Mark Gerstein Yale University Gerstein Lab orgcourses452 MG

Mark Gerstein Yale University Gerstein. Lab. org/courses/452 (MG lect. #3, last edit in spring '16) 1 (c) M Gerstein '14, Gerstein. Lab. org, Yale BIOINFORMATICS Multiple Sequences

Multiple Sequence Alignment Topics • Multiple Sequence Alignment • Motifs - Fast identification methods • Profile Patterns - Refinement via EM - Gibbs Sampling • HMMs • Applications Do not reproduce without permission 2 - Lectures. Gerstein. Lab. org (c) '09 - Protein Domain databases - Regression vs expression
It is widely used in: - Phylogenetic analysis - Prediction of protein secondary/tertiary structure - Finding diagnostic patterns to characterize protein families - Detecting new homologies between new genes and established sequence families Multiple Sequence Alignments - Practically useful methods only since 1987 - Before 1987 they were constructed by hand - The basic problem: no dynamic programming approach can be used - First useful approach by D. Sankoff (1987) based on phylogenetics (LEFT, adapted from Sonhammer et al. (1997). “Pfam, ” Proteins 28: 405 -20. ABOVE, G Barton AMAS web page) 3 (c) M Gerstein '14, Gerstein. Lab. org, Yale - One of the most essential tools in molecular biology

- Most multiple alignments based on this approach - Initial guess for a phylogenetic tree based on pairwise alignments - Built progressively starting with most closely related sequences - Follows branching order in phylogenetic tree - Sufficiently fast - Sensitive - Algorithmically heuristic, no mathematical property associated with the alignment - Biologically sound, it is common to derive alignments which are impossible to improve by eye (adapted from Sonhammer et al. (1997). “Pfam, ” Proteins 28: 405 -20) 4 (c) M Gerstein '14, Gerstein. Lab. org, Yale Progressive Multiple Alignments
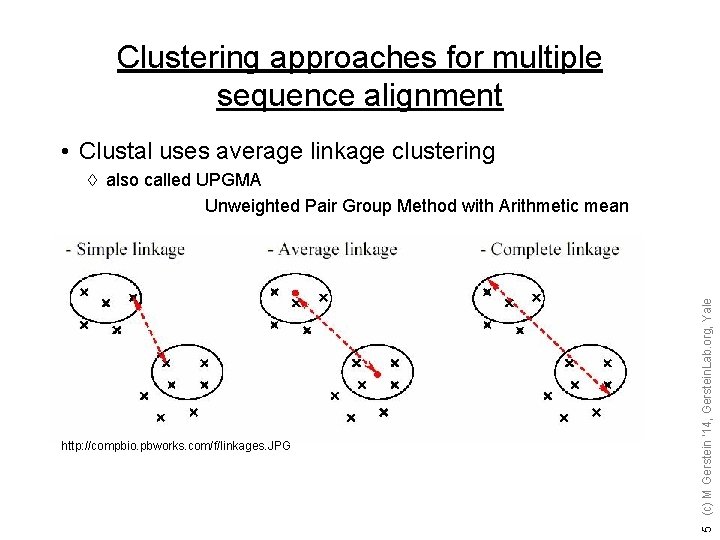
Clustering approaches for multiple sequence alignment • Clustal uses average linkage clustering http: //compbio. pbworks. com/f/linkages. JPG 5 (c) M Gerstein '14, Gerstein. Lab. org, Yale à also called UPGMA Unweighted Pair Group Method with Arithmetic mean

C 1 Q Example 6 (c) M Gerstein '14, Gerstein. Lab. org, Yale Ca 28_Human ELSAHATPAFTAVLTSPLPASGMPVKFDRTLYNGHSGYNPATGIFTCPVGGVYYFAYHVH VKGTNVWVALYKNNVPATYTYDEYKKGYLDQASGGAVLQLRPNDQVWVQIPSDQANGLYS TEYIHSSFSGFLLCPT C 1 qb_Human DYKATQKIAFSATRTINVPLRRDQTIRFDHVITNMNNNYEPRSGKFTCKVPGLYYFTYHA SSRGNLCVNLMRGRERAQKVVTFCDYAYNTFQVTTGGMVLKLEQGENVFLQATDKNSLLG MEGANSIFSGFLLFPD Cerb_Human VRSGSAKVAFSAIRSTNHEPSEMSNRTMIIYFDQVLVNIGNNFDSERSTFIAPRKGIYSF NFHVVKVYNRQTIQVSLMLNGWPVISAFAGDQDVTREAASNGVLIQMEKGDRAYLKLERG NLMGGWKYSTFSGFLVFPL COLE_LEPMA. 264 RGPKGPPGESVEQIRSAFSVGLFPSRSFPPPSLPVKFDKVFYNGEGHWDPTLNKFNVTYP GVYLFSYHITVRNRPVRAALVVNGVRKLRTRDSLYGQDIDQASNLALLHLTDGDQVWLET LRDWNGXYSSSEDDSTFSGFLLYPDTKKPTAM HP 27_TAMAS. 72 GPPGPPGMTVNCHSKGTSAFAVKANELPPAPSQPVIFKEALHDAQGHFDLATGVFTCPVP GLYQFGFHIEAVQRAVKVSLMRNGTQVMEREAEAQDGYEHISGTAILQLGMEDRVWLENK LSQTDLERGTVQAVFSGFLIHEN HSUPST 2_1. 95 GIQGRKGEPGEGAYVYRSAFSVGLETYVTIPNMPIRFTKIFYNQQNHYDGSTGKFHCNIP GLYYFAYHITVYMKDVKVSLFKKDKAMLFTYDQYQENNVDQASGSVLLHLEVGDQVWLQV YGEGERNGLYADNDNDSTFTGFLLYHDTN 2. HS 27109_1 ENALAPDFSKGSYRYAPMVAFFASHTYGMTIPGPILFNNLDVNYGASYTPRTGKFRIPYL GVYVFKYTIESFSAHISGFLVVDGIDKLAFESENINSEIHCDRVLTGDALLELNYGQEVW LRLAKGTIPAKFPPVTTFSGYLLYRT 4. YQCC_BACSU VVHGWTPWQKISGFAHANIGTTGVQYLKKIDHTKIAFNRVIKDSHNAFDTKNNRFIAPND GMYLIGASIYTLNYTSYINFHLKVYLNGKAYKTLHHVRGDFQEKDNGMNLGLNGNATVPM NKGDYVEIWCYCNYGGDETLKRAVDDKNGVFNFFD 5. BSPBSXSE_25 ADSGWTAWQKISGFAHANIGTTGRQALIKGENNKIKYNRIIKDSHKLFDTKNNRFVASHA GMHLVSASLYIENTERYSNFELYVYVNGTKYKLMNQFRMPTPSNNSDNEFNATVTGSVTV PLDAGDYVEIYVYVGYSGDVTRYVTDSNGALNYFD

