Structure Visualization UCSF Chimera Jos R Valverde CNBCSIC

Structure Visualization UCSF Chimera José R. Valverde CNB/CSIC jrvalverde@cnb. csic. es © José R. Valverde, 2014 CC-BY-NC-SA

Summary This is only a short introduction to visualization of macromolecular structures using UCSF Chimera There are many other tools that you can use Py. MOL Ras. Mol Jmol And many, many more. . . There are many more things that you can do with these tools

UCSF Chimera Interactive molecular graphics program Built on top of the Python programming language Can be extended using Python Provides Structure visualization Volume visualization Data analysis Data preparation for other computational tools Command-line interface for automating tasks
www. cgl. ucsf. edu/chimera Your first stop for downloads and information
Getting started Open UCSF Chimera (“chimera”)
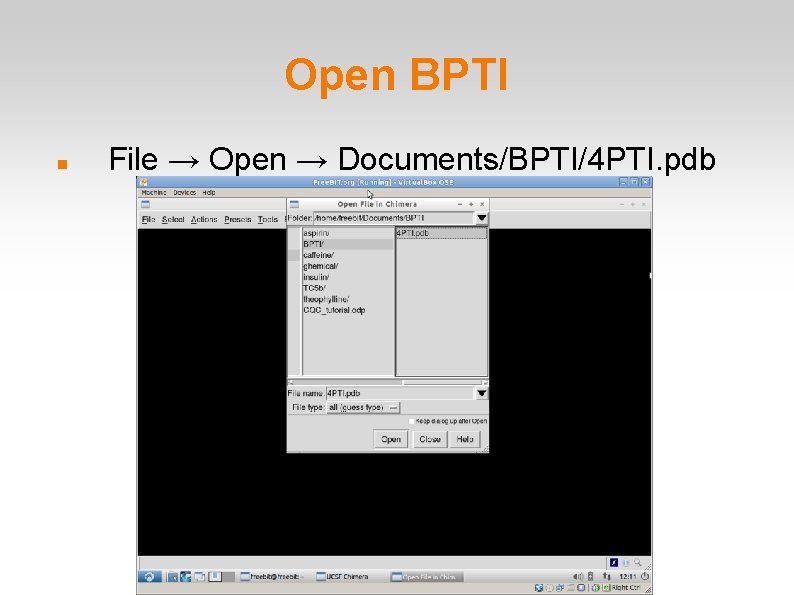
Open BPTI File → Open → Documents/BPTI/4 PTI. pdb

Ribbons view Experiment with the mouse buttons to move, rotate or zoom/pan the structure
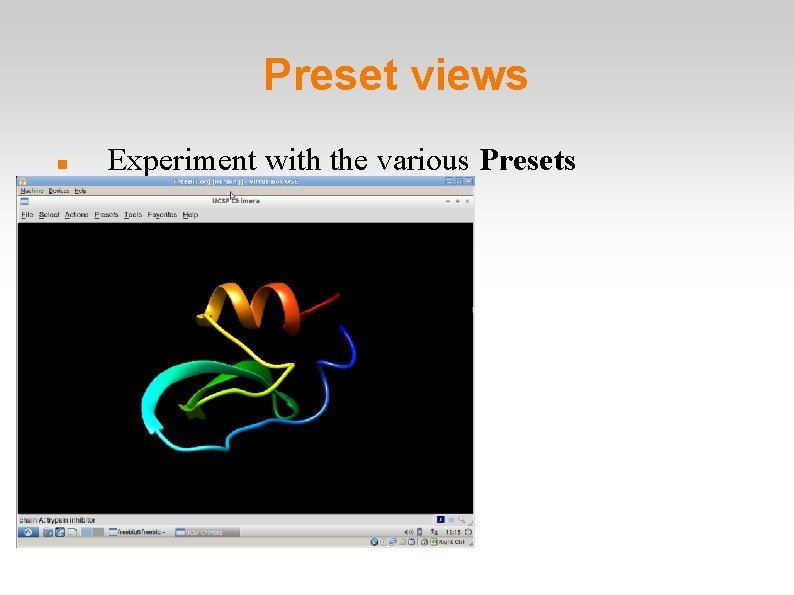
Preset views Experiment with the various Presets
Preset views Experiment with the various Presets
Preset views Experiment with the various Presets
Preset views Experiment with the various Presets
Preset views Experiment with the various Presets

