Mark Gerstein Yale University Gerstein Lab orgcourses452 last
Mark Gerstein, Yale University Gerstein. Lab. org/courses/452 (last edit in fall '05) 1 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu BIOINFORMATICS Sequences #1 (up to P-values)

• Basic Alignment via Dynamic Programming • Suboptimal Alignment • Gap Penalties • Similarity (PAM) Matrices • Multiple Alignment • Profiles, Motifs, HMMs • Local Alignment • Probabilistic Scoring Schemes • Rapid Similarity Search: Fasta • Rapid Similarity Search: Blast • Practical Suggestions on Sequence Searching • Transmembrane helix predictions • Secondary Structure Prediction: Basic GOR • Secondary Structure Prediction: Other Methods • Assessing Secondary Structure Prediction • Features of Genomic DNA sequences 2 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Sequence Topics (Contents)

Molecular Biology Information: Protein Sequence à ACDEFGHIKLMNPQRSTVWY but not BJOUXZ • Strings of ~300 aa in an average protein (in bacteria), ~200 aa in a domain • ~200 K known protein sequences d 1 dhfa_ d 8 dfr__ d 4 dfra_ d 3 dfr__ LNCIVAVSQNMGIGKNGDLPWPPLRNEFRYFQRMTTTSSVEGKQ-NLVIMGKKTWFSI LNSIVAVCQNMGIGKDGNLPWPPLRNEYKYFQRMTSTSHVEGKQ-NAVIMGKKTWFSI ISLIAALAVDRVIGMENAMPWN-LPADLAWFKRNTL----NKPVIMGRHTWESI TAFLWAQDRDGLIGKDGHLPWH-LPDDLHYFRAQTV----GKIMVVGRRTYESF d 1 dhfa_ d 8 dfr__ d 4 dfra_ d 3 dfr__ LNCIVAVSQNMGIGKNGDLPWPPLRNEFRYFQRMTTTSSVEGKQ-NLVIMGKKTWFSI LNSIVAVCQNMGIGKDGNLPWPPLRNEYKYFQRMTSTSHVEGKQ-NAVIMGKKTWFSI ISLIAALAVDRVIGMENAMPW-NLPADLAWFKRNTLD----KPVIMGRHTWESI TAFLWAQDRNGLIGKDGHLPW-HLPDDLHYFRAQTVG----KIMVVGRRTYESF d 1 dhfa_ d 8 dfr__ d 4 dfra_ d 3 dfr__ VPEKNRPLKGRINLVLSRELKEPPQGAHFLSRSLDDALKLTEQPELANKVDMVWIVGGSSVYKEAMNHP VPEKNRPLKDRINIVLSRELKEAPKGAHYLSKSLDDALALLDSPELKSKVDMVWIVGGTAVYKAAMEKP ---G-RPLPGRKNIILS-SQPGTDDRV-TWVKSVDEAIAACGDVP------EIMVIGGGRVYEQFLPKA ---PKRPLPERTNVVLTHQEDYQAQGA-VVVHDVAAVFAYAKQHLDQ----ELVIAGGAQIFTAFKDDV d 1 dhfa_ d 8 dfr__ d 4 dfra_ d 3 dfr__ -PEKNRPLKGRINLVLSRELKEPPQGAHFLSRSLDDALKLTEQPELANKVDMVWIVGGSSVYKEAMNHP -PEKNRPLKDRINIVLSRELKEAPKGAHYLSKSLDDALALLDSPELKSKVDMVWIVGGTAVYKAAMEKP -G---RPLPGRKNIILSSSQPGTDDRV-TWVKSVDEAIAACGDVPE-----. IMVIGGGRVYEQFLPKA -P--KRPLPERTNVVLTHQEDYQAQGA-VVVHDVAAVFAYAKQHLD----QELVIAGGAQIFTAFKDDV 3 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu • 20 letter alphabet
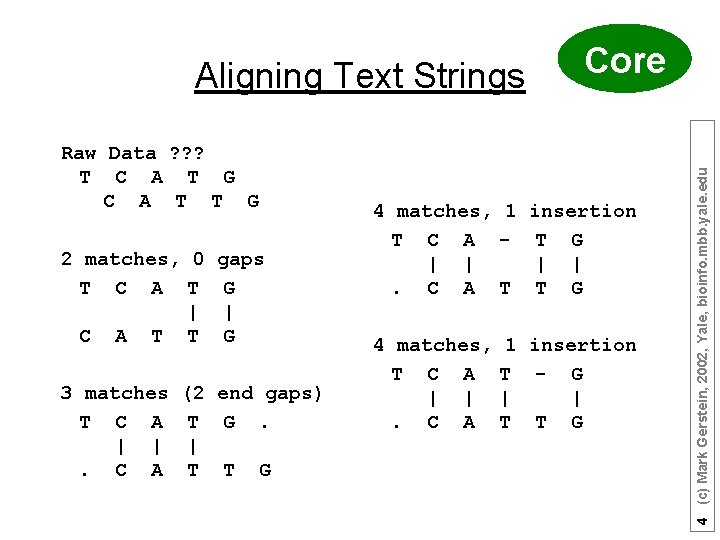
Raw Data ? ? ? T C A T G C A T T G 2 matches, 0 gaps T C A T G | | C A T T G 3 matches T C A | |. C A (2 end gaps) T G. | T T G 4 matches, 1 insertion T C A - T G | |. C A T T G 4 matches, T C A | |. C A 1 insertion T - G | | T T G 4 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Aligning Text Strings Core

• What to do for Bigger String? SSDSEREEHVKRFRQALDDTGMKVPMATTNLFTHPVFKDGGFTANDRDVRRYALRKTIRNIDLAVELGAETYVAWGGREGAESGGAKDVRDALDRMKEAFDLLGEYVTSQGYDIRFAIEP KPNEPRGDILLPTVGHALAFIERLERPELYGVNPEVGHEQMAGLNFPHGIAQALWAGKLFHIDLNGQNGIKYDQDLRFGAGDLRAAFWLVDLLESAGYSGPRHFDFKPPRTEDFDGVWAS • Needleman-Wunsch (1970) provided first automatic method à Dynamic Programming to Find Global Alignment • Their Test Data (J->Y) à ABCNYRQCLCRPM AYCYNRCKCRBP 5 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Dynamic Programming

Step 1 -- Make a Dot Plot (Similarity Matrix) Core 6 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Put 1's where characters are identical.
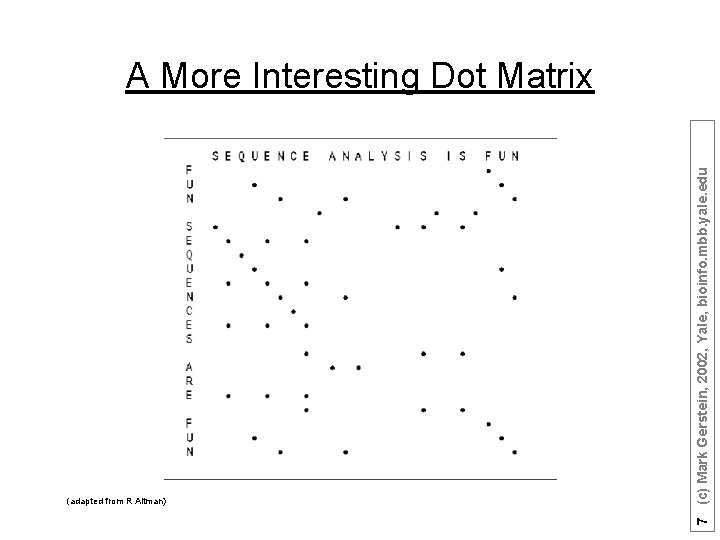
(adapted from R Altman) 7 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu A More Interesting Dot Matrix
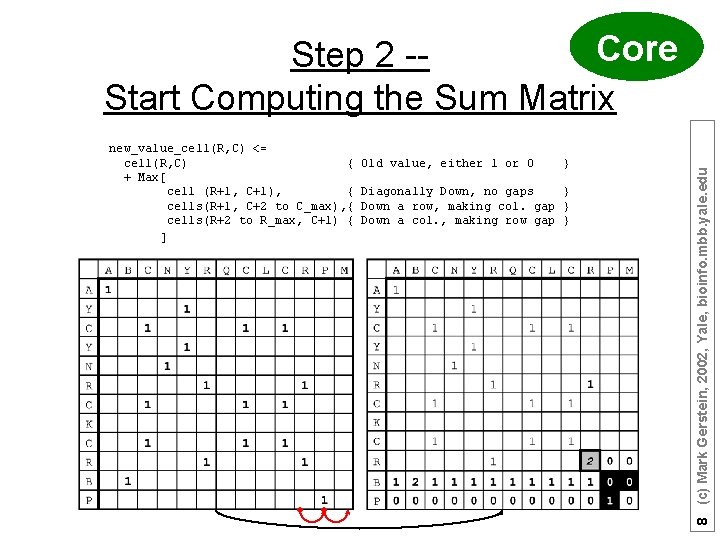
new_value_cell(R, C) <= cell(R, C) { + Max[ cell (R+1, C+1), { cells(R+1, C+2 to C_max), { cells(R+2 to R_max, C+1) { ] Old value, either 1 or 0 } Diagonally Down, no gaps } Down a row, making col. gap } Down a col. , making row gap } 8 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Core Step 2 -Start Computing the Sum Matrix
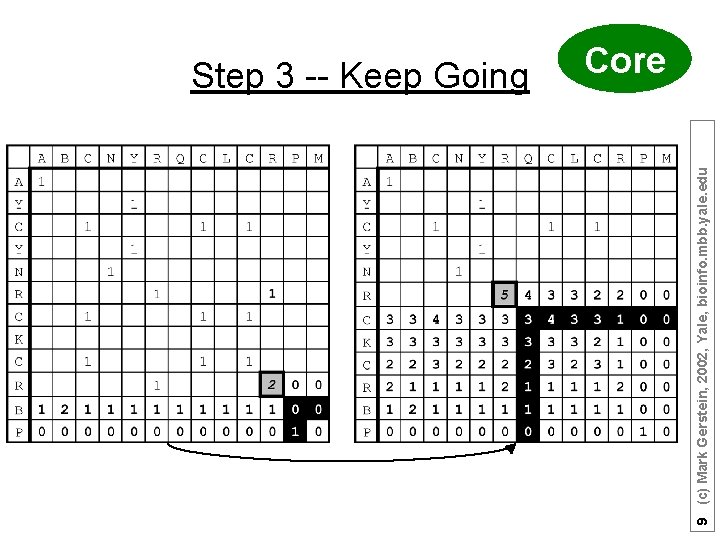
9 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Step 3 -- Keep Going Core
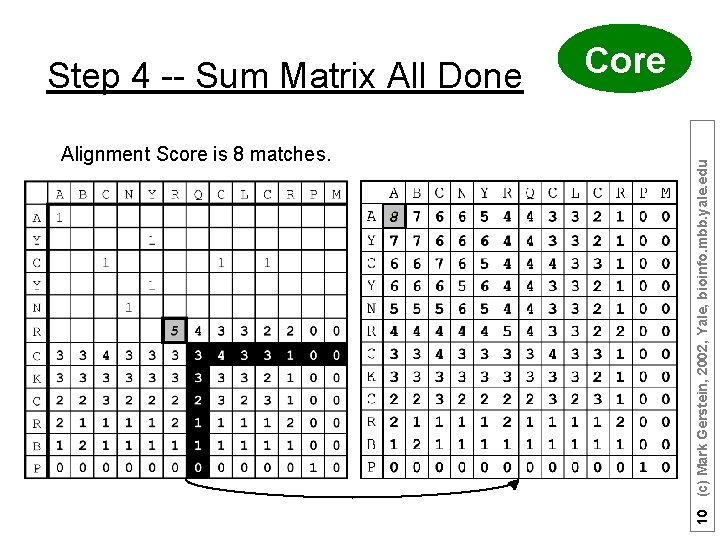
Alignment Score is 8 matches. 10 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Step 4 -- Sum Matrix All Done Core

