COMPLETE GENOME COMPARISON Dilvan Moreira Baseado em material

COMPLETE GENOME COMPARISON Dilvan Moreira (Baseado em material do prof. André Carvalho)

Reading Introduction to Computational Genomics: A Case Studies Approach Chapter 8

Conteúdo 3 Genome Comparison Symbiosis Chlamydias Genomics with a bag of genes Synteny Distance among homologous regions André de Carvalho - ICMC/USP 02/10/2020

Introduction 4 Humans have multiple bacteria species living inside them Most of them are no harmful (Symbiosis – most of times mutualism) Relationship mutually advantageous among two or more organisms from different species They help to digest food, or give us vitamins we do not produce by ourselves Ex. E. coli receives nutrients and produces K vitamine Its presence in our body prevents infections by other pathogens André de Carvalho - ICMC/USP 02/10/2020

Introduction 5 There are several examples of symbiotics relationships on nature Bees and flowers Cattle and cattle egret Shark and remora Fungi and plants (bracken, orchid) Crocodile and plover bird André de Carvalho - ICMC/USP 02/10/2020

Introduction 6 Some Symbionts are invisible, because they live inside the host E. coli They help the digestion of the host They benefit by living in the stomach of a mobile organism Termite and Tryconinpha Protozoa living in the digestive tract help the wood digestion (produces the celulase enzime) Its main source of nutrients André de Carvalho - ICMC/USP 02/10/2020

Introduction 7 Some species settled permanently in the host cells Becoming completely dependent for getting nutrients During the evolution process, big changes happened in the genome of these species André de Carvalho - ICMC/USP 02/10/2020

Introduction 8 Why does it occur? Natural selection does note promotes the maintenance of the function of these genes Mutations that destroy them are ignored Result: Symbiont intracellular genomes are among the smaller known genomes In size and number of genes André de Carvalho - ICMC/USP 02/10/2020

Chlamydias 9 Chlamydia trachomatis Symbiont intracellular bacteria intracelular in humans Parasite Does Main not beneficiate the host cause of STDs in USA 2 milions of new infections per year André de Carvalho - ICMC/USP 02/10/2020

Chlamydias 10 Chlamydia trachomatis Lost the capacity of generate several biochemical products They are able to live only in specific human cells Can not be cultivated in laboratory Present in urinary system André de Carvalho - ICMC/USP 02/10/2020
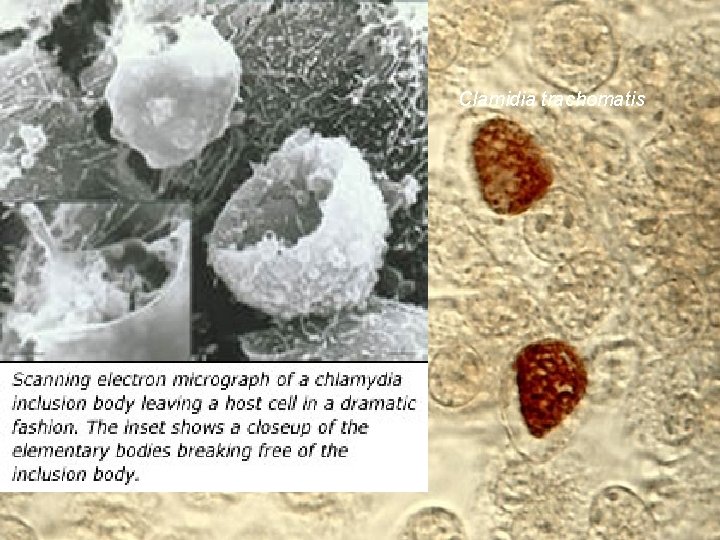
Clamidia trachomatis

