Research in EDS Genomics epigenetics metabolomics and the

Research in EDS- Genomics, epigenetics, metabolomics and the use of animal models of disease Roberto Mendoza-Londono MD, FCCMG, FRCPSC Associate Professor of Pediatrics and Molecular Genetics The Hospital for Sick Children and University of Toronto November 7, 2015

Ehlers-Danlos Syndrome �Spectrum of monogenic disorders �Wide range of phenotypic severity �Affecting joints, skin, blood vessels and internal organs to varying degrees �Most forms are caused by defects in one of the fibrillar collagens or of enzymes involved in fibrillar collagen processing �Recent research has identified defects in biosynthesis of other molecules in the extraceullar matrix and molecules involved in intracellular trafficking, secretion and assembly of ECM molecules
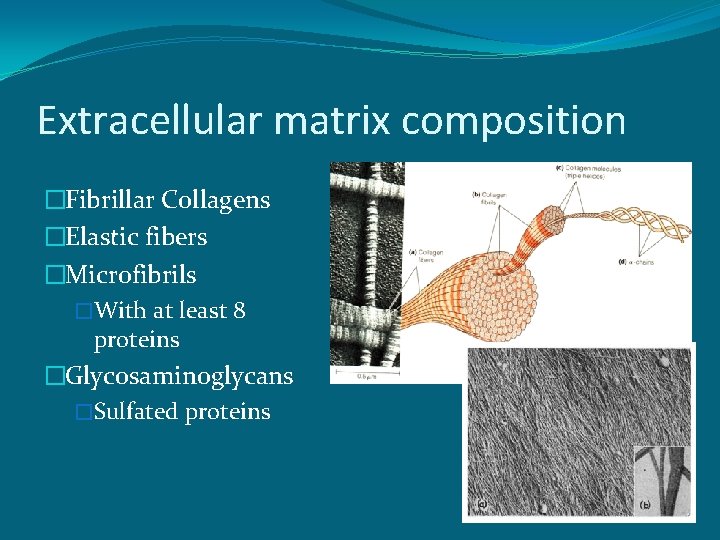
Extracellular matrix composition �Fibrillar Collagens �Elastic fibers �Microfibrils �With at least 8 proteins �Glycosaminoglycans �Sulfated proteins
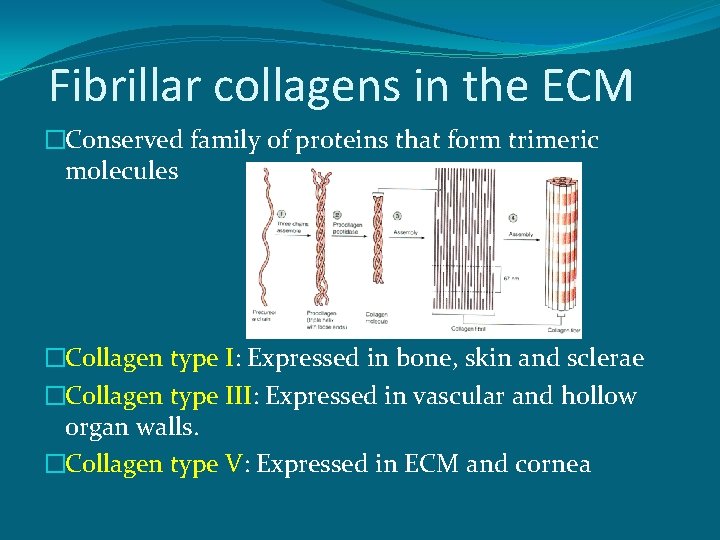
Fibrillar collagens in the ECM �Conserved family of proteins that form trimeric molecules �Collagen type I: Expressed in bone, skin and sclerae �Collagen type III: Expressed in vascular and hollow organ walls. �Collagen type V: Expressed in ECM and cornea

Proteins that modify or interact with ECM collagens �Lysyl hydroxylase 1 (PLOD 1 gene)- Adds a necessary modification to a lysine residue on collagen fibril �Procollagen I N-proteinase (ADAMTS 2 gene) is important in the maturation of the collagen �Tenascin X (TNXB gene) is secreted by matrix cells and is related to the formation of the elastic fiber network.