Step 5 -- Traceback Core A B C N Y - R Q C L C R - P M A Y C - Y N R - C K C R B P Hansel & Gretel 11 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Find Best Score (8) and Trace Back

A B C N Y - R Q C L C R - P M A Y C - Y N R - C K C R B P 12 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Step 5 -- Traceback Core
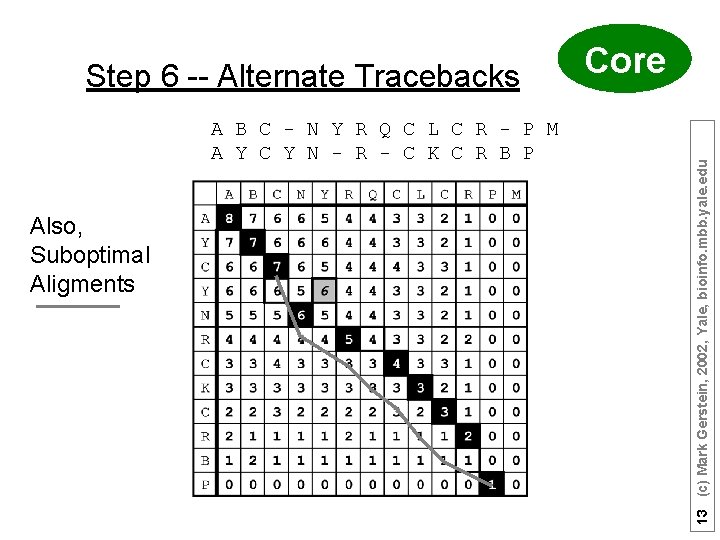
A B C - N Y R Q C L C R - P M A Y C Y N - R - C K C R B P Also, Suboptimal Aligments Core 13 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Step 6 -- Alternate Tracebacks

14 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Additional Aspects of Dynamic Programming Sequence Alignment: Suboptimals and Gaps

Suboptimal Alignments generated using the seed : : 1 AGCCGACGAC TCATTACAGC GGATCGCCTG TAACCCCTTA ATACCTTTAA GCTGCAGCAA GGCAGTCTAG GGGAGCAACC AGGTCTCCTG CGGTTATCGC CAGTCTCGGG TCCGTTCTTG CCGTAGGCTG GCGCCGTTAG CAAGTCGCAG GGCGTACAGA TGACTTGAGT AGGGACCTTA TCAATGCCGT AGCATCAAGA AGTAAGGAGA generated using the seed : : -453862491 1 : TTCGCTTGAG CTAGCTGAAC AACCAGATCG AGTCGTAATA AGCTGCAGTG AGACAAACAC CCGGGGGGCC CTAGCGCGCTGCGCC CTAAGGTTAC GATGGCAGCA AGCCTTCTGT TCCTCGCCTA GTGATGATAG 1573438385 AAGACATCTC GAGGGGATGG GGGCACGTAA GTCGTGAGGT GGAATTTCAC TGATAACTAC AGCACTCTGG 1 CGTGA TGCTAATCAC CTTTCTTCGT AAGGGTCCCG ATCGCAGAAC CGTGTGAAAG CGCAGACCTC ACTCTCTCCA ATTGCGAACA GTGGCGGCGC GTGCAAGAGT TTATAGGCAG ACCTGGCCCG GCAGAGGGAC GGCATATTAA TGTTTCGGTC GGCAACTAAA AGAGGAAGTA CCGTGTGCCT TTTTGTCCCT CGGCCTGACT TCGAAGATCC CAGACTCCAC GACGAAAGGA CGGGAGTACG GAGGCCGCTA GAGACTAATC TTTTCCGGCT Parameters: match weight = 10, transition weight = 1, transversion weight = -3 Gap opening penalty = 50 Gap continuation penalty = 1 Run as a local alignment (Smith-Waterman) (courtesy of Michael Zucker) 15 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu ; ; Random DNA sequence ; ; 500 nucleotides ; ; A: C: G: T = 1 : 1 ; RAN -453862491 AAATGCCAAA TCATACGAAC CCCACCGGGA TATACACTAA AATTCCAACT TCGGTATGAA GCTGGGGCAA TGATGT TTAATACCTT CGCCGTTAAT CACGGGCATA CCGCGGGGTA CCCCGGACAT CATATGACCA ATGGCGTGTT 1 ; ; Random DNA sequence ; ; 500 nucleotides ; ; A: C: G: T = 1 : 1 ; RAN 1573438385 CCCTCCATCG CCAGTTCCTGTCGTGA CGCGGATTAC CTATGGCATC TTCCGCTATA CCACAACGTG AATAGCCCGT TACGGGGCAT GACGCGGGCT GAACCTTCAA CGCTAACTAG GCTAGTTAGG CCCCATTTGT TCCTCTGAGG 1
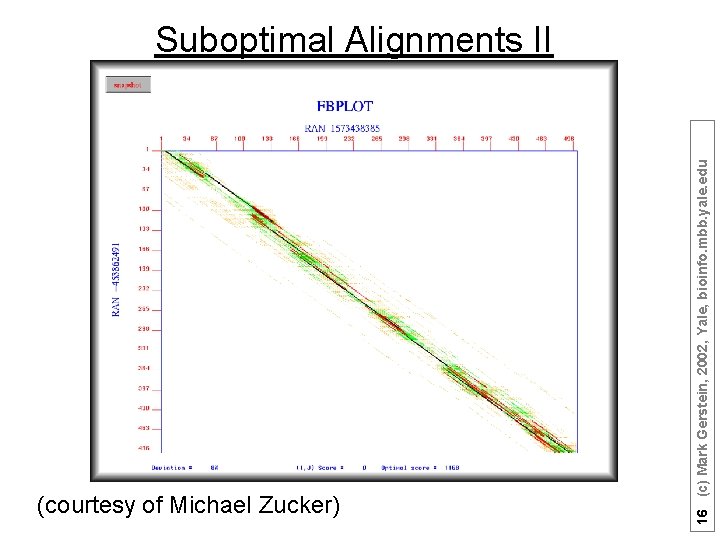
(courtesy of Michael Zucker) 16 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Suboptimal Alignments II

The score at a position can also factor in a penalty for introducing gaps (i. e. , not going from i, j to i- 1, j- 1). Gap penalties are often of linear form: GAP = a + b. N GAP is the gap penalty a = cost of opening a gap b = cost of extending the gap by one (affine) N = length of the gap (Here assume b=0, a=1/2, so GAP = 1/2 regardless of length. ) EX with a=. 5 & b=. 1 ATGCAAAAT ATG-AAAAT. 5 ATG--AAAT. 5 + (1)(. 1) ATG---AAT. 5 + (2)(. 1) =. 7 17 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Gap Penalties Core
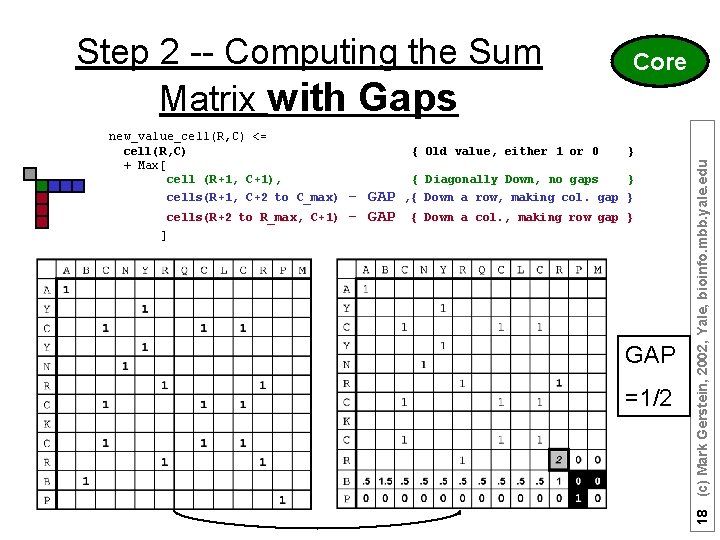
new_value_cell(R, C) <= cell(R, C) + Max[ cell (R+1, C+1), cells(R+1, C+2 to C_max) cells(R+2 to R_max, C+1) { Old value, either 1 or 0 - GAP Core } { Diagonally Down, no gaps } , { Down a row, making col. gap } { Down a col. , making row gap } ] GAP =1/2 18 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Step 2 -- Computing the Sum Matrix with Gaps
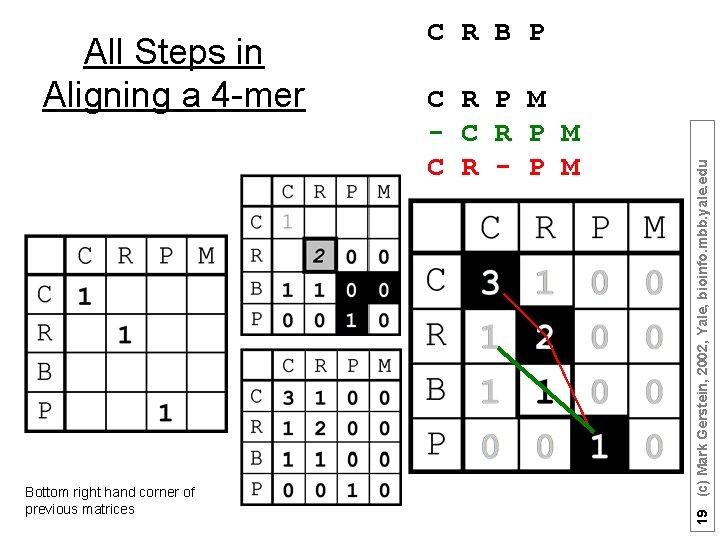
Bottom right hand corner of previous matrices C R P M - C R P M C R - P M 19 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu All Steps in Aligning a 4 -mer C R B P

à The best alignment that ends at a given pair of positions (i and j) in the 2 sequences is the score of the best alignment previous to this position PLUS the score for aligning those two positions. à An Example Below • Aligning R to K does not affect alignment of previous N-terminal residues. Once this is done it is fixed. Then go on to align D to E. • How could this be violated? Aligning R to K changes best alignment in box. ACSQRP--LRV-SH RSENCV A-SNKPQLVKLMTH VKDFCV Align ACSQRPLRVSHRSENCV & ASNKPQLVKLMTHVKDFCV ACSQRP--LRV-SH -R SENCV A-SNKPQLVKLMTH VK DFCV 20 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Key Idea in Dynamic Programming

