Mark Gerstein Yale University gersteinlab orgcourses452 last edit
Mark Gerstein, Yale University gersteinlab. org/courses/452 (last edit in spring '10) 1 (c) M Gerstein, 2010, Yale, gersteinlab. org BIOINFORMATICS Introduction

2 (c) M Gerstein, 2010, Yale, gersteinlab. org Start of class #1 [2006, 01. 13]

Bioinformatics + Computer Calculations 3 (c) M Gerstein, 2010, Yale, gersteinlab. org Biological Data

Core • (Molecular) Bio - informatics • One idea for a definition? Bioinformatics is conceptualizing biology in terms of molecules (in the sense of physical-chemistry) and then applying “informatics” techniques (derived from disciplines such as applied math, CS, and statistics) to understand organize the information associated with these molecules, on a large-scale. • Bioinformatics is a practical discipline with many applications. 4 (c) M Gerstein, 2010, Yale, gersteinlab. org What is Bioinformatics?

What is the Information? Molecular Biology as an Information Science DNA -> RNA -> Protein -> Phenotype -> DNA • Molecules à Sequence, Structure, Function • Processes à Mechanism, Specificity, Regulation • Genetic material • Information transfer (m. RNA) • Protein synthesis (t. RNA/m. RNA) • Some catalytic activity • Central Paradigm for Bioinformatics Genomic Sequence Information -> m. RNA (level) -> Protein Sequence -> Protein Structure -> Protein Function -> Phenotype • Large Amounts of Information à Standardized à Statistical • Most cellular functions are performed or facilitated by proteins. • Primary biocatalyst • Cofactor transport/storage • Mechanical motion/support • Immune protection • Control of growth/differentiation (idea from D Brutlag, Stanford, graphics from S Strobel) 5 (c) M Gerstein, 2010, Yale, gersteinlab. org • Central Dogma of Molecular Biology

Molecular Biology Information - DNA à Coding or Not? à Parse into genes? à 4 bases: AGCT à ~1 K in a gene, ~2 M in genome à ~3 Gb Human atggcaattaaaattggtatcaatggttttggtcgtatcggccgtattccgtgca gcacaacaccgtgatgacattgaagttgtaggtattaacgacttaatcgacgttgaatac atggcttatatgttgaaatatgattcaactcacggtcgtttcgacggcactgttgaagtg aaagatggtaacttagtggttaatggtaaaactatccgtgtaactgcagaacgtgatcca gcaaacttaaactggggtgcaatcggtgttgatatcgctgttgaagcgactggtttattc ttaactgatgaaactgctcgtaaacatatcactgcaggcgcaaaaaaagttgtattaact ggcccatctaaagatgcaacccctatgttcgtggtgtaaacttcaacgcatacgca ggtcaagatatcgtttctaacgcatcttgtacaacaaactgtttagctcctttagcacgt gttgttcatgaaactttcggtatcaaagatggtttaatgaccactgttcacgcaacgact gcaactcaaaaaactgtggatggtccatcagctaaagactggcgcggcggccgcggtgca tcacaaaacatcattccatcttcaacaggtgcagcgaaagcagtaggtaaagtattacct gcattaaacggtaaattaactggtatggctttccgtgttccaacgccaaacgtatctgtt gttgatttaacagttaatcttgaaaaaccagcttcttatgatgcaatcaaacaagcaatc aaagatgcagcggaaggtaaaacgttcaatggcgaattaaaaggcgtattaggttacact gaagatgctgttgtttctactgacttcaacggttgtgctttaacttctgtatttgatgca gacgctggtatcgcattaactgattctttcgttaaattggtatc. . . caaaaatagggttaatatgaatctcgatctccattttgttcatcgtattcaa caacaagccaaaactcgtacaaatatgaccgcacttcgctataaagaacacggcttgtgg cgagatatctcttggaaaaactttcaagagcaactcaactttctcgagcattgctt gctcacaatattgacgtacaagataaaatcgccatttttgcccataatatggaacgttgg gttgttcatgaaactttcggtatcaaagatggtttaatgaccactgttcacgcaacgact acaatcgttgacattgcgaccttacaaattcgagcaatcacagtgcctatttacgcaacc aatacagcccagcagaatttatcctaaatcacgccgatgtaaaaattctcttcgtc ggcgatcaagagcaatacgatcaaacattggaaattgctcatcattgtccaaaattacaa aaaattgtagcaatgaaatccaccattcaattacaacaagatcctcttgcacttgg 6 (c) M Gerstein, 2010, Yale, gersteinlab. org • Raw DNA Sequence

Molecular Biology Information: Protein Sequence • 20 letter alphabet but not BJOUXZ • Strings of ~300 aa in an average protein (in bacteria), ~200 aa in a domain • >4 M known protein sequences (uniprot, 2006) d 1 dhfa_ d 8 dfr__ d 4 dfra_ d 3 dfr__ LNCIVAVSQNMGIGKNGDLPWPPLRNEFRYFQRMTTTSSVEGKQ-NLVIMGKKTWFSI LNSIVAVCQNMGIGKDGNLPWPPLRNEYKYFQRMTSTSHVEGKQ-NAVIMGKKTWFSI ISLIAALAVDRVIGMENAMPWN-LPADLAWFKRNTL----NKPVIMGRHTWESI TAFLWAQDRDGLIGKDGHLPWH-LPDDLHYFRAQTV----GKIMVVGRRTYESF d 1 dhfa_ d 8 dfr__ d 4 dfra_ d 3 dfr__ LNCIVAVSQNMGIGKNGDLPWPPLRNEFRYFQRMTTTSSVEGKQ-NLVIMGKKTWFSI LNSIVAVCQNMGIGKDGNLPWPPLRNEYKYFQRMTSTSHVEGKQ-NAVIMGKKTWFSI ISLIAALAVDRVIGMENAMPW-NLPADLAWFKRNTLD----KPVIMGRHTWESI TAFLWAQDRNGLIGKDGHLPW-HLPDDLHYFRAQTVG----KIMVVGRRTYESF d 1 dhfa_ d 8 dfr__ d 4 dfra_ d 3 dfr__ VPEKNRPLKGRINLVLSRELKEPPQGAHFLSRSLDDALKLTEQPELANKVDMVWIVGGSSVYKEAMNHP VPEKNRPLKDRINIVLSRELKEAPKGAHYLSKSLDDALALLDSPELKSKVDMVWIVGGTAVYKAAMEKP ---G-RPLPGRKNIILS-SQPGTDDRV-TWVKSVDEAIAACGDVP------EIMVIGGGRVYEQFLPKA ---PKRPLPERTNVVLTHQEDYQAQGA-VVVHDVAAVFAYAKQHLDQ----ELVIAGGAQIFTAFKDDV d 1 dhfa_ d 8 dfr__ d 4 dfra_ d 3 dfr__ -PEKNRPLKGRINLVLSRELKEPPQGAHFLSRSLDDALKLTEQPELANKVDMVWIVGGSSVYKEAMNHP -PEKNRPLKDRINIVLSRELKEAPKGAHYLSKSLDDALALLDSPELKSKVDMVWIVGGTAVYKAAMEKP -G---RPLPGRKNIILSSSQPGTDDRV-TWVKSVDEAIAACGDVPE-----. IMVIGGGRVYEQFLPKA -P--KRPLPERTNVVLTHQEDYQAQGA-VVVHDVAAVFAYAKQHLD----QELVIAGGAQIFTAFKDDV 7 (c) M Gerstein, 2010, Yale, gersteinlab. org à ACDEFGHIKLMNPQRSTVWY

