GENE RECOGNITION Dilvan Moreira based on Prof Andr

GENE RECOGNITION Dilvan Moreira (based on Prof. André Carvalho presentation)

Reading Introduction to Computational Genomics: A Case Studies Approach Chapter 2

Content 3 Proteins Gene Expression Transcription Genetic Code Translation Genes discovery André de Carvalho - ICMC/USP 09/09/2020

Introduction 4 Gene Sweepstake Betting system generated in 2000 by the debate about the human genome number of genes Dr. Ewan Birney had thought at the bar of the Cold Spring Harbor Laboratory (CSHL) Each scientist participant genome CSHL Genome Meeting could bet a number Winning number to be announced in May 2003 at CSHL Genome Meeting Betting ranged from 26, 000 to more than 150, 000 André de Carvalho - ICMC/USP 09/09/2020
Introduction 5 Betting value: 2000: US$ 1 2001: US$ 5 2002: US$ 20 Until April 2003 165 bets in more than 50 countries Average: 61. 170 genes André de Carvalho - ICMC/USP 09/09/2020

Introduction 6 Lee Rowen was the winner Systems Biology Institute, Seattle Bet in 2001: 25. 949 genes He won half of US$ 1. 200 and an autographed book from James Watson ¼ for the bet of 27. 462, made in 2000 ¼ for the bet 26. 500, made in 2002 André de Carvalho - ICMC/USP 09/09/2020

Introduction 7 Number of gene estimates: 1990: ~300, 000 1995: ~100, 000 2000: ~30, 000 2004: ~25, 000 2007: Known Number ~19, 000 André de Carvalho - ICMC/USP 09/09/2020

Proteins 8 Big molecules Composed (AAs) by one or more amino acid chains Polypeptide Sizes chain may vary: 30 -40 AAs 200 -300 AAs (most common) Tens of thousands of AAs Define the organism structure and operation André de Carvalho - ICMC/USP 09/09/2020

Human Protein Functions 9 Estructural function Participate of tissue structure Ex. 1: Colagen Ex. 2: Actin and Myosin High resistance protein, found in skin, cartilage , bone and tendons Contractile proteins, abundant in the muscles where participate of the mechanism of muscle contraction Transport function Ex. : Hemoglobin Transports oxygen in the blood André de Carvalho - ICMC/USP 09/09/2020

Human Protein Functions 10 Enzymatic function Regulates the biological reactions Ex. : Lipases Transforms lipids in fatty acid and glycerol Hormonal function Stimulates Ex. : or inhibits the activity of certain organs Insulin Controls cells the sugar transport in the blood to inside de André de Carvalho - ICMC/USP 09/09/2020

Gene Expression 11 The process by which genes are used to produce of proteins Some genes do not encode proteins RNA is the final product Gene expression mechanisms are different for organisms: Prokaryotes Diffuse genetic material in cells (Ex. : bacteria) Eukaryotes Genetic material in a nucleus (Ex. : humans) André de Carvalho - ICMC/USP 09/09/2020
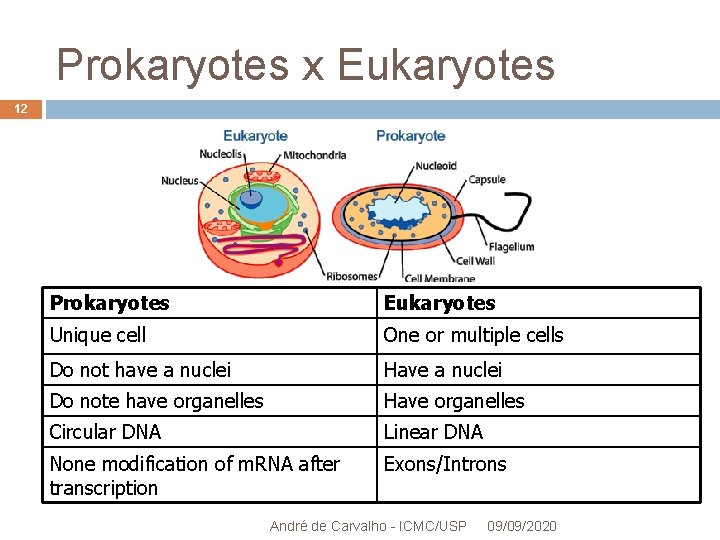
Prokaryotes x Eukaryotes 12 Prokaryotes Eukaryotes Unique cell One or multiple cells Do not have a nuclei Have a nuclei Do note have organelles Have organelles Circular DNA Linear DNA None modification of m. RNA after transcription Exons/Introns André de Carvalho - ICMC/USP 09/09/2020
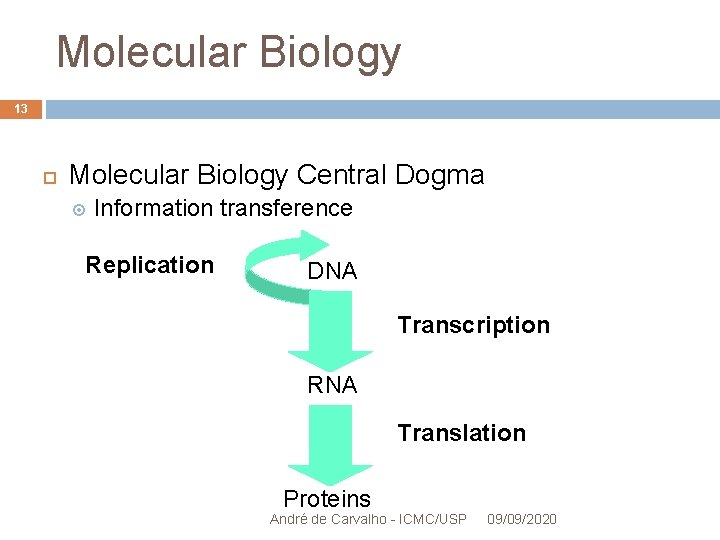
Molecular Biology 13 Molecular Biology Central Dogma Information transference Replication DNA Transcription RNA Translation Proteins André de Carvalho - ICMC/USP 09/09/2020
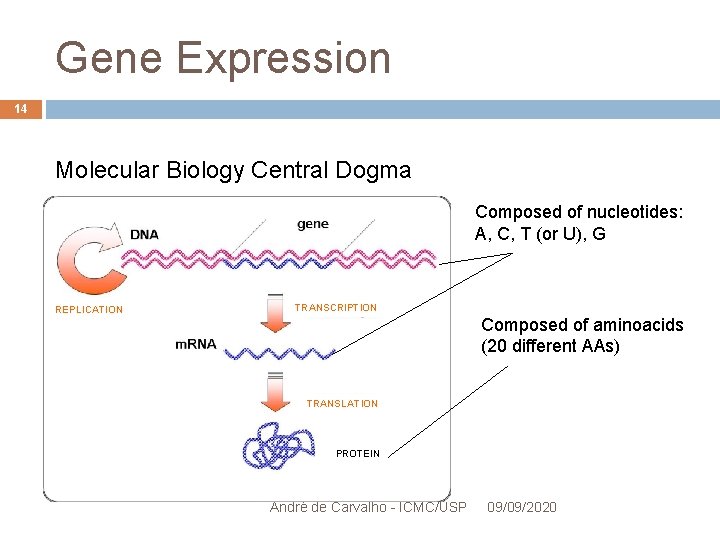
Gene Expression 14 Molecular Biology Central Dogma Composed of nucleotides: A, C, T (or U), G REPLICATION TRANSCRIPTION Composed of aminoacids (20 different AAs) TRANSLATION PROTEIN André de Carvalho - ICMC/USP 09/09/2020