SGMPLVSANHGVTG-------MPVSAFTVILS--KAYPA---VGCPHPIYEILYNRQQHY -----ALTG-------MPVSAFTVILS--KAYPG---ATVPIKFDKILYNRQQHY -----GGPA-------YEMPAFTAELT--APFPP---VGGPVKFNKLLYNGRQNY HAYAGKKGKHGGPA-------YEMPAFTAELT--VPFPP---VGAPVKFDKLLYNGRQNY -----ELSA-------HATPAFTAVLT--SPLPA---SGMPVKFDRTLYNGHSGY ----GTPGRKGEPGE---AAYMYRSAFSVGLETRVTVP-----NVPIRFTKIFYNQQNHY ------RGPKGPPGE---SVEQIRSAFSVGLFPSRSFPP---PSLPVKFDKVFYNGEGHW -------GPPGPPGMTVNCHSKGTSAFAVKAN--ELPPA---PSQPVIFKEALHDAQGHF -----NIRD-------QPRPAFSAIRQ---NPMT---LGNVVIFDKVLTNQESPY -------D---YRATQKVAFSALRTINSPLR----PNQVIRFEKVITNANENY -------D---YKATQKIAFSATRTINVPLR----RDQTIRFDHVITNMNNNY -------V---RSGSAKVAFSAIRSTNHEPSEMSNRTMIIYFDQVLVNIGNNF ---ENALAPDFSKGS---YRYAPMVAFFASHTYGMTIP------GPILFNNLDVNYGASY. *. : : MMCOL 10 A 1_1. 483 Ca 1 x_Chick S 15435 CA 18_MOUSE. 597 Ca 28_Human MM 37222_1. 98 COLE_LEPMA. 264 HP 27_TAMAS. 72 S 19018 C 1 qb_Mouse C 1 qb_Human Cerb_Human 2. HS 27109_1 DPRSGIFTCKIPGIYYFSYHVHVKGT--HVWVGLYKNGTP-TMYTY---DEYSKGYLDTA DPRTGIFTCRIPGLYYFSYHVHAKGT--NVWVALYKNGSP-VMYTY---DEYQKGYLDQA NPQTGIFTCEVPGVYYFAYHVHCKGG--NVWVALFKNNEP-VMYTY---DEYKKGFLDQA NPQTGIFTCEVPGVYYFAYHVHCKGG--NVWVALFKNNEP-MMYTY---DEYKKGFLDQA NPATGIFTCPVGGVYYFAYHVHVKGT--NVWVALYKNNVP-ATYTY---DEYKKGYLDQA DGSTGKFYCNIPGLYYFSYHITVYMK--DVKVSLFKKDKA-VLFTY---DQYQEKNVDQA DPTLNKFNVTYPGVYLFSYHITVRNR--PVRAALVVNGVR-KLRTR---DSLYGQDIDQA DLATGVFTCPVPGLYQFGFHIEAVQR--AVKVSLMRNGTQ-VMERE---AEAQDG-YEHI QNHTGRFICAVPGFYYFNFQVISKWD--LCLFIKSSSGGQ-PRDSLSFSNTNNKGLFQVL EPRNGKFTCKVPGLYYFTYHASSRGN---LCVNLVRGRDRDSMQKVVTFCDYAQNTFQVT EPRSGKFTCKVPGLYYFTYHASSRGN---LCVNLMRGRER--AQKVVTFCDYAYNTFQVT DSERSTFIAPRKGIYSFNFHVVKVYNRQTIQVSLMLNGWP----VISAFAGDQDVTREAA TPRTGKFRIPYLGVYVFKYTIESFSA--HISGFLVVDGIDKLAFESEN-INSEIHCDRVL. * * : MMCOL 10 A 1_1. 483 Ca 1 x_Chick S 15435 CA 18_MOUSE. 597 Ca 28_Human MM 37222_1. 98 COLE_LEPMA. 264 HP 27_TAMAS. 72 S 19018 C 1 qb_Mouse C 1 qb_Human Cerb_Human 2. HS 27109_1 SGSAIMELTENDQVWLQLPNA-ESNGLYSSEYVHSSFSGFLVAPM------SGSAVIDLMENDQVWLQLPNS-ESNGLYSSEYVHSSFSGFLFAQI------SGSAVLLLRPGDRVFLQMPSE-QAAGLYAGQYVHSSFSGYLLYPM------SGSAVLLLRPGDQVFLQNPFE-QAAGLYAGQYVHSSFSGYLLYPM------SGGAVLQLRPNDQVWVQIPSD-QANGLYSTEYIHSSFSGFLLCPT------SGSVLLHLEVGDQVWLQVYGDGDHNGLYADNVNDSTFTGFLLYHDTN----SNLALLHLTDGDQVWLETLR--DWNGXYSSSEDDSTFSGFLLYPDTKKPTAM SGTAILQLGMEDRVWLENKL--SQTDLERG-TVQAVFSGFLIHEN------AGGTVLQLRRGDEVWIEKDP--AKGRIYQGTEADSIFSGFLIFPS------TGGVVLKLEQEEVVHLQATD---KNSLLGIEGANSIFTGFLLFPD------TGGMVLKLEQGENVFLQATD---KNSLLGMEGANSIFSGFLLFPD------SNGVLIQMEKGDRAYLKLER---GN-LMGG-WKYSTFSGFLVFPL------TGDALLELNYGQEVWLRLAK----GTIPAKFPPVTTFSGYLLYRT------. : : . : * *: *. Clustal Alignment 7 (c) M Gerstein '14, Gerstein. Lab. org, Yale MMCOL 10 A 1_1. 483 Ca 1 x_Chick S 15435 CA 18_MOUSE. 597 Ca 28_Human MM 37222_1. 98 COLE_LEPMA. 264 HP 27_TAMAS. 72 S 19018 C 1 qb_Mouse C 1 qb_Human Cerb_Human 2. HS 27109_1

