MARKOV MODELS Dilvan Moreira based on Prof Andr

MARKOV MODELS Dilvan Moreira (based on Prof. André Carvalho presentation)

Reading Introduction to Computational Genomics: A Case Studies Approach Chapter 4

Topics 3 Introduction Odor receptors Hidden Markov Chains Applications in Bioinformatics Case Study Algorithms André de Carvalho - ICMC/USP 15/12/2021

Introduction 4 In 2004 , Richard Axel and Linda Buck won the Nobel prize Elucidation of the olfactory system It includes a large family of proteins Odor receptors (OR) Odorants or olfactory Combination allows feel more than 10, 000 different odors Located on the cell surface of the nasal passage Detect odor molecules when they are inhaled and pass information to the brain André de Carvalho - ICMC/USP 15/12/2021

Olfactory Receptors 5 They must be able to cross the cell membrane to carry information to the brain It contains 7 transmembrane domains Parts of highly hydrophobic AAs interact with the fat of the membrane Protein has alternating sections of hydrophobic and hydrophilic AAs Discovery allowed the description of similar receptors Detection of taste and pheromone André de Carvalho - ICMC/USP 15/12/2021

Olfactory Receptors 6 Largest family of genes in the human genome 1000 600 genes, ~40 % of them are functional are called pseudogenes Inactive functional genes or defective Result of natural selection Predominance of other sense (vision) Usually similar to the functional genes Dogs ( ~1500 genes) and rats (~1000 genes) have ~80 % of functional genes André de Carvalho - ICMC/USP 15/12/2021

Olfactory Receptors 7 OR analysis requires more sophisticated computational tools Similarity between multiple ORs genes is low to detect with pair alignment Relevant signals can be detected with multiple alignment of all ORs Genome has much noise Ex. : regions with high amount of GC can have long sections of As and Ts André de Carvalho - ICMC/USP 15/12/2021
Modeling Sequences 8 Multinomial models Nucleotides are independent and identically distributed p = (p. A, p. C, p. G, p. T) Markov models Probability of a symbol depends on the previous symbols André de Carvalho - ICMC/USP 15/12/2021

Hidden Markov Chains 9 Multinomial and Markov models Isolated, they fail to capture many of the sequences properties Hidden Markov Chains (HMM) One of the main Algorithms in Bioinformatics Hidden Markov Models Combinam modelos de sequências multinomiais e de Markov Combine multinomial sequences model and Markov model André de Carvalho - ICMC/USP 15/12/2021

Hidden Markov Chains 10 Models a sequence as being indirectly generated by a Markov chain Each position in the sequence has a hidden state The sequence is modeled as a double random process Generate the HMM Transform the hidden chain in the observed sequence Using a different multinomial distribution for each state of the Markov chain André de Carvalho - ICMC/USP 15/12/2021

Hidden Markov Chains 11 Widely used for voice recognition Used in 1989 to Segmentation of DNA sequences Churchill segmented DNA sequences in regions with similar nucleotide uses Later used for other applications: Identification (recognition) gene Prediction of protein structure Identifies patterns that do not have a rigidly defined structure André de Carvalho - ICMC/USP 15/12/2021

Hidden Markov Chains 12 Finite automata Set of N states, H (hidden alphabet) An alphabet of M observable symbols , S Probabilities of transition between states , T Probabilities of symbols emission in each state , and André de Carvalho - ICMC/USP 15/12/2021

Hidden Markov Chains 13 Central ideas: A string is represented by a system A system can have different states A system can switch between states with probability T of transition In each state, the system sends symbols to a string with probability of emission E André de Carvalho - ICMC/USP 15/12/2021