Gene Expression 15 Some later findings contradict this dogma: RNA can replicates in some viruses and plants Viral RNA can be transcribed in DNA (enzyme reverse transcriptase) DNA can directly translate specific proteins - without transcription Some proteins can self-replicate (Prions) Causer of mad cow disease André de Carvalho - ICMC/USP 09/09/2020

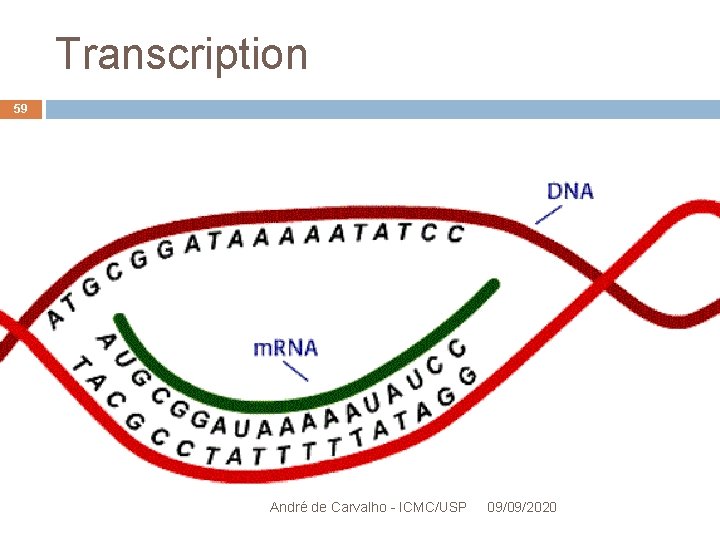
Transcription 16 Movie: Transcription de DNA Performed by RNA polymerase enzyme RNA polymerase begins the transcription after binding to a DNA regulatory signal Promoter or promoter region Produces messenger RNA molecules (m. RNA) Part of DNA transcribed into RNA = transcription unit André de Carvalho - ICMC/USP 09/09/2020

Transcription 17 Transcription process depends on the organism Eukaryote organisms Genes are transcribed independently There is a promoter before to each gene Prokaryote organisms Several consecutive genes can be transcribed into a single RNA molecule There is not necessarily a promoter before to each gene André de Carvalho - ICMC/USP 09/09/2020
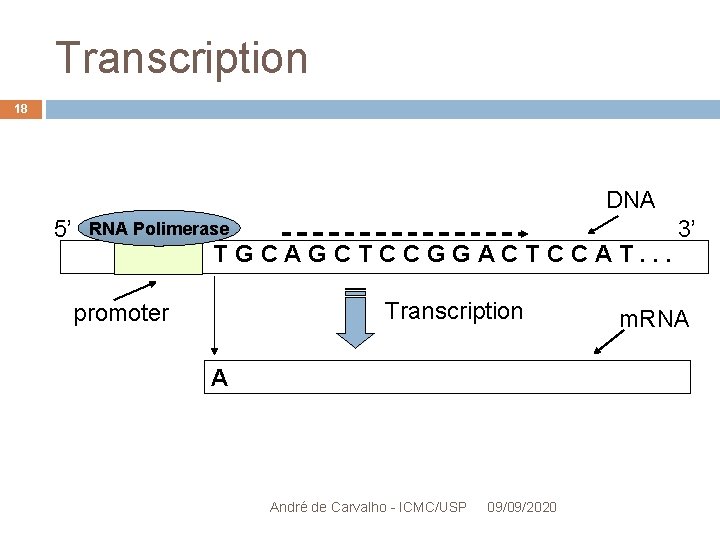
Transcription 18 DNA 5’ RNA Polimerase TGCAGCTCCGGACTCCAT. . . Transcription promoter A André de Carvalho - ICMC/USP 09/09/2020 3’ m. RNA
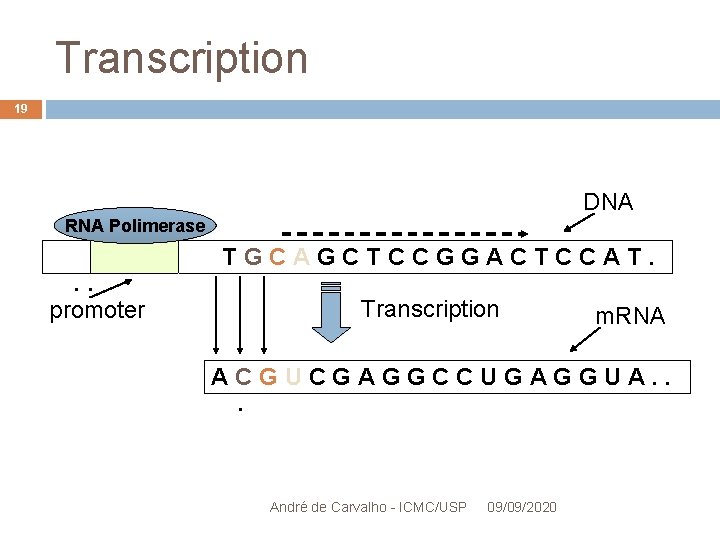
Transcription 19 DNA RNA Polimerase . . promoter TGCAGCTCCGGACTCCAT. Transcription m. RNA ACGUCGAGGCCUGAGGUA. . . André de Carvalho - ICMC/USP 09/09/2020

