R programming Jos R Valverde jrvalverdecnb csic es
R programming José R. Valverde jrvalverde@cnb. csic. es CNB/CSIC Flow control 8 1 20 Statistical Facts Flying on a plane is indeed secure: almost all deceased in plane accidents died after reaching the floor.

Preamble Getting data into R
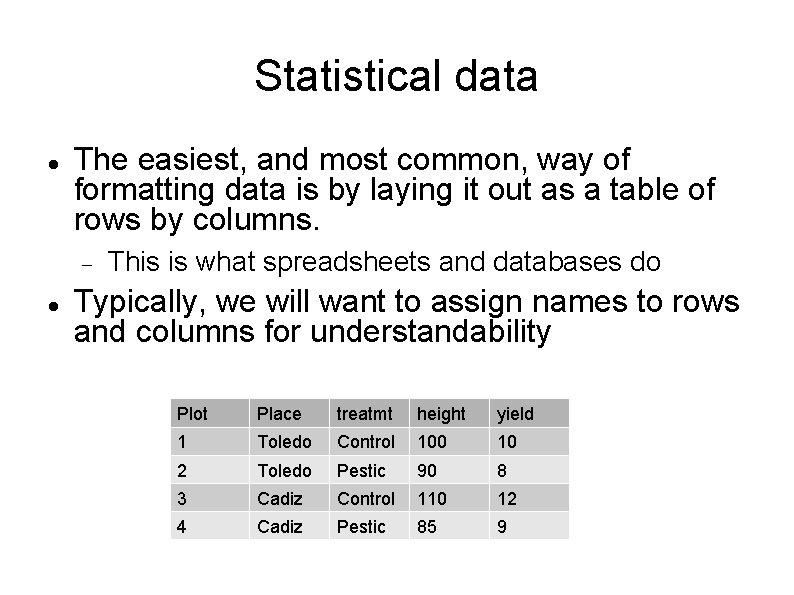
Statistical data The easiest, and most common, way of formatting data is by laying it out as a table of rows by columns. This is what spreadsheets and databases do Typically, we will want to assign names to rows and columns for understandability Plot Place treatmt height yield 1 Toledo Control 100 10 2 Toledo Pestic 90 8 3 Cadiz Control 110 12 4 Cadiz Pestic 85 9

Exchanging data There are many ways, but some formtas have become standards supported by almost everyone: CSV: each row occupies one line, with values separated by commas ', '. To avoid some problems, it is common to enclose each field in double quotes. TAB: or tab separated, with each row taking one line and fields separated by a tabulator character

Variants If you use CSV, then none of your fields should contain a ', ' because then it would be interpreted as two fields instead of one. This is specially troublesome under some localities where ', ' is used as a decimal or thousands separator Enclosing fields in double quotes fixes this issue But then fields cannot contain double quotes Usingle quotes instead of double quotes fixes this issue But then fields cannot contain single quotes. . . Check your data!

Getting data from a file You will need to specify The separation character between fields (TAB, comma, white space, etc. . . ' The delimiter character for fields (single or double quotes, etc. . . ) Any localization information, such as the character used to separate the integer and decimal parts in numbers. Whether the file includes a header with column names and possibly row names in one of the columns.

A sample file id 1 2 3 4 5 6 group time 1 time 2 time 3 time 4 A 31 29 15 26 A 24 28 20 32 A 14 20 28 30 B 38 34 30 34 B 25 29 25 N/A B 30 28 16 34 Copy and paste this and save it to a file 'twisk. txt'

Reading a table of data twisk <- read. table('twisk. txt', header=TRUE, row. names=1) twisk This reads a table from the specified file, using the first line in the table as a header that contains all the column names, and the first column as a list of row names By default, we assume that fields are separated by white space.

Dealing with missing data Often, when collecting data, we find situation were an observation cannot be collected. This observation must be identified as 'Not Available'. Different experimenters may decide to encode non-availability of data in different ways (e. g. as an impossible or negative value or as a specific string like 'N/A') We can identify these values with 'na. strings=' twisk <- read. table('twisk. txt', header=TRUE, row. names=1, na. strings='N/A')

Using CSV data Most spreadsheets will export data for use in other programs as 'CSV' comma-delimited files Id, group, time 1, time 2, time 3, time 4 1, A, 31, 29, 15, 26 2, A, 24, 28, 20, 32 3, A, 14, 20, 28, 30 4, B, 38, 34, 30, 34 5, B, 25, 29, 25, N/A 6, B, 30, 28, 16, 34 twisk <- read. csv('twisk. csv', header=TRUE, na. strings='NA')

Reading 'headless' data In cases where the data contains no 'header', i. e. it lacks the names of the columns, we can read it and add the names aftwerwards with names( ) We must assign to names(table) a vector with the names of the columns (and which, thus, must have as many names as columns are there) names(twisk) <- c("GRP", "T 1", "T 2", "T 3", "T 4") twisk rownames(table) allows us to set row names.