Molecular Biology Information: Macromolecular Structure • DNA/RNA/Protein à Almost all protein 8 (c) M Gerstein, 2010, Yale, gersteinlab. org (RNA Adapted From D Soll Web Page, Right Hand Top Protein from M Levitt web page)
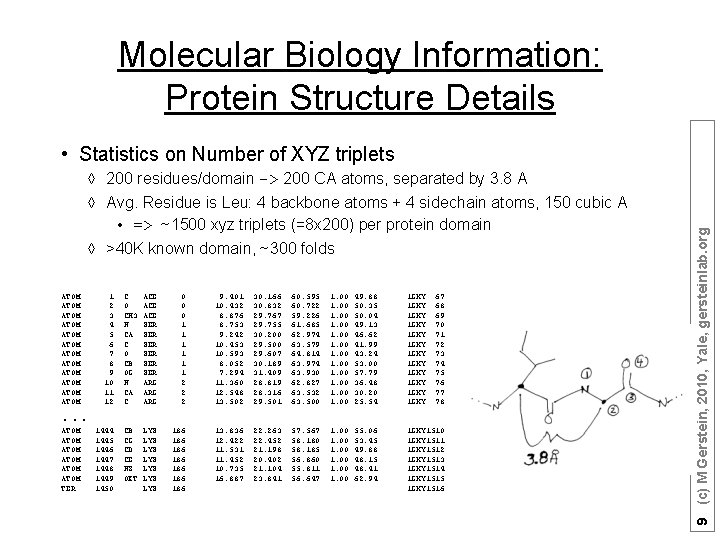
Molecular Biology Information: Protein Structure Details à 200 residues/domain -> 200 CA atoms, separated by 3. 8 A à Avg. Residue is Leu: 4 backbone atoms + 4 sidechain atoms, 150 cubic A • => ~1500 xyz triplets (=8 x 200) per protein domain à >40 K known domain, ~300 folds ATOM ATOM ATOM 1 2 3 4 5 6 7 8 9 10 11 12 C O CH 3 N CA C O CB OG N CA C ACE ACE SER SER SER ARG ARG 0 0 0 1 1 1 2 2 2 9. 401 10. 432 8. 876 8. 753 9. 242 10. 453 10. 593 8. 052 7. 294 11. 360 12. 548 13. 502 30. 166 30. 832 29. 767 29. 755 30. 200 29. 500 29. 607 30. 189 31. 409 28. 819 28. 316 29. 501 60. 595 60. 722 59. 226 61. 685 62. 974 63. 579 64. 814 63. 974 63. 930 62. 827 63. 532 63. 500 1. 00 1. 00 49. 88 50. 35 50. 04 49. 13 46. 62 41. 99 43. 24 53. 00 57. 79 36. 48 30. 20 25. 54 1 GKY 1 GKY 1 GKY 67 68 69 70 71 72 73 74 75 76 77 78 1444 1445 1446 1447 1448 1449 1450 CB CG CD CE NZ OXT LYS LYS 186 186 13. 836 12. 422 11. 531 11. 452 10. 735 16. 887 22. 263 22. 452 21. 198 20. 402 21. 104 23. 841 57. 567 58. 180 58. 185 56. 860 55. 811 56. 647 1. 00 55. 06 53. 45 49. 88 48. 15 48. 41 62. 94 1 GKY 1510 1 GKY 1511 1 GKY 1512 1 GKY 1513 1 GKY 1514 1 GKY 1515 1 GKY 1516 . . . ATOM ATOM TER 9 (c) M Gerstein, 2010, Yale, gersteinlab. org • Statistics on Number of XYZ triplets

Molecular Biology Information: Whole Genomes • The Revolution Driving Everything Kerlavage, A. R. , Bult, C. J. , Tomb, J. F. , Dougherty, B. A. , Merrick, J. M. , Mc. Kenney, K. , Sutton, G. , Fitzhugh, W. , Fields, C. , Gocayne, J. D. , Scott, J. , Shirley, R. , Liu, L. I. , Glodek, A. , Kelley, J. M. , Weidman, J. F. , Phillips, C. A. , Spriggs, T. , Hedblom, E. , Cotton, M. D. , Utterback, T. R. , Hanna, M. C. , Nguyen, D. T. , Saudek, D. M. , Brandon, R. C. , Fine, L. D. , Fritchman, J. L. , Fuhrmann, J. L. , Geoghagen, N. S. M. , Gnehm, C. L. , Mc. Donald, L. A. , Venter, J. C. (1995). "Wholegenome random sequencing and assembly of Haemophilus influenzae rd. " Genome sequence now Science 269: 496 -512. accumulate so quickly that, (Picture adapted from TIGR website, in less than a week, a single http: //www. tigr. org) laboratory can produce • Integrative Data more bits of data than 1995, HI (bacteria): 1. 6 Mb & 1600 genes done Shakespeare managed in a 1997, yeast: 13 Mb & ~6000 genes for yeast lifetime, although the latter 1998, worm: ~100 Mb with 19 K genes make better reading. Small, K. V. , Fraser, C. M. , Smith, H. O. & 1999: >30 completed genomes! 2003, human: 3 Gb & 100 K genes. . . -- G A Pekso, Nature 401: 115 -116 (1999) 10 (c) M Gerstein, 2010, Yale, gersteinlab. org Fleischmann, R. D. , Adams, M. D. , White, O. , Clayton, R. A. , Kirkness, E. F. ,
![Bacteria, 1. 6 Mb, ~1600 genes [Science 269: 496] 1997 Eukaryote, 13 Mb, ~6 Bacteria, 1. 6 Mb, ~1600 genes [Science 269: 496] 1997 Eukaryote, 13 Mb, ~6](http://slidetodoc.com/presentation_image_h/0a40b064f93c920f7b7e2d2843b60107/image-11.jpg)
Bacteria, 1. 6 Mb, ~1600 genes [Science 269: 496] 1997 Eukaryote, 13 Mb, ~6 K genes [Nature 387: 1] Genomes highlight the Finiteness of the “Parts” in Biology 1998 real thing, Apr ‘ 00 Animal, ~100 Mb, ~20 K genes [Science 282: 1945] 2000? Human, ~3 Gb, ~100 K genes [? ? ? ] ‘ 98 spoof 11 (c) M Gerstein, 2010, Yale, gersteinlab. org 1995