21 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Additional Aspects of Dynamic Programming Sequence Alignment: Substitution Matrices

• Identity Matrix à Match L with L => 1 Match L with D => 0 Match L with V => 0? ? • S(aa-1, aa-2) à Match L with L => 1 Match L with D => 0 Match L with V =>. 5 • Number of Common Ones à PAM à Blossum à Gonnet A R N D C Q E G H I L K M F P S T W Y V Core A 4 -1 -2 -2 0 -1 -1 0 -2 -1 -1 -2 -1 1 0 -3 -2 0 R -1 5 0 -2 -3 1 0 -2 0 -3 -2 2 -1 -3 -2 -1 -1 -3 -2 -3 N -2 0 6 1 -3 0 0 0 1 -3 -3 0 -2 -3 -2 1 0 -4 -2 -3 D -2 -2 1 6 -3 0 2 -1 -1 -3 -4 -1 -3 -3 -1 0 -1 -4 -3 -3 C 0 -3 -3 -3 8 -3 -4 -3 -3 -1 -1 -3 -1 -2 -3 -1 -1 -2 -2 -1 Q -1 1 0 0 -3 5 2 -2 0 -3 -2 1 0 -3 -1 0 -1 -2 E -1 0 0 2 -4 2 5 -2 0 -3 -3 1 -2 -3 -1 0 -1 -3 -2 -2 G 0 -2 0 -1 -3 -2 -2 6 -2 -4 -4 -2 -3 -3 -2 0 -2 -2 -3 -3 H -2 0 1 -1 -3 0 0 -2 7 -3 -3 -1 -2 -2 2 -3 I -1 -3 -3 -3 -1 -3 -3 -4 -3 4 2 -3 1 0 -3 -2 -1 -3 -1 3 L -1 -2 -3 -4 -3 2 4 -2 2 0 -3 -2 -1 1 K -1 2 0 -1 -3 1 1 -2 -1 -3 -2 5 -1 -3 -1 0 -1 -3 -2 -2 M -1 -1 -2 -3 -1 0 -2 -3 -2 1 2 -1 5 0 -2 -1 -1 1 F -2 -3 -3 -3 -1 0 0 -3 0 6 -4 -2 -2 1 3 -1 P -1 -2 -2 -1 -3 -1 -1 -2 -2 -3 -3 -1 -2 -4 6 -1 -1 -4 -3 -2 S 1 -1 1 0 -1 0 0 0 -1 -2 -2 0 -1 -2 -1 4 1 -3 -2 -2 T 0 -1 -1 -2 -2 -1 -1 -2 -1 1 5 -2 -2 0 W -3 -3 -4 -4 -2 -2 -3 -1 1 -4 -3 -2 10 2 -3 Y -2 -2 -2 -3 -2 -1 -2 -3 2 -1 -1 -2 -1 3 -3 -2 -2 2 6 -1 V 0 -3 -3 -3 -1 -2 -2 -3 -3 3 1 -2 1 -1 -2 -2 0 -3 -1 4 22 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Similarity (Substitution) Matrix

+ —> More likely than random 0 —> At random base rate - —> Less likely than random 1 Manually align protein structures (or, more risky, sequences) 2 Look at frequency of a. a. substitutions at structurally constant sites. -- i. e. pair i-j exchanges 3 Compute log-odds S(aa-1, aa-2) = log 2 ( freq(O) / freq(E) ) O = observed exchanges, E = expected exchanges • odds = freq(observed) / freq(expected) • Sij = log odds • freq(expected) = f(i)*f(j) = is the chance of getting amino acid i in a column and then having it change to j • e. g. A-R pair observed only a tenth as often as expected AAVLL… AAVQI… AVVQL… ASVLL… 90% 45% Core 23 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Where do matrices come from?
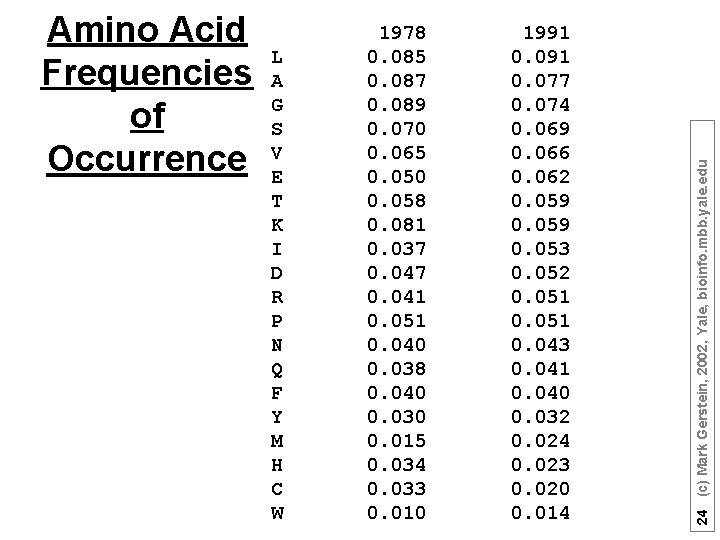
L A G S V E T K I D R P N Q F Y M H C W 1978 0. 085 0. 087 0. 089 0. 070 0. 065 0. 050 0. 058 0. 081 0. 037 0. 041 0. 051 0. 040 0. 038 0. 040 0. 030 0. 015 0. 034 0. 033 0. 010 1991 0. 077 0. 074 0. 069 0. 066 0. 062 0. 059 0. 053 0. 052 0. 051 0. 043 0. 041 0. 040 0. 032 0. 024 0. 023 0. 020 0. 014 24 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Amino Acid Frequencies of Occurrence
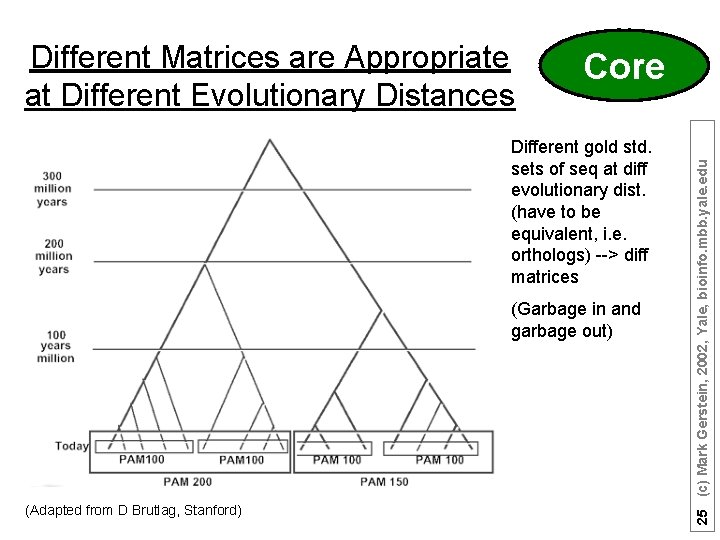
Core Different gold std. sets of seq at diff evolutionary dist. (have to be equivalent, i. e. orthologs) --> diff matrices (Garbage in and garbage out) (Adapted from D Brutlag, Stanford) 25 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Different Matrices are Appropriate at Different Evolutionary Distances

26 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu End of class (m-3) 2005, 10. 24
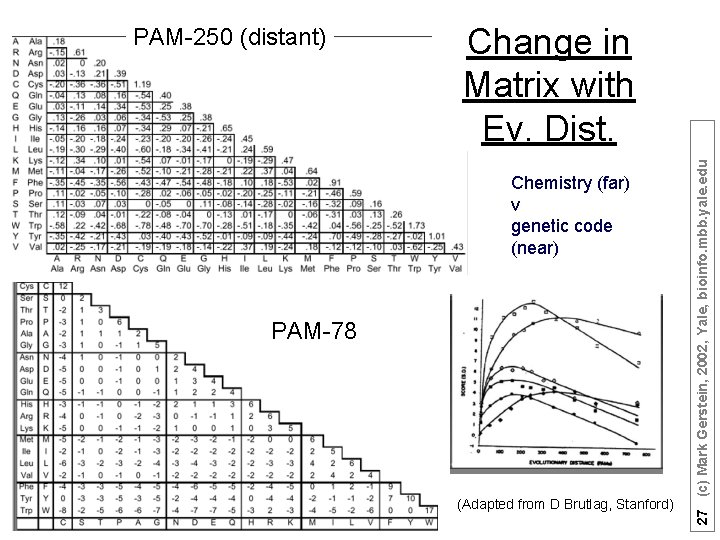
Change in Matrix with Ev. Dist. Chemistry (far) v genetic code (near) PAM-78 (Adapted from D Brutlag, Stanford) 27 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu PAM-250 (distant)

The BLOSUM matrices: Henikoff & Henikoff (Henikoff, S. & Henikoff J. G. (1992) PNAS 89: 10915 -10919). The BLOSUM Matrices This leads to a series of matrices, analogous to the PAM series of matrices. BLOSUM 80: derived at the 80% identity level. BLOSUM 62 is the BLAST default 28 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Some concepts challenged: Are the evolutionary rates uniform over the whole of the protein sequence? (No. )

29 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Additional Aspects of Dynamic Programming Sequence Alignment: Local Alignment

1 The scoring system uses negative scores for mismatches 2 The minimum score for at a matrix element is zero 3 Find the best score anywhere in the matrix (not just last column or row) • These three changes cause the algorithm to seek high scoring subsequences, which are not penalized for their global effects (mod. 1), which don’t include areas of poor match (mod. 2), and which can occur anywhere (mod. 3) Core (Adapted from R Altman) 30 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Modifications for Local Alignment
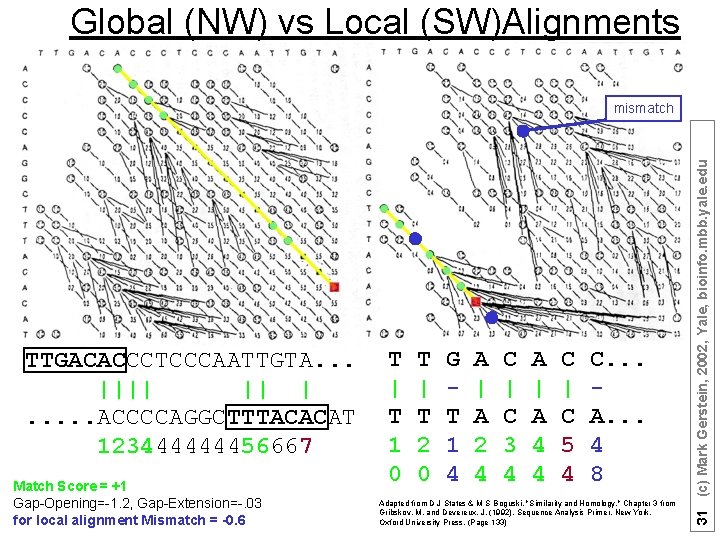
Global (NW) vs Local (SW)Alignments TTGACACCCTCCCAATTGTA. . . |||| || |. . . ACCCCAGGCTTTACACAT 123444444456667 Match Score = +1 Gap-Opening=-1. 2, Gap-Extension=-. 03 for local alignment Mismatch = -0. 6 T | T 1 0 T | T 2 0 G T 1 4 A | A 2 4 C | C 3 4 A | A 4 4 C | C 5 4 C. . . A. . . 4 8 Adapted from D J States & M S Boguski, "Similarity and Homology, " Chapter 3 from Gribskov, M. and Devereux, J. (1992). Sequence Analysis Primer. New York, Oxford University Press. (Page 133) 31 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu mismatch