Problems with Progressive Alignments - Local Minimum Problem - Parameter Choice Problem - It stems from greedy nature of alignment (mistakes made early in alignment cannot be corrected later) - A better tree gives a better alignment (UPGMA neighbour-joining tree method) 2. Parameter Choice Problem • - It stems from using just one set of parameters (and hoping that they will do for all) 8 (c) M Gerstein '14, Gerstein. Lab. org, Yale 1. Local Minimum Problem

9 (c) M Gerstein '14, Gerstein. Lab. org, Yale Domain Problem in Multiple Alignment

Fuse multiple alignment into: - Motif: a short signature pattern identified in the conserved region of the multiple alignment - Profile: frequency of each amino acid at each position is estimated Profiles Motifs HMMs Can get more sensitive searches with these multiple alignment representations (Run the profile against the DB. ) 10 (c) M Gerstein '14, Gerstein. Lab. org, Yale - HMM: Hidden Markov Model, a generalized profile in rigorous mathematical terms

Multiple Alignment Do not reproduce without permission 11 - Lectures. Gerstein. Lab. org (c) '09 MOTIFS

• Given a collection of binding sites (or protein sequences with binding motifs), develop a representation of those sites that can be used to search new sites and reliably predict where additional binding sites occur. • Given a set of sequences known to contain binding sites for a common factor, but not knowing where the sites are, discover the location of the sites in each sequence and a representation of the protein. [Adapted from C Bruce, CBB 752 '09] 12 (c) M Gerstein '14, Gerstein. Lab. org, Yale 2 different applications for motif analysis

13 (c) M Gerstein '14, Gerstein. Lab. org, Yale Motifs - several proteins are grouped together by similarity searches - they share a conserved motif - motif is stringent enough to retrieve the family members from the complete protein database - PROSITE: a collection of motifs (1135 different motifs)

Prosite Pattern -- EGF like pattern A sequence of about thirty to forty amino-acid residues long found in the sequence of epidermal growth factor (EGF) has been shown [1 to 6] to be present, in a more or less conserved form, in a large number of other, mostly animal proteins. The proteins currently known to contain one or more copies of an EGF-like pattern are listed below. - Bone morphogenic protein 1 (BMP-1), a protein which induces cartilage and bone formation. Caenorhabditis elegans developmental proteins lin-12 (13 copies) and glp-1 (10 copies). Calcium-dependent serine proteinase (CASP) which degrades the extracellular matrix proteins type …. Cell surface antigen 114/A 10 (3 copies). Cell surface glycoprotein complex transmembrane subunit. Coagulation associated proteins C, Z (2 copies) and S (4 copies). Coagulation factors VII, IX, X and XII (2 copies). Complement C 1 r/C 1 s components (1 copy). Complement-activating component of Ra-reactive factor (RARF) (1 copy). Complement components C 6, C 7, C 8 alpha and beta chains, and C 9 (1 copy). Epidermal growth factor precursor (7 -9 copies). 'C': conserved cysteine involved in a disulfide bond. 'G': often conserved glycine 'a': often conserved aromatic amino acid '*': position of both patterns. 'x': any residue -Consensus pattern: C-x-C-x(5)-G-x(2)-C [The 3 C's are involved in disulfide bonds] http: //www. expasy. ch/sprot/prosite. html 14 (c) M Gerstein '14, Gerstein. Lab. org, Yale +----------+ +-------------+ | | x(4)-C-x(0, 48)-C-x(3, 12)-C-x(1, 70)-C-x(1, 6)-C-x(2)-G-a-x(0, 21)-G-x(2)-C-x | | ****************** +----------+

Multiple Alignment Do not reproduce without permission 15 - Lectures. Gerstein. Lab. org (c) '09 PROFILES