Current Questions for Research in Genetics in EDS �What are the other genes responsible for EDS? �EDS hypermobility type �What other genes or environmental factors alter the clinical presentation in EDS? �Modifier genes �Epigenetic factors �What treatments may modify the manifestations of EDS? �Drug screening strategies

Research areas of interest Definitions �Genomics �The study of genes and their functions, and related techniques. �genomics vs genetics � Genetics scrutinizes the functioning and composition of the single gene � Genomics addresses all genes and their inter relationships in order to identify their combined influence on the growth and development of the organism �Epigenetics �The study of changes in organisms caused by modification of gene expression rather than alteration of the genetic code itself. May be influenced by the environment �Metabolomics �the systematic study of the unique chemical fingerprints that specific cellular processes leave behind

Genomics: Molecular Methods Single Gene Analysis Increasing data Gene Panels Exome Genome Increasing tests • Molecular diagnostics = testing genes known to be associated with a disorder • Increasing complexity with exomes and genomes, more data + need for interpretation • 2000+ single gene tests being offered diagnostically and many combinations of panels • Many disorders are genetically heterogeneous with ‘new’ gene discoveries occurring

Why Exome Sequencing? Sanger + Targeted Panel • Very accurate • Optimization possible • No bias for genes • inexpensive per exon • Low chance of incidental findings • Standardized workflow • “Easy” analysis • Re-use of exomes to interpret new ones • Quick turn-around • “Easy” interpretation - Exome • Low diagnostic yield for genetically heterogeneous diseases • Low throughput • Design and re-design required • Different designs for different disorders • Sufficient cases required + logistics of runs • Simple to add new genes • Sequencing bias • No non-coding regions • Secondary findings Genome • No bias in what you sequence • Little technical biases • Allows detection of SVs and SNVs in one experiment • Data analysis bottleneck • Interpretation of noncoding variants • Expensive, timeconsuming • More streamlined, reduced cost, higher yield in heterogeneous disorders • WES is currently more feasible to do ‘in house’ than WGS
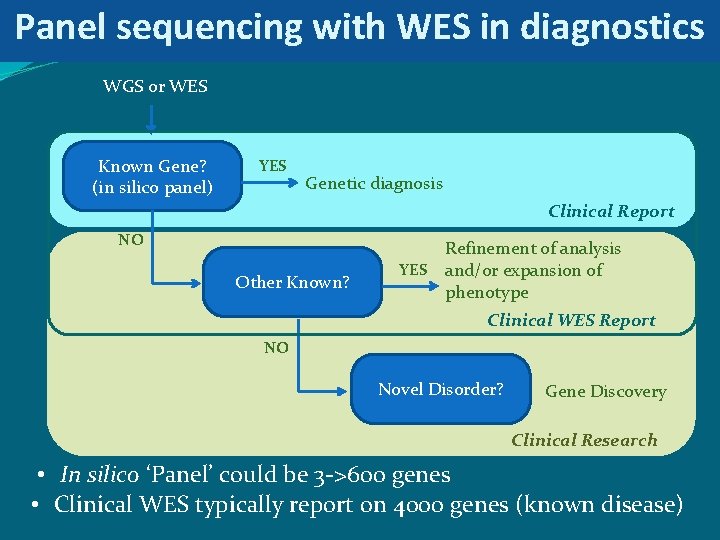
Panel sequencing with WES in diagnostics WGS or WES Known Gene? (in silico panel) YES Genetic diagnosis Clinical Report NO Other Known? YES Refinement of analysis and/or expansion of phenotype Clinical WES Report NO Novel Disorder? Gene Discovery Clinical Research • In silico ‘Panel’ could be 3 ->600 genes • Clinical WES typically report on 4000 genes (known disease)