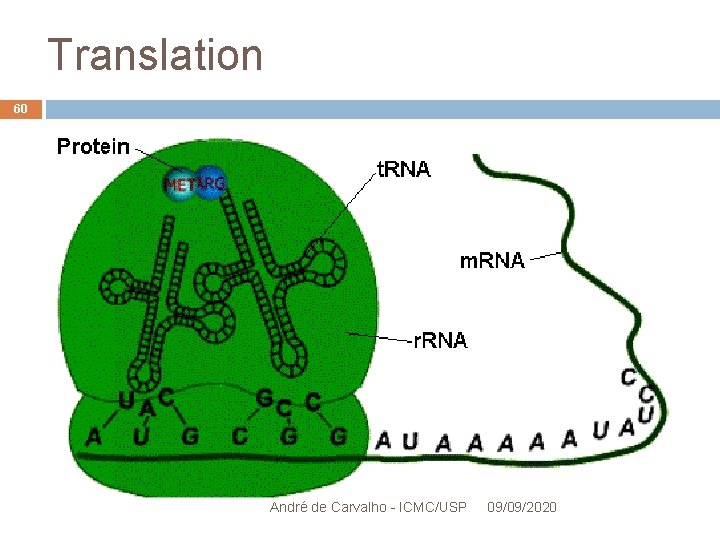
Translation 20 Movie: Translation RNA m. RNA reading is performed by a molecule of ribosome Read message is used to set up a chain of protein Genetic Code: set of rules for mapping DNA (RNA) into proteins André de Carvalho - ICMC/USP 09/09/2020