Hidden Markov Models 14 Model parameters: Transition probabilities: define the probabilities to change state of the Markov chain André de Carvalho - ICMC/USP 15/12/2021
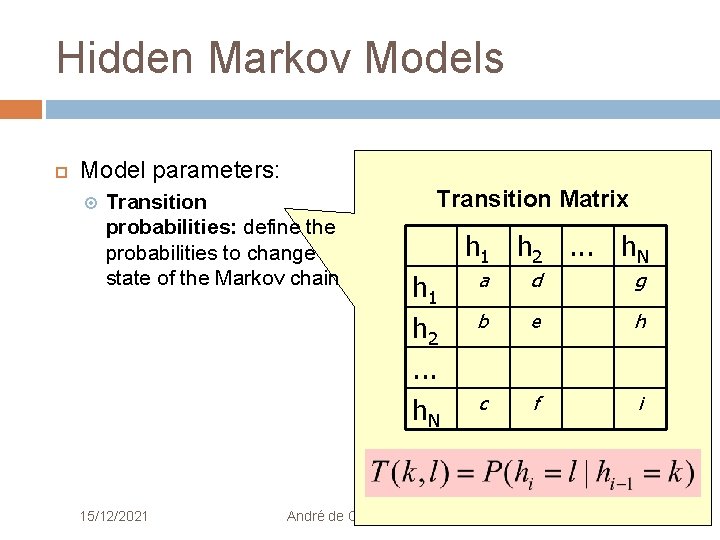
Hidden Markov Models Model parameters: Transition probabilities: define the probabilities to change state of the Markov chain 15/12/2021 Transition Matrix h 1 h 2. . . h. N a d g b e h c f i André de Carvalho - ICMC/USP 15

Hidden Markov Models Model parameters: Transition probabilities: define the probabilities to change state of the Markov chain Emission probabilities: probability of the symbols of observable sequence being produced in each of the states 15/12/2021 André de Carvalho - ICMC/USP 16

Hidden Markov Models Model parameters: Transition probabilities: define the probabilities to change state of the Markov chain Emission probabilities: probability of the symbols of observable sequence being produced in each of the states 15/12/2021 Matriz de Emissão h 1 h 2. . . h. N s 1 s 2 . . . s. M a d g b e h c f i André de Carvalho - ICMC/USP 17

Hidden Markov Models 18 h: sequence of hidden states s: symbol sequence Initial probability for states: The probabilities of the symbols in each state and the transitions between states must be 1. André de Carvalho - ICMC/USP 15/12/2021

Hidden Markov Models 19 The model parameters can be estimated using the Expectation-Maximization (EM) algorithm, based on known data André de Carvalho - ICMC/USP 15/12/2021

Example 20 Two things: a fair and an addict Given a sequence of entries, can you guess which data originated each value of the sequence? André de Carvalho - ICMC/USP 15/12/2021
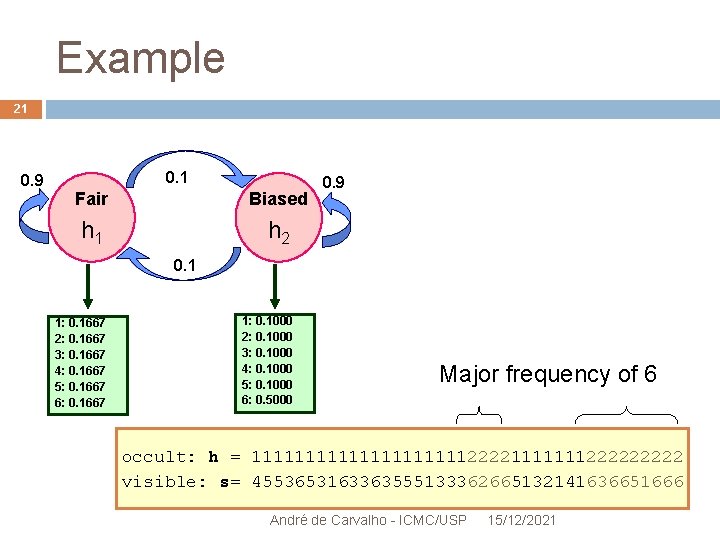
Example 21 0. 9 0. 1 Fair Biased h 1 h 2 0. 9 0. 1 1: 0. 1667 2: 0. 1667 3: 0. 1667 4: 0. 1667 5: 0. 1667 6: 0. 1667 1: 0. 1000 2: 0. 1000 3: 0. 1000 4: 0. 1000 5: 0. 1000 6: 0. 5000 Major frequency of 6 occult: h = 11111111112222111111122222 visible: s= 4553653163363555133362665132141636651666 André de Carvalho - ICMC/USP 15/12/2021
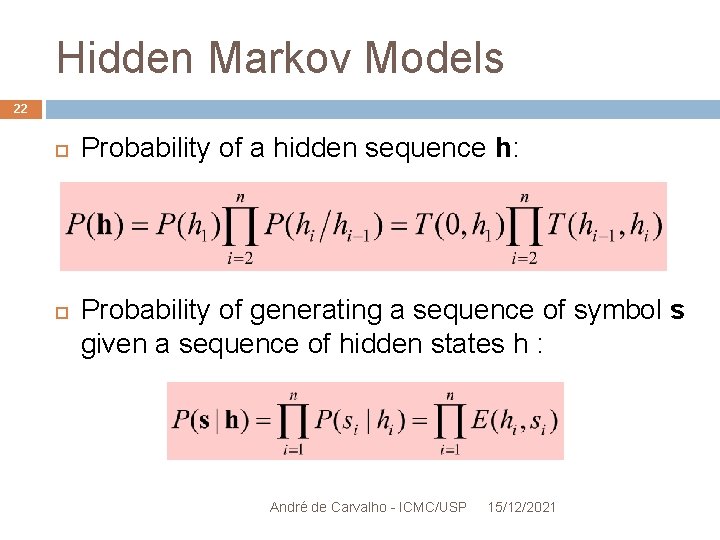
Hidden Markov Models 22 Probability of a hidden sequence h: Probability of generating a sequence of symbol s given a sequence of hidden states h : André de Carvalho - ICMC/USP 15/12/2021