Reading in factor variables read. table() will treat any column with literal (non-numeric) values as composed of 'factors' or categorical variables. It makes more sense to collect categorical variables by name If you didn't, and encoded a factor using numbers (e. g. 1 / 2 for sex M / F) you will need to tell R using factor( ) converts its argument into a factor

A numerical factor example id 1 2 3 4 5 6 group 1 1 1 2 2 2 time 1 31 24 14 38 25 30 time 2 29 28 20 34 29 28 time 3 15 20 28 30 25 16 time 4 26 32 30 34 29 34

Convert numbers to factors We can select values by number and re-assign them a textual value t <- read. table("twisk 2. txt", header=TRUE) t t$group t$id[ t$group ==1 ] <- 'A' t$group[ t$group ==2 ] <- 'B' t$group <- factor(t$group)

Scripting

Why We have already used a lot of commands Do you remember each one and all of them? If you don't, then welcome to the selected group of advanced programmers If you do, don't worry, you will forget too soon and then you will be welcome to the selected group of advanced programmers Advanced programmers do not remember everything, nor do they like to write commands

Putting commands in a file If you do not want to repeat all the commands each time, then there is a simple solution Collect all the commands in a file This must be a text file (not a word processor file) Whenever you need to, ask R to repeat the work in that file Instead of typing the commands all over again

Example data(iris) dim(iris) names(iris) row. names(iris) with(iris, qqnorm(Petal. Length, ylab="Petal length") qqline(iris$Petal. Length) shapiro. test(iris$Petal. Length) bartlett. test(Petal. Length ~ Species, data=iris)

Running a script It is easy: use the 'source( )', Luke. . . If you saved the former commands in a text file and named the file, for example, 'script. R', then it suffices to type within R source('script. R') Or you can directly use it with Rscript. R

Reinforcing feeble memories If you still remember what this script did, you are lucky Even so, you will not remember one week, one month or one year from now. For this reason it is highly recommended that you add notes to your commands explaining what they do These notes are preceded by a # and are called 'comments'

Comments R will ignore your commens (i. e. whenever R finds a # in a line, it will ignore all the rest of the line) This allows you to document your code (the commands you write for R) You are strongly encouraged to extensively document everything you do. Best done while you write it, before you forget. .

# R comes with a number of standard datasets for us to try # we will make use of one of them in this script # iris gives the measurements in centimeters of the variables # sepal length and width and petal length and width, # for 50 flowers from each of 3 species of iris. data(iris) dim(iris) # get the dimensions of the data set names(iris) # get the column (variable) names row. names(iris) # get the row names # test for normality shapiro. test(iris$Petal. Length) # test for homogeneity of variances bartlett. test(Petal. Length ~ Species, data=iris) # draw a Q-Q normality plot with(iris, qqnorm(Petal. Length, ylab="Petal length")) qqline(iris$Petal. Length)

Flow control

Logical operations == != >< >= <= equal & | ! equal not equal greater/less than or and or not

Conditional execution if (cond == true) cmd 1 else cmd 2 Example If (1 == 0) { print(1) } else { print(2) } If operates on length-one logical vectors (i. e. tests a single value)

Statements and compound statements A single command is called a statement You can group several commands together as if they were a single one with { and } This new, multi-command, group is called a compound statement Thus if (condition == true) stmt 1 else stmt 2 is conceptually the same as If (condition == true) { cmd 1, cmd 2. . . } else { cmd 3, cmd 4. . . }

Vectorial conditions ifelse(test, true_value, false_value) This function operates on vectors x <- 1: 10 # Creates sample data ifelse(x<5 | x>8, x, 0) We have already seen short-hand versions of this method