Chlamydias 12 Chlamydia pneumoniae Symbiont intracellular bacteria in humans Parasite, infects cells from the superior and inferior Respiratory tract Causes pneumonia and bronchitis Possible association with the artherosclerosis process in heart arteries Hardened, narrowed and no elastic myocardial ischemia André de Carvalho - ICMC/USP 02/10/2020
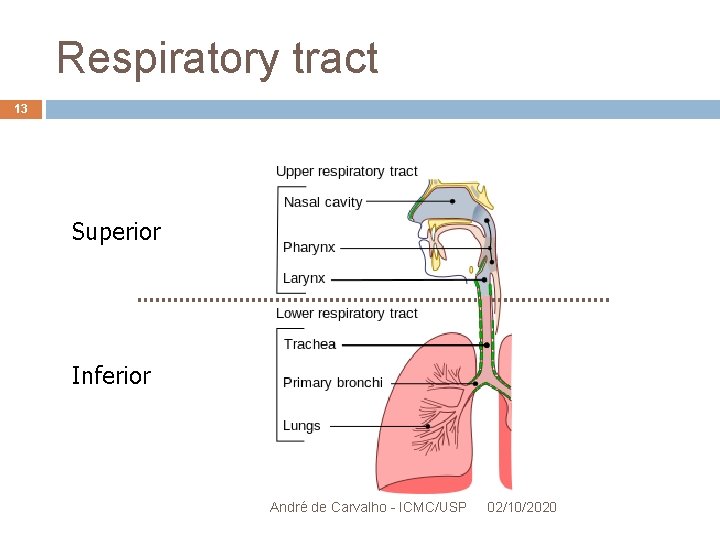
Respiratory tract 13 Superior Inferior André de Carvalho - ICMC/USP 02/10/2020
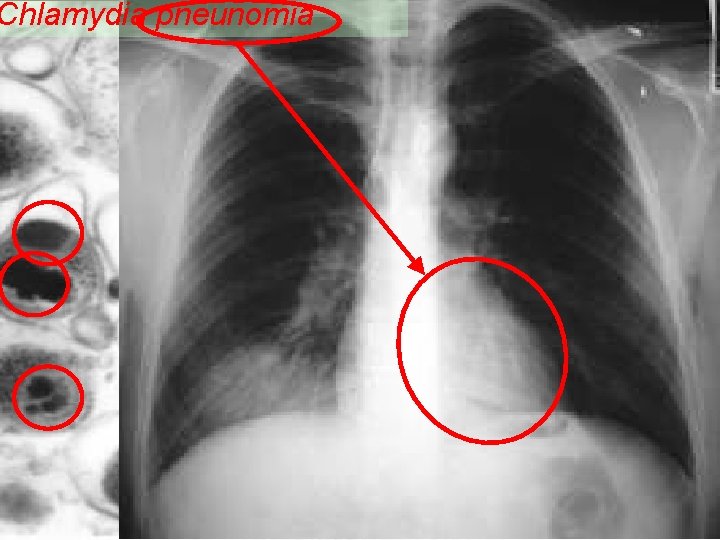
Chlamydia pneunomia Human Respiratory Tract

Chlamydias 15 Both C. pneumoniae and C. trachomatis have Metabolic and biosynthetic functions reduced Very small genome (both have 1 Mb in size) André de Carvalho - ICMC/USP 02/10/2020

Chlamydias 16 Six different species have already been completely sequenced They But They are all mandatorily intracelular symbiont not all of them are harmful to the host have: Small differences in the number and identity of lost genes since the common ancestral Apparently, the intracellular life of Chlamydia started 700 million years ago With the arising of the first eukaryote organisms André de Carvalho - ICMC/USP 02/10/2020

Chlamydias 17 Mitochondria and chloroplasts may originate from symbiont intracellular bacteria They lost so many genes that they became part of eukariotic cells Chlamydias Also have a symbiotic relationship with eukaryote hosts They seem to be strongly related with the chloroplast ancestors André de Carvalho - ICMC/USP 02/10/2020

Genome Comparison 18 Challenges related to the Complete Genome Comparison Chlamydias species genomes are used as study cases Small sizes Relatively low rated of genome evolution They allow to answer possible questions only with the complete comparison André de Carvalho - ICMC/USP 02/10/2020

Genome Comparison 19 Analyze differences among the whole set of genes from two genomes Generates new knowledge in: Genome evolution Sequence function Gene function Comparison of DNA sequences using different resolution comparing to the previously used André de Carvalho - ICMC/USP 02/10/2020