Profile : a position-specific scoring matrix composed of 21 columns and N rows (N=length of sequences in multiple alignment) What happens with gaps? 5 16 (c) M Gerstein '14, Gerstein. Lab. org, Yale Profiles
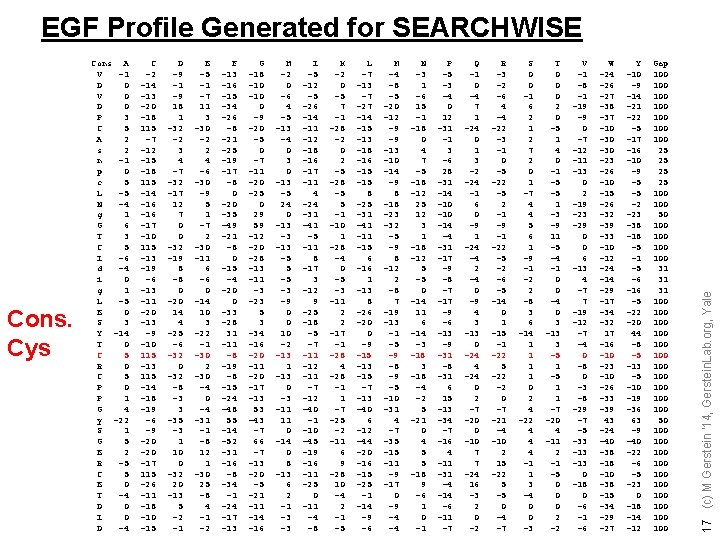
Cons. Cys Cons A V -1 D 0 V 0 D 0 P 3 C 5 A 2 s 2 n -1 p 0 c 5 L -5 N -4 g 1 G 6 T 3 C 5 I -6 d -4 i 0 g 1 L -5 E 0 S 3 Y -14 T 0 C 5 R 0 C 5 P 0 P 1 G 4 y -22 S 1 G 5 E 2 R -5 C 5 E 0 T -4 D 0 I 0 D -4 C -2 -14 -13 -20 -18 115 -7 -12 -15 -18 115 -14 -16 -17 -10 115 -13 -19 -6 -13 -11 -20 -13 -9 -10 115 -13 115 -14 -18 -19 -6 -9 -20 -17 115 -26 -11 -18 -10 -15 D -9 -1 -9 18 1 -32 -2 3 4 -7 -32 -17 12 7 0 0 -32 -19 8 -8 0 -20 14 4 -25 -6 -32 0 -32 -8 -3 3 -35 -3 1 10 0 -32 20 -13 5 -2 -1 E -5 -1 -7 11 3 -30 -2 2 4 -6 -30 -9 5 1 -7 2 -30 -11 6 -6 0 -14 10 3 -22 -1 -30 2 -30 -4 -31 -1 -8 12 1 -30 25 -8 4 -1 -2 F -13 -16 -15 -34 -26 -8 -21 -25 -19 -17 -8 0 -20 -35 -49 -21 -8 0 -15 -4 -20 0 -33 -28 31 -11 -8 -19 -8 -15 -24 -48 55 -14 -52 -31 -16 -8 -34 -1 -24 -17 -13 G -18 -10 0 -9 -20 -5 0 -7 -11 -20 -25 0 29 59 -12 -20 -28 -13 -11 -3 -23 5 3 -34 -16 -20 -11 -20 -17 -13 53 -43 -7 66 -7 -13 -20 -5 -21 -14 -16 H -2 0 -6 4 -5 -13 -4 0 3 0 -13 -5 24 0 -13 -3 -13 -5 5 -5 -3 -9 0 0 10 -2 -13 1 -13 0 -3 -11 11 0 -14 0 8 -13 6 2 -1 -3 -3 I -5 -12 -5 -26 -14 -11 -12 -18 -16 -17 -11 4 -24 -31 -41 -5 -11 8 -17 3 -12 9 -25 -18 -5 -7 -11 -12 -11 -7 -12 -40 -1 -10 -45 -19 -16 -11 -25 0 -11 -4 -8 K -2 0 -5 7 -1 -28 -2 0 2 -5 -28 -5 5 -1 -10 1 -28 -4 0 -5 -3 -11 2 2 -17 -1 -28 4 -28 -1 1 -7 -25 -2 -11 6 9 -28 10 -4 2 -1 -5 L -7 -13 -7 -27 -14 -15 -13 -18 -16 -15 8 -25 -31 -41 -15 6 -16 1 -13 8 -26 -20 0 -9 -15 -13 -15 -7 -13 -40 6 -12 -44 -20 -16 -15 -25 -1 -14 -9 -6 M -4 -8 -5 -20 -12 -9 -9 -13 -10 -14 -9 8 -18 -23 -32 -5 -9 8 -12 2 -8 7 -19 -13 -1 -5 -9 -8 -9 -5 -10 -31 4 -7 -35 -11 -9 -17 0 -9 -4 -4 N -3 1 -6 15 -1 -18 0 4 7 -5 -18 -12 25 12 3 1 -18 -12 5 -5 0 -14 11 6 -14 -3 -18 -4 -2 5 -21 0 4 5 5 -18 9 -6 1 0 -1 P -5 -3 -4 0 12 -31 -1 3 -6 28 -31 -14 -10 -14 -4 -31 -17 -9 -8 -7 -17 -9 -6 -13 -9 -31 -8 -31 6 15 -13 -34 -7 -16 4 -11 -31 -4 -14 -6 -11 -7 Q -1 0 -4 7 1 -24 0 1 3 -2 -24 -1 6 0 -9 1 -24 -4 2 -4 0 -9 4 3 -13 0 -24 4 -24 0 2 -7 -20 0 -10 7 7 -24 16 -3 2 0 -2 R -3 -2 -6 4 -4 -22 -3 -1 0 -5 -22 -5 2 -1 -9 -1 -22 -5 -2 -6 -5 -14 0 1 -15 -1 -22 5 -22 -2 0 -7 -21 -4 -10 2 15 -22 5 -5 0 -4 -7 S 0 0 -1 6 2 1 2 7 2 0 1 -7 4 4 5 6 1 -9 -1 -2 2 -8 3 6 -14 1 1 0 2 4 -22 4 4 4 -1 1 3 -4 0 0 -3 T 0 0 0 2 0 -5 1 4 0 -1 -5 -5 1 -3 -9 11 -5 -4 -1 0 0 -4 0 3 -13 3 -5 1 1 -7 -20 4 -11 2 -1 -5 0 0 0 2 -2 V -1 -8 -1 -19 -9 0 -7 -12 -11 -13 0 2 -19 -23 -29 0 0 6 -13 4 -7 7 -19 -12 -7 -4 0 -8 0 -3 -8 -29 -7 -5 -33 -13 0 -18 0 -6 -1 -6 W -24 -26 -27 -38 -37 -10 -30 -23 -26 -10 -15 -26 -32 -39 -33 -10 -12 -24 -14 -29 -17 -34 -32 17 -16 -10 -23 -10 -26 -33 -39 43 -24 -40 -38 -10 -38 -15 -34 -29 -27 Y -10 -9 -14 -21 -22 -5 -17 -16 -10 -9 -5 -5 -2 -23 -38 -18 -5 -1 -5 -6 -16 -5 -22 -20 44 -8 -5 -13 -5 -10 -19 -36 63 -9 -40 -22 -6 -5 -23 0 -18 -14 -12 Gap 100 100 25 25 100 50 100 100 31 31 31 100 100 100 50 100 100 100 17 (c) M Gerstein '14, Gerstein. Lab. org, Yale EGF Profile Generated for SEARCHWISE

M(p, a) = chance of finding amino acid a at position p Msimp(p, a) = number of times a occurs at p divided by number of sequences However, what if don’t have many sequences in alignment? Msimp(p, a) might be baised. Zeros for rare amino acids. Thus: Mcplx(p, a)= b=1 to 20 Msimp(p, b) x Y(b, a): Dayhoff matrix for a and b amino acids S(p, a) ~ a=1 to 20 Msimp(p, a) ln Msimp(p, a) 18 (c) M Gerstein '14, Gerstein. Lab. org, Yale Profiles formula for position M(p, a)