Translation 21 To encode 20 AAs 3 nucleotides are needed (codon): 41 = 4 AAs 42 = 16 AAs 43 = 64 AAs Genetic Code defines the amino acid codons mapping Nearly all living things use the same code (default code) Few organisms use a slightly different code André de Carvalho - ICMC/USP 09/09/2020
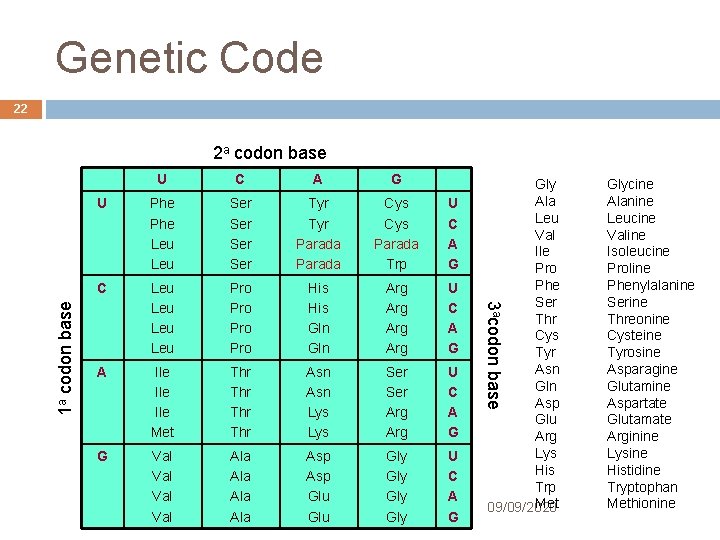
Genetic Code 22 U C A G U Phe Leu Ser Ser Tyr Parada Cys Parada Trp U C A G C Leu Leu Pro Pro His Gln Arg Arg U C A G A Ile Ile Met Thr Thr Asn Lys Ser Arg U C A G G Val Val Ala Ala Asp Gly U Asp Gly C Glu Gly A André de Carvalho - ICMC/USP Glu Gly G Gly Ala Leu Val Ile Pro Phe Ser Thr Cys Tyr Asn Gln Asp Glu Arg Lys His Trp Met 09/09/2020 3 acodon base 1 a codon base 2 a codon base Glycine Alanine Leucine Valine Isoleucine Proline Phenylalanine Serine Threonine Cysteine Tyrosine Asparagine Glutamine Aspartate Glutamate Arginine Lysine Histidine Tryptophan Methionine
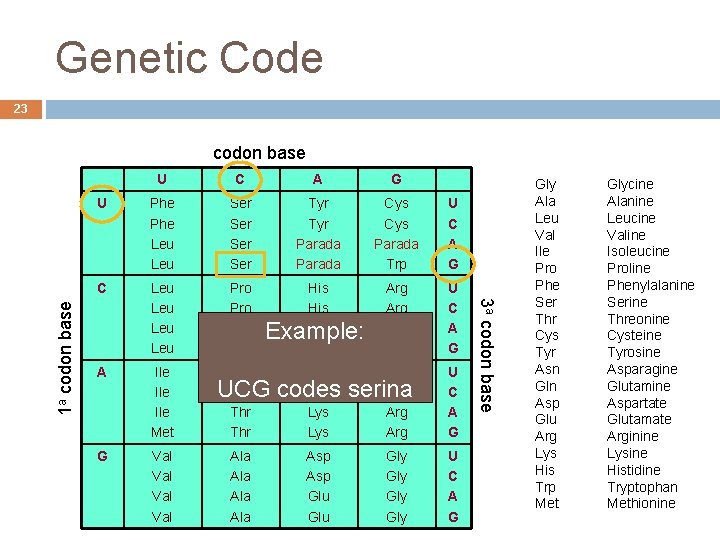
Genetic Code 23 U C A G U Phe Leu Ser Ser Tyr Parada Cys Parada Trp U C A G C Leu Leu Pro Pro His Gln Arg Arg U C A G A Ile Ile Met Thr Thr Asn Lys Ser Arg U C A G G Val Val Example: UCG codes serina Ala Ala Asp Gly U Asp Gly C Glu Gly A André de Carvalho - ICMC/USP Glu Gly G 3 a codon base 1 a codon base Gly Ala Leu Val Ile Pro Phe Ser Thr Cys Tyr Asn Gln Asp Glu Arg Lys His Trp Met Glycine Alanine Leucine Valine Isoleucine Proline Phenylalanine Serine Threonine Cysteine Tyrosine Asparagine Glutamine Aspartate Glutamate Arginine Lysine Histidine Tryptophan Methionine
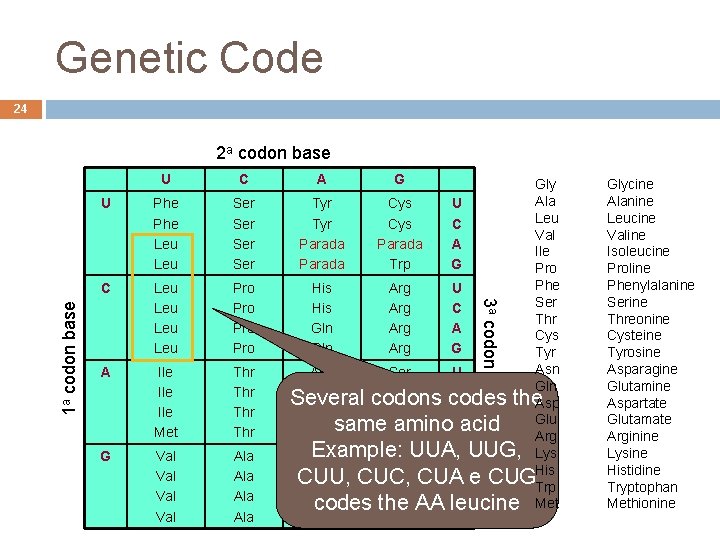
Genetic Code 24 U C U Phe Leu Ser Ser C Leu Leu Pro Pro A Ile Ile Met Thr Thr G Val Val Ala Ala A G Gly Ala Tyr Cys U Leu Tyr Cys C Val Parada A Ile Parada Trp G Pro Phe His Arg U Ser His Arg C Thr Gln Arg A Cys Gln Arg G Tyr Asn Ser U Gln Asn Ser C Several codons codes the. Asp Lys Arg A Glu same amino Lys Arg Gacid Arg Example: Asp Gly. UUA, U UUG, Lys Asp Gly CUA C e CUGHis CUU, CUC, Trp Glu Gly A Met codes the AA leucine André de Carvalho - ICMC/USP 09/09/2020 Glu Gly G 3 a codon base 1 a codon base 2 a codon base Glycine Alanine Leucine Valine Isoleucine Proline Phenylalanine Serine Threonine Cysteine Tyrosine Asparagine Glutamine Aspartate Glutamate Arginine Lysine Histidine Tryptophan Methionine
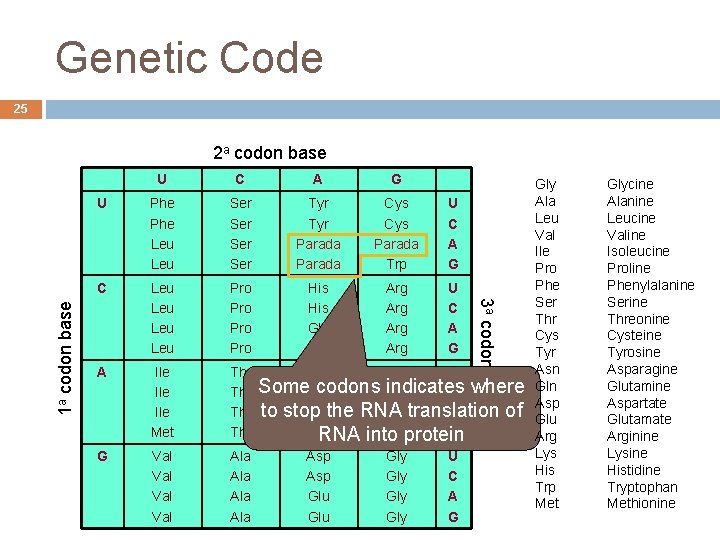
Genetic Code 25 U C A G U Phe Leu Ser Ser Tyr Parada Cys Parada Trp U C A G C Leu Leu Pro Pro His Gln Arg Arg U C A G A Ile Ile Met Thr Thr G Val Val Ala Ala 3 a codon base 1 a codon base 2 a codon base Asn Ser U Some. Asn codons Ser indicates C where to stop of Lysthe RNA Argtranslation A Lys Argprotein G RNA into Asp Gly U Asp Gly C Glu Gly A André de Carvalho - ICMC/USP Glu Gly G Gly Ala Leu Val Ile Pro Phe Ser Thr Cys Tyr Asn Gln Asp Glu Arg Lys His Trp Met Glycine Alanine Leucine Valine Isoleucine Proline Phenylalanine Serine Threonine Cysteine Tyrosine Asparagine Glutamine Aspartate Glutamate Arginine Lysine Histidine Tryptophan Methionine
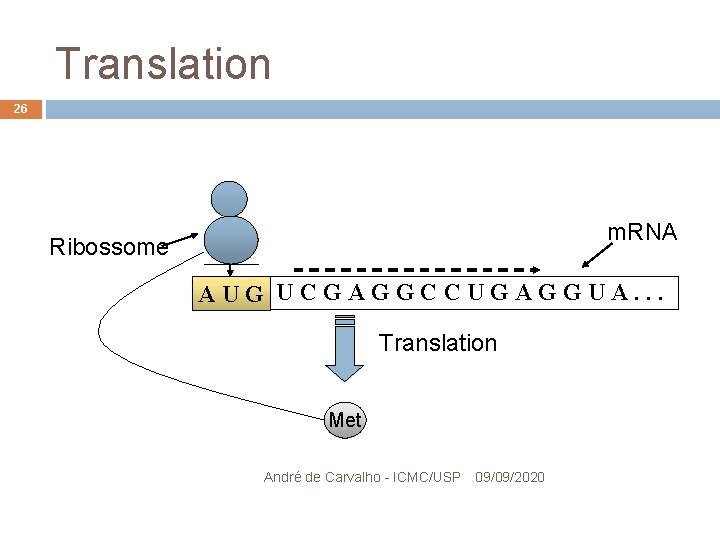
Translation 26 m. RNA Ribossome CG UCGAGGCCUGAGGUA. . . AU Translation Met André de Carvalho - ICMC/USP 09/09/2020
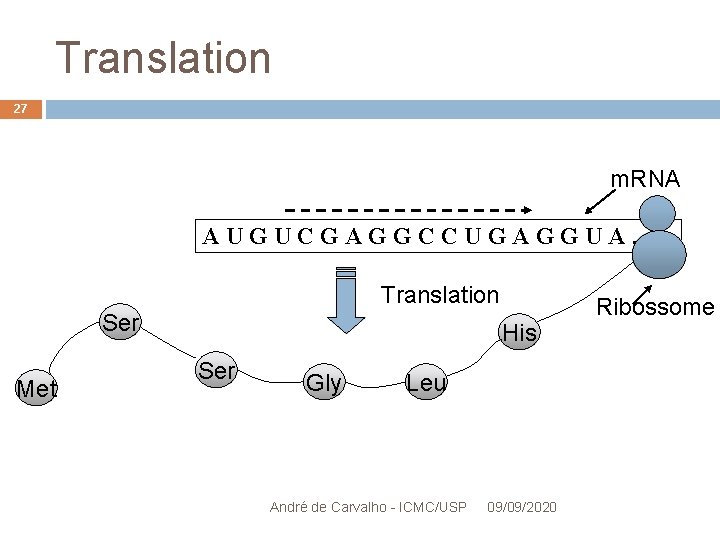
Translation 27 m. RNA AUGUCGAGGCCUGAGGUA. . . Translation Ser Met His Ser Gly Leu André de Carvalho - ICMC/USP 09/09/2020 Ribossome