Genome Comparison 20 Genetic variability Most of the time, due to a nucleotide polymorphism Insertions, deletions and substitutions Frequently, non local transformations are observed inside of and between species Transfer of long parts among species is more frequent than we imagined 20% of E. coli genome is due to horizontal transfer André de Carvalho - ICMC/USP 02/10/2020

Genome Comparison 21 Sequence rearrange by shuffling inside a genome is very common too Inversions: inversion of a whole DNA segment Transpositions: DNA is cut from a genome’s place and pasted on another (cut and paste) Chromosomes may break and the pieces may rearrange in new combinations Complete genomes may be duplicated, generating polyploidy individuals André de Carvalho - ICMC/USP 02/10/2020

Genome Comparison 22 Set of genes in a genome may change without the drastic effects of big Deletions Duplications Horizontal transfer Simple genes are frequently acquired or lost by Duplications and Deletions in small scale André de Carvalho - ICMC/USP 02/10/2020
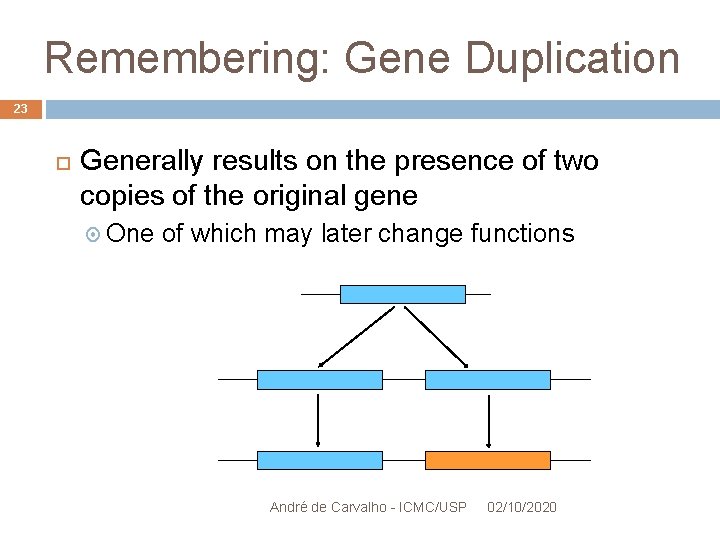
Remembering: Gene Duplication 23 Generally results on the presence of two copies of the original gene One of which may later change functions André de Carvalho - ICMC/USP 02/10/2020

Effects of Gene Duplication Normal fly Fly with an extra pair of wings Fly with a pair of antenna changes into an extra pair of legs 02/10/2020 André de Carvalho - ICMC/USP 24

Gene Deletion 25 Small deletions may transform functional genes into pseudogenes As the structure shift This lost is very common Genes have redundant functions Changes in the environment made the gene unnecessary Also the opposite Gain of genes André de Carvalho - ICMC/USP 02/10/2020

Genome Comparison 26 Lost and gain of genes may result in big difference among genomes Number of genes Role of the genes in the genomes Use of the “Genomics with a bag of genes” approach André de Carvalho - ICMC/USP 02/10/2020

Genomics with a bag of genes 27 Variety of rearrange due to evolution difficults the alignment of two genomes Inversions, Transpositions, Duplications, . . . Most of variations: Do not bring relevant information Will be difficult to interpretate Most of times, the alignment will be impossible André de Carvalho - ICMC/USP 02/10/2020

Genomics with a bag of genes 28 Complete genome alignment does not work GAC ACTTTTTGG GGG TATATA CATGT AAATAAT CG AACCCCCG GAC ACTTTTTGG TATATA CATGT GGG CATGT AAATAAT CG AACCCCG Inversion Transposition Duplication André de Carvalho - ICMC/USP Deletion 02/10/2020

Genomics with a bag of genes 29 Division into new chromosomes Gene shuffling among the chromosomes GAC ACTTTTTGG GGG TATATA CATGT AAATAAT CG AACCCCCG GAC ACTTTTTGG TATATA CATGT GGG CATGT AAATAAT CG AACCCCG Inversion Transposition Duplication André de Carvalho - ICMC/USP Deletion 02/10/2020