Hidden Markov Models 23 If h is known (rare), joint probability of s and h : André de Carvalho - ICMC/USP 15/12/2021

Hidden Markov Models 24 If h is unknown (common) one can use theorem of total probability to calculate the probability of s: Todas as cadeias ocultas de tamanho n Grows exponentially with n André de Carvalho - ICMC/USP 15/12/2021

Hidden Markov Models 25 Most likely sequence Found with the Viterbi Algorithm (dynamic programming) André de Carvalho - ICMC/USP 15/12/2021

26 HMM – Applications in Bioinformatics Gary Churchill was the first to use HMM in genomics, in 1989 Segmentation of DNA sequences in zones of similar use of nucleotides Today: Segmentation Multiple Alignment Prediction of protein function Gene Discovery André de Carvalho - ICMC/USP 15/12/2021

Segmentation 27 Most common task The sequences (genes or proteins) may contain regions with distinct properties Infer the hidden states representing these regions, as well as determine its limits in order to: better annotation better comprehension of the dynamics of the sequence André de Carvalho - ICMC/USP 15/12/2021

Segmentation- Example 28 Genomic of the lambda bacteriophage It has long sections of sequence that are: Rich in GC Rich in AT HMM to segment the genome in regions with these characteristics André de Carvalho - ICMC/USP 15/12/2021