H(p, a) = - a=1 to 20 f(p, a) log 2 f(p, a), where f(p, a) = frequency of amino acid a occurs at position p ( Msimp(p, a) ) Say column only has one aa (AAAAA): H(p, a) = 1 log 2 1 + 0 log 2 0 + … = 0 + 0 + … = 0 Say column is random with all aa equiprobable (ACD. . ): Hrand(p, a) =. 05 log 2. 05 + … = -. 22 + … = -4. 3 Say column is random with aa occurring according to probability found in the sequence databases (ACAAAADAADDDDAAAA…. ): Hdb(a) = - a=1 to 20 F(a) log 2 F(a), where F(a) is freq. of occurrence of a in DB Hcorrected(p, a) = H(p, a) – Hdb(a) 19 (c) M Gerstein '14, Gerstein. Lab. org, Yale Profiles formula for entropy H(p, a)
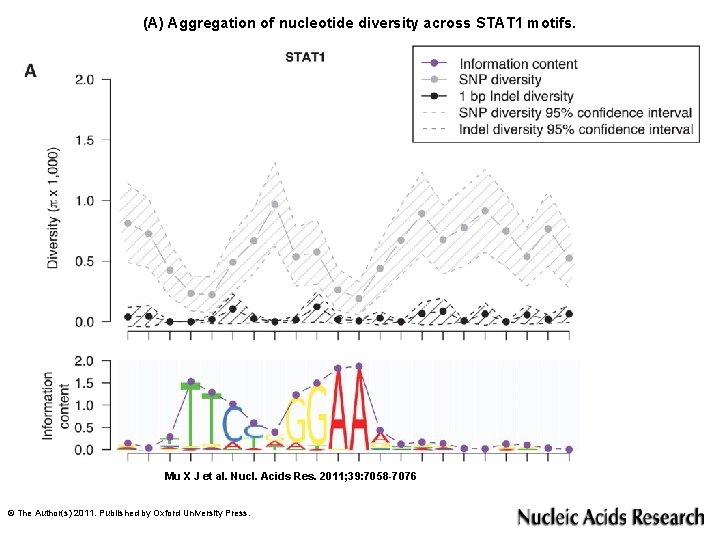
(A) Aggregation of nucleotide diversity across STAT 1 motifs. Mu X J et al. Nucl. Acids Res. 2011; 39: 7058 -7076 © The Author(s) 2011. Published by Oxford University Press.
![[Adapted from C Bruce, CBB 752 '09] Mac. Isaac & Fraenkel, 2006 21 (c) [Adapted from C Bruce, CBB 752 '09] Mac. Isaac & Fraenkel, 2006 21 (c)](http://slidetodoc.com/presentation_image_h2/038750f27aaa6090fa623720b1b4a17f/image-21.jpg)
[Adapted from C Bruce, CBB 752 '09] Mac. Isaac & Fraenkel, 2006 21 (c) M Gerstein '14, Gerstein. Lab. org, Yale Scanning for Motifs with PWMs

22 (c) M Gerstein '14, Gerstein. Lab. org, Yale Y-Blast Parameters: overall • Automatically builds profile and then searches with this threshold, inclusion • Also PHI-blast threshold, interations
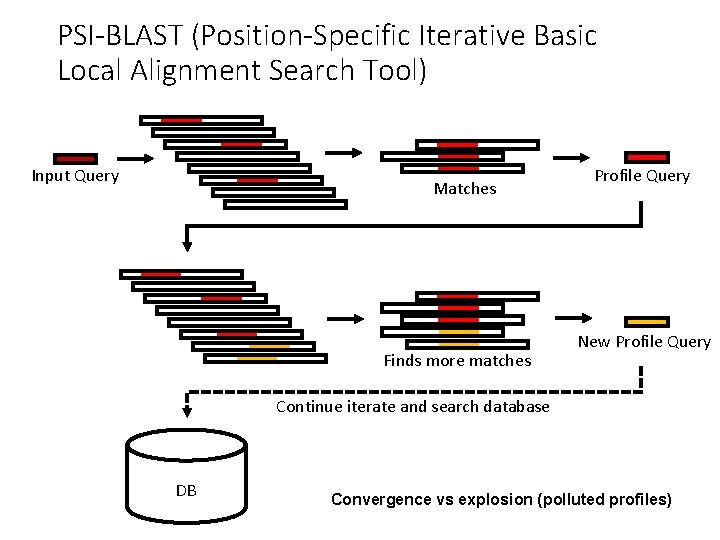
PSI-BLAST (Position-Specific Iterative Basic Local Alignment Search Tool) Input Query Matches Finds more matches Profile Query New Profile Query Continue iterate and search database DB Convergence vs explosion (polluted profiles)

Speed v Sensitivity Tradeoff Blast FASTA Smith. Waterman PSI-Blast Profiles HMMs 24 (c) M Gerstein '14, Gerstein. Lab. org, Yale Speed Sensitivity PSI Blast as a form of Semi-supervised learning

Multiple Alignment: Probabilistic Approaches for Determining PWMs • Expectation Maximization: Search the PWM space randomly • Gibbs sampling: Search sequence space randomly. [Adapted from C Bruce, CBB 752 '09]

Expectation-Maximization (EM) algorithm • • Used in statistics for finding maximum likelihood estimates of parameters in probabilistic models, where the model depends on unobserved latent variables. EM alternates between performing – an expectation (E) step, which computes an expectation of the likelihood by including the latent variables as if they were observed, and – a maximization (M) step, which computes the maximum likelihood estimates of the parameters by maximizing the expected likelihood found on the E step. • The parameters found on the M step are then used to begin another E step, and the process is repeated. [Adapted from C Bruce, CBB 752 '09]