Next generation sequencing gene panels �The Hospital for Sick Children- Available Ehlers Danlos Panel (17 genes) �ACTA 2, ADAMTS 2, ATP 7 A, CHST 14, COL 1 A 1, COL 1 A 2, COL 3 A 1, COL 5 A 2, FBN 2, FKBP 14, PLOD 1, SLC 39 A 13, SMAD 3, TGFB 2, TGFBR 1, TNXB �CTGT-Ehlers-Danlos syndrome NGS panel – Dominant (5 genes) �COL 1 A 1 / COL 1 A 2 / COL 3 A 1 / COL 5 A 2 �CTGT-Ehlers-Danlos syndrome NGS panel – Recessive (5 genes) �ATP 7 A / CHST 14 / FKBP 14 / PLOD 1 / SLC 39 A 13 http: //www. ctgt. net/
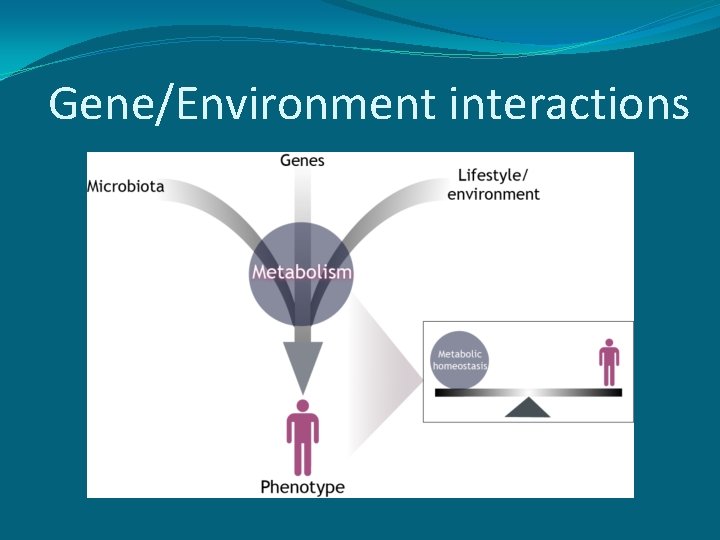
Gene/Environment interactions
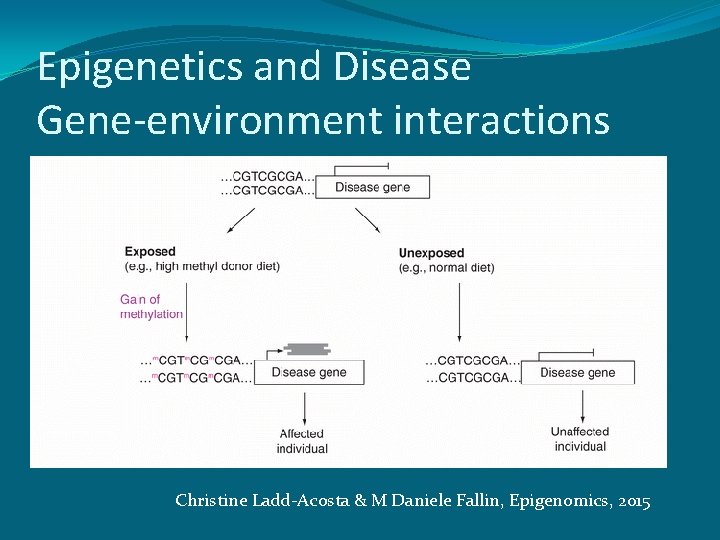
Epigenetics and Disease Gene-environment interactions Christine Ladd-Acosta & M Daniele Fallin, Epigenomics, 2015

Epigenetics and Disease

Epigenetics as a Biomarker

Epigenetics and epidemiology �Epigenetic data in epidemiology may further our understanding of disease mechanisms �Understanding epigenetics as a modifier of genetic or environmental risk for disease can expand our understanding of disease mechanisms. �Epigenetics changes provide a potential biological mechanism to explain gene–environment interactions associated with disease. �Epigenetic data may be used clinically as a biomarker for diagnosis, prognosis or response to treatment. �Epigenetics has the potential to serve as a robust biomarker of exposure for epidemiology studies that seek to examine environmental risk factors for disease. Christine Ladd-Acosta & M Daniele Fallin, Epigenomics, 2015
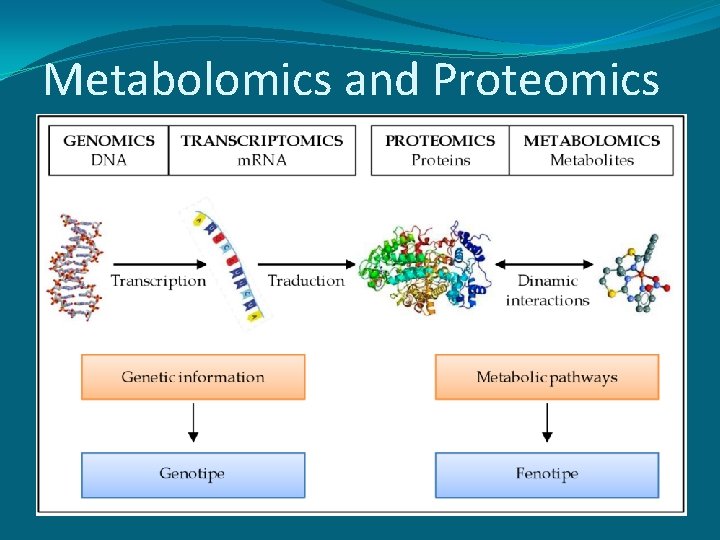
Metabolomics and Proteomics