Other Types of Data à Early experiments yeast • Complexity at 10 time points, 6000 x 10 = 60 K floats à Now tiling array technology • 50 M data points to tile the human genome at ~50 bp res. à Can only sequence genome once but can do an infinite variety of array experiments • Phenotype Experiments à KOs, transposons • Protein Interactions à For yeast: 6000 x 6000 / 2 ~ 18 M possible interactions à maybe 30 K real • Regulatory Networks 12 (c) M Gerstein, 2010, Yale, gersteinlab. org • Gene Expression
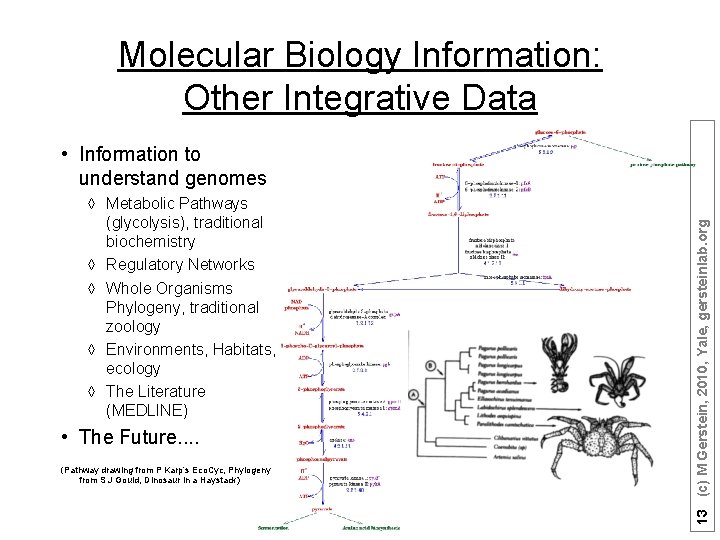
Molecular Biology Information: Other Integrative Data à Metabolic Pathways (glycolysis), traditional biochemistry à Regulatory Networks à Whole Organisms Phylogeny, traditional zoology à Environments, Habitats, ecology à The Literature (MEDLINE) • The Future. . (Pathway drawing from P Karp’s Eco. Cyc, Phylogeny from S J Gould, Dinosaur in a Haystack) 13 (c) M Gerstein, 2010, Yale, gersteinlab. org • Information to understand genomes

• (Molecular) Bio - informatics • One idea for a definition? Bioinformatics is conceptualizing biology in terms of molecules (in the sense of physical-chemistry) and then applying “informatics” techniques (derived from disciplines such as applied math, CS, and statistics) to understand organize the information associated with these molecules, on a large-scale. • Bioinformatics is a practical discipline with many applications. 14 (c) M Gerstein, 2010, Yale, gersteinlab. org What is Bioinformatics?
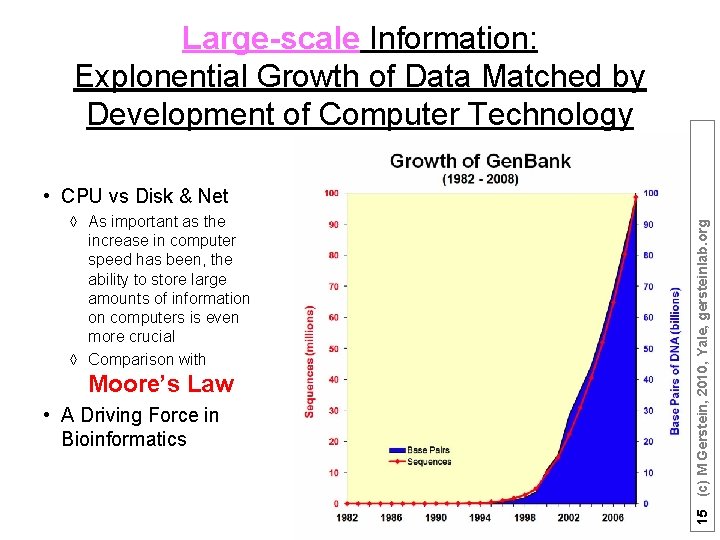
Large-scale Information: Explonential Growth of Data Matched by Development of Computer Technology à As important as the increase in computer speed has been, the ability to store large amounts of information on computers is even more crucial à Comparison with Moore’s Law • A Driving Force in Bioinformatics 15 (c) M Gerstein, 2010, Yale, gersteinlab. org • CPU vs Disk & Net
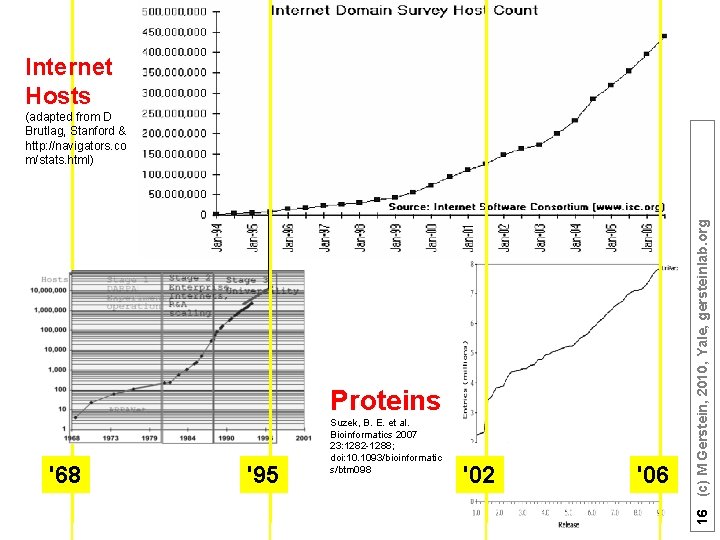
Internet Hosts Proteins '68 '95 Suzek, B. E. et al. Bioinformatics 2007 23: 1282 -1288; doi: 10. 1093/bioinformatic s/btm 098 '02 '06 16 (c) M Gerstein, 2010, Yale, gersteinlab. org (adapted from D Brutlag, Stanford & http: //navigators. co m/stats. html)
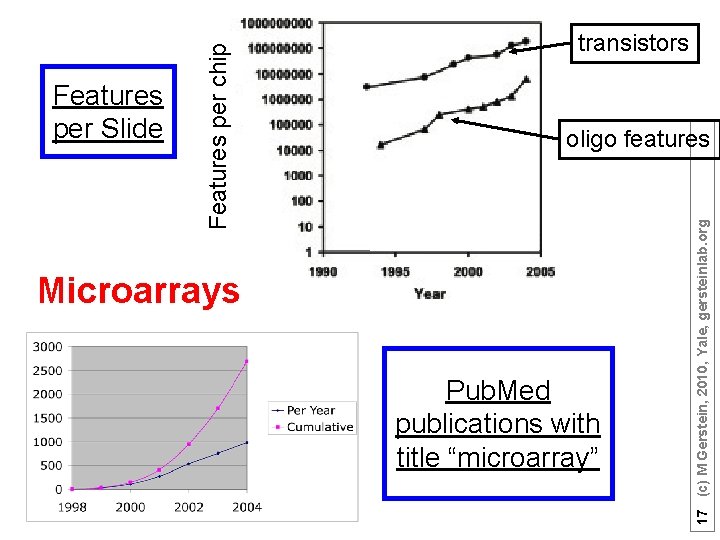
oligo features Microarrays Pub. Med publications with title “microarray” 17 (c) M Gerstein, 2010, Yale, gersteinlab. org Features per chip Features per Slide transistors
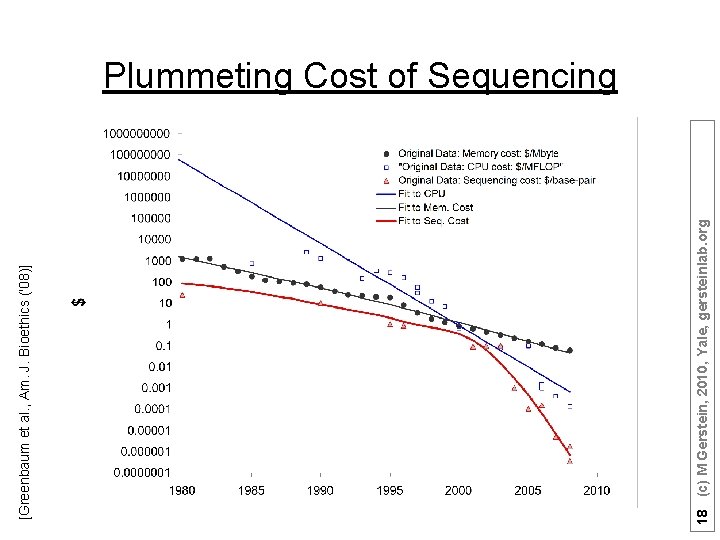
18 (c) M Gerstein, 2010, Yale, gersteinlab. org [Greenbaum et al. , Am. J. Bioethics ('08)] Plummeting Cost of Sequencing