• GLOBAL = best alignment of entirety of both sequences à For optimum global alignment, we want best score in the final row or final column à Are these sequences generally the same? à Needleman Wunsch à find alignment in which total score is highest, perhaps at expense of areas of great local similarity • LOCAL = best alignment of segments, without regard to rest of sequence à For optimum local alignment, we want best score anywhere in matrix (will discuss) à Do these two sequences contain high scoring subsequences à Smith Waterman à find alignment in which the highest scoring subsequences are identified, at the expense of the overall score (Adapted from R Altman) 32 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Local vs. Global Alignment Core

33 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Multiple Alignment
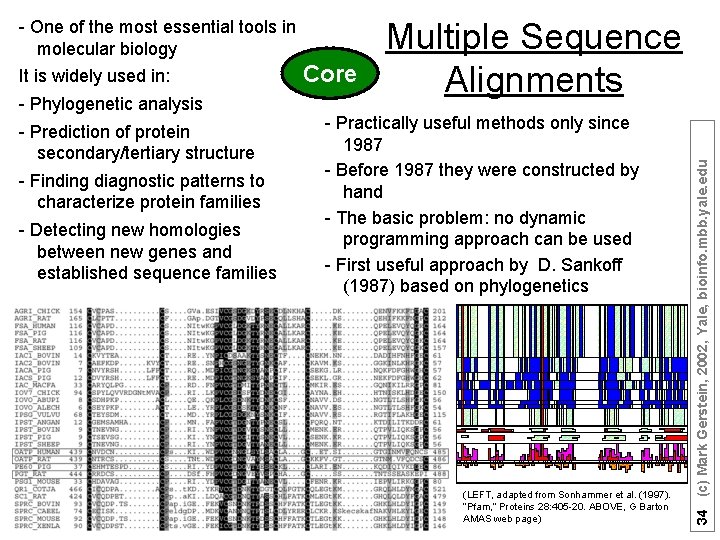
It is widely used in: - Phylogenetic analysis - Prediction of protein secondary/tertiary structure - Finding diagnostic patterns to characterize protein families - Detecting new homologies between new genes and established sequence families Core Multiple Sequence Alignments - Practically useful methods only since 1987 - Before 1987 they were constructed by hand - The basic problem: no dynamic programming approach can be used - First useful approach by D. Sankoff (1987) based on phylogenetics (LEFT, adapted from Sonhammer et al. (1997). “Pfam, ” Proteins 28: 405 -20. ABOVE, G Barton AMAS web page) 34 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu - One of the most essential tools in molecular biology

- Most multiple alignments based on this approach - Initial guess for a phylogenetic tree based on pairwise alignments - Built progressively starting with most closely related sequences - Follows branching order in phylogenetic tree - Sufficiently fast - Sensitive - Algorithmically heuristic, no mathematical property associated with the alignment - Biologically sound, it is common to derive alignments which are impossible to improve by eye (adapted from Sonhammer et al. (1997). “Pfam, ” Proteins 28: 405 -20) 35 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Progressive Multiple Alignments
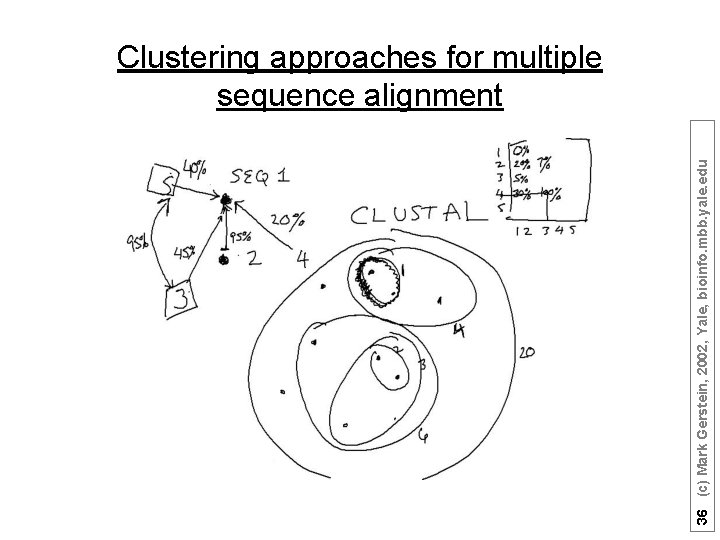
36 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Clustering approaches for multiple sequence alignment

- Local Minimum Problem - Parameter Choice Problem 1. Local Minimum Problem - It stems from greedy nature of alignment (mistakes made early in alignment cannot be corrected later) - A better tree gives a better alignment (UPGMA neighbour-joining tree method) 2. Parameter Choice Problem • - It stems from using just one set of parameters (and hoping that they will do for all) 37 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Problems with Progressive Alignments
38 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Domain Problem in Sequence Clustering

39 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Patterns: Motifs and Profiles
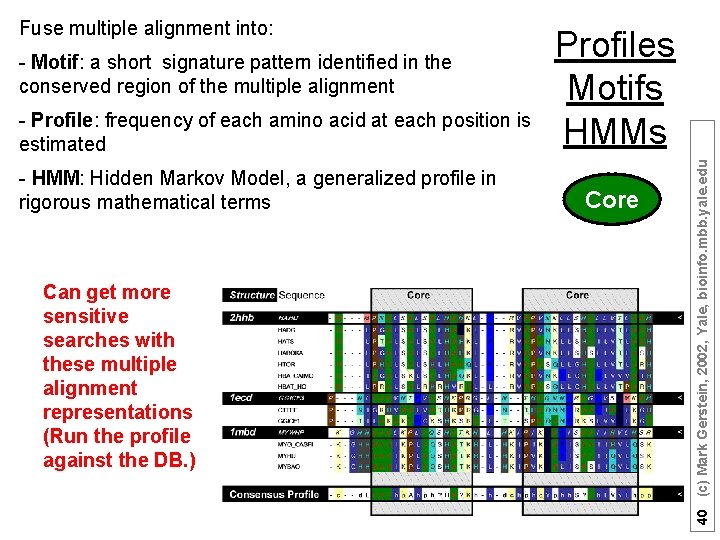
- Motif: a short signature pattern identified in the conserved region of the multiple alignment - Profile: frequency of each amino acid at each position is estimated - HMM: Hidden Markov Model, a generalized profile in rigorous mathematical terms Can get more sensitive searches with these multiple alignment representations (Run the profile against the DB. ) Profiles Motifs HMMs Core 40 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Fuse multiple alignment into:

Core 41 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Motifs - several proteins are grouped together by similarity searches - they share a conserved motif - motif is stringent enough to retrieve the family members from the complete protein database - PROSITE: a collection of motifs (1135 different motifs)
![PKC_PHOSPHO_SIT E Protein kinase C phosphorylation site [ST]-x-[RK] Post-translational modifications RGD Cell attachment sequence PKC_PHOSPHO_SIT E Protein kinase C phosphorylation site [ST]-x-[RK] Post-translational modifications RGD Cell attachment sequence](http://slidetodoc.com/presentation_image_h2/84cd8b00bd54d40c816274cfdc7bf6b2/image-42.jpg)
PKC_PHOSPHO_SIT E Protein kinase C phosphorylation site [ST]-x-[RK] Post-translational modifications RGD Cell attachment sequence R-G-D Domains SOD_CU_ZN_1 Copper/Zinc superoxide dismutase [GA]-[IMFAT]-H-[LIVF]-Hx(2)-[GP]-[SDG]-x-[STAGDE] Enzymes_Oxidoreduc tases THIOL_PROTEASE_ ASN Eukaryotic thiol (cysteine) proteases active site [FYCH]-[WI]-[LIVT]-x-[KRQAG]-N -[ST]-W-x(3)-[FYW]-G-x(2)-G[LFYW]-[LIVMFYG]-x-[LIVMF] Enzymes_Hydrolases TNFR_NGFR_1 TNFR/CD 27/30/4 0/95 cysteine-rich region C-x(4, 6)-[FYH]-x(5, 10)-C-x(0, 2)C-x(2, 3)-C-x(7, 11)-C-x(4, 6)[DNEQSKP]-x(2)-C Receptors 42 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Motifs · Each element in a pattern is separated from its neighbor by a “-”. · The symbol “x” is used for a position where any amino acid is accepted. · Ambiguities are indicated by listing the acceptable amino acids for a given position, between brackets “[]”. · Ambiguities are also indicated by listing between a pair of braces “{}” the amino acids that are not accepted at a given position. · Repetition of an element of the pattern is indicated by with a numerical value or a numerical range between parentheses following that element.
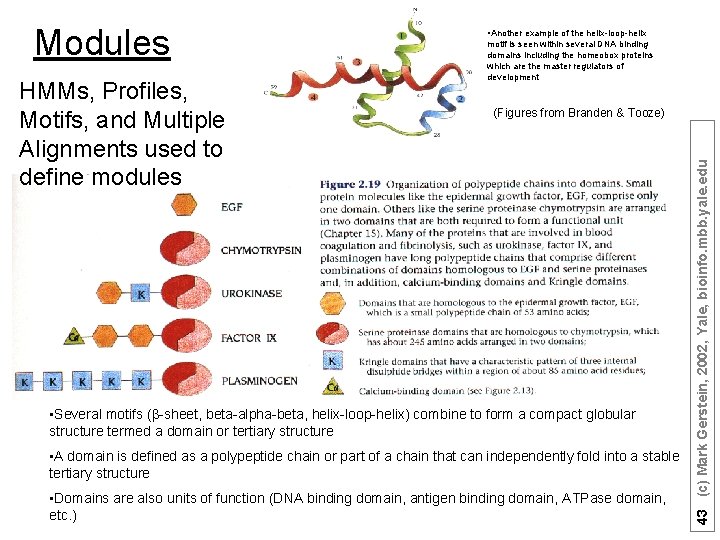
HMMs, Profiles, Motifs, and Multiple Alignments used to define modules motif is seen within several DNA binding domains including the homeobox proteins which are the master regulators of development (Figures from Branden & Tooze) • Several motifs (b-sheet, beta-alpha-beta, helix-loop-helix) combine to form a compact globular structure termed a domain or tertiary structure • A domain is defined as a polypeptide chain or part of a chain that can independently fold into a stable tertiary structure • Domains are also units of function (DNA binding domain, antigen binding domain, ATPase domain, etc. ) 43 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Modules • Another example of the helix-loop-helix