Alternating approach 1. 2. 3. 4. Guess an initial weight matrix Use weight matrix to predict instances in the input sequences Use instances to predict a weight matrix Repeat 2 [E-step] & 3 [M-step] until satisfied. Examples: Gibbs sampler (Lawrence et al. ) MEME (expectation max. / Bailey, Elkan) ANN-Spec (neural net / Workman, Stormo) Another good source is Wes Craven’s 776 course: https: //www. biostat. wisc. edu/~craven/776/lecture 9. pdf [Adapted from B Noble, GS 541 at UW, http: //noble. gs. washington. edu/~wnoble/genome 541/]

EM (again!) foreach subsequence of width W convert subsequence to a matrix do { re-estimate motif occurrences from matrix EM re-estimate matrix model from motif occurrences } until (matrix model stops changing) end select matrix with highest score [Adapted from B Noble, GS 541 at UW, http: //noble. gs. washington. edu/~wnoble/genome 541]

Sample DNA sequences >ce 1 cg TAATGTTTGTGCTGGTTTTTGTGGCATCGGGCGAGAATA GCGCGTGGTGTGAAAGACTGTTTTTTTGATCGTTTTCAC AAAAATGGAAGTCCACAGTCTTGACAG >ara GACAAAAACGCGTAACAAAAGTGTCTATAATCACGGCAG AAAAGTCCACATTGATTATTTGCACGGCGTCACACTTTG CTATGCCATAGCATTTTTATCCATAAG >bglr 1 ACAAATCCCAATAACTTAATTATTGGGATTTGTTATATA TAACTTTATAAATTCCTAAAATTACACAAAGTTAATAAC TGTGAGCATGGTCATATTTTTATCAAT >crp CACAAAGCGAAAGCTATGCTAAAACAGTCAGGATGCTAC AGTAATACATTGATGTACTGCATGTATGCAAAGGACGTC ACATTACCGTGCAGTACAGTTGATAGC [Adapted from B Noble, GS 541 at UW, http: //noble. gs. washington. edu/~wnoble/genome 541/]

Motif occurrences >ce 1 cg taatgtttgtgctggtttttgtggcatcgggcgagaata gcgcgtggtgtgaaagactgtttt. TTTGATCGTTTTCAC aaaaatggaagtccacagtcttgacag >ara gacaaaaacgcgtaacaaaagtgtctataatcacggcag aaaagtccacattgatta. TTTGCACGGCGTCACactttg ctatgccatagcatttttatccataag >bglr 1 acaaatcccaataacttaattattgggatttgttatata taactttataaattcctaaaattacacaaagttaataac TGTGAGCATGGTCATatttttatcaat >crp cacaaagcgaaagctatgctaaaacagtcaggatgctac agtaatacattgatgtactgcatgta. TGCAAAGGACGTC ACattaccgtgcagtacagttgatagc [Adapted from B Noble, GS 541 at UW, http: //noble. gs. washington. edu/~wnoble/genome 541/]
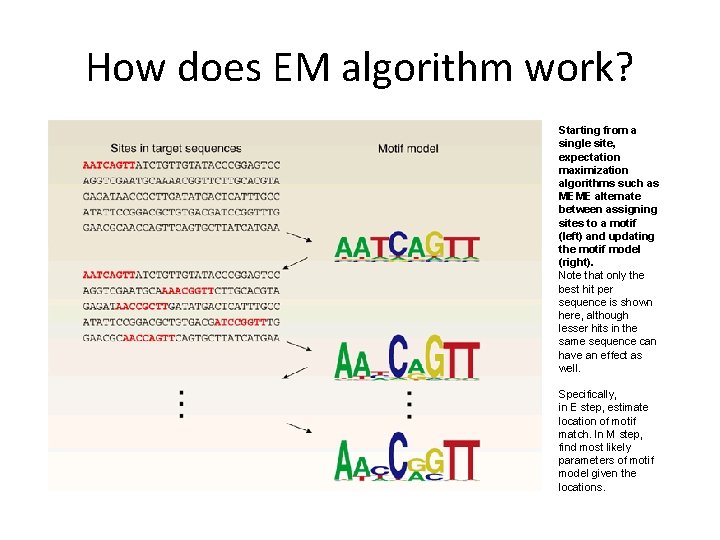
How does EM algorithm work? Starting from a single site, expectation maximization algorithms such as MEME alternate between assigning sites to a motif (left) and updating the motif model (right). Note that only the best hit per sequence is shown here, although lesser hits in the same sequence can have an effect as well. Specifically, in E step, estimate location of motif match. In M step, find most likely parameters of motif model given the locations.

MEME - a practical program using EM • Subsequences which occur in the input DNA sequence are used as the starting points from which EM converges iteratively to locally optimal motifs. This increases the likelihood of finding globally optimal motifs. • Multiple occurrences of a motif are allowed. Algorithm is allowed to ignore sequences with no appearance of a shared motif. So, more resistance to noisy data. • Motifs are probabilistically erased after they are found, so more than one motif can be found. [Adapted from C Bruce, CBB 752 '09]

Multiple Alignment Do not reproduce without permission 33 - Lectures. Gerstein. Lab. org (c) '09 Gibbs Sampling

Initialization • Step 1: Randomly guess an instance si from each of t input sequences {S 1, . . . , St}. sequence 1 sequence 2 sequence 3 sequence 4 ACAGTGT TTAGACC GTGACCA ACCCAGGTTT sequence 5 [Adapted from B Noble, GS 541 at UW, http: //noble. gs. washington. edu/~wnoble/genome 541/]