Animal Models and Genetic Disease �Animal models have greatly improved our understanding of the cause and progression of human genetic diseases. �Allow investigation of environmental effects and modifier genes in a controlled manner �Useful tool for discovering targets for therapeutic drugs. �Caveat: The same treatments do not always translate to human clinical trials. �Most available animal models are made in mice, and they recreate some aspects of the particular disease. However, few, if any, replicate all the symptoms.

Animal models available Simmons, D. (2008) Nature Education 1(1): 70
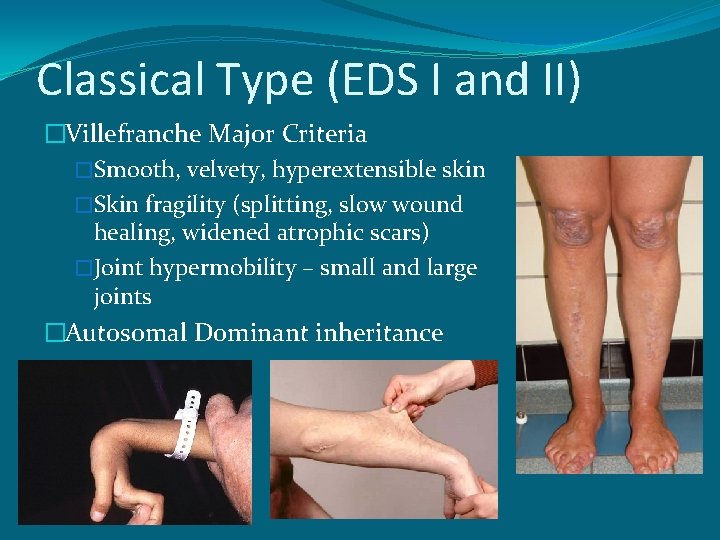
Classical Type (EDS I and II) �Villefranche Major Criteria �Smooth, velvety, hyperextensible skin �Skin fragility (splitting, slow wound healing, widened atrophic scars) �Joint hypermobility – small and large joints �Autosomal Dominant inheritance

Classical Type (EDS I and II) Ultrastructural findings Irregular, abnormally-shaped collagen fibers Electron Microscopy-Full thickness biopsy Cauliflower deformity of collagen fibers From: Hausser & Anton-Lamprecht 1994

Classical Type (EDS I and II) Molecular genetics �Heterogeneous �Mutations in COL 5 A 1 and COL 5 A 2 (50 -90%) �To date no deletions or duplications in COL 5 A 1 or COL 5 A 2 have been identified in patients with EDS. �Null mutations in COL 1 A 2 �Homozygous mutations of Tenascin X (TNXB)

Collagen type V knock out mouse Collagen content of dermis in col 5 a 1+/+ and +/minus animals as a function of age. col 5 a 1+/-mice have hyperextensible skin and demonstrate decreased tensile strength in normal and wounded skin. The dermis of col 5 a 1+/-contains poorly organized and less densely packed fibers. © 2006 by American Society for Biochemistry and Molecular Biology Richard J. Wenstrup et al. J. Biol. Chem. 2006; 281: 1288812895