Selection: display water Select → Residue → HOH Actions → Atoms/Bonds → Show Note that H atoms are not present in the PDB file
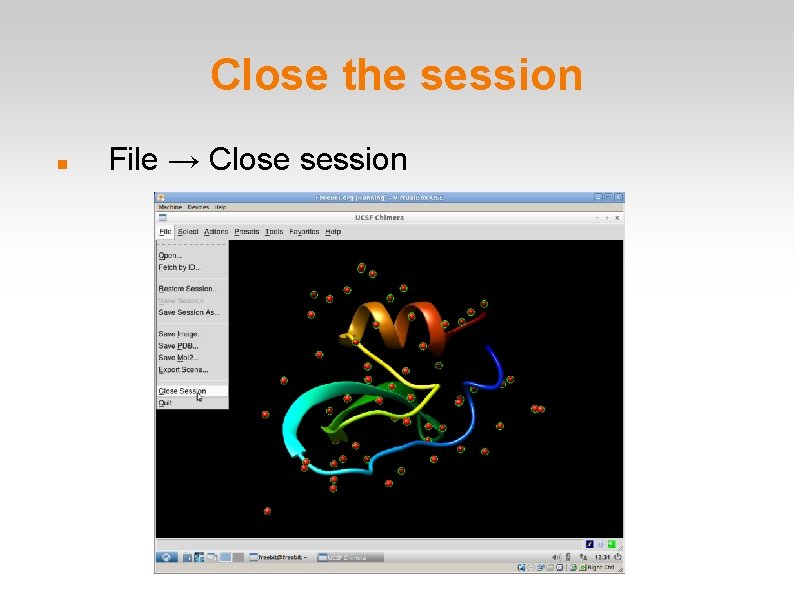
Close the session File → Close session
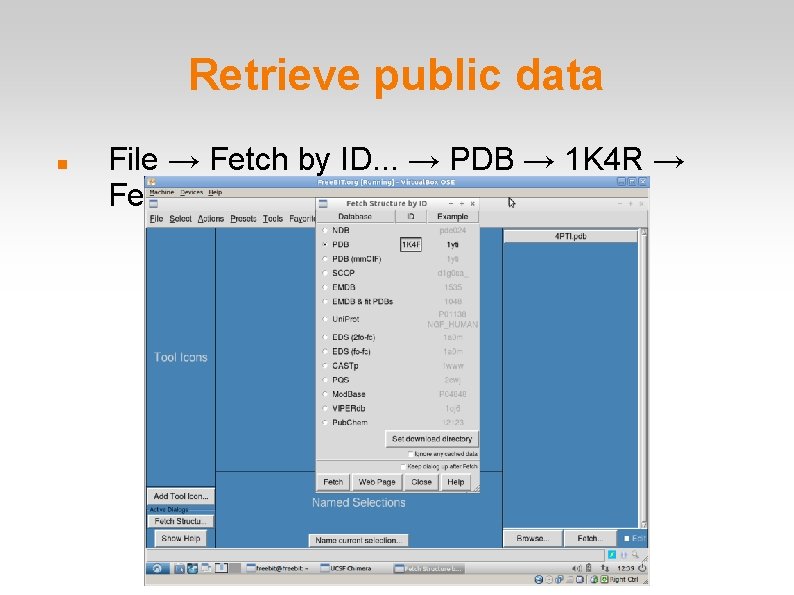
Retrieve public data File → Fetch by ID. . . → PDB → 1 K 4 R → Fetch
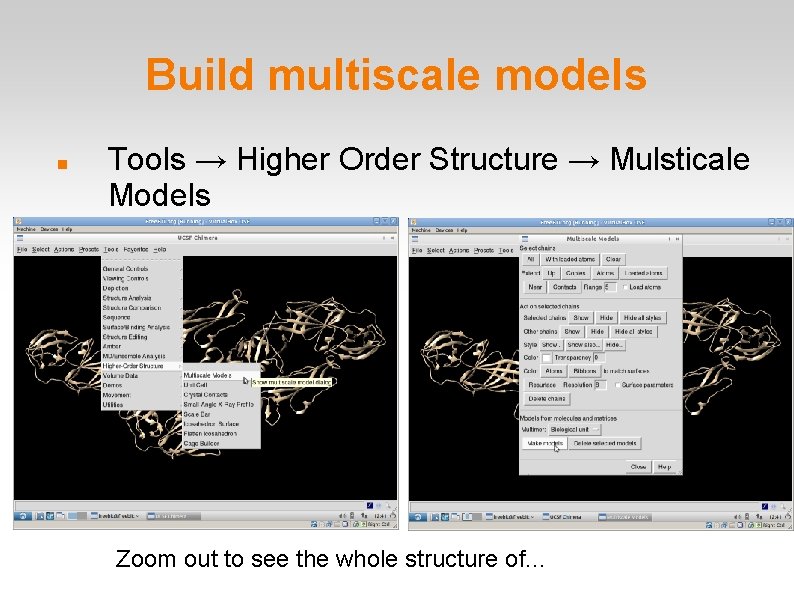
Build multiscale models Tools → Higher Order Structure → Mulsticale Models Zoom out to see the whole structure of. . .
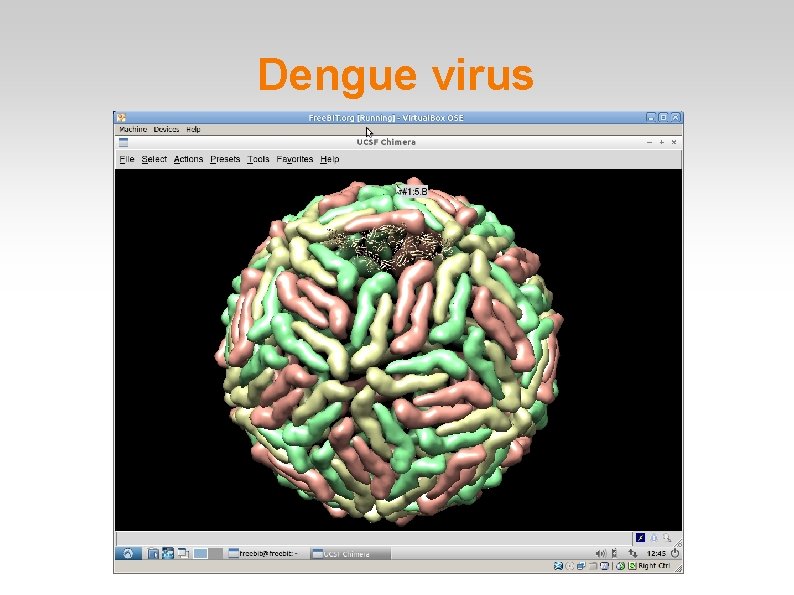
Dengue virus