(courtesy of Finn Drablos) 19 (c) M Gerstein, 2010, Yale, gersteinlab. org Jobs: Bioinformatics is born!

• (Molecular) Bio - informatics • One idea for a definition? Bioinformatics is conceptualizing biology in terms of molecules (in the sense of physical-chemistry) and then applying “informatics” techniques (derived from disciplines such as applied math, CS, and statistics) to understand organize the information associated with these molecules, on a large-scale. • Bioinformatics is a practical discipline with many applications. 20 (c) M Gerstein, 2010, Yale, gersteinlab. org What is Bioinformatics?
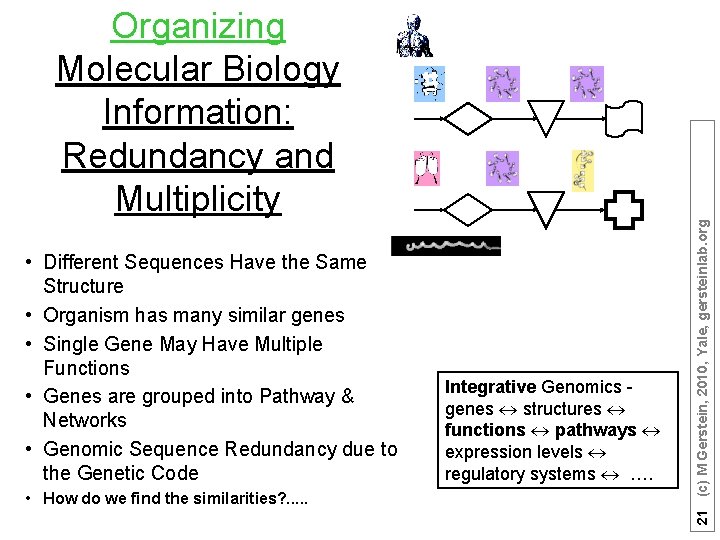
• Different Sequences Have the Same Structure • Organism has many similar genes • Single Gene May Have Multiple Functions • Genes are grouped into Pathway & Networks • Genomic Sequence Redundancy due to the Genetic Code • How do we find the similarities? . . . Integrative Genomics genes structures functions pathways expression levels regulatory systems …. 21 (c) M Gerstein, 2010, Yale, gersteinlab. org Organizing Molecular Biology Information: Redundancy and Multiplicity
22 (c) M Gerstein, 2010, Yale, gersteinlab. org Molecular Parts = Conserved Domains, Folds, &c
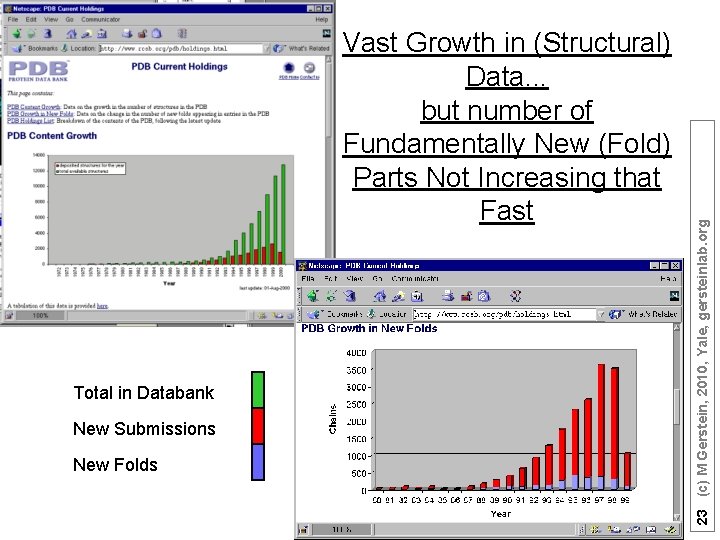
Total in Databank New Submissions New Folds 23 (c) M Gerstein, 2010, Yale, gersteinlab. org Vast Growth in (Structural) Data. . . but number of Fundamentally New (Fold) Parts Not Increasing that Fast
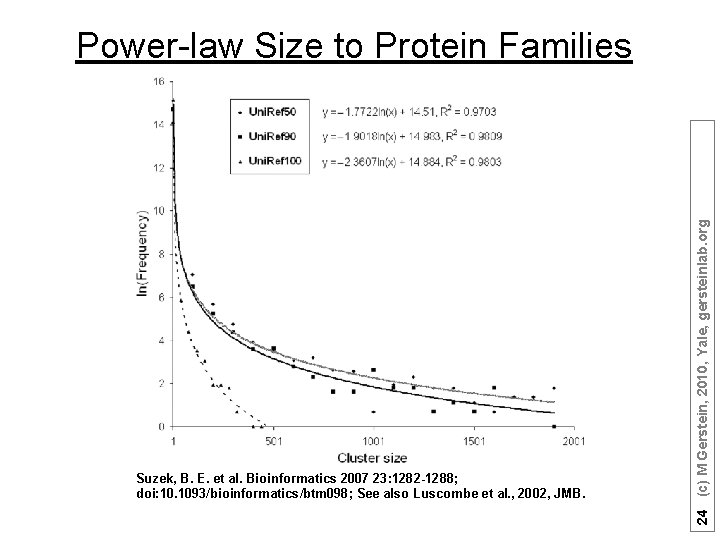
Suzek, B. E. et al. Bioinformatics 2007 23: 1282 -1288; doi: 10. 1093/bioinformatics/btm 098; See also Luscombe et al. , 2002, JMB. 24 (c) M Gerstein, 2010, Yale, gersteinlab. org Power-law Size to Protein Families