C 1 Q Example Extra 44 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Ca 28_Human ELSAHATPAFTAVLTSPLPASGMPVKFDRTLYNGHSGYNPATGIFTCPVGGVYYFAYHVH VKGTNVWVALYKNNVPATYTYDEYKKGYLDQASGGAVLQLRPNDQVWVQIPSDQANGLYS TEYIHSSFSGFLLCPT C 1 qb_Human DYKATQKIAFSATRTINVPLRRDQTIRFDHVITNMNNNYEPRSGKFTCKVPGLYYFTYHA SSRGNLCVNLMRGRERAQKVVTFCDYAYNTFQVTTGGMVLKLEQGENVFLQATDKNSLLG MEGANSIFSGFLLFPD Cerb_Human VRSGSAKVAFSAIRSTNHEPSEMSNRTMIIYFDQVLVNIGNNFDSERSTFIAPRKGIYSF NFHVVKVYNRQTIQVSLMLNGWPVISAFAGDQDVTREAASNGVLIQMEKGDRAYLKLERG NLMGGWKYSTFSGFLVFPL COLE_LEPMA. 264 RGPKGPPGESVEQIRSAFSVGLFPSRSFPPPSLPVKFDKVFYNGEGHWDPTLNKFNVTYP GVYLFSYHITVRNRPVRAALVVNGVRKLRTRDSLYGQDIDQASNLALLHLTDGDQVWLET LRDWNGXYSSSEDDSTFSGFLLYPDTKKPTAM HP 27_TAMAS. 72 GPPGPPGMTVNCHSKGTSAFAVKANELPPAPSQPVIFKEALHDAQGHFDLATGVFTCPVP GLYQFGFHIEAVQRAVKVSLMRNGTQVMEREAEAQDGYEHISGTAILQLGMEDRVWLENK LSQTDLERGTVQAVFSGFLIHEN HSUPST 2_1. 95 GIQGRKGEPGEGAYVYRSAFSVGLETYVTIPNMPIRFTKIFYNQQNHYDGSTGKFHCNIP GLYYFAYHITVYMKDVKVSLFKKDKAMLFTYDQYQENNVDQASGSVLLHLEVGDQVWLQV YGEGERNGLYADNDNDSTFTGFLLYHDTN 2. HS 27109_1 ENALAPDFSKGSYRYAPMVAFFASHTYGMTIPGPILFNNLDVNYGASYTPRTGKFRIPYL GVYVFKYTIESFSAHISGFLVVDGIDKLAFESENINSEIHCDRVLTGDALLELNYGQEVW LRLAKGTIPAKFPPVTTFSGYLLYRT 4. YQCC_BACSU VVHGWTPWQKISGFAHANIGTTGVQYLKKIDHTKIAFNRVIKDSHNAFDTKNNRFIAPND GMYLIGASIYTLNYTSYINFHLKVYLNGKAYKTLHHVRGDFQEKDNGMNLGLNGNATVPM NKGDYVEIWCYCNYGGDETLKRAVDDKNGVFNFFD 5. BSPBSXSE_25 ADSGWTAWQKISGFAHANIGTTGRQALIKGENNKIKYNRIIKDSHKLFDTKNNRFVASHA GMHLVSASLYIENTERYSNFELYVYVNGTKYKLMNQFRMPTPSNNSDNEFNATVTGSVTV PLDAGDYVEIYVYVGYSGDVTRYVTDSNGALNYFD

SGMPLVSANHGVTG-------MPVSAFTVILS--KAYPA---VGCPHPIYEILYNRQQHY -----ALTG-------MPVSAFTVILS--KAYPG---ATVPIKFDKILYNRQQHY -----GGPA-------YEMPAFTAELT--APFPP---VGGPVKFNKLLYNGRQNY HAYAGKKGKHGGPA-------YEMPAFTAELT--VPFPP---VGAPVKFDKLLYNGRQNY -----ELSA-------HATPAFTAVLT--SPLPA---SGMPVKFDRTLYNGHSGY ----GTPGRKGEPGE---AAYMYRSAFSVGLETRVTVP-----NVPIRFTKIFYNQQNHY ------RGPKGPPGE---SVEQIRSAFSVGLFPSRSFPP---PSLPVKFDKVFYNGEGHW -------GPPGPPGMTVNCHSKGTSAFAVKAN--ELPPA---PSQPVIFKEALHDAQGHF -----NIRD-------QPRPAFSAIRQ---NPMT---LGNVVIFDKVLTNQESPY -------D---YRATQKVAFSALRTINSPLR----PNQVIRFEKVITNANENY -------D---YKATQKIAFSATRTINVPLR----RDQTIRFDHVITNMNNNY -------V---RSGSAKVAFSAIRSTNHEPSEMSNRTMIIYFDQVLVNIGNNF ---ENALAPDFSKGS---YRYAPMVAFFASHTYGMTIP------GPILFNNLDVNYGASY. *. : : MMCOL 10 A 1_1. 483 Ca 1 x_Chick S 15435 CA 18_MOUSE. 597 Ca 28_Human MM 37222_1. 98 COLE_LEPMA. 264 HP 27_TAMAS. 72 S 19018 C 1 qb_Mouse C 1 qb_Human Cerb_Human 2. HS 27109_1 DPRSGIFTCKIPGIYYFSYHVHVKGT--HVWVGLYKNGTP-TMYTY---DEYSKGYLDTA DPRTGIFTCRIPGLYYFSYHVHAKGT--NVWVALYKNGSP-VMYTY---DEYQKGYLDQA NPQTGIFTCEVPGVYYFAYHVHCKGG--NVWVALFKNNEP-VMYTY---DEYKKGFLDQA NPQTGIFTCEVPGVYYFAYHVHCKGG--NVWVALFKNNEP-MMYTY---DEYKKGFLDQA NPATGIFTCPVGGVYYFAYHVHVKGT--NVWVALYKNNVP-ATYTY---DEYKKGYLDQA DGSTGKFYCNIPGLYYFSYHITVYMK--DVKVSLFKKDKA-VLFTY---DQYQEKNVDQA DPTLNKFNVTYPGVYLFSYHITVRNR--PVRAALVVNGVR-KLRTR---DSLYGQDIDQA DLATGVFTCPVPGLYQFGFHIEAVQR--AVKVSLMRNGTQ-VMERE---AEAQDG-YEHI QNHTGRFICAVPGFYYFNFQVISKWD--LCLFIKSSSGGQ-PRDSLSFSNTNNKGLFQVL EPRNGKFTCKVPGLYYFTYHASSRGN---LCVNLVRGRDRDSMQKVVTFCDYAQNTFQVT EPRSGKFTCKVPGLYYFTYHASSRGN---LCVNLMRGRER--AQKVVTFCDYAYNTFQVT DSERSTFIAPRKGIYSFNFHVVKVYNRQTIQVSLMLNGWP----VISAFAGDQDVTREAA TPRTGKFRIPYLGVYVFKYTIESFSA--HISGFLVVDGIDKLAFESEN-INSEIHCDRVL. * * : MMCOL 10 A 1_1. 483 Ca 1 x_Chick S 15435 CA 18_MOUSE. 597 Ca 28_Human MM 37222_1. 98 COLE_LEPMA. 264 HP 27_TAMAS. 72 S 19018 C 1 qb_Mouse C 1 qb_Human Cerb_Human 2. HS 27109_1 SGSAIMELTENDQVWLQLPNA-ESNGLYSSEYVHSSFSGFLVAPM------SGSAVIDLMENDQVWLQLPNS-ESNGLYSSEYVHSSFSGFLFAQI------SGSAVLLLRPGDRVFLQMPSE-QAAGLYAGQYVHSSFSGYLLYPM------SGSAVLLLRPGDQVFLQNPFE-QAAGLYAGQYVHSSFSGYLLYPM------SGGAVLQLRPNDQVWVQIPSD-QANGLYSTEYIHSSFSGFLLCPT------SGSVLLHLEVGDQVWLQVYGDGDHNGLYADNVNDSTFTGFLLYHDTN----SNLALLHLTDGDQVWLETLR--DWNGXYSSSEDDSTFSGFLLYPDTKKPTAM SGTAILQLGMEDRVWLENKL--SQTDLERG-TVQAVFSGFLIHEN------AGGTVLQLRRGDEVWIEKDP--AKGRIYQGTEADSIFSGFLIFPS------TGGVVLKLEQEEVVHLQATD---KNSLLGIEGANSIFTGFLLFPD------TGGMVLKLEQGENVFLQATD---KNSLLGMEGANSIFSGFLLFPD------SNGVLIQMEKGDRAYLKLER---GN-LMGG-WKYSTFSGFLVFPL------TGDALLELNYGQEVWLRLAK----GTIPAKFPPVTTFSGYLLYRT------. : : . : * *: *. Extra Clustal Alignment 45 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu MMCOL 10 A 1_1. 483 Ca 1 x_Chick S 15435 CA 18_MOUSE. 597 Ca 28_Human MM 37222_1. 98 COLE_LEPMA. 264 HP 27_TAMAS. 72 S 19018 C 1 qb_Mouse C 1 qb_Human Cerb_Human 2. HS 27109_1

Prosite Pattern -- EGF like pattern - Bone morphogenic protein 1 (BMP-1), a protein which induces cartilage and bone formation. Caenorhabditis elegans developmental proteins lin-12 (13 copies) and glp-1 (10 copies). Calcium-dependent serine proteinase (CASP) which degrades the extracellular matrix proteins type …. Cell surface antigen 114/A 10 (3 copies). Cell surface glycoprotein complex transmembrane subunit. Coagulation associated proteins C, Z (2 copies) and S (4 copies). Coagulation factors VII, IX, X and XII (2 copies). Complement C 1 r/C 1 s components (1 copy). Complement-activating component of Ra-reactive factor (RARF) (1 copy). Complement components C 6, C 7, C 8 alpha and beta chains, and C 9 (1 copy). Epidermal growth factor precursor (7 -9 copies). Extra 46 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu A sequence of about thirty to forty amino-acid residues long found in the sequence of epidermal growth factor (EGF) has been shown [1 to 6] to be present, in a more or less conserved form, in a large number of other, mostly animal proteins. The proteins currently known to contain one or more copies of an EGF-like pattern are listed below. +----------+ +-------------+ | | x(4)-C-x(0, 48)-C-x(3, 12)-C-x(1, 70)-C-x(1, 6)-C-x(2)-G-a-x(0, 21)-G-x(2)-C-x | | ****************** +----------+ 'C': conserved cysteine involved in a disulfide bond. 'G': often conserved glycine 'a': often conserved aromatic amino acid '*': position of both patterns. 'x': any residue -Consensus pattern: C-x-C-x(5)-G-x(2)-C [The 3 C's are involved in disulfide bonds] http: //www. expasy. ch/sprot/prosite. html
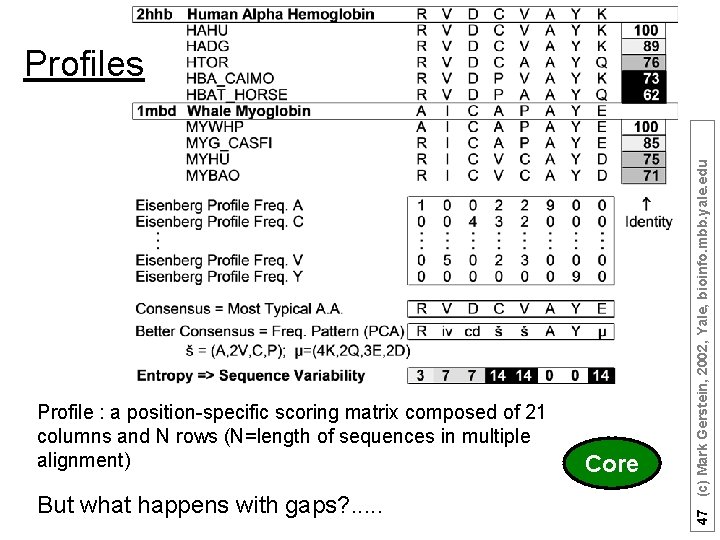
Profile : a position-specific scoring matrix composed of 21 columns and N rows (N=length of sequences in multiple alignment) 5 But what happens with gaps? . . . Core 47 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Profiles