Translation 28 m. RNA has, at both ends, regions which are not to be translated Untranslated The final position of translation is given by one of the termination codons Do regions – UTRs not encode amino acids But where to start translating ? André de Carvalho - ICMC/USP 09/09/2020
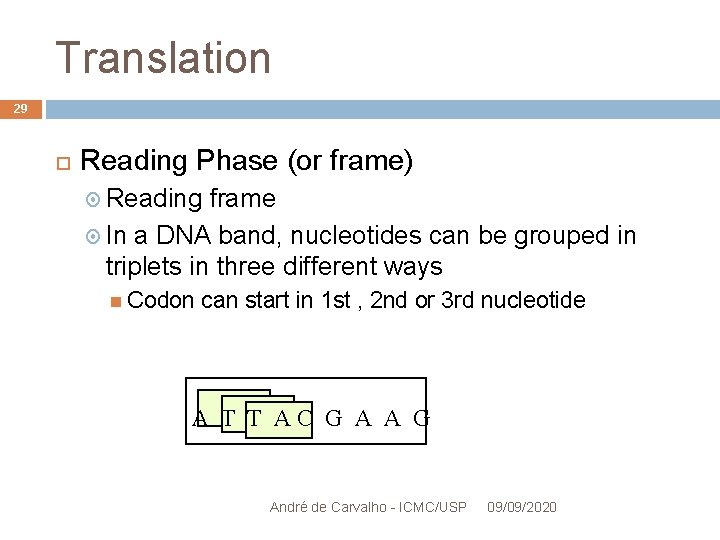
Translation 29 Reading Phase (or frame) Reading frame In a DNA band, nucleotides can be grouped in triplets in three different ways Codon can start in 1 st , 2 nd or 3 rd nucleotide A T T AC G A A G André de Carvalho - ICMC/USP 09/09/2020
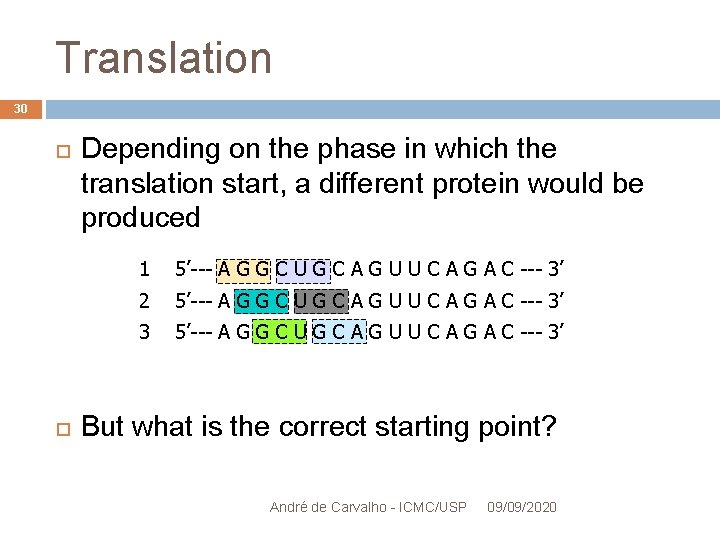
Translation 30 Depending on the phase in which the translation start, a different protein would be produced 1 5’--- A G G C U G C A G U U C A G A C --- 3’ 2 5’--- A G G C U G C A G U U C A G A C --- 3’ 3 5’--- A G G C U G C A G U U C A G A C --- 3’ But what is the correct starting point? André de Carvalho - ICMC/USP 09/09/2020

Translation 31 AUG codon Encodes methionine Specifies the beginning of translation and phase Reading Start codon In general, protein starts with a methionine Exception: GUG (valine) It occurs less frequently Protein synthesis in bacteria André de Carvalho - ICMC/USP 09/09/2020
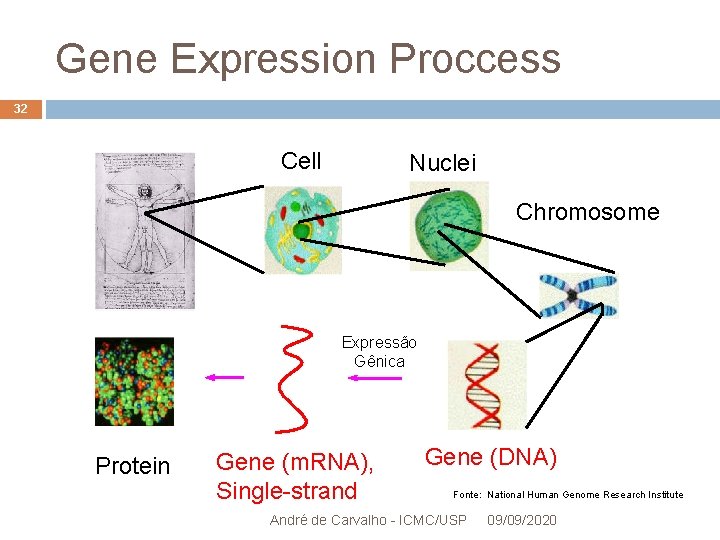
Gene Expression Proccess 32 Cell Nuclei Chromosome Expressão Gênica Protein Gene (m. RNA), Single-strand Gene (DNA) Fonte: National Human Genome Research Institute André de Carvalho - ICMC/USP 09/09/2020
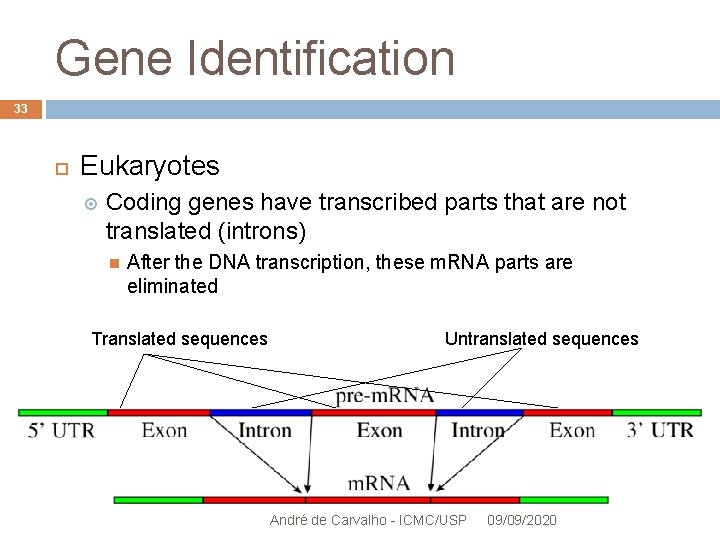
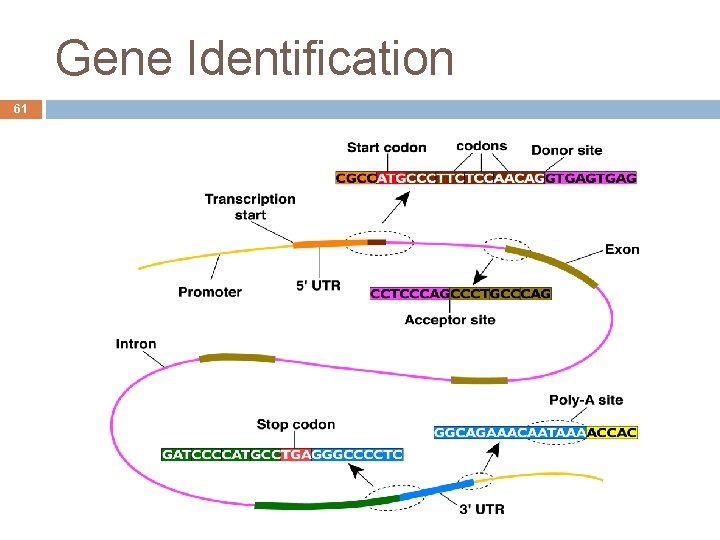
Gene Identification 33 Eukaryotes Coding genes have transcribed parts that are not translated (introns) After the DNA transcription, these m. RNA parts are eliminated Translated sequences Untranslated sequences André de Carvalho - ICMC/USP 09/09/2020