Genomics with a bag of genes 30 Complete Genome Comparison does not work Approach more manageable and informative: Comparison of the individual genes of each genome Break the genomes into pieces (genes) Combine the comparisons of the pieces André de Carvalho - ICMC/USP 02/10/2020

Genomics with a bag of genes 31 Comparison of two genomes Find genes that are in both of them Use ORF-finder with threshold of 100 codons Ex. : Organismo C. trachomatis C. pneumonia E. coli Tamanho 1 042 519 1 229 853 - André de Carvalho - ICMC/USP ORFs 916 1048 5000 02/10/2020

Genomics with a bag of genes 32 To descover genes and routes that were lost in the Chlamydia evolution: Identify the genes in each species CP and CT lost several genes Parasite the host and steal several products of its genes It is expected that different biological processes for life in respiratory and urinary systems Difference in genomes can identify the function of the remaining genes André de Carvalho - ICMC/USP 02/10/2020

Genomics with a bag of genes 33 What is the most appropriate method for comparing acquired and lost genes? Identify genes in each genome and calculate similarity in relation to the other genes Information about the similarity among all pairs of genes is necessary to: Study nucleotide substitutions among orthologous genes Find blocks of genes with conserved order Analyze changes in the size of the gene families André de Carvalho - ICMC/USP 02/10/2020

Genomics with a bag of genes 34 To generate similarity scores Use amino acid sequences of all genes from both genomes Create n x m matrix with alignment value between each pair of genes NW Algorithm may be used, by normalizing the result using the sequences size For major sequences, use faster algorithms but less accurate (BLAST) André de Carvalho - ICMC/USP 02/10/2020

Genomics with a bag of genes 35 Duplications can generate large families of related genes Relationship between these homologous genes can be quite complex Two main types of relationship: paralogs orthologs Matrix can be used to distinguish paralogs and orthologs genes André de Carvalho - ICMC/USP 02/10/2020

Genomics with a bag of genes 36 paralogs: Genes that arise from Duplication and subsequent specialization All genes in a family of genes from the same genome orthologs Genes are not related by Duplication only by specialization Used to estimate the number of replacements And time between species for reconstructing phylogenetic trees André de Carvalho - ICMC/USP 02/10/2020

Identification of orthologs 37 BRHs (Best Reciprocal Hits) Better mutual similarity Most common method ORFs pair is BRH if each one is the best match for each other ORF may not belong to any pair BHR ORF may have ortholog in other species and a paralog in the same genome André de Carvalho - ICMC/USP 02/10/2020

Identification of orthologs 38 Similarity matrix BRH orthologs Using threshold equal to 100 codons 1964 ORFs (916 CT and 1048 CP) 728 orthologous pairs 126 paralogs pairs (56 CT and 70 CP) Paralogs pairs are more similar to each other that pairs of orthologous Result of duplication of genes after the species split 253 remaining ORFs can be paralogs or older genes lost in the other species by specialization André de Carvalho - ICMC/USP 02/10/2020

Gene Family Identification 39 At first, all genes have a common ancestor and are from the same family Family: group of related genes that probably have similar functions It has a common ancestor that is fairly recent There is no specific rule Rule used: genes belong to the same family have identity 50% Depending on the species which are found, these genes are considered paralogs or orthologs André de Carvalho - ICMC/USP 02/10/2020

Gene Family Identification 40 Changes in the size of the gene family are useful for studying gene function and evolution Expansion and contraction of gene families helps determine differences between species Gene Family Identification equivalent between genomes Group any gene, regardless of the genome After defining clusters (families), count the gene number in each family that came from each genome André de Carvalho - ICMC/USP 02/10/2020