• (Molecular) Bio - informatics • One idea for a definition? Bioinformatics is conceptualizing biology in terms of molecules (in the sense of physical-chemistry) and then applying “informatics” techniques (derived from disciplines such as applied math, CS, and statistics) to understand organize the information associated with these molecules, on a large-scale. • Bioinformatics is a practical discipline with many applications. 25 (c) M Gerstein, 2010, Yale, gersteinlab. org What is Bioinformatics?

General Types of “Informatics” techniques in Bioinformatics à Building, Querying à Representing Complex data • Datamining à Machine Learning à Clustering à Text String Comparison • • • Text Search 1 D Alignment Tree construction à Significance Statistics • Network Analysis • Structure Analysis & Geometry à Graphics (Surfaces, Volumes) à Comparison and 3 D Matching (Vision, recognition) • Physical Simulation à à à Newtonian Mechanics Electrostatics Numerical Algorithms Simulation Modelling Chemical Reactions & Cellular Processes 26 (c) M Gerstein, 2010, Yale, gersteinlab. org • Databases

Data Mining as New Paradigm for Scientific Computing à Prediction based on physical principles à EX: Exact Determination of Rocket Trajectory à Emphasizes: Supercomputer, CPU • Biology à Classifying information and discovering unexpected relationships à EX: Gene Expression Network à Emphasizes: networks, “federated” database 27 (c) M Gerstein, 2010, Yale, gersteinlab. org • Physics

28 (c) M Gerstein, 2010, Yale, gersteinlab. org NY Times 14 Dec-09 article on Jim Gray's 4 th Paradigm

Vs. Chemical Understanding, Mechanism, Molecular Biology How Does Prediction Fit into the Definition? 29 (c) M Gerstein, 2010, Yale, gersteinlab. org Statistical Analysis vs. Classical Physics Bioinformatics, Genomic Surveys

Practical Stuff 30 (c) M Gerstein, 2010, Yale, gersteinlab. org Bioinformatics: Practical Application of Simulation and Data Mining

People & Times à MW 1. 00 -2. 15 BASS 305 à Two classes in Bass 405 (25 Jan. and 1 Feb. ) à Discussion sect. each week (TBD) • Instructors à Mark Gerstein (in charge), data mining lectures • TFs à Raymond Auerbach à Lukas Habegger • Office Hours for Mark à 15' after this class à 30' after class on Wed. à By appointment • James Noonan + others à Corey S. O'Hern, simulation lectures • Steven Kleinstein • No Fri. class this week 31 (c) M Gerstein, 2010, Yale, gersteinlab. org • Timing

gersteinlab. org/courses/452 • Will post exact schedule soon! • PDFs and PPTs of lectures • Previous year's courses (more than 10 years!) • cbb 752@gersteinlab. org 32 (c) M Gerstein, 2010, Yale, gersteinlab. org http: //www.
![• Quizzes (~4) [1 st probably on 2 Feb. ] • No Final • Quizzes (~4) [1 st probably on 2 Feb. ] • No Final](http://slidetodoc.com/presentation_image_h/0a40b064f93c920f7b7e2d2843b60107/image-33.jpg)
• Quizzes (~4) [1 st probably on 2 Feb. ] • No Final Exam • Discussion Section Participation • Final Project* • Homework* * CBB/CS students will do programming here and MBB will do writing • Prerequisites à MB&B 301 b and MATH 115 a or b, or permission of instructor 33 (c) M Gerstein, 2010, Yale, gersteinlab. org Grading

Two courses à Programming Assignments, Extra computation à Programming final project • MBB 452/752 à No programming à Written Final Project 34 (c) M Gerstein, 2010, Yale, gersteinlab. org • CBB 752/CS 752

Techniques in data mining & simulation applied to bioinformatics, the computational analysis of gene sequences, macromolecular structures, and functional genomics data on a large scale. (Some topics include: ) Sequence alignment, comparative genomics and phylogenetics, biological databases, geometric analysis of protein structure, molecular-dynamics simulation, biological networks, microarray normalization, and machinelearning approaches to data integration. Not the same as Genomics & Bioinformatics in previous years contains all of the "Bioinformatics" and then more (!) with less "Genomics". Previous years slides are a rough guide to the bioinformatics part. 35 (c) M Gerstein, 2010, Yale, gersteinlab. org Course Catalog Description

Bioinformatics Topics -Genome Sequence à introns à exons à promotors • Characterizing Repeats in Genomic DNA à Statistics à Patterns • Duplications in the Genome à Large scale genomic alignment • Whole-Genome Comparisons • Finding Structural RNAs 36 (c) M Gerstein, 2010, Yale, gersteinlab. org • Finding Genes in Genomic DNA