Gene Identification 34 Exons: Part of the gene that is transcribed and eventually translated Coding regions that can be translated into proteins 5' and 3' UTRs regions are exons, but are not translated They are about 2 % of the human genome They can consider like the data of a program André de Carvalho - ICMC/USP 09/09/2020

Gene Identification 35 Introns: Interspersed sequences that are eliminated in translation Non-coding regions Have regulatory functions (control) and structural integrity It can be seen as the logic of a program Human genome have much more control structure that the rice genome André de Carvalho - ICMC/USP 09/09/2020

Open Reading Frame 36 Open Reading Frame (ORF) DNA sequence of any size (multiple of 3) It begins with an initiation codon May have internal initiation codons Ends with a stop codon It has no internal termination codons It has the potential to encode a protein André de Carvalho - ICMC/USP 09/09/2020

Open Reading Frame 37 As the chromosomes are double-stranded, the gene may be in any one of the tapes Always from 5’ a 3’ 3 Reading Frames can be identified on each strand = 6 frames Identification Algorithm in searches for ORFs on both strands A strand can be easily obtained from the other DNA databases store only one strand André de Carvalho - ICMC/USP 09/09/2020

Gene Identification 38 Can an ORF be used to find potential genes? Only in Prokaryotes Because only the genes of Prokaryotes are unique continuous ORFs In Eukaryotes. . . It's complicated Researchers find candidate genes Potential Genes André de Carvalho - ICMC/USP 09/09/2020

Gene Identification 39 Methods that can be used to find genes : In Prokaryotes : Simple methods based on statistical properties of the sequence In Eukaryotes: Methods based on alignment of sequences or Markov sequence models André de Carvalho - ICMC/USP 09/09/2020

Gene Identification André de Carvalho ICMC/USP Prokaryotes Small genomes: 0. 5 – 10· 106 bp High density of codification (> 90%) Easy identification of genes Accuracy 99% Problems Eukaryotes Overlap of ORFs Short genes Large genomes: 107 – 1010 bp Low density of codification (< 50%) Complex identification of genes Accuracy 50% Problems Several 40 09/09/2020

Gene Identification 41 Algorithm to find ORFs: Given a string s and a positive value k For each phase Reading Dividing the DNA sequence into sections with 3 bases Find all triples excerpts that: Starting with a start codon and ending with a termination codon Repeat for the reverse complement of the sequence Back ORFs of greater length k André de Carvalho - ICMC/USP 09/09/2020

Example 42 ORFs in M. genitalium genome Algorithm with different values for k K = 90 Find 543 ORFs in the genome K = 100 Only accepted candidate genes with more than k Aas Find 471 ORFs in the genome Original paper of the genome cites 470 ORFs Including untranslated RNA genes (not detected by previous algorithm) André de Carvalho - ICMC/USP 09/09/2020

Gene Identification 43 There may be many parts of DNA containing ORF features randomly How to know if an ORF is a good candidate gene? André de Carvalho - ICMC/USP 09/09/2020

Gene Identification 44 Hypothesis Test Calculate the probability of finding a L length ORF in a random sequence Make inferences based on that probability An ORF is significant when it is highly unlikely under a null model (Null Model) André de Carvalho - ICMC/USP 09/09/2020

Gene Identification 45 Hypotheses: H 0: ORF was generated by a random process H 1: ORF was generated by a biologically relevant process André de Carvalho - ICMC/USP 09/09/2020

Gene Identification 46 p-value: Probability of obtaining a value of the tested statistic (L, for example) that is the value randomly observed, if H 0 is true L = length of an ORF It is compared with a significance level chosen If less than , H 1 is accepted and the ORF is considered significant Otherwise it is rejected André de Carvalho - ICMC/USP 09/09/2020

Gene Identification 47 Determine a minimum length L needed to have an ORF to be a candidate gene What is the probability of an ORF length greater than or equal to L arise randomly? Which is the threshold for L such that 95% of the random ORFs are smaller than L? André de Carvalho - ICMC/USP 09/09/2020

Gene Identification 48 Probability of obtaining a termination codon (uniform distribution of codons ): 3/64 Probability of codons that are not stop codon: 61/64 Probability of a succession of L or more nonterminal codons after a start codon: (61/64)L André de Carvalho - ICMC/USP 09/09/2020

Gene Identification 49 Using =0, 05 we can estimate the minimum acceptable size of an ORF: Like (61/64)62 = 0. 051 95% of spurious ORFs will be removed if ORFs with L 64 are discarded 62 + 64 Initiation codon + stop codon André de Carvalho - ICMC/USP 09/09/2020

Gene Identification 50 However, the distribution of bases in most organisms is not uniform Estimates the frequencies starting from the same sequence: Ptermination = P(TAA) + P(TAG) + P(TGA) P(L no stop codons) = (1 - Ptermination)L For a given , estimate L in the same manner described above André de Carvalho - ICMC/USP 09/09/2020

Gene Identification 51 And if it is not possible to calculate the exact pvalue? Theoretical or computational reasons Generate sequences with the same properties of the data using a technique of " randomization": Permutation of the original sequence Bootstrapping (sample with replacement) Using these sequences to calculate a null distribution The p-value is calculated by finding the rank of L in the null distribution André de Carvalho - ICMC/USP 09/09/2020

