomic sequence is obtained, it is subjected to bioinformatic analysis to determine structure and function • Identify apparent ORFs and consensus regulatory sequences to identify potential genes • Identify corresponding c. DNA (and EST) sequences to identify genuine coding regions • Polypeptide similarity analysis (similarity to polypeptides encoded in other genomes)

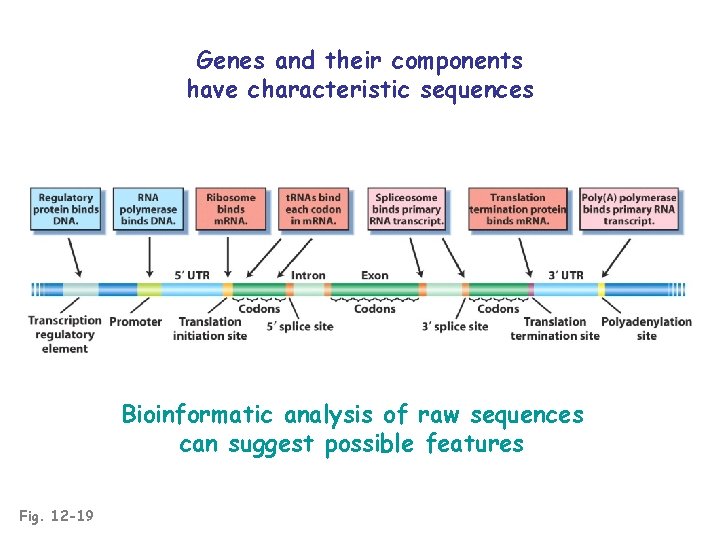
Genes and their components have characteristic sequences Bioinformatic analysis of raw sequences can suggest possible features Fig. 12 -19

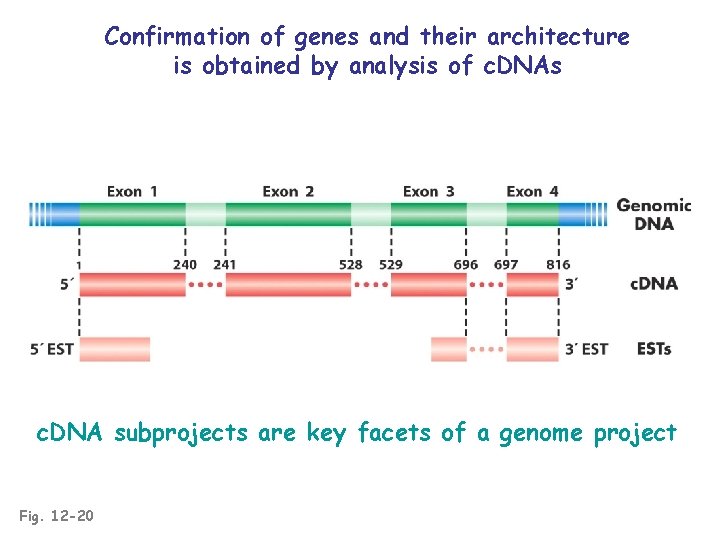
Confirmation of genes and their architecture is obtained by analysis of c. DNAs c. DNA subprojects are key facets of a genome project Fig. 12 -20

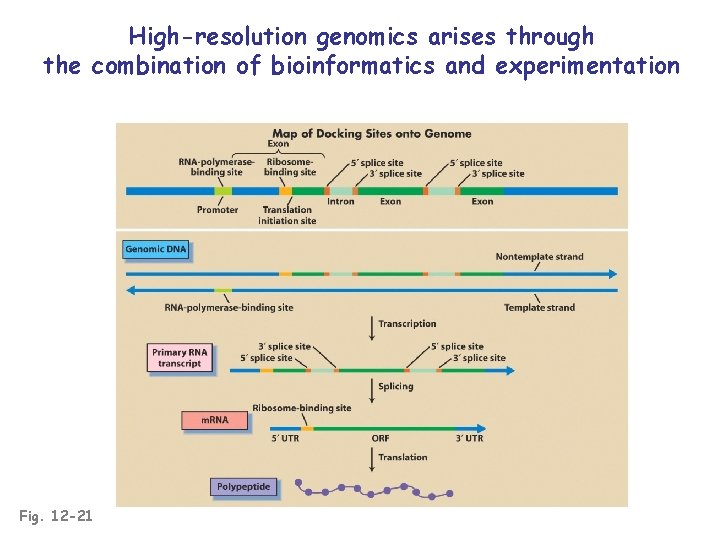
High-resolution genomics arises through the combination of bioinformatics and experimentation Fig. 12 -21