COL 5 A 1 KO mouse model conclusions �COl 5 A 1 mouse model recapitulates the clinical features seen in patients with Ehlers-Danlos syndrome classic type. �col 5 a 1+/-mice have decreased compliance and tensile strength of the aorta �Hyperextensible skin with decreased tensile strength of both normal and wounded skin. �Joint laxity appears to be absent in col 5 a 1+/-mice; �The most likely explanation for this surprising observation is that type XI collagen functions in dosage compensation in tendon. 3
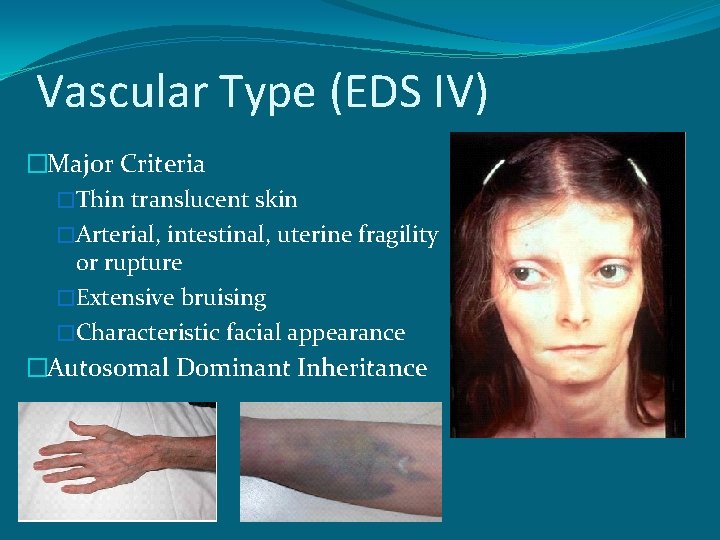
Vascular Type (EDS IV) �Major Criteria �Thin translucent skin �Arterial, intestinal, uterine fragility or rupture �Extensive bruising �Characteristic facial appearance �Autosomal Dominant Inheritance
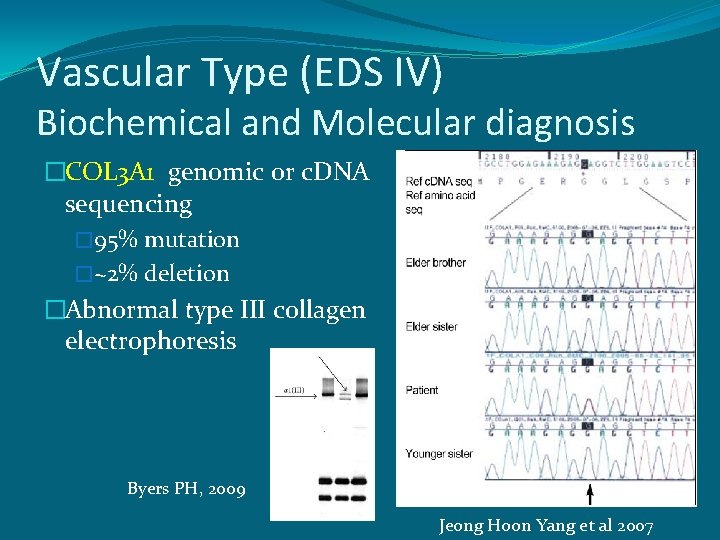
Vascular Type (EDS IV) Biochemical and Molecular diagnosis �COL 3 A 1 genomic or c. DNA sequencing � 95% mutation �~2% deletion �Abnormal type III collagen electrophoresis Byers PH, 2009 Jeong Hoon Yang et al 2007