52 Example: Mycoplasma genitalium Original sequence : 11. 922 ORFs Method used : base permutation Makes permutation, search the ORFs and stores their lengths On the "randomized“ sequence: 17. 367 ORFs H 0 = "randomized“ sequence André de Carvalho - ICMC/USP 09/09/2020

53 Example: Mycoplasma genitalium Maintain as candidate genes the ORFs of the real sequence major than the ORFs of random sequence The maximum length of the ORFs in random sequence was 402 bp Estimated number of ORFs in the real sequence (> 402 bp) was 326 Next the number of really existing genes, 470 André de Carvalho - ICMC/USP 09/09/2020
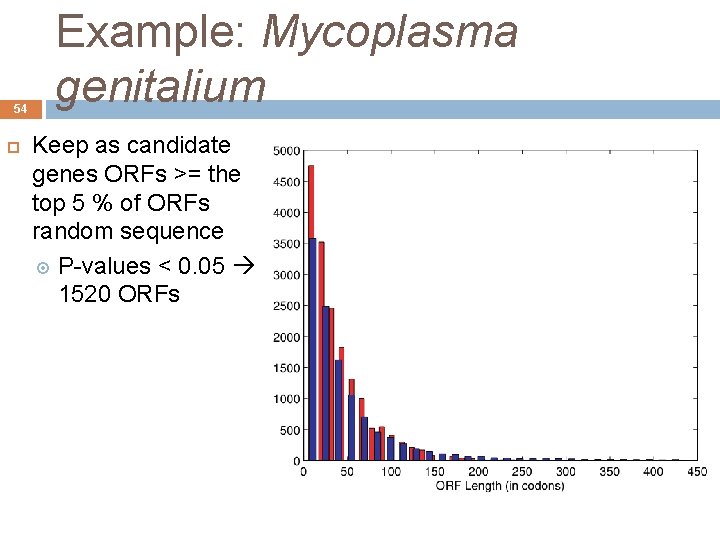
54 Example: Mycoplasma genitalium Keep as candidate genes ORFs >= the top 5 % of ORFs random sequence P-values < 0. 05 1520 ORFs

Conclusion 55 Recognition of genes in DNA sequences Costly process in laboratories Simple Technique Prokaryotes X Eukaryotes More sophisticated techniques confirm or reject candidates Comparison with known sequences André de Carvalho - ICMC/USP 09/09/2020
Questions?
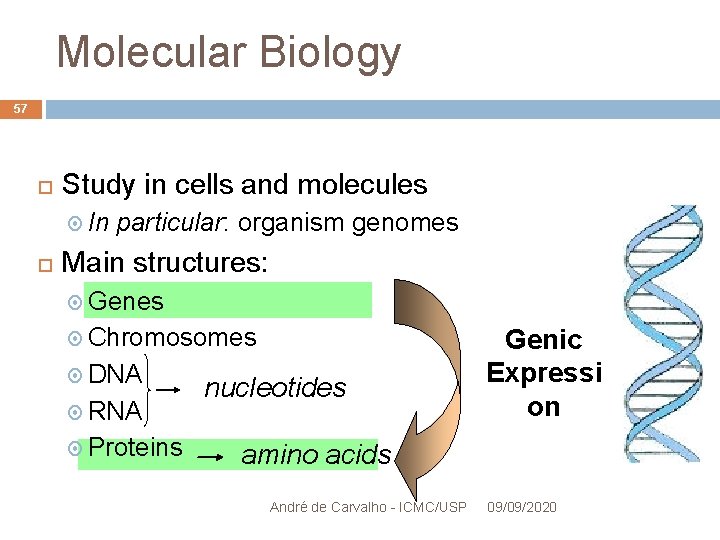
Molecular Biology 57 Study in cells and molecules In particular: organism genomes Main structures: Genes Chromosomes DNA RNA Proteins nucleotides Genic Expressi on amino acids André de Carvalho - ICMC/USP 09/09/2020
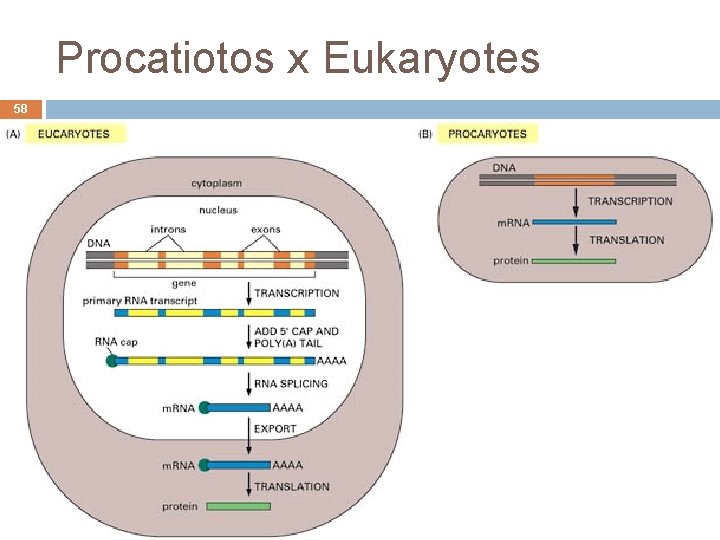
Procatiotos x Eukaryotes 58 André de Carvalho - ICMC/USP 09/09/2020
Transcription 59 André de Carvalho - ICMC/USP 09/09/2020
Translation 60 André de Carvalho - ICMC/USP 09/09/2020
Gene Identification 61 André de Carvalho - ICMC/USP 09/09/2020
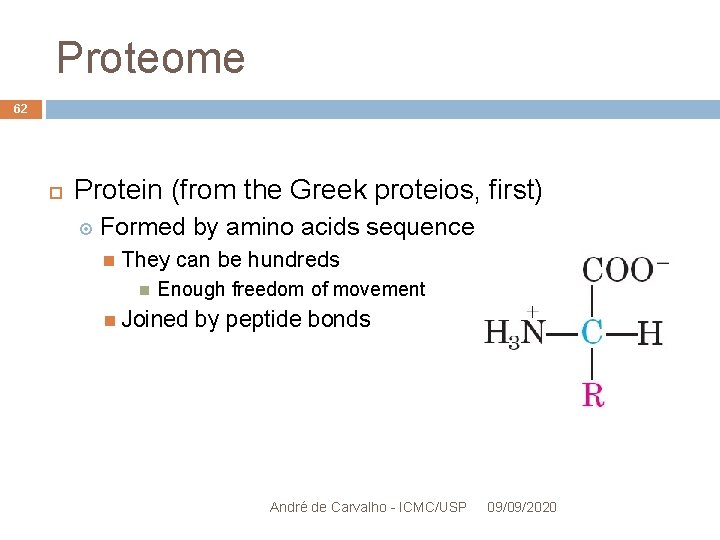
Proteome 62 Protein (from the Greek proteios, first) Formed by amino acids sequence They can be hundreds Enough freedom of movement Joined by peptide bonds André de Carvalho - ICMC/USP 09/09/2020
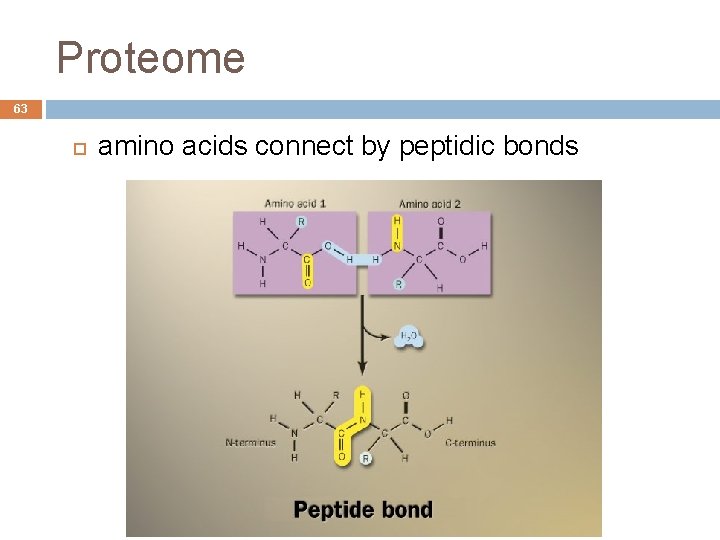
Proteome 63 amino acids connect by peptidic bonds André de Carvalho - ICMC/USP 09/09/2020