à non-exact string matching, gaps à How to align two strings optimally via Dynamic Programming à Local vs Global Alignment à Suboptimal Alignment à Hashing to increase speed (BLAST, FASTA) à Amino acid substitution scoring matrices • Multiple Alignment and Consensus Patterns à How to align more than one sequence and then fuse the result in a consensus representation à Transitive Comparisons à HMMs, Profiles à Motifs Bioinformatics Topics -Protein Sequence • Scoring schemes and Matching statistics à How to tell if a given alignment or match is statistically significant à A P-value (or an e-value)? à Score Distributions (extreme val. dist. ) à Low Complexity Sequences • Evolutionary Issues à Rates of mutation and change 37 (c) M Gerstein, 2010, Yale, gersteinlab. org • Sequence Alignment

• Secondary Structure “Prediction” à à à via Propensities Neural Networks, Genetic Alg. • Simple Statistics TM-helix finding Assessing Secondary Structure Prediction • Structure Prediction: Protein v RNA Tertiary Structure Prediction à à Fold Recognition Threading Ab initio (Quaternary structure prediction) • Direct Function Prediction à Active site identification • Relation of Sequence Similarity to Structural Similarity 38 (c) M Gerstein, 2010, Yale, gersteinlab. org Bioinformatics Topics -Sequence / Structure

Topics -- Structures à Basic Protein Geometry and Least-Squares Fitting • Distances, Angles, Axes, Rotations à Calculating a helix axis in 3 D via fitting a line à LSQ fit of 2 structures à Molecular Graphics • Calculation of Volume and Surface à à à How to represent a plane How to represent a solid How to calculate an area Hinge prediction Packing Measurement • Structural Alignment à Aligning sequences on the basis of 3 D structure. à DP does not converge, unlike sequences, what to do? à Other Approaches: Distance Matrices, Hashing • Fold Library • Docking and Drug Design as Surface Matching 39 (c) M Gerstein, 2010, Yale, gersteinlab. org • Structure Comparison
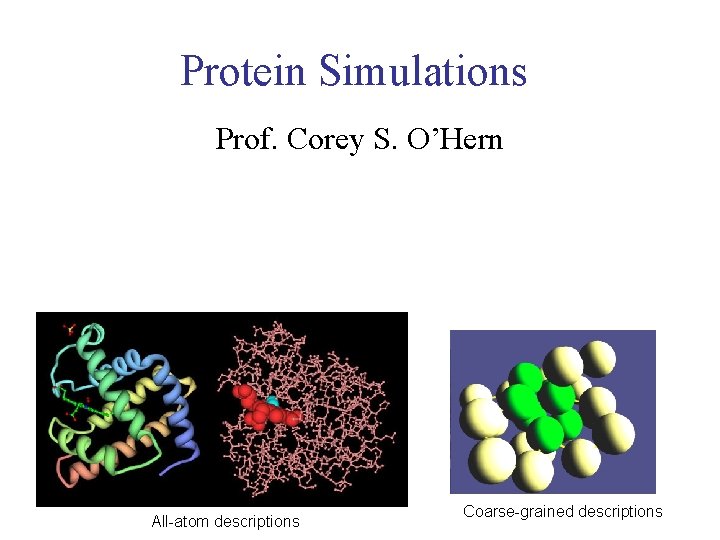
Protein Simulations Prof. Corey S. O’Hern All-atom descriptions Coarse-grained descriptions

Your model is oversimplified and has nothing to do with biology! Molecular biologist Your model is too complicated and has no predictive power! Biological Physicist
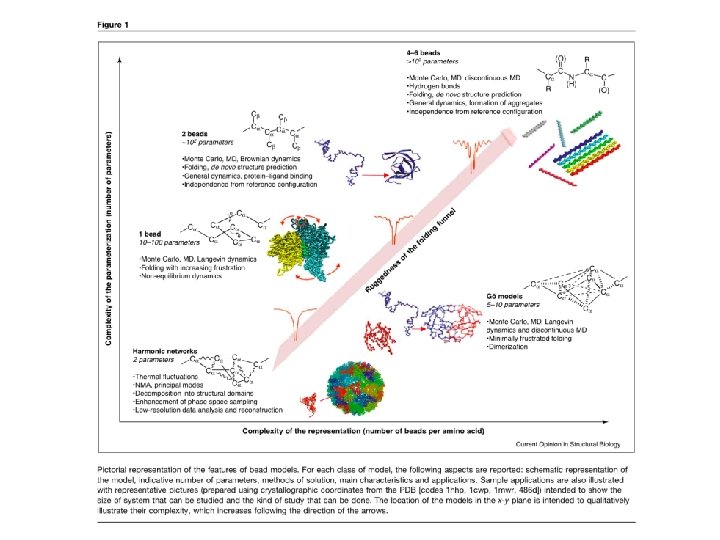
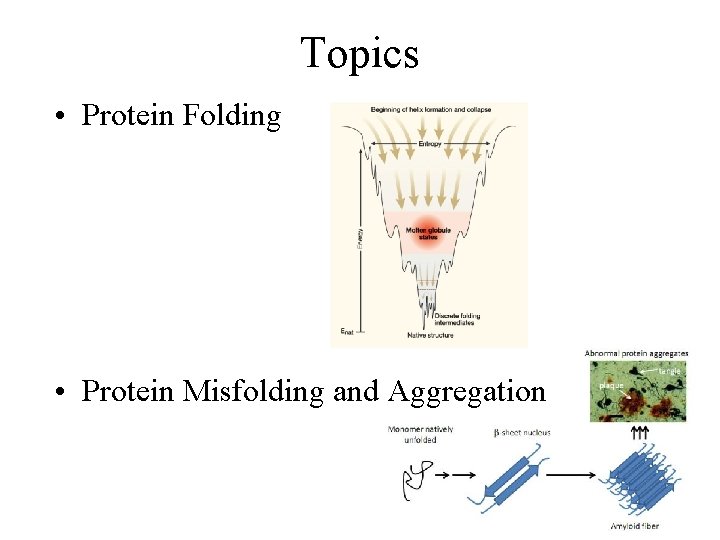
Topics • Protein Folding • Protein Misfolding and Aggregation

Journal Articles 1. J. D. Honeycutt and D. Thirumalai, “The nature of folded states of globular proteins, ” Biopolymers 32 (1992) 695. 2. W. C. Swope and J. W. Pitera, “Describing protein folding kinetics By molecular dynamics simulations. 1. Theory, ” J. Phys. Chem. B 108 (2004) 6571. 3. D. Bratko, T. Cellmer, J. M. Prausnitz, and H. W. Blanch, “Molecular Simulation of protein aggregation, ” Biotechnology and Bioengineering 96 (2007) 1.