Gene Family Identification 41 Hierarchical clustering algorithm Uses similarity matrix to define distance between genes NJ algorithm or UPGMA Clusters found depend on the values of the algorithm parameters André de Carvalho - ICMC/USP 02/10/2020
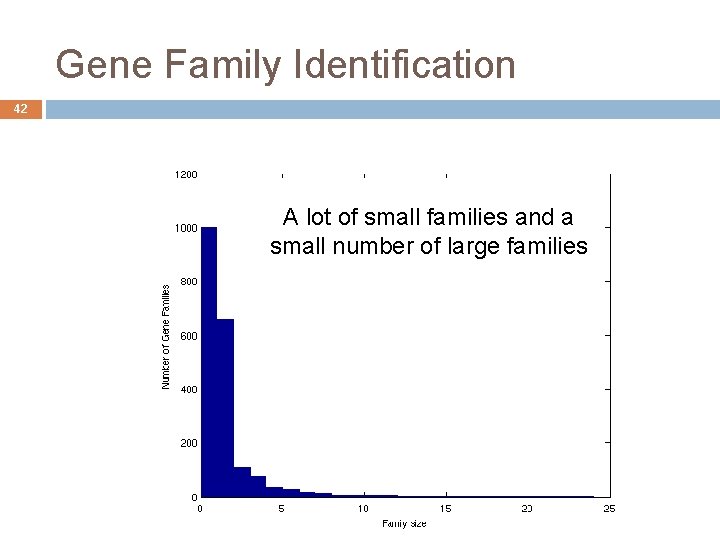
Gene Family Identification 42 A lot of small families and a small number of large families André de Carvalho - ICMC/USP 02/10/2020

Gene Family Identification 43 The largest families found to CT and CP were: CT 12 6 CP 12 15 9 9 10 10 Function ABC transporters Outer membrane G protein family Unknown André de Carvalho - ICMC/USP 02/10/2020

Gene Family Identification 44 ABC transformers are transmembrane proteins with binding region at both sites ATP-binding cassette They have an important role one transportation inside and outside the cell All cells acquire molecules and ions that they need in the external environment through the membrane Very old, they are almost identical in all organisms Family includes more than 30 proteins André de Carvalho - ICMC/USP 02/10/2020
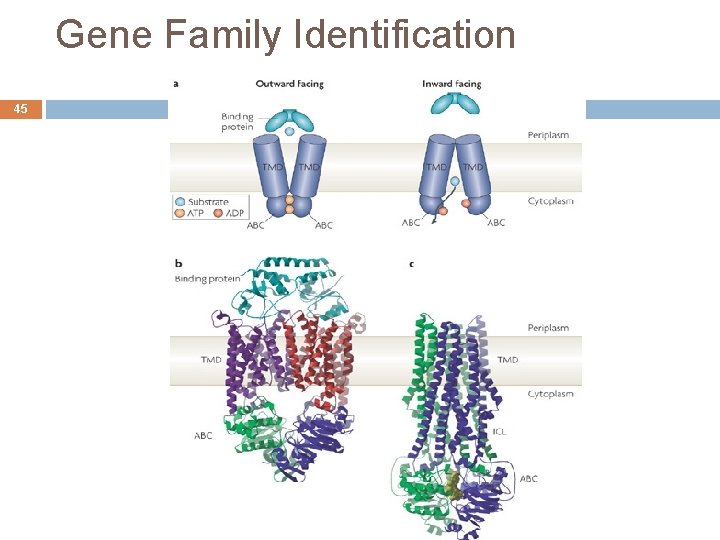
Gene Family Identification 45 André de Carvalho - ICMC/USP 02/10/2020

Gene Family Identification 46 Alternative approaches to find orthologous genes between species Construction of a phylogenetic tree with all genes of a family of both species It is expected that orthologs one-to-one appear as siblings in the leaf nodes Since these genes are separated only by a specialized event between the two genomes compared More reliable but more difficult automation André de Carvalho - ICMC/USP 02/10/2020

Synteny 47 Relative order of genes on the same chromosome Greek origin: syn = together, taenia = tape Once the orthologous genes have been identified Examine changes in their physical position in chromosomes over time See if syntenic relationships are preserved across species André de Carvalho - ICMC/USP 02/10/2020

Synteny 48 Systemic relationships are usually shuffled by: Inversions Transpositions Noise can be added to the study of synteny because of: Deletions Duplications Inserções André de Carvalho - ICMC/USP 02/10/2020