Epsilon/Zeta TA Close session and retrieve PDB ID 3 Q 8 X Select → Structure → Ligand Note that complex atoms are shown in wire mode Ligand Amino acids

Show ligand (or protein) surface Select the molecule you want Select menu Ctrl+Click on atom and use up/down arrows Ctrl-Click → select atom Up/Down arrow → expand/reduce selection Residue Molecule Ensemble Model … Actions → Surface → Show

Select Active Site Select → Structure → Ligand Select → Zone Actions → Atom/Bonds → Show
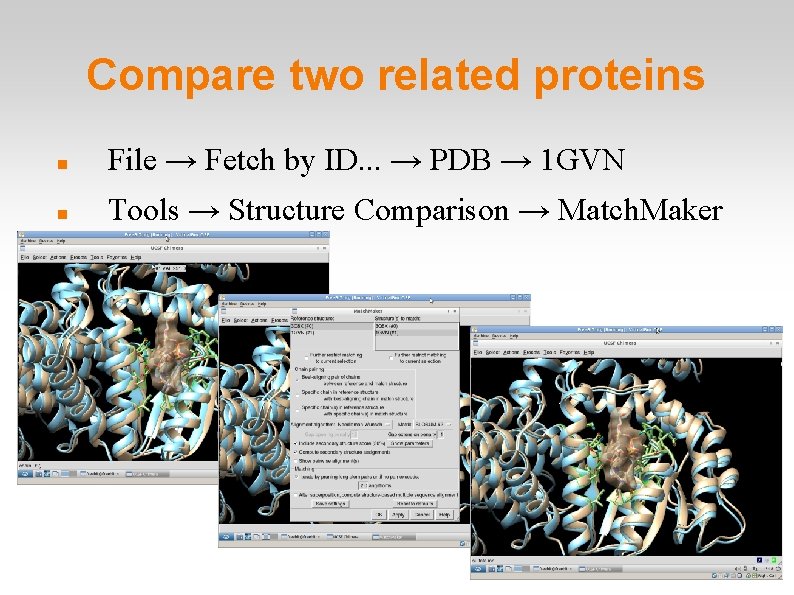
Compare two related proteins File → Fetch by ID. . . → PDB → 1 GVN Tools → Structure Comparison → Match. Maker