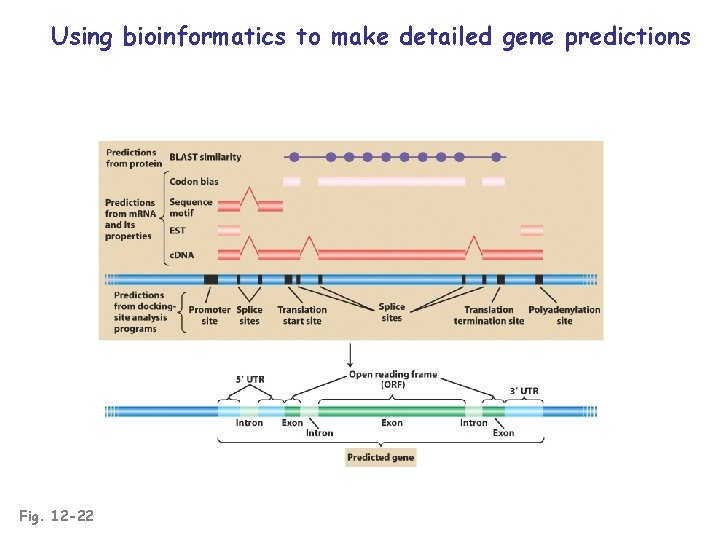
Using bioinformatics to make detailed gene predictions Fig. 12 -22

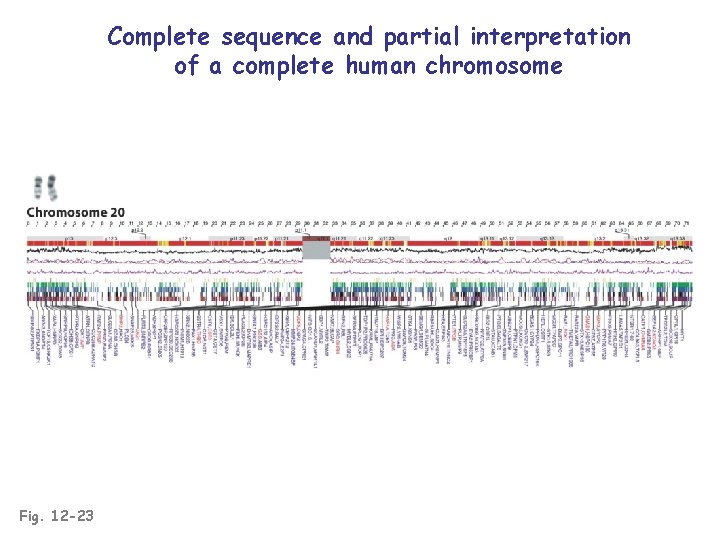
Complete sequence and partial interpretation of a complete human chromosome Fig. 12 -23

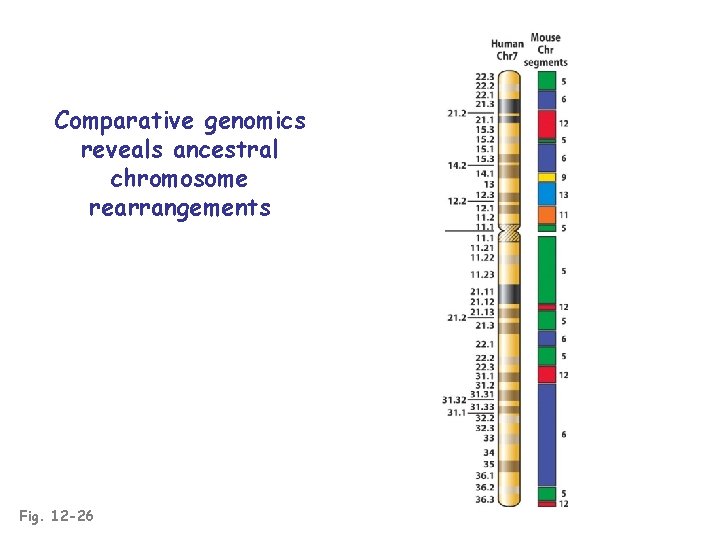
Comparative genomics reveals ancestral chromosome rearrangements Fig. 12 -26

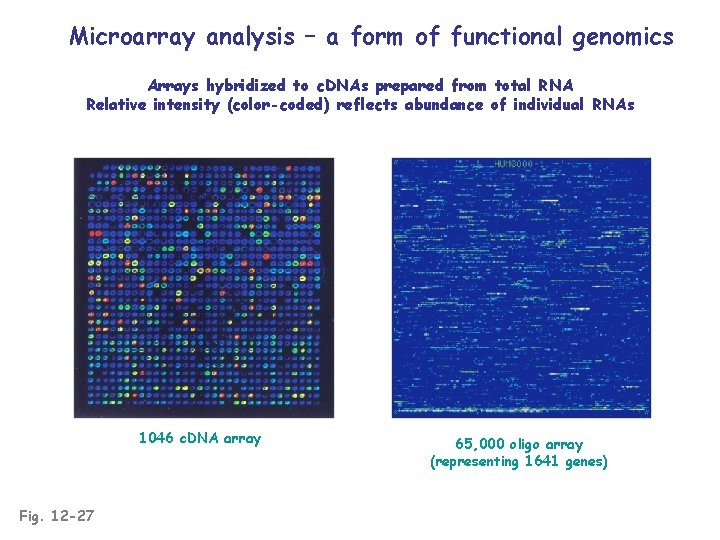
Microarray analysis – a form of functional genomics Arrays hybridized to c. DNAs prepared from total RNA Relative intensity (color-coded) reflects abundance of individual RNAs 1046 c. DNA array Fig. 12 -27 65, 000 oligo array (representing 1641 genes)

Fig. 12 -

Fig. 12 -