Chemical nature biosynthesis degradation of purines Rosalind Elsie

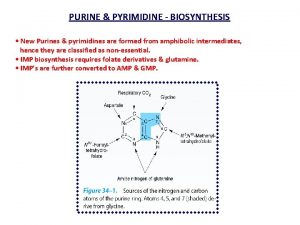
Chemical nature, biosynthesis & degradation of purines

Rosalind Elsie Franklin was an English chemist and X-ray crystallographer who made critical contributions to the understanding of the fine molecular structures of DNA, RNA Maurice Wilkins studied biological molecules like DNA and viruses using a variety of microscopes and spectrophotometers. He eventually began using X-rays to produce diffraction images of DNA molecules. James Dewey Watson, is an American molecular biologist, geneticist and zoologist, best known as a co-discoverer of the structure of DNA in 1953 with Francis Crick Wilkins shared the 1962 Nobel Prize in Physiology or Medicine with Watson and Crick.
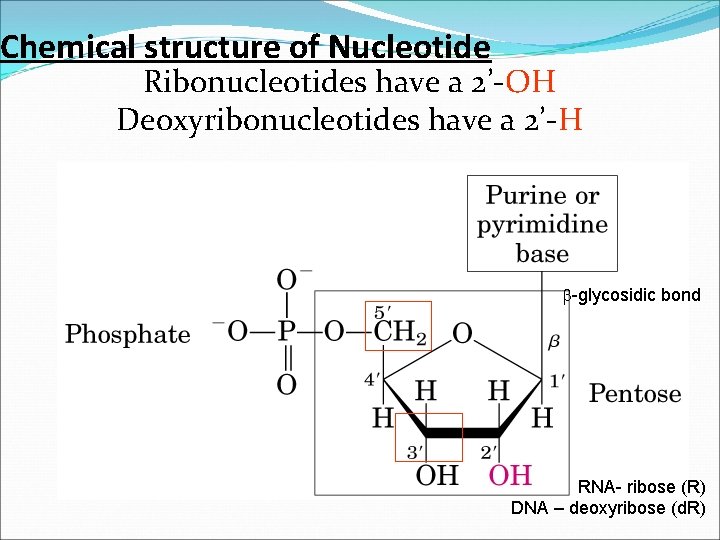
Chemical structure of Nucleotide Ribonucleotides have a 2’-OH Deoxyribonucleotides have a 2’-H b-glycosidic bond RNA- ribose (R) DNA – deoxyribose (d. R)
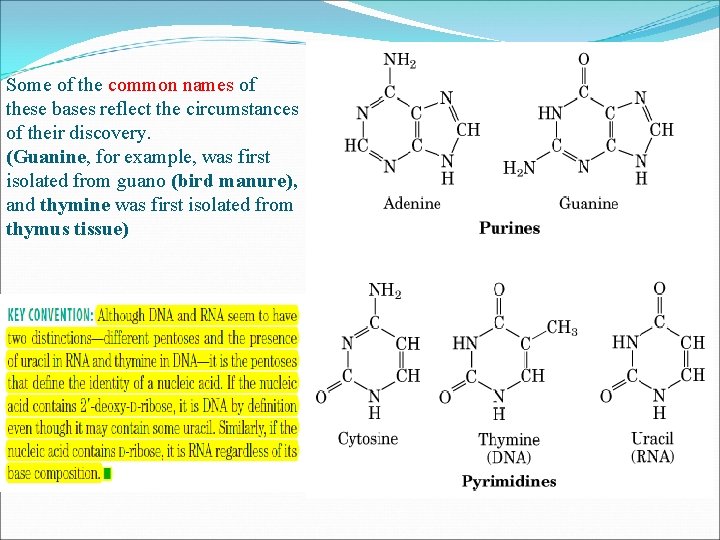
Some of the common names of these bases reflect the circumstances of their discovery. (Guanine, for example, was first isolated from guano (bird manure), and thymine was first isolated from thymus tissue)
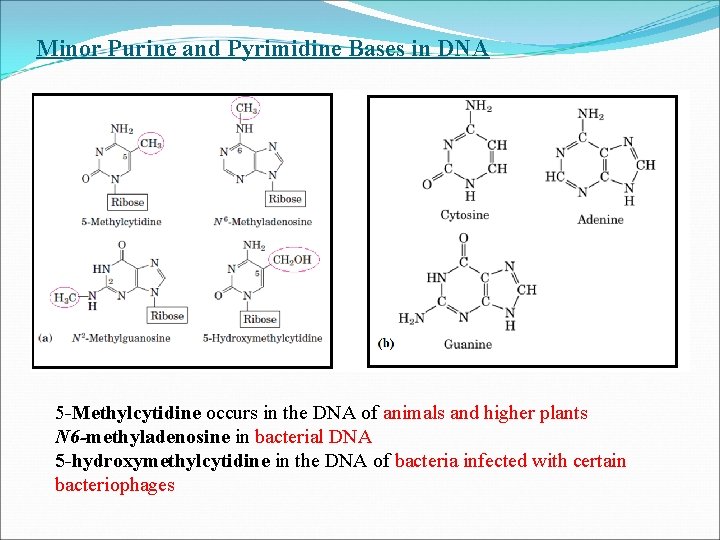
Minor Purine and Pyrimidine Bases in DNA 5 -Methylcytidine occurs in the DNA of animals and higher plants N 6 -methyladenosine in bacterial DNA 5 -hydroxymethylcytidine in the DNA of bacteria infected with certain bacteriophages
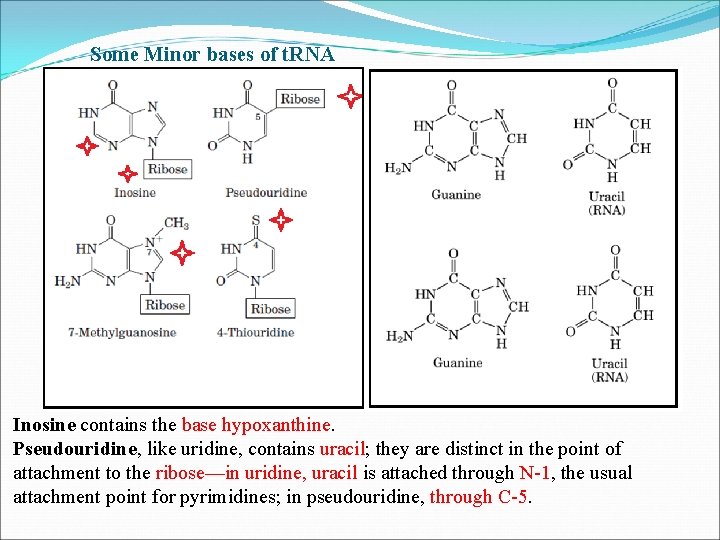
Some Minor bases of t. RNA Inosine contains the base hypoxanthine. Pseudouridine, like uridine, contains uracil; they are distinct in the point of attachment to the ribose—in uridine, uracil is attached through N-1, the usual attachment point for pyrimidines; in pseudouridine, through C-5.