Segmentation- Example 29 Hidden states: " GC-rich" and "AT rich " Observable symbols: A, C , G and T Estimates the parameters: EM Algorithm Transition matrices and random initial emission Assuming 2 hidden states and four visible symbols André de Carvalho - ICMC/USP 15/12/2021
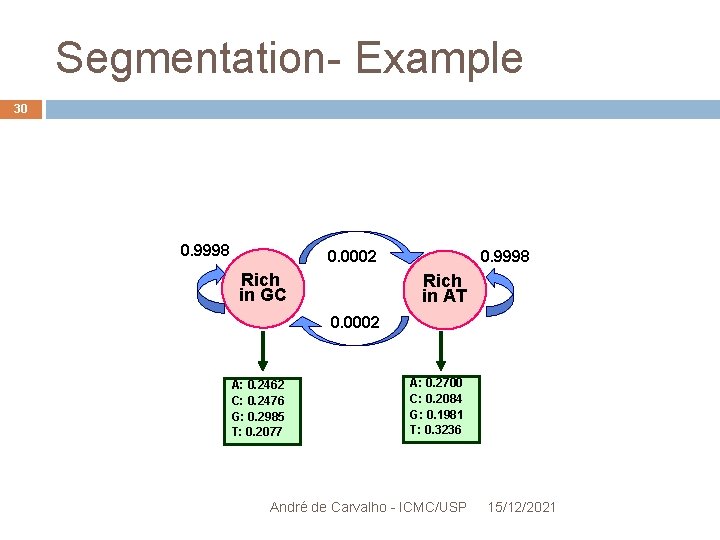
Segmentation- Example 30 0. 9998 0. 0002 Rich in GC 0. 9998 Rich in AT 0. 0002 A: 0. 2462 C: 0. 2476 G: 0. 2985 T: 0. 2077 A: 0. 2700 C: 0. 2084 G: 0. 1981 T: 0. 3236 André de Carvalho - ICMC/USP 15/12/2021
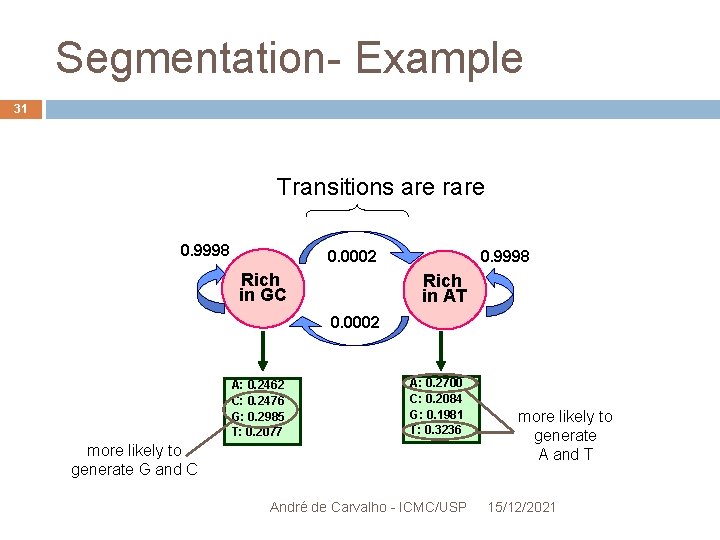
Segmentation- Example 31 Transitions are rare 0. 9998 0. 0002 Rich in GC 0. 9998 Rich in AT 0. 0002 A: 0. 2462 C: 0. 2476 G: 0. 2985 T: 0. 2077 A: 0. 2700 C: 0. 2084 G: 0. 1981 T: 0. 3236 more likely to generate G and C André de Carvalho - ICMC/USP more likely to generate A and T 15/12/2021

Segmentation- Example 32 Using the Viterbi Algorithm and the estimated model, we obtain the Segmentation of the sequence in the shown points: André de Carvalho - ICMC/USP 15/12/2021

Multiple Alignment Prediction of protein function 33 Uses HMM Profile (p. HMM) p. HMMs can be seen as: Abstract descriptions of a family of proteins Statistical summaries of a multiple alignment André de Carvalho - ICMC/USP 15/12/2021

HMM Profile 34 Encodes information on the frequency of residue and also insertions and deletions of each column of the multiple alignment Created from the multiple alignment of homologous sequences For each column in the alignment, the model has: State of Match (match) - distribution of residue State of insertion State deletion Each state of Match and Insertion has the Emission Matrix with the emission probabilities of each residue (amino acids or nucleotides) André de Carvalho - ICMC/USP 15/12/2021 Residues are not emitted in the states of deletion
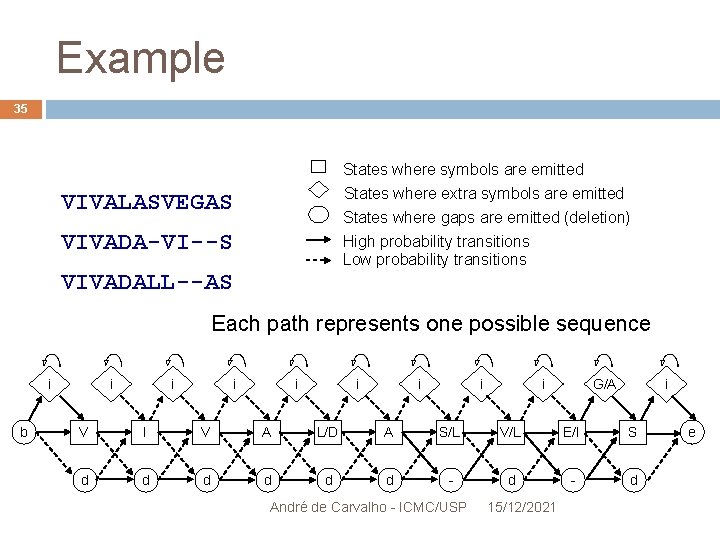
Example 35 States where symbols are emitted States where extra symbols are emitted VIVALASVEGAS States where gaps are emitted (deletion) VIVADA-VI--S High probability transitions Low probability transitions VIVADALL--AS Each path represents one possible sequence i b i i i i G/A i V I V A L/D A S/L V/L E/I S d d d - d André de Carvalho - ICMC/USP 15/12/2021 e