Iteration over a list of values For (variable in sequence) statement Example mydf <- iris Myve <- NULL # Creates an empty vector container for(i in seq(along=mydf[, 1])) { myve <- c(myve, mean(as. numeric(mydf[i, 1: 3]))) } myve
![For (2) x <- 1: 10 z <- NULL for(i in seq(along=x)) { if(x[i] For (2) x <- 1: 10 z <- NULL for(i in seq(along=x)) { if(x[i]](http://slidetodoc.com/presentation_image_h/d096a9b89eae5d4dc55f952a6692dd1d/image-29.jpg)
For (2) x <- 1: 10 z <- NULL for(i in seq(along=x)) { if(x[i] < 5) { z <- c(z, x[i] - 1) } else { z <- c(z, x[i] / x[i]) } } z

For (3) x <- 1: 10 z <- NULL for(i in seq(along=x)) { if (x[i]<5) { z <- c(z, x[i]-1) } else { stop("values need to be <5") } } z

Iterate while a condition holds while (condition) statement Example z <- 0 while(z < 5) { z <- z + 2 print(z) } Notice the output and think about it.

Iterate indefinitely repeat statement The statements are repeated for ever unless break() is called Example z <- 0 repeat { z <- z + 1 print(z) if(z > 100) break() } You better set a condition to stop the loop!

Plotting revisited data(iris) par(mfrow=c(4, 4)) for ( i in 1: 4 ) { for (j in 1: 4 ) { plot(iris[ , i], iris[ , j], xlab=names(iris)[i], ylab=names(iris)[j]) } } That makes quite a difference, doesn't it?

Plotting revisited (2) Try now pairs(iris) If you were wondering, “how comes nobody else has tried to simplify this, if it is so easy? ”, now you know the answer. Yes, indeed, someone realized that making multiple plots was that easy and made a special function to facilitate that.

Working with collections of values We can process all the values in a collection using a loop x <- rnorm(10) for (i in 1: 10) print(x[i]) But R provides a more convenient way to work with collections of data values: apply( ) apply(object, dim, function) This will apply function to the values in the specified object (a matrix or data. frame). dim can be 1 or 2 depending on whether we want to act on rows or columns.
![Working with collections means <- apply( iris[ , 1: 4], 2, mean) means Here, Working with collections means <- apply( iris[ , 1: 4], 2, mean) means Here,](http://slidetodoc.com/presentation_image_h/d096a9b89eae5d4dc55f952a6692dd1d/image-36.jpg)
Working with collections means <- apply( iris[ , 1: 4], 2, mean) means Here, we select columns 1 to 4 of the dataset iris, and tell R to apply the function mean to each column (2). medians <- apply( iris[ , 1: 4], 2, median) variances <- apply( iris[ , 1: 4], 2, var) medians ; variances ^ 0. 5 # sd

Polymorphism Some functions may have more than one role, i. e. we can do several different things using the same name, like with print( ). var(iris$Sepal. Length) var(iris) When applied to a matrix or data. frame, var gives the variance/covariance matrix cor(iris$Sepal. Length, iris$Petal. Length) cor(iris[, 1: 4]) # correlation matrix

Processing collections by factor tapply(vector, factor, function) aggregate(x, by, function) Example ## Computes mean values of vector agregates # defined by factor for (i in 1: 4) tapply(as. vector(iris[, i]), iris$Species, mean) ## The aggregate function provides related # functionality

For vectors and lists lapply returns a list sapply returns the simplest object possible: a vector or a matrix Examples ## Creates a sample list mylist <- as. list(iris[1: 3, 1: 3]) mylist ## Compute sum of each list component and return result # as list lapply(mylist, sum)

Sorting data Sort a vector x <- c(1. 1, 5. 5, 2. 2, 9. 9, 7. 7, 4. 4, 6. 6, 5. 0) sort(x) Get the indexes that will result in a sorted list order(x) in_order <- order(x) x[in_order]

Sorting matrices and data frames To sort a matrix or data frame according to some columns, we can use order( ) Example: to sort according to sex and age we could use index <- order(data$sex, data$age) sorted_data <- data[order, ] idx <- order(iris$Sepal. length, iris$Sepal. Width) iris[idx, ]

Example

A proteomic dataset We have a strain that produces an interesting product and three mutant alternatives that we expect to be more efficient Are there differences between the strands? If so, which are the differences? We'll use the data in file “proteomics. tab” data < - read. table(“proteomics. tab”, sep='t' header=TRUE, na. strings='NA', dec=”. ”, strip. white=TRUE, row. names=1)

The dataset We have data from 10 control experiments (control. 1 to control. 10) 2 experiments on mutant 1 (mutant. 1. A and mutant. 1. B) 1 experiment from mutant. 2 and another from mutant. 3 For each strain, we have measured peptides and assigned them to 312 proteins. We collect the number of observations of each protein in each experiment.