Gibbs sampler • Steps 2 & 3 (search): – Throw away an instance si: remaining (t - 1) instances define weight matrix. – Weight matrix defines instance probability at each position of input string Si – Pick new si according to probability distribution (not necessarily always the si giving the highest prob. ) • Return highest-scoring motif seen [Adapted from B Noble, GS 541 at UW, http: //noble. gs. washington. edu/~wnoble/genome 541/]
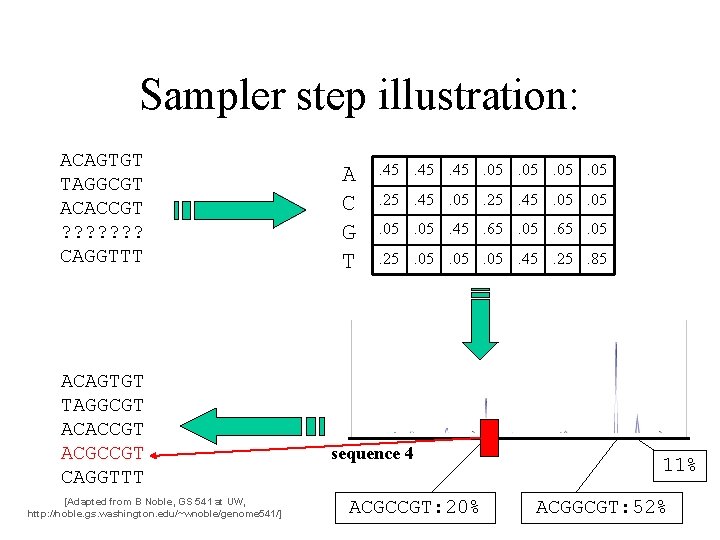
Sampler step illustration: ACAGTGT TAGGCGT ACACCGT ? ? ? ? CAGGTTT ACAGTGT TAGGCGT ACACCGT ACGCCGT CAGGTTT [Adapted from B Noble, GS 541 at UW, http: //noble. gs. washington. edu/~wnoble/genome 541/] A C G T . 45. 45. 05. 05. 25. 45. 05. 05. 05. 45. 65. 05. 25. 05. 05. 45. 25. 85 sequence 4 ACGCCGT: 20% 11% ACGGCGT: 52%

Comparison • Both EM and Gibbs sampling involve iterating over two steps • Convergence: – EM converges when the PSSM stops changing. – Gibbs sampling runs until you ask it to stop. • Solution: – EM may not find the motif with the highest score. – Gibbs sampling will provably find the motif with the highest score, if you let it run long enough. [Adapted from B Noble, GS 541 at UW, http: //noble. gs. washington. edu/~wnoble/genome 541/]

Multiple Alignment Do not reproduce without permission 38 - Lectures. Gerstein. Lab. org (c) '09 HMMs
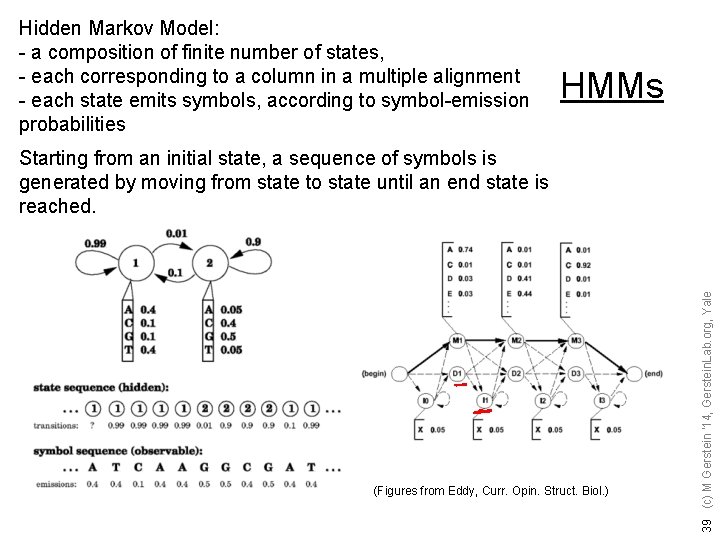
Hidden Markov Model: - a composition of finite number of states, - each corresponding to a column in a multiple alignment - each state emits symbols, according to symbol-emission probabilities HMMs (Figures from Eddy, Curr. Opin. Struct. Biol. ) 39 (c) M Gerstein '14, Gerstein. Lab. org, Yale Starting from an initial state, a sequence of symbols is generated by moving from state to state until an end state is reached.
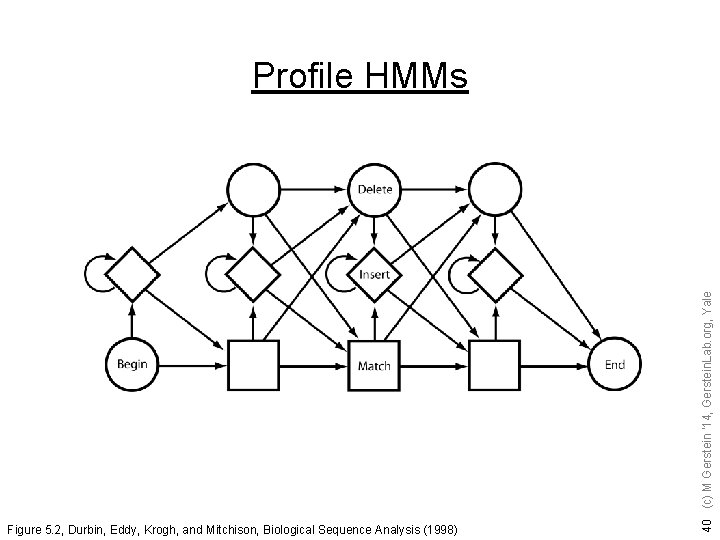
Figure 5. 2, Durbin, Eddy, Krogh, and Mitchison, Biological Sequence Analysis (1998) 40 (c) M Gerstein '14, Gerstein. Lab. org, Yale Profile HMMs
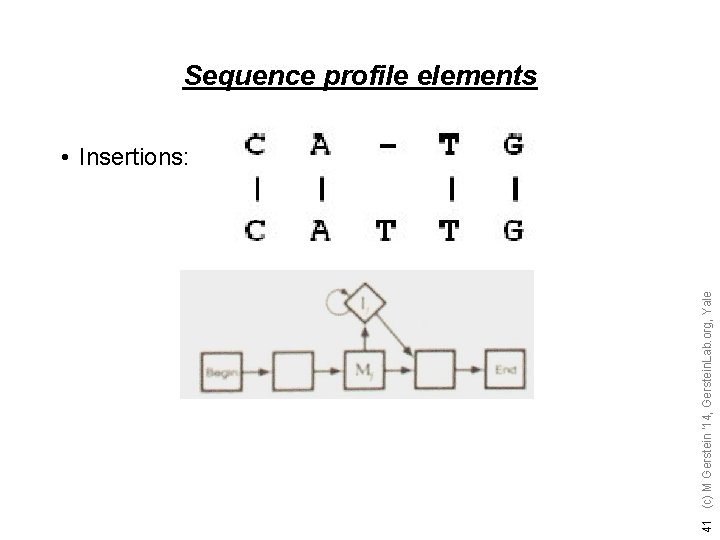
41 (c) M Gerstein '14, Gerstein. Lab. org, Yale Sequence profile elements • Insertions:

