1 Mark Gerstein Yale University gersteinlab orgcourses452 last

1 - Mark Gerstein, Yale University gersteinlab. org/courses/452 (last edit in Spring '19, pack #12) Lectures. Gerstein. Lab. org Biomedical Data Science Hi-C Analysis

Hi-C analysis illustrates much of the material in the class 2 - • Illustrates the evolution of the problem of annotating active & repressed regions in the genome - Original formulation in terms of “peak calling” on the linear genome - Revision of the original work, now at multi-scale - Recent radical change: now thinking of the genome as a 3 D folded molecule Lectures. Gerstein. Lab. org • Provides an illustration of - How machine learning functions to make sense of large, complex datasets - Network topology - Aggregation plots - Spectral methods (SVD)
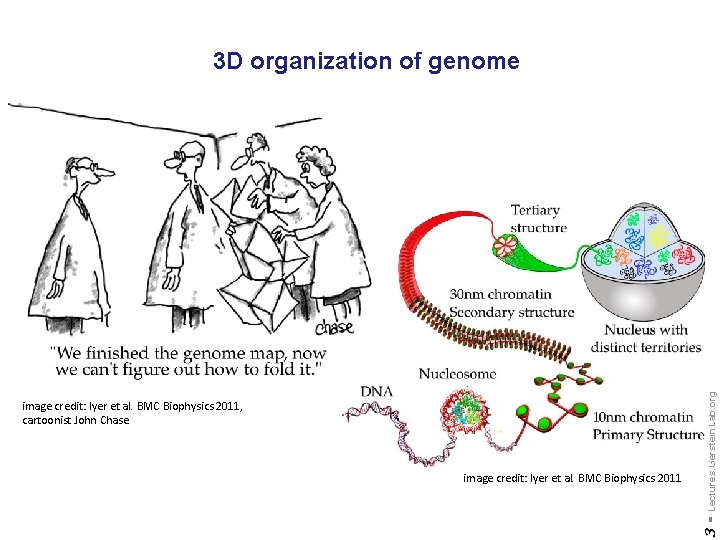
image credit: Iyer et al. BMC Biophysics 2011 3 - image credit: Iyer et al. BMC Biophysics 2011, cartoonist John Chase Lectures. Gerstein. Lab. org 3 D organization of genome
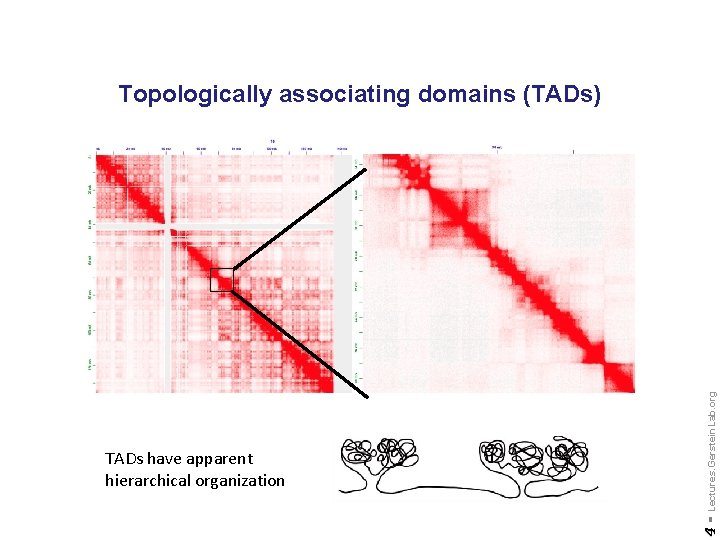
4 - TADs have apparent hierarchical organization Lectures. Gerstein. Lab. org Topologically associating domains (TADs)

5 - Lectures. Gerstein. Lab. org Modularity
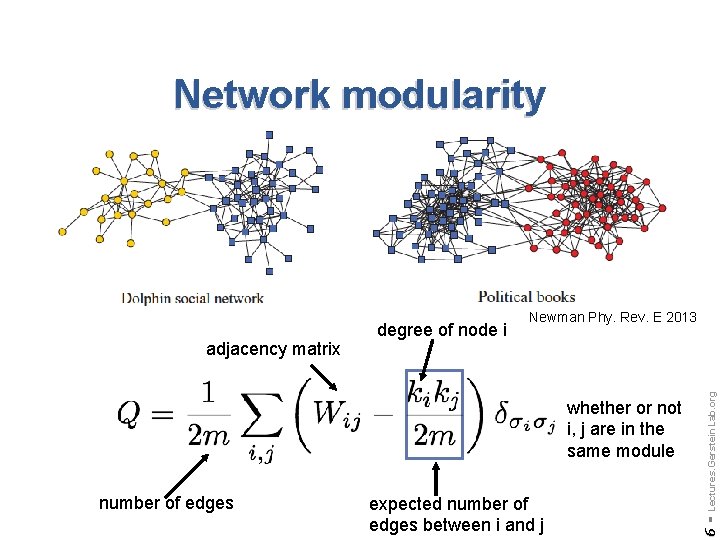
Network modularity whether or not i, j are in the same module number of edges expected number of edges between i and j Lectures. Gerstein. Lab. org Newman Phy. Rev. E 2013 6 - adjacency matrix degree of node i