H(p, a) = - a=1 to 20 f(p, a) log 2 f(p, a), where f(p, a) = frequency of amino acid a occurs at position p ( Msimp(p, a) ) Say column only has one aa (AAAAA): H(p, a) = 1 log 2 1 + 0 log 2 0 + … = 0 + 0 + … = 0 Say column is random with all aa equiprobable (ACD. . ): Hrand(p, a) =. 05 log 2. 05 + … = -. 22 + … = -4. 3 Say column is random with aa occurring according to probability found in the sequence databases (ACAAAADAADDDDAAAA…. ): Hdb(a) = - a=1 to 20 F(a) log 2 F(a), where F(a) is freq. of occurrence of a in DB Core Hcorrected(p, a) = H(p, a) – Hdb(a) 48 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Profiles formula for entropy H(p, a)
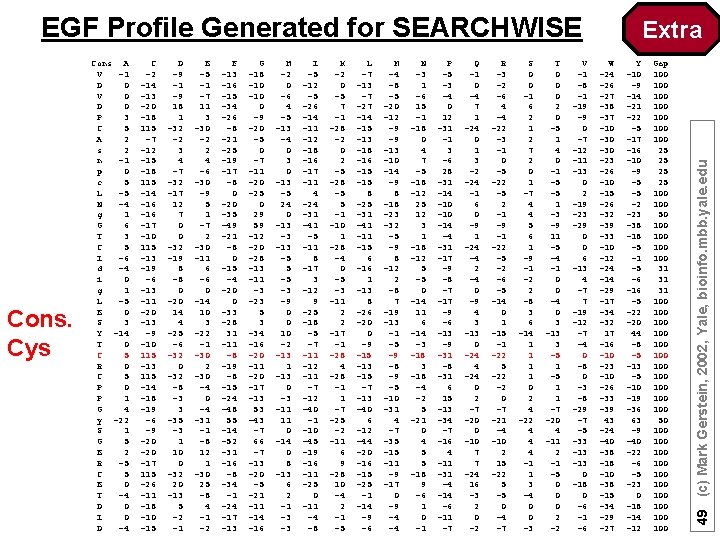
EGF Profile Generated for SEARCHWISE C -2 -14 -13 -20 -18 115 -7 -12 -15 -18 115 -14 -16 -17 -10 115 -13 -19 -6 -13 -11 -20 -13 -9 -10 115 -13 115 -14 -18 -19 -6 -9 -20 -17 115 -26 -11 -18 -10 -15 D -9 -1 -9 18 1 -32 -2 3 4 -7 -32 -17 12 7 0 0 -32 -19 8 -8 0 -20 14 4 -25 -6 -32 0 -32 -8 -3 3 -35 -3 1 10 0 -32 20 -13 5 -2 -1 E -5 -1 -7 11 3 -30 -2 2 4 -6 -30 -9 5 1 -7 2 -30 -11 6 -6 0 -14 10 3 -22 -1 -30 2 -30 -4 -31 -1 -8 12 1 -30 25 -8 4 -1 -2 F -13 -16 -15 -34 -26 -8 -21 -25 -19 -17 -8 0 -20 -35 -49 -21 -8 0 -15 -4 -20 0 -33 -28 31 -11 -8 -19 -8 -15 -24 -48 55 -14 -52 -31 -16 -8 -34 -1 -24 -17 -13 G -18 -10 0 -9 -20 -5 0 -7 -11 -20 -25 0 29 59 -12 -20 -28 -13 -11 -3 -23 5 3 -34 -16 -20 -11 -20 -17 -13 53 -43 -7 66 -7 -13 -20 -5 -21 -14 -16 H -2 0 -6 4 -5 -13 -4 0 3 0 -13 -5 24 0 -13 -3 -13 -5 5 -5 -3 -9 0 0 10 -2 -13 1 -13 0 -3 -11 11 0 -14 0 8 -13 6 2 -1 -3 -3 I -5 -12 -5 -26 -14 -11 -12 -18 -16 -17 -11 4 -24 -31 -41 -5 -11 8 -17 3 -12 9 -25 -18 -5 -7 -11 -12 -11 -7 -12 -40 -1 -10 -45 -19 -16 -11 -25 0 -11 -4 -8 K -2 0 -5 7 -1 -28 -2 0 2 -5 -28 -5 5 -1 -10 1 -28 -4 0 -5 -3 -11 2 2 -17 -1 -28 4 -28 -1 1 -7 -25 -2 -11 6 9 -28 10 -4 2 -1 -5 L -7 -13 -7 -27 -14 -15 -13 -18 -16 -15 8 -25 -31 -41 -15 6 -16 1 -13 8 -26 -20 0 -9 -15 -13 -15 -7 -13 -40 6 -12 -44 -20 -16 -15 -25 -1 -14 -9 -6 M -4 -8 -5 -20 -12 -9 -9 -13 -10 -14 -9 8 -18 -23 -32 -5 -9 8 -12 2 -8 7 -19 -13 -1 -5 -9 -8 -9 -5 -10 -31 4 -7 -35 -11 -9 -17 0 -9 -4 -4 N -3 1 -6 15 -1 -18 0 4 7 -5 -18 -12 25 12 3 1 -18 -12 5 -5 0 -14 11 6 -14 -3 -18 -4 -2 5 -21 0 4 5 5 -18 9 -6 1 0 -1 P -5 -3 -4 0 12 -31 -1 3 -6 28 -31 -14 -10 -14 -4 -31 -17 -9 -8 -7 -17 -9 -6 -13 -9 -31 -8 -31 6 15 -13 -34 -7 -16 4 -11 -31 -4 -14 -6 -11 -7 Q -1 0 -4 7 1 -24 0 1 3 -2 -24 -1 6 0 -9 1 -24 -4 2 -4 0 -9 4 3 -13 0 -24 4 -24 0 2 -7 -20 0 -10 7 7 -24 16 -3 2 0 -2 R -3 -2 -6 4 -4 -22 -3 -1 0 -5 -22 -5 2 -1 -9 -1 -22 -5 -2 -6 -5 -14 0 1 -15 -1 -22 5 -22 -2 0 -7 -21 -4 -10 2 15 -22 5 -5 0 -4 -7 S 0 0 -1 6 2 1 2 7 2 0 1 -7 4 4 5 6 1 -9 -1 -2 2 -8 3 6 -14 1 1 0 2 4 -22 4 4 4 -1 1 3 -4 0 0 -3 T 0 0 0 2 0 -5 1 4 0 -1 -5 -5 1 -3 -9 11 -5 -4 -1 0 0 -4 0 3 -13 3 -5 1 1 -7 -20 4 -11 2 -1 -5 0 0 0 2 -2 V -1 -8 -1 -19 -9 0 -7 -12 -11 -13 0 2 -19 -23 -29 0 0 6 -13 4 -7 7 -19 -12 -7 -4 0 -8 0 -3 -8 -29 -7 -5 -33 -13 0 -18 0 -6 -1 -6 W -24 -26 -27 -38 -37 -10 -30 -23 -26 -10 -15 -26 -32 -39 -33 -10 -12 -24 -14 -29 -17 -34 -32 17 -16 -10 -23 -10 -26 -33 -39 43 -24 -40 -38 -10 -38 -15 -34 -29 -27 Y -10 -9 -14 -21 -22 -5 -17 -16 -10 -9 -5 -5 -2 -23 -38 -18 -5 -1 -5 -6 -16 -5 -22 -20 44 -8 -5 -13 -5 -10 -19 -36 63 -9 -40 -22 -6 -5 -23 0 -18 -14 -12 Gap 100 100 25 25 100 50 100 100 31 31 31 100 100 100 50 100 100 100 49 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Cons. Cys Cons A V -1 D 0 V 0 D 0 P 3 C 5 A 2 s 2 n -1 p 0 c 5 L -5 N -4 g 1 G 6 T 3 C 5 I -6 d -4 i 0 g 1 L -5 E 0 S 3 Y -14 T 0 C 5 R 0 C 5 P 0 P 1 G 4 y -22 S 1 G 5 E 2 R -5 C 5 E 0 T -4 D 0 I 0 D -4 Extra
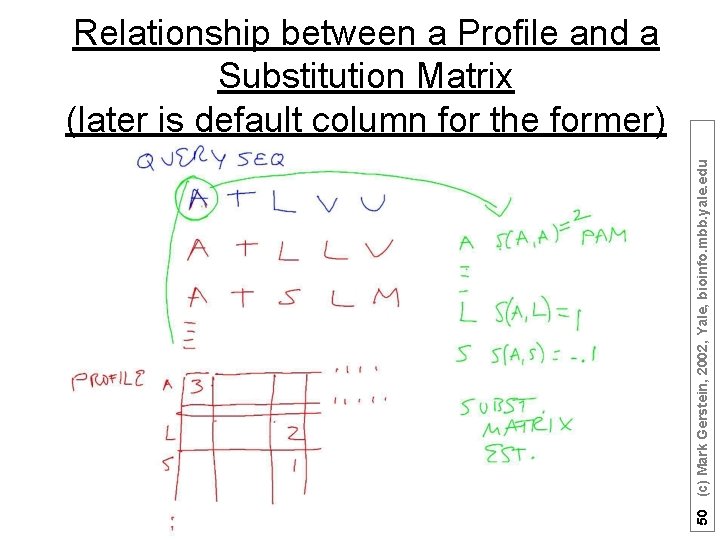
50 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Relationship between a Profile and a Substitution Matrix (later is default column for the former)