Example 36 p. HMM permite calcular o grau com que uma seqüência se ajusta ao modelo: p. HMM lets you calculate the degree to which a sequence fits the model: For each sequence passing through the model, can be assigned a probability or score Para cada sequência que passa pelo modelo, pode ser atribuída uma probabilidade ou pontuação Pontuação alta significa ajuste ao modelo High score means adjustment to the model Used to search. André a database by other members de Carvalho - ICMC/USP 15/12/2021

Example 37 How is it done? Using Blast for separating a protein database in families of related proteins Build a multiple alignment for each family Builds a HMM Profile and optimizes its parameters for each of the Multiple Alignment Aligns the target sequence with each of the p. HMM to find the best fit between the target sequence and one of p. HMM The family of the target sequence is the one used in p. HMM with best fit (most likely) Viterbi algorithm : computes the best path through the model Forward Algorithm: Calculates the sum of the André de Carvalho - ICMC/USP 15/12/2021 probabilities of all possible alignments

Gene Discovery 38 HMM allows to integrate several different signals, of probabilistic nature, in the search for genes Transcription factors binding sites ORFs Splice sites Start and stop codons With the constructed model, it uses the Viterbi algorithm to find genes (nucleotide sequence comprising the gene) André de Carvalho - ICMC/USP 15/12/2021

Case Study 39 Olfactory Receptors (OR) They belong to the family of proteins "7 -TM receptors" Para perceber as moléculas fora da célula e sinalizar sua descoberta dentro da célula os OR precisam atravessar a membrana da célula For recognizing the molecules out of the cell and signaling its discovery inside the cell, the ORs need to cross the cell membrane OR has 7 sections with highly hydrophobic amino acids alternating with hydrophilic regions that characterize the function of these proteins

Case Study 40 How to identify if a X protein belongs to a particular known family? Comparison of sequences of pairs is not sufficient to identify family members of the "7 -TM receptors" Alignment with the family p. HMM is more appropriate André de Carvalho - ICMC/USP 15/12/2021

Case Study 41 Comparison of protein X With a typical sequence of the "7 -TM receptors" family using global alignment Score : 54. 8 With the constructed p. HMM using thousands of aligned sequences Score : 154. 6 The signal indicating that the protein belongs to the family "7 -TM receptors" is much stronger using p. HMM André de Carvalho - ICMC/USP 15/12/2021