Algorithms Probability of a path through the model Viterbi maximizes for seq Forward Algorithm – finds probability P that a model l emits a given sequence O by summing over all paths that emit the sequence the probability of that path Viterbi Algorithm – finds the most probable path through the model for a given sequence (both usually just boil down to simple applications of dynamic programming) 42 (c) M Gerstein '14, Gerstein. Lab. org, Yale Forward sums of all possible paths
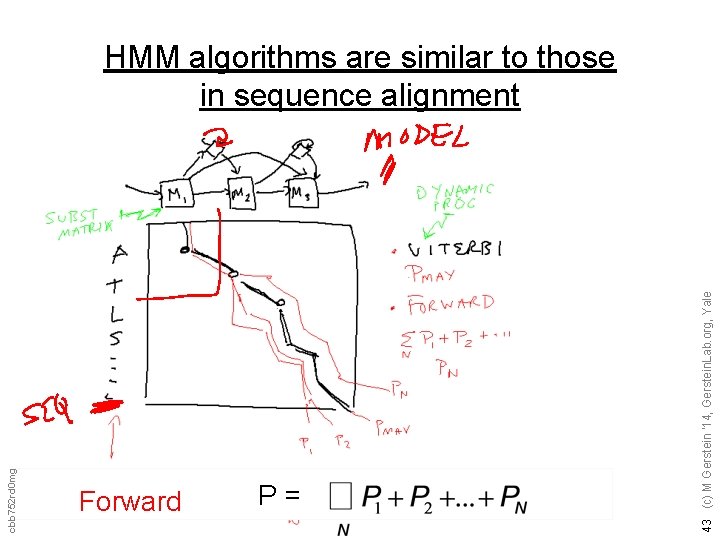
Forward P= 43 (c) M Gerstein '14, Gerstein. Lab. org, Yale cbb 752 rd 0 mg HMM algorithms are similar to those in sequence alignment
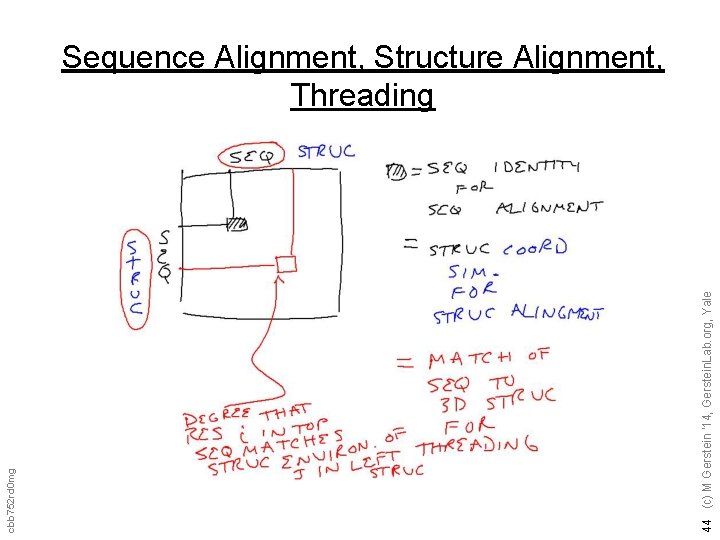
44 (c) M Gerstein '14, Gerstein. Lab. org, Yale cbb 752 rd 0 mg Sequence Alignment, Structure Alignment, Threading
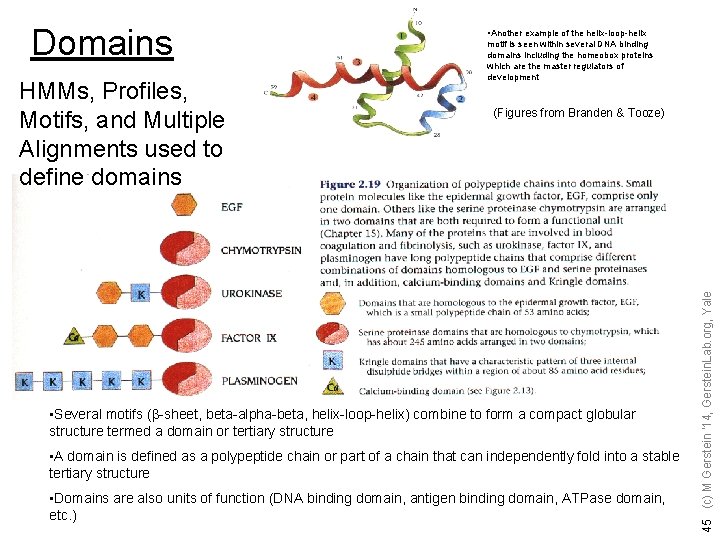
HMMs, Profiles, Motifs, and Multiple Alignments used to define domains motif is seen within several DNA binding domains including the homeobox proteins which are the master regulators of development (Figures from Branden & Tooze) • Several motifs (b-sheet, beta-alpha-beta, helix-loop-helix) combine to form a compact globular structure termed a domain or tertiary structure • A domain is defined as a polypeptide chain or part of a chain that can independently fold into a stable tertiary structure • Domains are also units of function (DNA binding domain, antigen binding domain, ATPase domain, etc. ) 45 (c) M Gerstein '14, Gerstein. Lab. org, Yale Domains • Another example of the helix-loop-helix

Applications of HMMs: Protein Domain Databases Pfam http: //pfam. sanger. ac. uk/ SMART http: //smart. embl-heidelberg. de/ CDD Interpro 46 (c) M Gerstein '14, Gerstein. Lab. org, Yale etc.
- Slides: 46