Protein 64 Can bend in different 3 -dimensional shapes Folding is fast (about 2 seconds) and consistent Structure of a protein determines what it does Enzymes Cell signaling Antibodies André de Carvalho - ICMC/USP 09/09/2020

Protein 65 Structure can be described according to 4 levels or structures Primary Secondary Tertiary Quaternary André de Carvalho - ICMC/USP 09/09/2020

Protein 66 Primary structure Amino acids sequence comprise the polypeptide chain Exact order of the amino acids in a protein constitutes its primary structure F–P–A–V–A–F Proteins spontaneously fold itself Assuming a three -dimensional shape Form depends on the sequence of amino acids Form of protein defines function André de Carvalho - ICMC/USP 09/09/2020

Proteins 67 Secondary structure Represents the regular patterns and local repetitive Found Two in the protein folding common local arrangements in proteins: Alpha helix Beta-sheet André de Carvalho - ICMC/USP 09/09/2020

Proteins André de Carvalho ICMC/USP Alpha helix Beta-sheet 68 09/09/2020

Proteins 69 Tertiary structure Sequential combination of secondary structures It describes how the protein folding occurs in 3 dimensional space Folding global result of all the polypeptide chain Defines the protein format Enzymes usually have a compact globular form André de Carvalho - ICMC/USP 09/09/2020
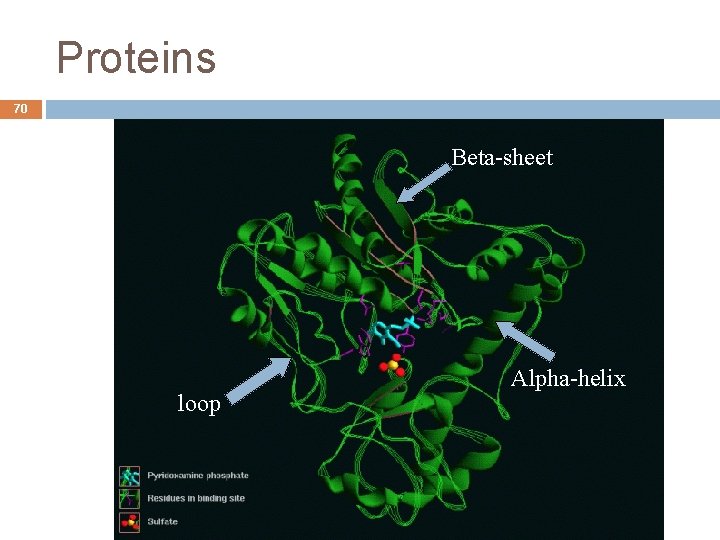
Proteins 70 Beta-sheet Alpha-helix loop André de Carvalho - ICMC/USP 09/09/2020

Proteins 71 Quaternary structure Many proteins are formed of more than one polypeptide chain The quaternary structure describes how the different subunits joined and fit To form the complete structure of the protein Ex. : the molecule of human hemoglobin is composed of four subunits André de Carvalho - ICMC/USP 09/09/2020
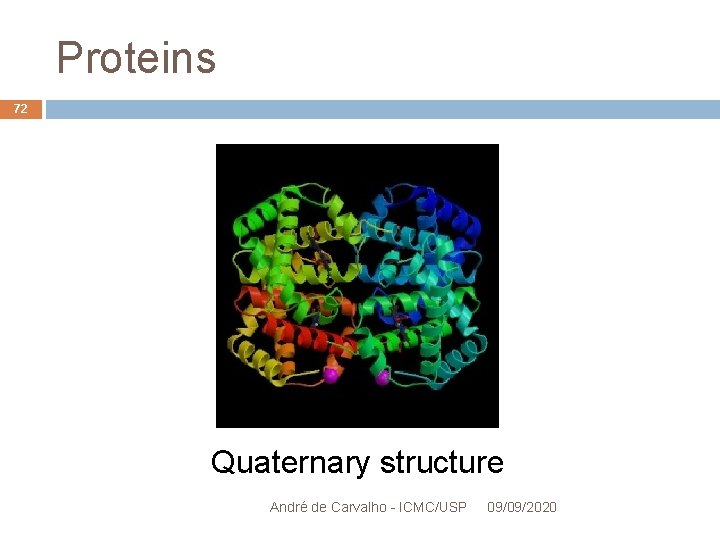
Proteins 72 Quaternary structure André de Carvalho - ICMC/USP 09/09/2020

Transposons 73 Mobile DNA Segments They may move to different regions or replicate within the genome Jumping genes Occupy a large portion of the genome Present in almost all organisms Effects: Cause mutations Increase or reduce the amount of DNA in the genome André de Carvalho - ICMC/USP 09/09/2020
Questions?
- Slides: 74