Synteny 49 Even so, it is usually easy to identify blocks of synteny Long sections wherein the relative order of orthologous genes have been preserved Find synteny blocks is important to: Annotation of non-coding sequences Set homologous intergenic regions Which may have little or no similarity Never would be identified by alignment André de Carvalho - ICMC/USP 02/10/2020
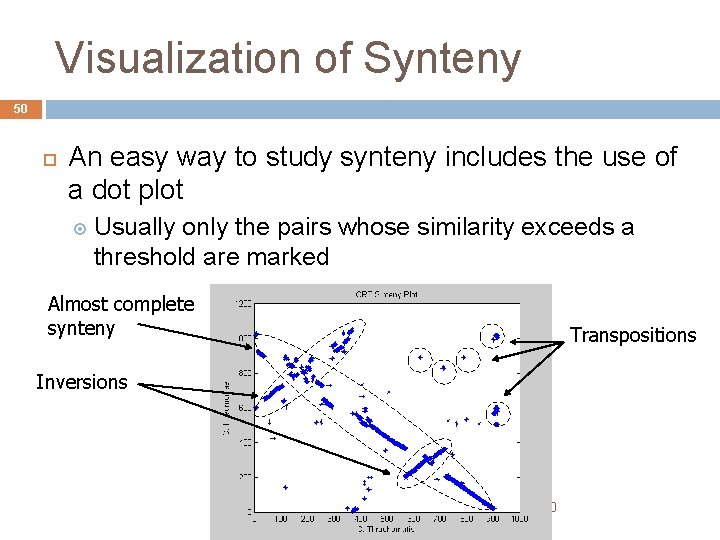
Visualization of Synteny 50 An easy way to study synteny includes the use of a dot plot Usually only the pairs whose similarity exceeds a threshold are marked Almost complete synteny Transpositions Inversions André de Carvalho - ICMC/USP 02/10/2020
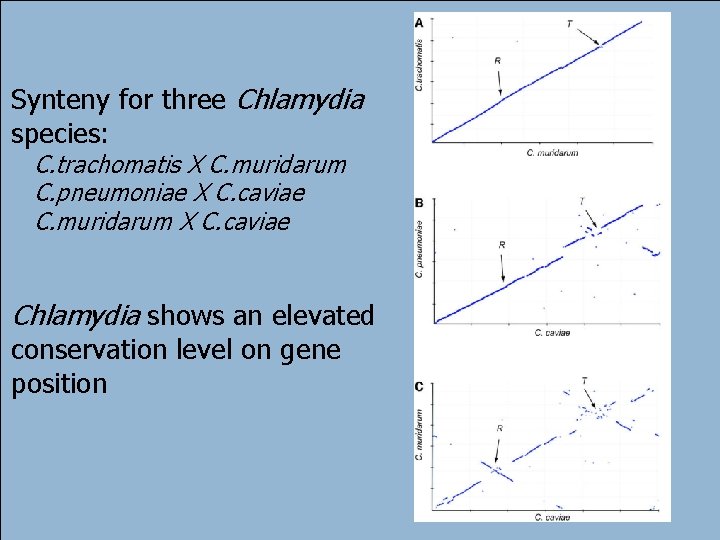
Synteny for three Chlamydia species: C. trachomatis X C. muridarum C. pneumoniae X C. caviae C. muridarum X C. caviae Chlamydia shows an elevated conservation level on gene position 51 André de Carvalho - ICMC/USP 02/10/2020
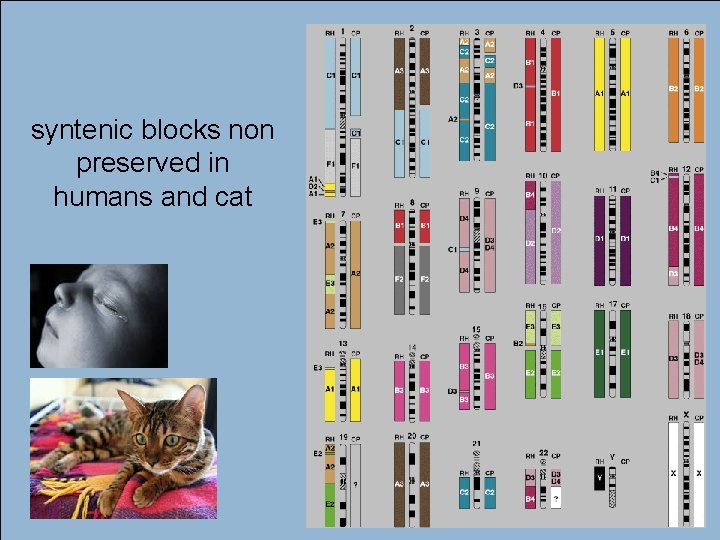
52 syntenic blocks non preserved in humans and cat André de Carvalho - ICMC/USP 02/10/2020