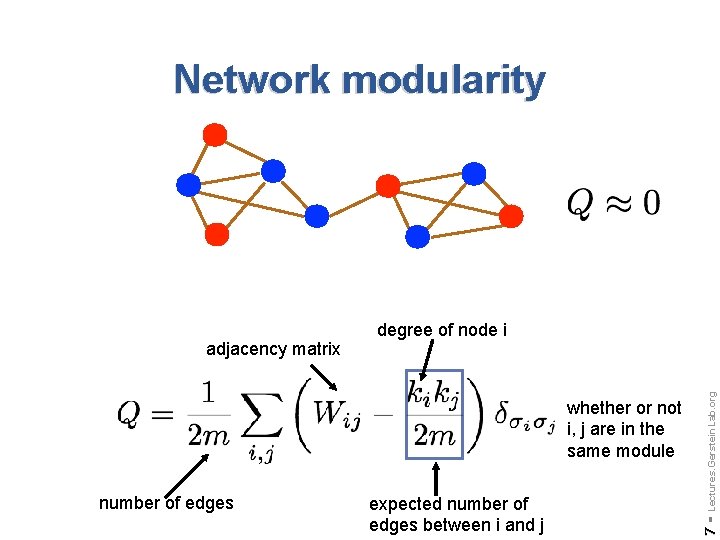
Network modularity number of edges expected number of edges between i and j Lectures. Gerstein. Lab. org whether or not i, j are in the same module 7 - adjacency matrix degree of node i
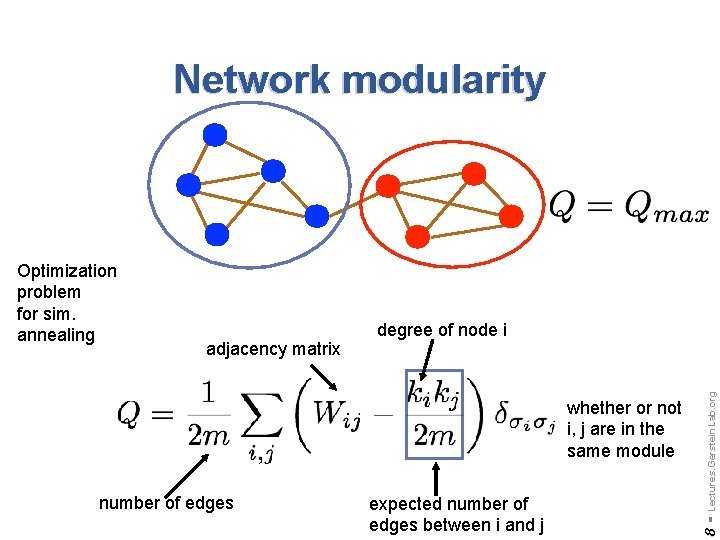
Network modularity whether or not i, j are in the same module number of edges expected number of edges between i and j Lectures. Gerstein. Lab. org adjacency matrix degree of node i 8 - Optimization problem for sim. annealing

9 - Lectures. Gerstein. Lab. org TAD Finding
![[Yan et al. , PLOS Comp. Bio. (in revision, ‘ 17); bio. Rxiv 097345] [Yan et al. , PLOS Comp. Bio. (in revision, ‘ 17); bio. Rxiv 097345]](http://slidetodoc.com/presentation_image_h/192f3fbc10b1c3fa1a054654e02a3c28/image-10.jpg)
[Yan et al. , PLOS Comp. Bio. (in revision, ‘ 17); bio. Rxiv 097345] 10 - Lectures. Gerstein. Lab. org Identifying TADs in multiple resolutions
![[Yan et al. , PLOS Comp. Bio. (in revision, ‘ 17); bio. Rxiv 097345] [Yan et al. , PLOS Comp. Bio. (in revision, ‘ 17); bio. Rxiv 097345]](http://slidetodoc.com/presentation_image_h/192f3fbc10b1c3fa1a054654e02a3c28/image-11.jpg)
[Yan et al. , PLOS Comp. Bio. (in revision, ‘ 17); bio. Rxiv 097345] 11 - Lectures. Gerstein. Lab. org Identifying TADs in multiple resolutions