51 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Patterns: HMMs
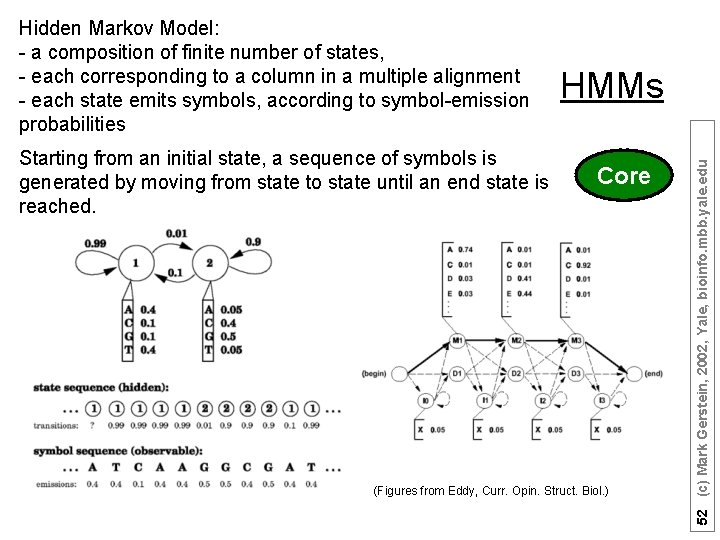
Starting from an initial state, a sequence of symbols is generated by moving from state to state until an end state is reached. HMMs Core (Figures from Eddy, Curr. Opin. Struct. Biol. ) 52 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Hidden Markov Model: - a composition of finite number of states, - each corresponding to a column in a multiple alignment - each state emits symbols, according to symbol-emission probabilities
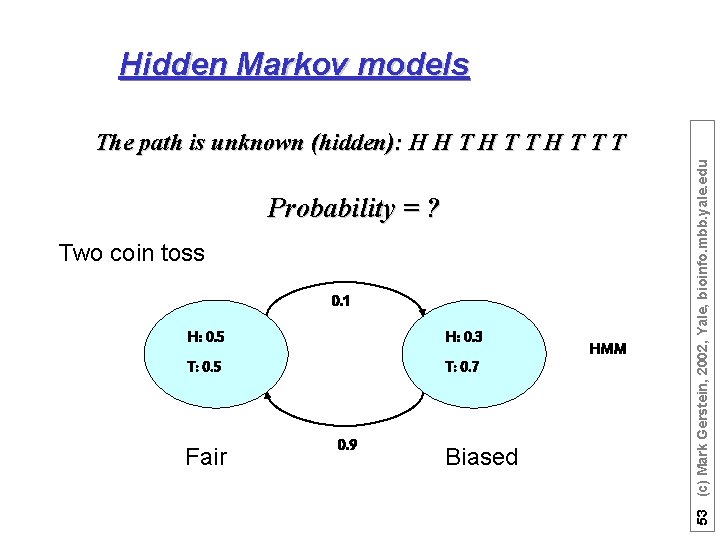
Hidden Markov models Probability = ? Two coin toss 0. 1 H: 0. 5 H: 0. 3 T: 0. 5 T: 0. 7 Fair 0. 9 Biased HMM 53 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu The path is unknown (hidden): H H T T T
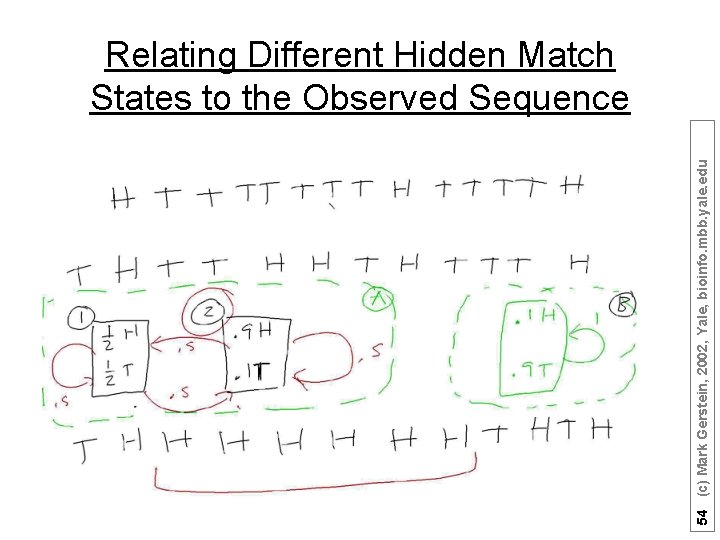
54 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Relating Different Hidden Match States to the Observed Sequence

55 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu End of class (m-4) 2005, 10. 26
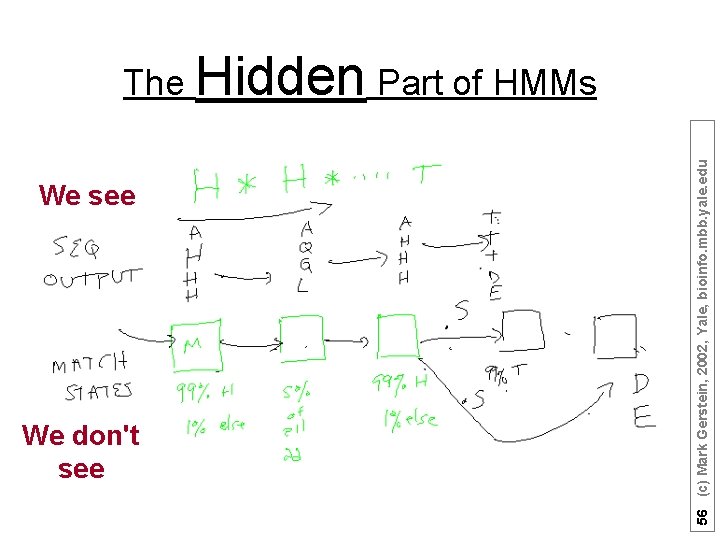
We see We don't see 56 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu The Hidden Part of HMMs
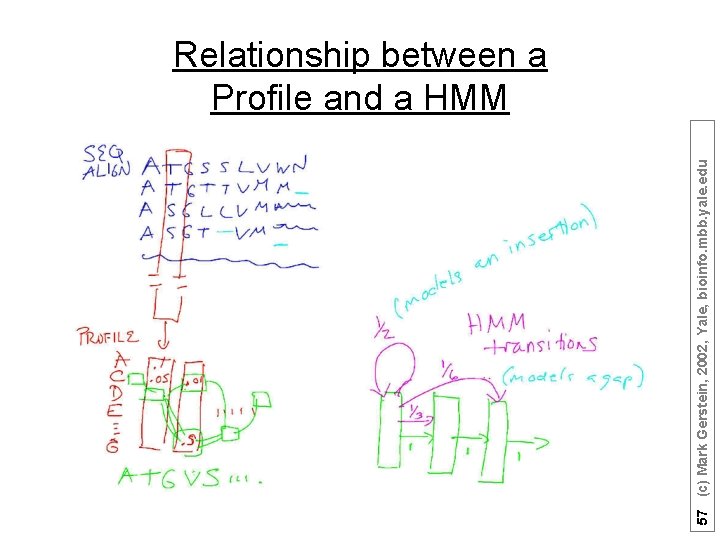
57 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Relationship between a Profile and a HMM

HMMs allow the calculation of Probability of a path through the model (Viterbi gives best path for a given seq, whereas forward sums of all possible paths) Forward Algorithm – finds probability P that a model l emits a given sequence O by summing over all paths that emit the sequence the probability of that path Viterbi Algorithm – finds the most probable path through the model for a given sequence (both usually just boil down to simple applications of dynamic programming) 58 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Algorithms
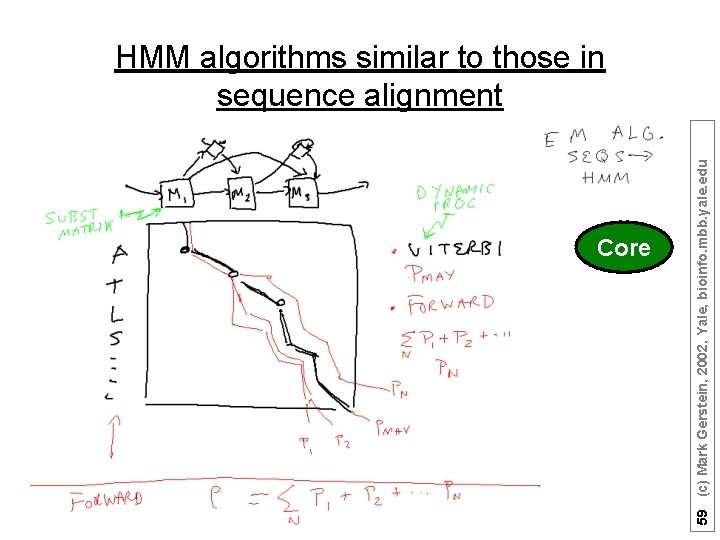
Core 59 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu HMM algorithms similar to those in sequence alignment
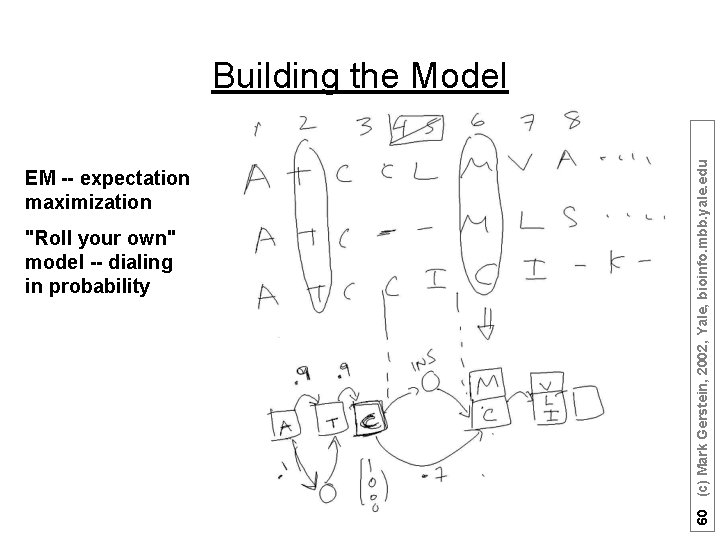
EM -- expectation maximization "Roll your own" model -- dialing in probability 60 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Building the Model
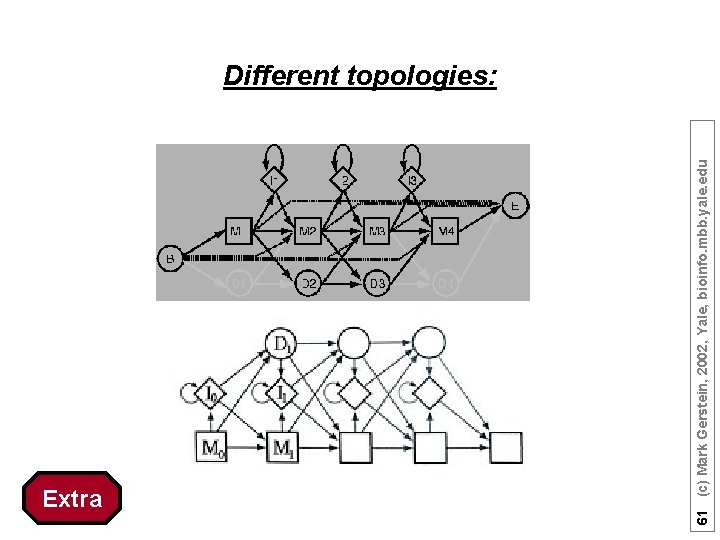
Extra 61 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Different topologies:
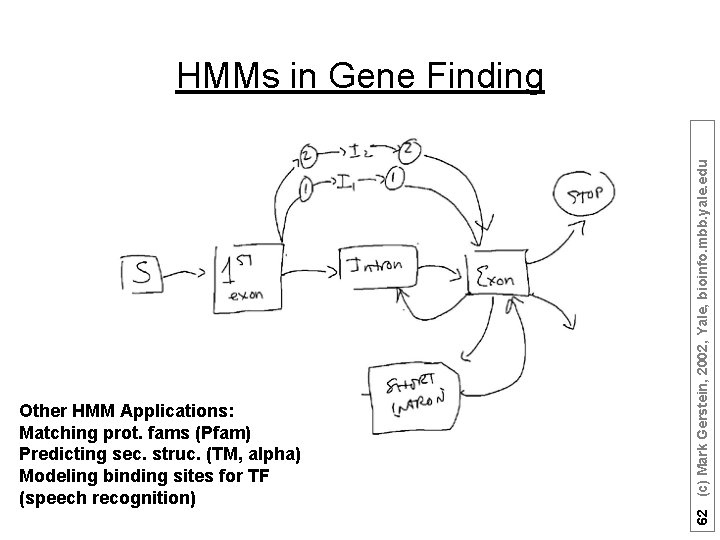
Other HMM Applications: Matching prot. fams (Pfam) Predicting sec. struc. (TM, alpha) Modeling binding sites for TF (speech recognition) 62 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu HMMs in Gene Finding
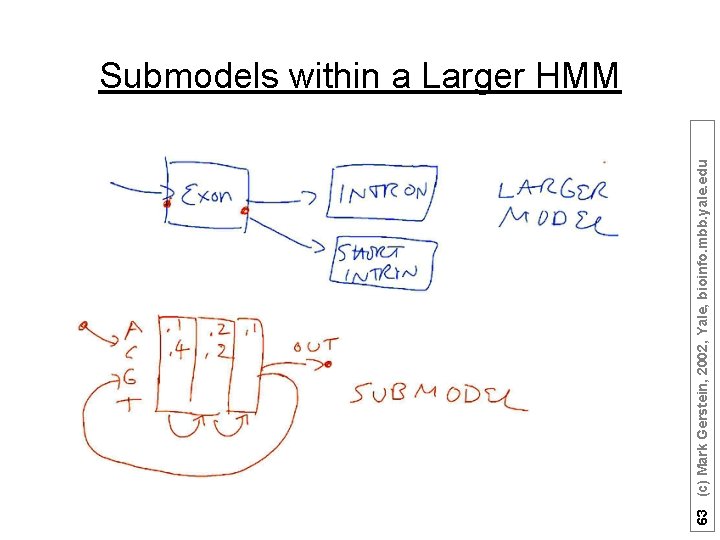
63 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Submodels within a Larger HMM