Model Panel Favorites → Model Panel Hide (unmark “Shown”) surface and 3 Q 8 X Hide 1 GVN and Show 3 Q 8 X. . . play

Save selected atoms Notice that 1 GVN does not contain a ligand After aligning them, we can “transfer” the ligand from 3 Q 8 X to 1 GVN Select → Selection Mode → Append Select → Clear selection Select → Chain → A, B, C, D → 1 GVN Select → Residue → UD 1 Actions → Write. PDB → Save both models Save selected atoms only + Save relative to model 3 Q 8 X + File name: 1 GVN+UNAG + Save multiple models in a single file
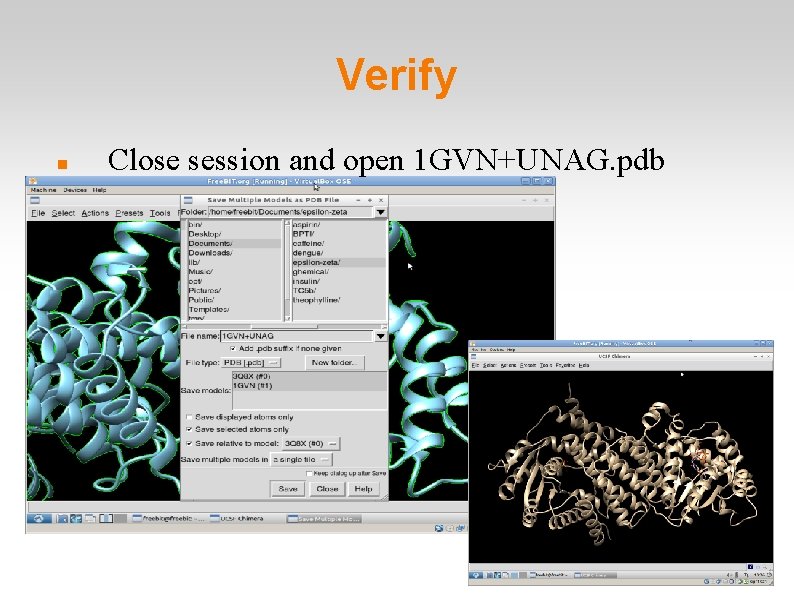
Verify Close session and open 1 GVN+UNAG. pdb

There is a lot more. . . Do not hesitate to check the UCSF Chimera web site for more information.

mov/subunit. mpeg

The magic garden You may be wondering where did all those structures came from. . Open a web browser and go to PDB web site: www. ebi. ac. uk/pdbe www. rcsb. org This is the database repository of known 3 D macromolecular structures. You can search it. . . and use the PDB ID to load structures in Chimera. File → Fetch by ID

Some interesting entries 309 D (DNA) 173 D (DNA bound to anticancer drug) 3 WN 8 (collagen) 3 J 6 R (Human papillomavirus) Multiscale Model In the main page, click on “Learn” (left pane) to see a wealth of educational resources from PDB (posters, paper models, animations. . . )

EMDB http: //www. ebi. ac. uk/pdbe/emdb/ This is the Electron Microscopy databank You can search or browse it, e. g. : 1321 (bacteriophage T 7) 6243 (poliovirus) 2240 (human myosin from heart muscle) … move around.

Questions? Original picture by muzree, license CC 0 http: //pixabay. com/en/mosquito-bite-decease-malaria-213806/
- Slides: 30