Synteny 53 Syntenic blocks detection helps identify intergenic homologous regions These regions usually evolve more rapidly than the rest of the genome Harder Genes to align and allocate homology are used as anchors To ensure that the examined intergenic regions are homologous Allows examine the evolution of these sequences André de Carvalho - ICMC/USP 02/10/2020

Synteny 54 There noncoding regions of proteins that are conserved Parts that are transcribed into RNA Regulatory sequences Phylogenetic footprint Anchor alignment with syntenic coding regions Make it easier to find short sequences that are conserved in non-coding stretches of DNA André de Carvalho - ICMC/USP 02/10/2020

Synteny 55 Phylogenetic footprint has been used for: Find conserved regulatory sequences Coding regions Example of the region with 3500 with bp with 3 ORFs preserving synteny between 2 species Analysis André de Carvalho - ICMC/USP 02/10/2020
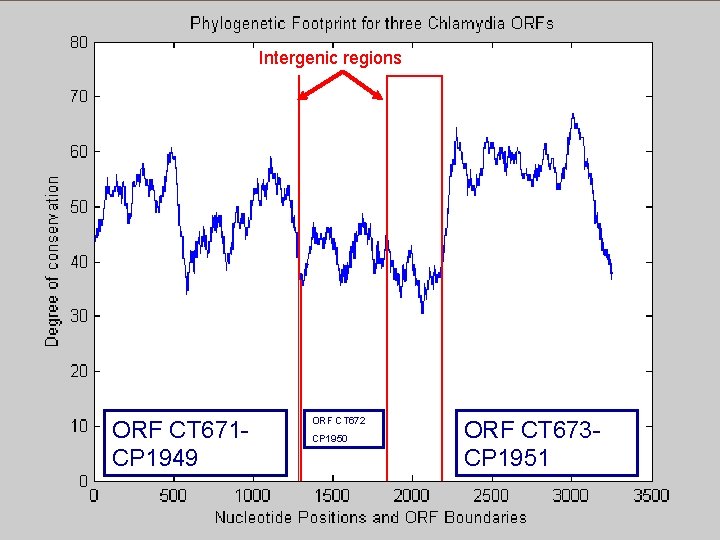
Intergenic regions ORF CT 671 CP 1949 ORF CT 672 CP 1950 ORF CT 673 CP 1951

Metric for Syntenic Distance 57 Systemic blocks are rearranged by Inversions or Transpositions between two genomes Alignment of pairs does not work The goal is to calculate the number of genomic rearrangements that separate two species Not the number of different nucleotides Finding fewer Inversions carrying from a genome to another Transposition inclusion complicates the problem André de Carvalho - ICMC/USP 02/10/2020

Metric for Syntenic Distance 58 Finding fewer Inversions carrying from a genome to another Strings of ORFs Noncoding homologous regions Order by Inversion Várias possíveis séries de Inversions entre suas sequencias com mesma distância Several possible inversions series among their sequences with the same distance Different sequence of Inversions André de Carvalho - ICMC/USP 02/10/2020