Methods I Coarse-grained Brownian dynamics simulations of proteins
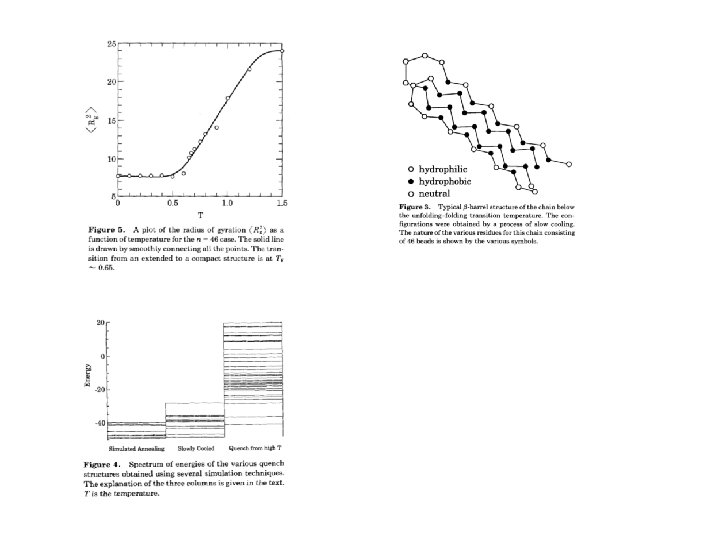

Methods II …energy minimization, simulated annealing, Markov models, Master-equation approaches, Monte Carlo simulations, advanced sampling techniques…
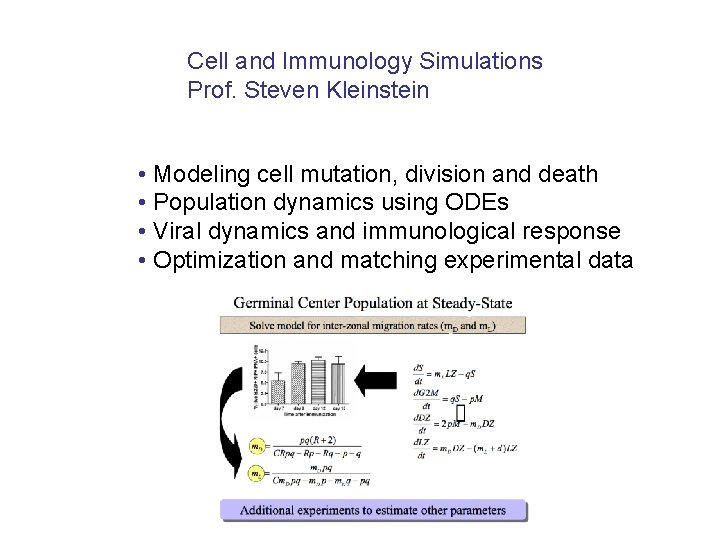
Cell and Immunology Simulations Prof. Steven Kleinstein • Modeling cell mutation, division and death • Population dynamics using ODEs • Viral dynamics and immunological response • Optimization and matching experimental data
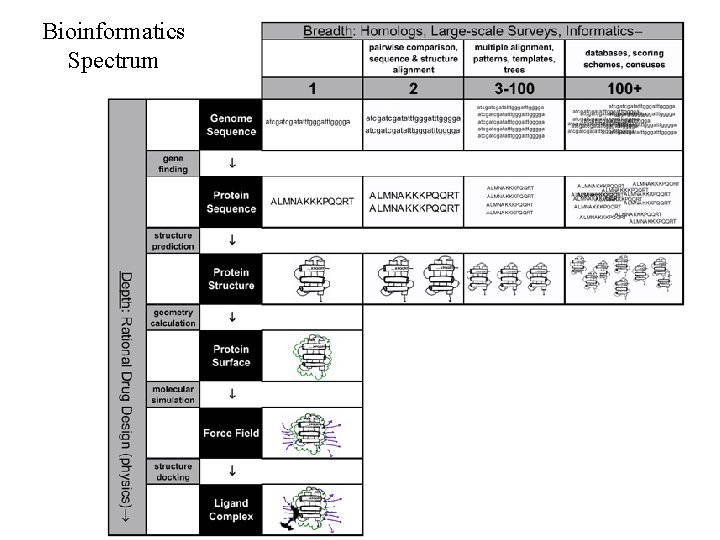
Bioinformatics Spectrum

Next Generation Sequencing & Big Data • • • Seq. Tech Assembly RNA-seq Ch. IP-seq Metagenomics

Topics – (Func) Genomics • Expression Analysis – – Time Courses clustering Measuring differences Identifying Regulatory Regions Normalization and scoring of arrays • Function Classification and Orthologs • The Genomic vs. Singlemolecule Perspective • Genome Comparisons – – Large-scale censuses Frequent Words Analysis Genome Annotation Identification of interacting proteins • Structural Genomics – Folds in Genomes, shared & common folds – Bulk Structure Prediction

• Relational Database Concepts and how they interface with Biological Information • DB interoperation • What are the Units ? – What are the units of biological information for organization? • sequence, structure • motifs, modules, domains – How classified: folds, motions, pathways, functions? Topics – DBs/Surveys • Clustering & Trees – Basic clustering • UPGMA • single-linkage • multiple linkage – Other Methods • Parsimony, Maximum likelihood – Evolutionary implications • Visualization of Large Amounts of Information • The Bias Problem – sequence weighting – sampling

Mining • Information integration and fusion – Dealing with heterogeneous data • Dimensionality Reduction (PCA etc) • Networks – Topology Analysis – Prediction – Global structure and local motifs


• (Molecular) Bio - informatics • One idea for a definition? Bioinformatics is conceptualizing biology in terms of molecules (in the sense of physical-chemistry) and then applying “informatics” techniques (derived from disciplines such as applied math, CS, and statistics) to understand organize the information associated with these molecules, on a large-scale. • Bioinformatics is a practical discipline with many applications. 55 (c) M Gerstein, 2010, Yale, gersteinlab. org What is Bioinformatics?