COL 3 A 1 Mouse model T. K. Cooper et al. Vet Pathol 2010; 47: 1028 -1039

Hypermobility Type (EDS III) �Major Criteria �Generalized joint hypermobility �Significant joint pain �Less skin involvement (lax but not as overtly fragile) �Autosomal Dominant inheritance* �Molecular Basis: Mostly unknown �Heterozygosity for TNXB null mutations �Rarely COL 3 A 1 G 637 S mutation
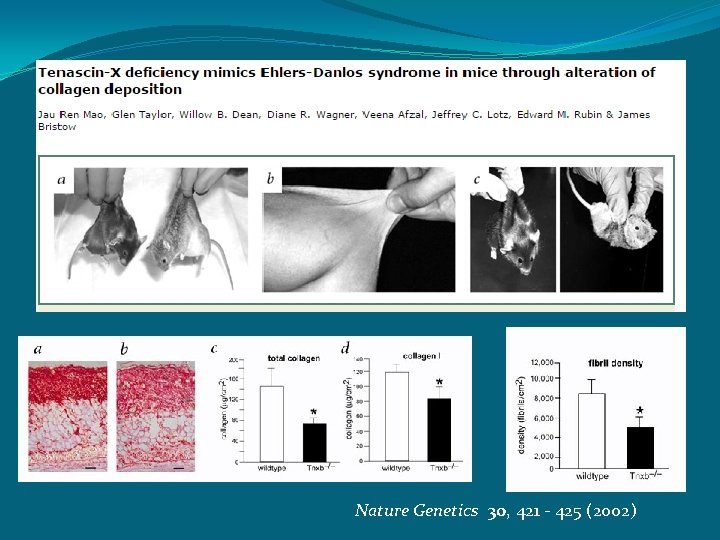
Nature Genetics 30, 421 - 425 (2002)
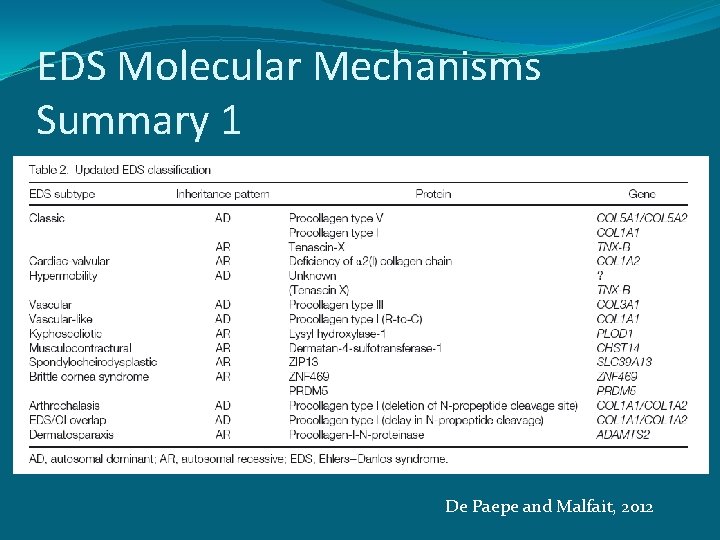
EDS Molecular Mechanisms Summary 1 De Paepe and Malfait, 2012
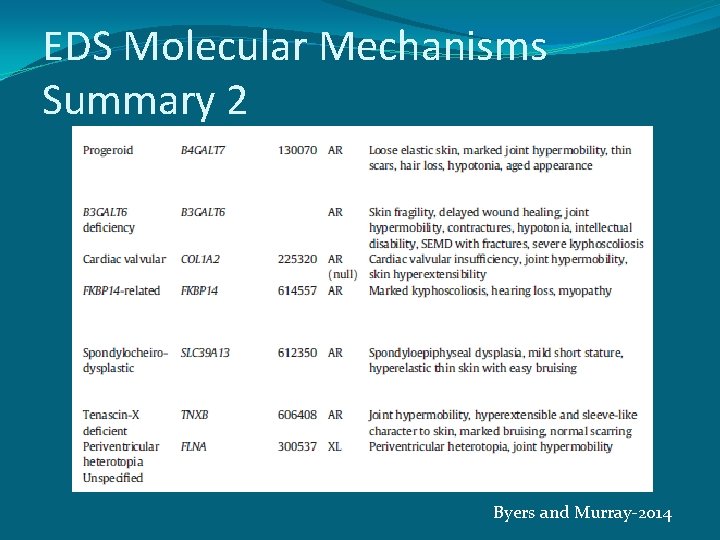
EDS Molecular Mechanisms Summary 2 Byers and Murray-2014
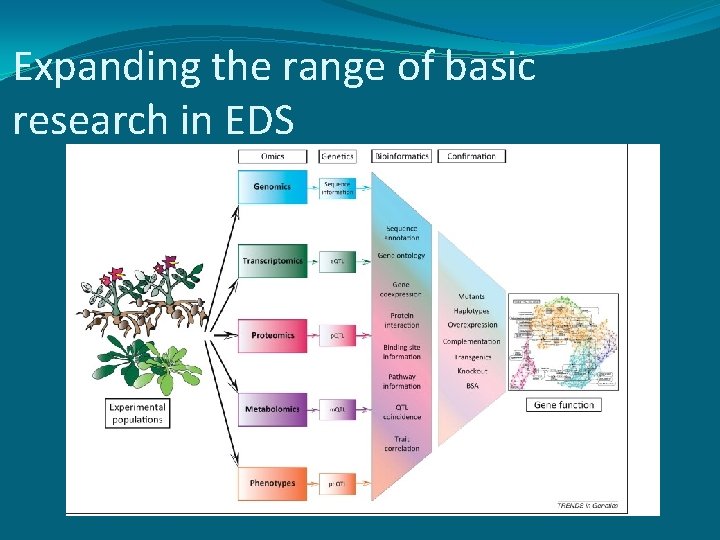
Expanding the range of basic research in EDS

Future Directions �Identify molecular basis of other forms of EDS �Study the role of the environment on disease manifestation (Epigenetics) �Identify biomarkers for diagnosis, monitoring and prognosis. �Elucidate common final pathways that may be targets for treatment for multiple groups of patients. �Develop treatment strategies that address the molecular defect.
- Slides: 33