Greedy Algorithm 59 1 Assign one of the sequences as Pattern (s); Inversions will only be applied to the other sequence (t); 2 Start from one end of the Pattern sequence and move until you find the position where the symbols are different, si ti; 3 Perform the necessary Inversions, so the symbol in tj match the symbol sj. When this occurs, invert ti: tj; 4 Continue the movement along the string applying Inversions when necessary until all symbols match André de Carvalho - ICMC/USP 02/10/2020
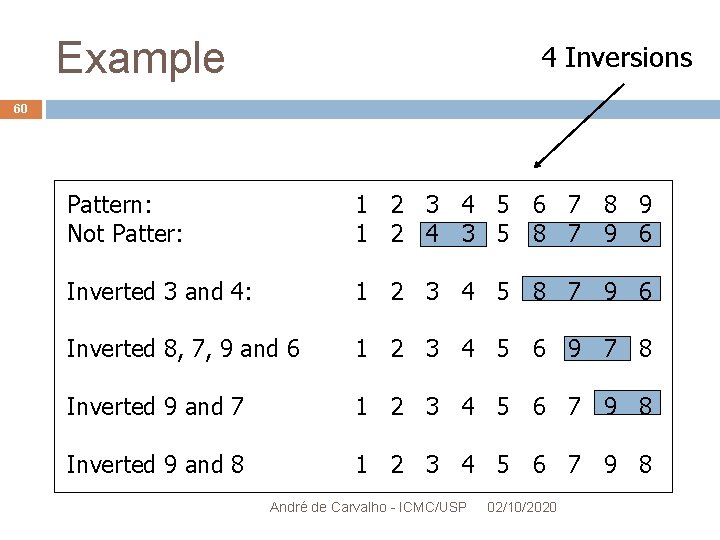
Example 4 Inversions 60 Pattern: Not Patter: 1 2 3 4 5 6 7 8 9 1 2 4 3 5 8 7 9 6 Inverted 3 and 4: 1 2 3 4 5 8 7 9 6 Inverted 8, 7, 9 and 6 1 2 3 4 5 6 9 7 8 Inverted 9 and 7 1 2 3 4 5 6 7 9 8 Inverted 9 and 8 1 2 3 4 5 6 7 9 8 André de Carvalho - ICMC/USP 02/10/2020

Analysis of Results 61 4 inversion operations occurred Distance of Inversion = 4 If operations were carried out following Pattern, distance would be 3 Algorithm does not generate minimum distance when Inversions overlap Other algorithms provide better estimates With increased computational complexity André de Carvalho - ICMC/USP 02/10/2020

Conclusion 62 Genome Comparison Chlamydias Genomics with a bag of genes Synteny Distance among homologous regions André de Carvalho - ICMC/USP 02/10/2020
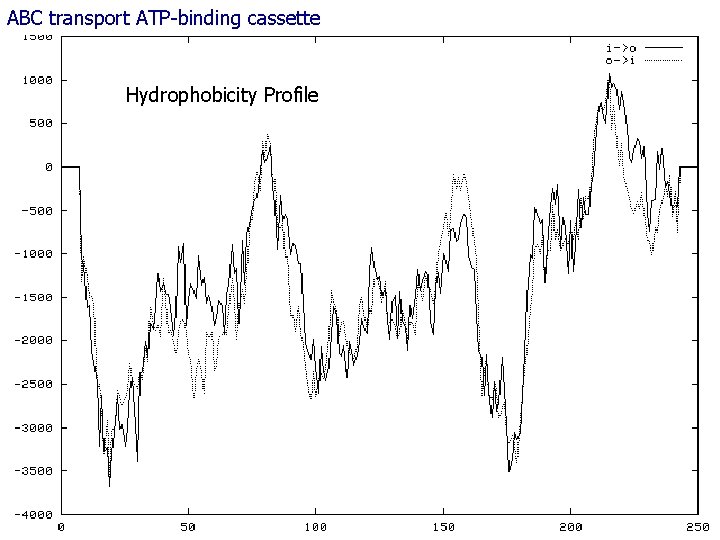
ABC transport ATP-binding cassette 63 Hydrophobicity Profile Identification Gene Family André de Carvalho - ICMC/USP 02/10/2020
Questions?
- Slides: 64