![[Yan et al. , PLOS Comp. Bio. (in revision, ‘ 17); bio. Rxiv 097345] [Yan et al. , PLOS Comp. Bio. (in revision, ‘ 17); bio. Rxiv 097345]](http://slidetodoc.com/presentation_image_h/192f3fbc10b1c3fa1a054654e02a3c28/image-12.jpg)
[Yan et al. , PLOS Comp. Bio. (in revision, ‘ 17); bio. Rxiv 097345] Identifying TADs in multiple resolutions Lectures. Gerstein. Lab. org in equations 12 - Numerically solve for
![[Yan et al. , PLOS Comp. Bio. (in revision, ‘ 17); bio. Rxiv 097345] [Yan et al. , PLOS Comp. Bio. (in revision, ‘ 17); bio. Rxiv 097345]](http://slidetodoc.com/presentation_image_h/192f3fbc10b1c3fa1a054654e02a3c28/image-13.jpg)
[Yan et al. , PLOS Comp. Bio. (in revision, ‘ 17); bio. Rxiv 097345] 13 - Lectures. Gerstein. Lab. org Identifying TADs in multiple resolutions
![[Yan et al. , PLOS Comp. Bio. (in revision, ‘ 17); bio. Rxiv 097345] [Yan et al. , PLOS Comp. Bio. (in revision, ‘ 17); bio. Rxiv 097345]](http://slidetodoc.com/presentation_image_h/192f3fbc10b1c3fa1a054654e02a3c28/image-14.jpg)
[Yan et al. , PLOS Comp. Bio. (in revision, ‘ 17); bio. Rxiv 097345] Identifying TADs in multiple resolutions 14 - Lectures. Gerstein. Lab. org a modified Louvain algorithm
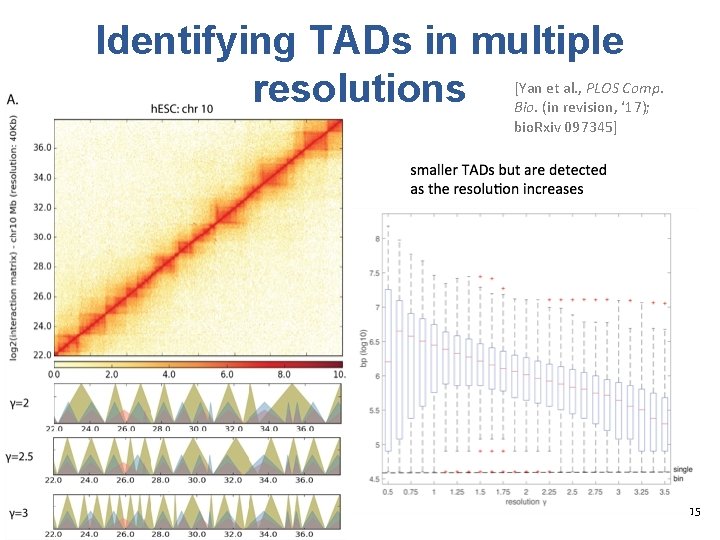
Identifying TADs in multiple resolutions [Yan et al. , PLOS Comp. Bio. (in revision, ‘ 17); bio. Rxiv 097345] 15