Parametric or not Our first decision is which kind of tests shall we apply. A first hypothesis may be that protein observation follows a normal distribution We need to check each protein and verify that its observations in the control group do indeed follow a normal distribution.

Transforming data We currently have data organized in columns by strain We want to analyze it by protein: we need to transpose the dataset tdata < - t(data) Now, each column corresponds to one protein and we can use shapiro. test(tdata[1])

Exercice Do all 312 tests formality for each protein using the shapiro. test( ) function.
![Doing it all at once for (i in 1: 312) print(shapiro. test(tdata[ , i])) Doing it all at once for (i in 1: 312) print(shapiro. test(tdata[ , i]))](http://slidetodoc.com/presentation_image_h/d096a9b89eae5d4dc55f952a6692dd1d/image-48.jpg)
Doing it all at once for (i in 1: 312) print(shapiro. test(tdata[ , i])) Or even better yet for (i in 1: 312) { print(paste(“Analyzing normality of protein”, i)) print(shapiro. test(tdata[ , i]) }

Compare strains To compare strains we'll use the data arranged by strains in columns. We will use as reference the observations of each protein across the control group data$control. sum <- apply(data[ , 1: 10], 1, sum) data$control. mean <- apply(data[ , 1: 10], 1, mean) data$control. p <- data$control. sum / sum(data$control. sum)

Comparing two frequency tables We'll use the Chi-square Goodness-of-fit test. We want to compare a data distribution with the expected frequencies/probabilities and see if the observed distribution may have been drawn from the expected distribution. chisq. test(data$control. 1, p=data$control. p) Very often, we will want to compare a control and a test distribution. R offers a shortcut for this chisq. test(data$control. 1, p=data$control. sum, rescale. p=TRUE)

Exercice Compare all strains using the chisq. test( ) function to perform a goodness-of-fit analysis.

A possible solution for (i in c(11, 12, 13, 14) ) { # names(data) produces a vector with # all the column names in 'data' # which we use to obtain the name of # this “i” column print( paste('Testing', names(data)[i]) ) chisq. test(data[ , i], p=data$control. p) }

Manipulating data The data we are using is not really amenable to standard statistical tests. We have too many zeros for χ² based analyses We may prefer to use proportions or transformed data Some proteins appear in such a small proportion that they may have been missed in the analysis purely by chance, we may want to exclude them from the analysis. We could consider only the topmost significant proteins

Dealing with the most significant proteins Sort the dataset according to protein frequency. Since it needs not be the same in all strains, we will sort by the frequency in the control group. Then compare each mutant strain to the control considering the topmost 10, 50, 100, 150 and 200 proteins. Can you do it?
![Solution idx ← order(data$control. sum) sdata ← data[idx, ] for (j in c(10, 50, Solution idx ← order(data$control. sum) sdata ← data[idx, ] for (j in c(10, 50,](http://slidetodoc.com/presentation_image_h/d096a9b89eae5d4dc55f952a6692dd1d/image-55.jpg)
Solution idx ← order(data$control. sum) sdata ← data[idx, ] for (j in c(10, 50, 100, 150, 200)) { print(chisq. test(data$control. 1[1: j], p=data$control. sum[1: j], rescale. p=TRUE)) print(chisq. test(data$mutant. 1 A[1: j], p=data$control. sum[1: j], rescale. p=TRUE)) print(chisq. test(data$mutant. 1 B[1: j], p=data$control. sum[1: j],

Printing important information Using the test directly may provide too much (or too little) information, or just not the information we really want. We can tell R to print exactly what we want. Each test returns a result. The result contains a lot of information such as the test score, the P value, residuals, etc. . . We can select which information is printed using $ notation: chisq. test(mutant. 1, p=control. mean, rescale. p=T)$p. value

Exercice Repeat the analyses, but this time printing only the p. value obtained in each test. You will want to also print the test name and the parameters.

Removing values We now want to remove all rows where the protein count (for any strain) is 0 and repeat the analysis. You can look it up on google if you want Generate a new dataset without rows containing zeros (removing proteins with a count of zero en any of the strains)

Solution Remove rows with any value == 0 data[ apply(data, 1, function(x) all(x != 0)), ] data[ ! (apply( data, 1, function(x) any(x == 0)), ] Assign to a new variable and repeat the analysis

Thanks Questions?
- Slides: 60