Case Study 42 Segmenting the OR in hydrophobic and hydrophilic regions HMM with 2 states: "within the membrane" and "outside the membrane" André de Carvalho - ICMC/USP 15/12/2021
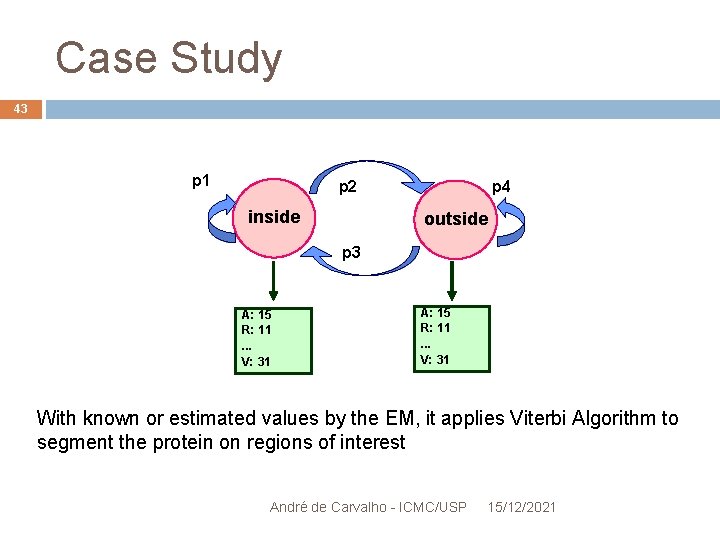
Case Study 43 p 1 p 2 inside p 4 outside p 3 A: 15 R: 11. . . V: 31 With known or estimated values by the EM, it applies Viterbi Algorithm to segment the protein on regions of interest André de Carvalho - ICMC/USP 15/12/2021
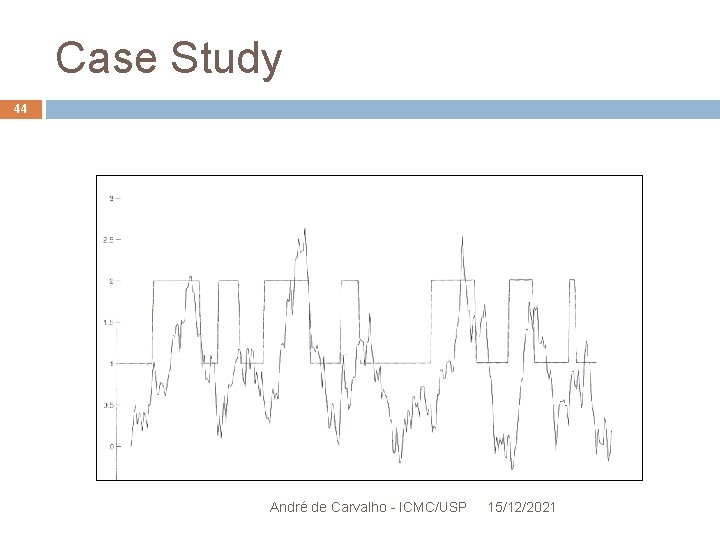
Case Study 44 André de Carvalho - ICMC/USP 15/12/2021

Viterbi Algorithm 45 Given a sequence s of size n and an HMM with parameters T and E, find the most likely hidden sequence Create a table V of size |H|x(n+1); Initialize i = 0; V(0, 0) = 1; V(k, 0) = 0 for k > 0; For i = 1: n, calculate each entry using this recursive relation V(j, i) = E(j, s(i)) * maxk {V(k, i-1) * T(k, j) } pointer(i, j) = arg maxk {V(k, i-1)*T(k, j) } Output: P(s, h*) = maxk {V(k, n)} Trace-back: i = n: 1, using: h*i-1 = pointer(i, h*i) Output: h*(n) = arg maxk {V(k, n)} André de Carvalho - ICMC/USP 15/12/2021

Forward Algorithm 46 Given a sequence s with size n and HMM parameters T and E, calculate the likelihood of a sequence Creates a F table with size |H|x(n+1); Initialize i = 0; F(0, 0) = 1; F(k, 0) = 0 for k > 0; For i = 1: n, calculates each entry using this recursive relation F(j, i) = E(j, s(i)) * ∑k {F(k, i-1) * T(k, j) } Output: P(s) = ∑k F(k, n) André de Carvalho - ICMC/USP 15/12/2021

47 EM Algorithm Expectation Maximization Given a sequence s of size n and an HMM with unknown parameters T and E , estimate the parameters 1. Initialize h, E and T; 2. Given s and h, estimate E and T only counting the symbols; 3. Given s, E and T, estimates h with the Viterbi algorithm, for example; 4. Repeat the steps 2 and 3 until any stop criterion is satisfied André de Carvalho - ICMC/USP 15/12/2021

Conclusion 48 Importance of the sequence alignment André de Carvalho - ICMC/USP 15/12/2021
Questions?
- Slides: 49