Enrichment of histone features at different resolution [Yan et al. , PLOS Comp. Bio. (in revision, ‘ 17); bio. Rxiv 097345] 16

17 - Lectures. Gerstein. Lab. org Using Matrix Decomposition for Hi-C Contact Matrices
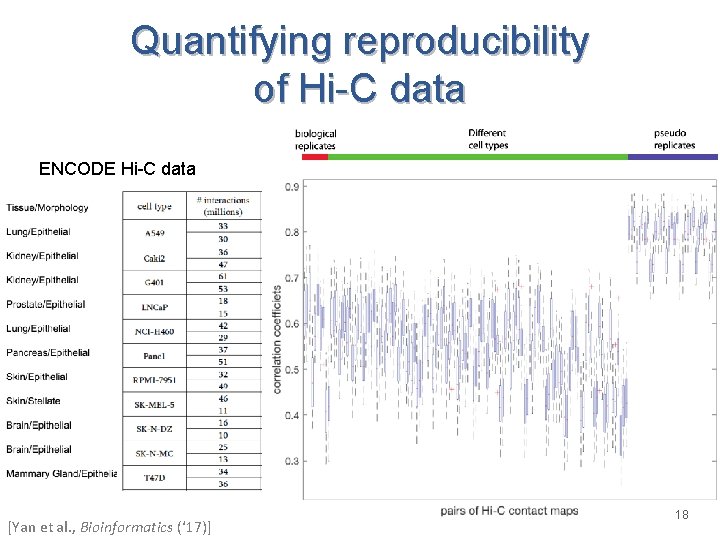
Quantifying reproducibility of Hi-C data ENCODE Hi-C data [Yan et al. , Bioinformatics (‘ 17)] 18
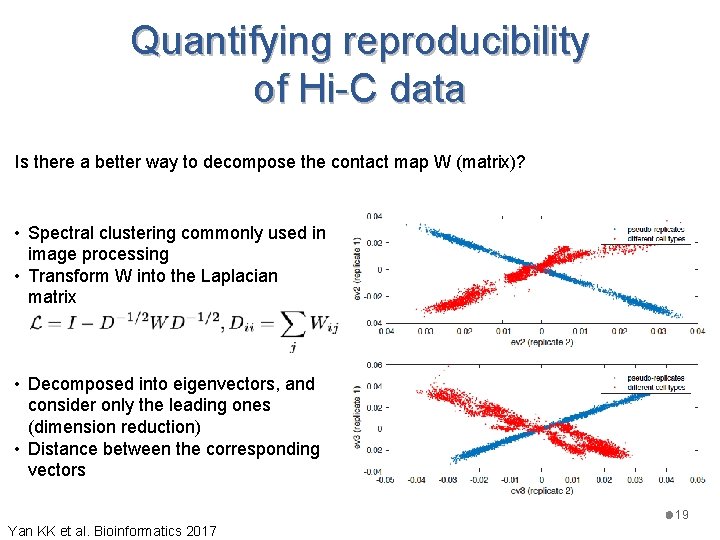
Quantifying reproducibility of Hi-C data Is there a better way to decompose the contact map W (matrix)? • Spectral clustering commonly used in image processing • Transform W into the Laplacian matrix • Decomposed into eigenvectors, and consider only the leading ones (dimension reduction) • Distance between the corresponding vectors 19 Yan KK et al. Bioinformatics 2017
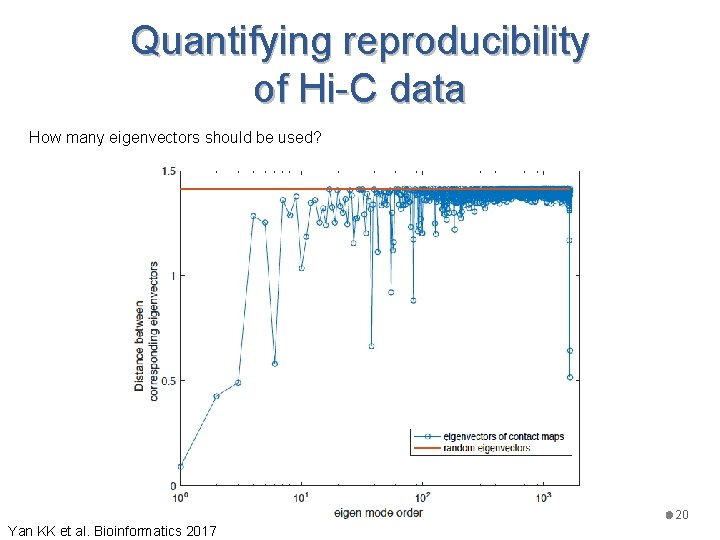
Quantifying reproducibility of Hi-C data How many eigenvectors should be used? 20 Yan KK et al. Bioinformatics 2017
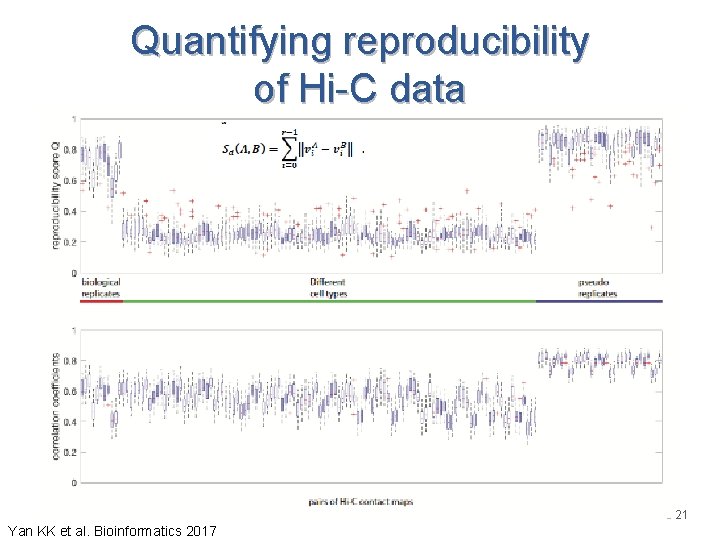
Quantifying reproducibility of Hi-C data 21 Yan KK et al. Bioinformatics 2017
![A distance measure between two contact maps [Yan et al. , Bioinformatics (‘ 17)] A distance measure between two contact maps [Yan et al. , Bioinformatics (‘ 17)]](http://slidetodoc.com/presentation_image_h/192f3fbc10b1c3fa1a054654e02a3c28/image-22.jpg)
A distance measure between two contact maps [Yan et al. , Bioinformatics (‘ 17)] 22
- Slides: 22