Core • Understanding How Structures Bind Other Molecules (Function) • Designing Inhibitors • Docking, Structure Modeling (From left to right, figures adapted from Olsen Group Docking Page at Scripps, Dyson NMR Group Web page at Scripps, and from Computational Chemistry Page at Cornell Theory Center). 56 (c) M Gerstein, 2010, Yale, gersteinlab. org Major Application I: Designing Drugs
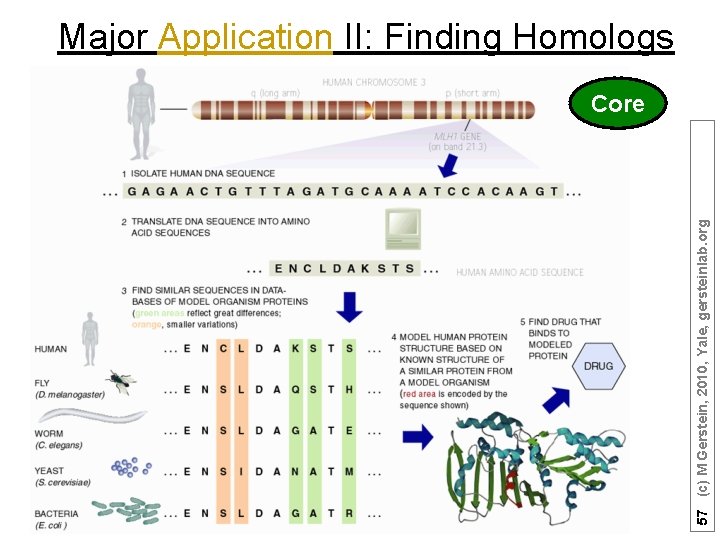
57 (c) M Gerstein, 2010, Yale, gersteinlab. org Major Application II: Finding Homologs Core

• Overall Occurrence of a Certain Feature in the Genome à e. g. how many kinases in Yeast • Compare Organisms and Tissues à Expression levels in Cancerous vs Normal Tissues • Databases, Statistics • Using this for picking drug targets (Clock figures, yeast v. Synechocystis, adapted from Gene. Quiz Web Page, Sander Group, EBI) 58 (c) M Gerstein, 2010, Yale, gersteinlab. org Major Application I|I: Overall Genome Characterization

59 (c) M Gerstein, 2010, Yale, gersteinlab. org End of class #1 [2010, 01. 11] Start of class #2 [2006, 01. 13]

• (Molecular) Bio - informatics • One idea for a definition? Bioinformatics is conceptualizing biology in terms of molecules (in the sense of physical-chemistry) and then applying “informatics” techniques (derived from disciplines such as applied math, CS, and statistics) to understand organize the information associated with these molecules, on a large-scale. • Bioinformatics is a practical discipline with many applications. 60 (c) M Gerstein, 2010, Yale, gersteinlab. org What is Bioinformatics?

61 (c) M Gerstein, 2010, Yale, gersteinlab. org Defining the Boundaries of the Field

Are They or Aren’t They Bioinformatics? (#1) à Automated Bibliographic Search of the biological literature and Textual Comparison à Knowledge bases for biological literature • Motif Discovery Using Gibb's Sampling • Methods for Structure Determination à Computational Crystallography • Refinement à NMR Structure Determination • Distance Geometry • Metabolic Pathway Simulation • The DNA Computer 62 (c) M Gerstein, 2010, Yale, gersteinlab. org • Digital Libraries

Are They or Aren’t They Bioinformatics? (#1, Answers) à Automated Bibliographic Search and Textual Comparison à Knowledge bases for biological literature • (YES) Motif Discovery Using Gibb's Sampling • (NO? ) Methods for Structure Determination à Computational Crystallography • Refinement à NMR Structure Determination • (YES) Distance Geometry • (YES) Metabolic Pathway Simulation • (NO) The DNA Computer 63 (c) M Gerstein, 2010, Yale, gersteinlab. org • (YES? ) Digital Libraries

Are They or Aren’t They Bioinformatics? (#2) • Gene identification by sequence inspection • DNA methods in forensics • Modeling of Populations of Organisms à Ecological Modeling • Genomic Sequencing Methods à Assembling Contigs à Physical and genetic mapping • Linkage Analysis à Linking specific genes to various traits 64 (c) M Gerstein, 2010, Yale, gersteinlab. org à Prediction of splice sites

Are They or Aren’t They Bioinformatics? (#2, Answers) • (YES) Gene identification by sequence inspection • (YES) DNA methods in forensics • (NO) Modeling of Populations of Organisms à Ecological Modeling • (NO? ) Genomic Sequencing Methods à Assembling Contigs à Physical and genetic mapping • (YES) Linkage Analysis à Linking specific genes to various traits 65 (c) M Gerstein, 2010, Yale, gersteinlab. org à Prediction of splice sites

• RNA structure prediction Identification in sequences • Radiological Image Processing à Computational Representations for Human Anatomy (visible human) • Artificial Life Simulations à Artificial Immunology / Computer Security à Genetic Algorithms in molecular biology • Homology modeling • Determination of Phylogenies Based on Nonmolecular Organism Characteristics • Computerized Diagnosis based on Genetic Analysis (Pedigrees) 66 (c) M Gerstein, 2010, Yale, gersteinlab. org Are They or Aren’t They Bioinformatics? (#3)

• (YES) RNA structure prediction Identification in sequences • (NO) Radiological Image Processing à Computational Representations for Human Anatomy (visible human) • (NO) Artificial Life Simulations à Artificial Immunology / Computer Security à (NO? ) Genetic Algorithms in molecular biology • (YES) Homology modeling • (NO) Determination of Phylogenies Based on Nonmolecular Organism Characteristics • (NO) Computerized Diagnosis based on Genetic Analysis (Pedigrees) 67 (c) M Gerstein, 2010, Yale, gersteinlab. org Are They or Aren’t They Bioinformatics? (#3, Answers)

Further Thoughts in 2005 on the "Boundary of Bioinformatics" à Does topic stand alone? à Is bioinformatics acting as tool? à How does it relate to lab work? à Prediction? • Relationship to other disciplines à Medical informatics à Genomics and Comp. Bioinformatics à Systems biology • Biological question is important, not the specific technique -- but it has to be computational à Using computers to understand biology vs using biology to inspire computation • Some new ones (2005) à Disease modeling [are you modeling molecules? ] à Enzymology (kinetics and rates? ) [is it a simulation or is it interpreting 1 expt. ? ] à Genetic algs used in gene finding HMMs used in gene finding • vs. Genetic algs used in speech recognition HMMs used in speech recognition à Semantic web used for representing biological information 68 (c) M Gerstein, 2010, Yale, gersteinlab. org • Issues that were uncovered

Some Further Boundary Examples in 2006 à What if it incluced non-molecular data such as age ? • Use of whole genome sequences to create phylogenies [YES] • Integration and organization of biological databases [YES] 69 (c) M Gerstein, 2010, Yale, gersteinlab. org • Char. drugs and other small molecules (cheminformatics or bioinformatics? ) [YES] • Molecular phenotype discovery – looking for gene expression signatures of cancer [YES]

Some Further Boundary Examples in 2010…. 70 (c) M Gerstein, 2010, Yale, gersteinlab. org • Processing of Next. Gen sequencing image files [? ? ? ]

71 (c) M Gerstein, 2010, Yale, gersteinlab. org class #2 continues in next ppt
- Slides: 71