64 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu P-values

Core Simplest score (for identity matrix) is S = # matches What does a Score of 10 mean? What is the Right Cutoff? S = Total Score S(i, j) = similarity matrix score for aligning i and j Sum is carried out over all aligned i and j n = number of gaps (assuming no gap ext. penalty) G = gap penalty 65 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu The Score

• How does Score Rank Relative to all the Other Core Possible Scores à P-value à Percentile Test Score Rank • All-vs-All comparison of the Database (100 K x 100 K) à Graph Distribution of Scores à ~1010 scores much smaller number of true positives à N dependence 66 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Score in Context of Other Scores
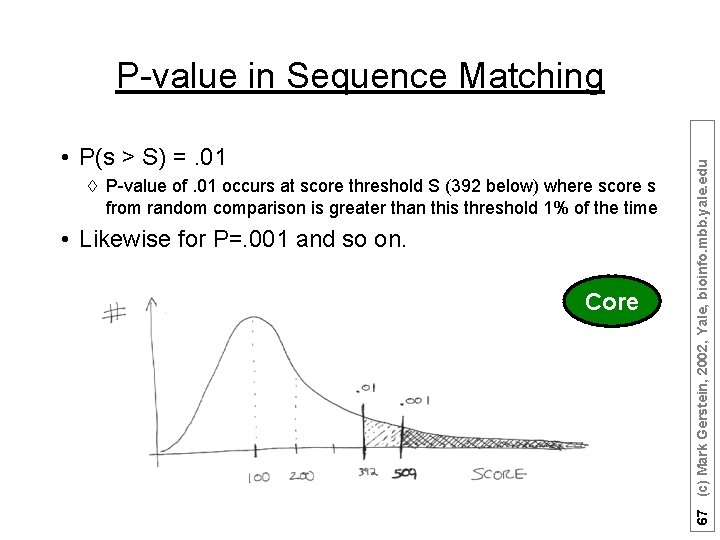
• P(s > S) =. 01 à P-value of. 01 occurs at score threshold S (392 below) where score s from random comparison is greater than this threshold 1% of the time • Likewise for P=. 001 and so on. Core 67 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu P-value in Sequence Matching
![1 Core 2 [ e. g. P(score s>392) = 1% chance] 3 à For 1 Core 2 [ e. g. P(score s>392) = 1% chance] 3 à For](http://slidetodoc.com/presentation_image_h2/84cd8b00bd54d40c816274cfdc7bf6b2/image-68.jpg)
1 Core 2 [ e. g. P(score s>392) = 1% chance] 3 à For sequences, originally used in Blast (Karlin-Altschul). Then in FASTA, &c. à Extrapolated Percentile Rank: How does a Score Rank Relative to all Other Scores? • One Strategy: Fit to Observed Distribution 1)All-vs-All comparison 2)Graph Distribution of Scores in 2 D (N dependence); 1 K x 1 K families -> ~1 M scores; ~2 K included TPs 3)Fit a function r(S) to TN distribution (TNs from scop); Integrating r gives P(s>S), the CDF, chance of getting a score better than threshold S randomly 4) Use same formalism for sequence & structure 68 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu P-values • Significance Statistics
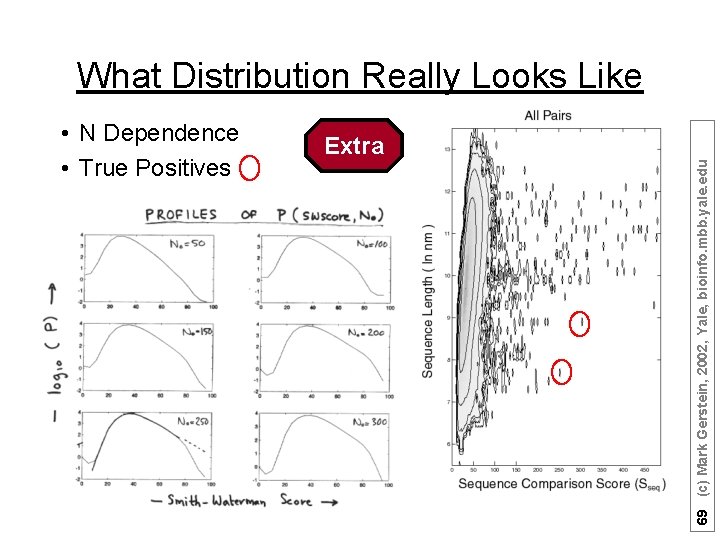
What Distribution Really Looks Like Extra 69 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu • N Dependence • True Positives
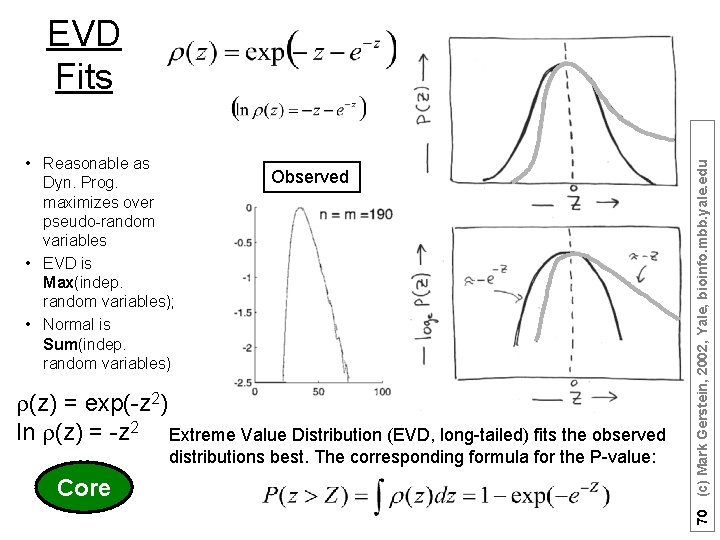
• Reasonable as Dyn. Prog. maximizes over pseudo-random variables • EVD is Max(indep. random variables); • Normal is Sum(indep. random variables) Observed r(z) = exp(-z 2) ln r(z) = -z 2 Extreme Value Distribution (EVD, long-tailed) fits the observed distributions best. The corresponding formula for the P-value: Core 70 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu EVD Fits
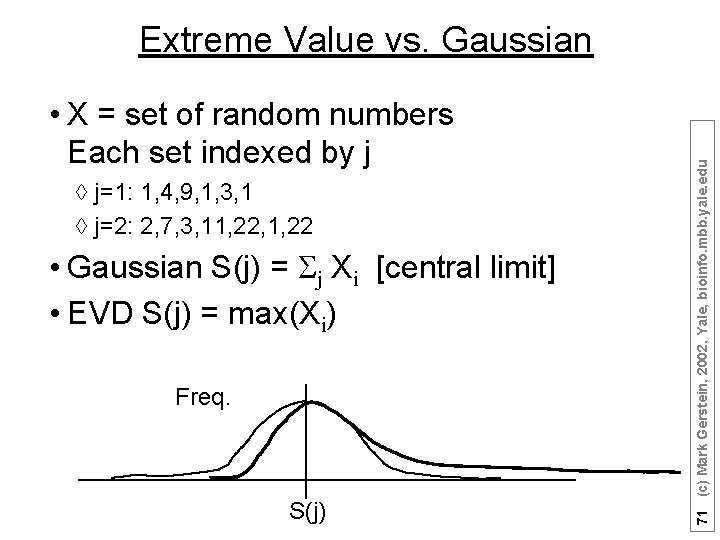
• X = set of random numbers Each set indexed by j à j=1: 1, 4, 9, 1, 3, 1 à j=2: 2, 7, 3, 11, 22, 1, 22 • Gaussian S(j) = j Xi [central limit] • EVD S(j) = max(Xi) Freq. S(j) 71 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Extreme Value vs. Gaussian

72 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu End of class (m-5) 2005, 10. 31 (quiz-1 is up to here)

• Score (Significance) Threshold • Maximize Coverage with an Acceptable Error Rate (graphic adapted from M Levitt) • TP, TN, FP, FN; TP and TN are good! • We get *P and *N from our program • We get T* and F* from a gold-standard • Want Max(TP, TN) vs (FP, FN) 73 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Objective is to Find Distant Homologues
![100% Coverage (roughly, fraction of sequences that one confidently “says something” about) [sensitivity=tp/p=tp/(tp+fn)] Core 100% Coverage (roughly, fraction of sequences that one confidently “says something” about) [sensitivity=tp/p=tp/(tp+fn)] Core](http://slidetodoc.com/presentation_image_h2/84cd8b00bd54d40c816274cfdc7bf6b2/image-74.jpg)
100% Coverage (roughly, fraction of sequences that one confidently “says something” about) [sensitivity=tp/p=tp/(tp+fn)] Core Thresh=10 Thresh=20 Thresh=30 Different score thresholds 100% Two “methods” (red is more effective) Error rate (fraction of the “statements” that are false positives) [Specificity = tn/n =tn/(tn+fp)] error rate = 1 -specificity = fp/n 74 (c) Mark Gerstein, 2002, Yale, bioinfo. mbb. yale. edu Coverage v Error Rate (ROC Graph)
- Slides: 74