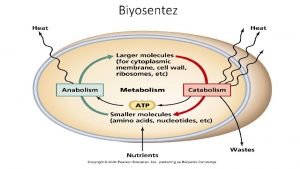
Nucleotides actively participate in many biochemical reactions ATP and GTP are energy sources Uridine derivatives of sugars participate in carbohydrate metabolism Coenzymes (NAD, FAD, Co-A) are nucleotide derivatives [ATP], [ADP], [AMP] act as allosteric (Activation of protein from other than active site) regulators of key enzymes

Biochemical Importance o Diet Nucleoprotein, Purine and Pyrimidine bases do not directly incorporate in Nucleic acids synthesis o Human body synthesize the nitrogenous bases for nucleic acid, ATP, NAD, Co-A from Amphibolic intermediate o Nucleoprotein digested in intestinal tract o Nucleic acids is degraded with help of Ribonucleases, Deoxyribonucleases, Polynucleotidases to nucleotides o Mononucleotides hydrolysed nucleoside by Nucleotidases, Phosphatases o Furthermore, it produce uric acid by help of phsophorylase enzymes

Biosynthesis of Purine Nucleotide o All form of life synthesize the Purine even Pyrimidine nucleotide except parasitic protozoa o Nucleotides can synthesize by o De Novo pathway: Nucleotides are constructed from simple precursors from metabolic process o Salvage pathways: Recovery and recycling of nucleotides obtained in the diet

De Novo versus salvage pathways De novo Pathway Activated PRPP and aminoacid + ATP + Co 2 + ---To ------Nucleotide Salvage Pathway Activated Ribose (5 -phosphoribosyl-1 -Pyrophosphate) also called PRPP and Base ------ Nucleotide

John Buchanan in 1948 Found initial clues about the synthesis of Purine by de novo by feeding a variety of isotopically labeled compounds to pigeons and chemically determining the positions of the labeled atoms in their excreted uric acid (a Purine). Used birds for experiments as they excrete waste nitrogen almost entirely as uric acid ( water insoluble)
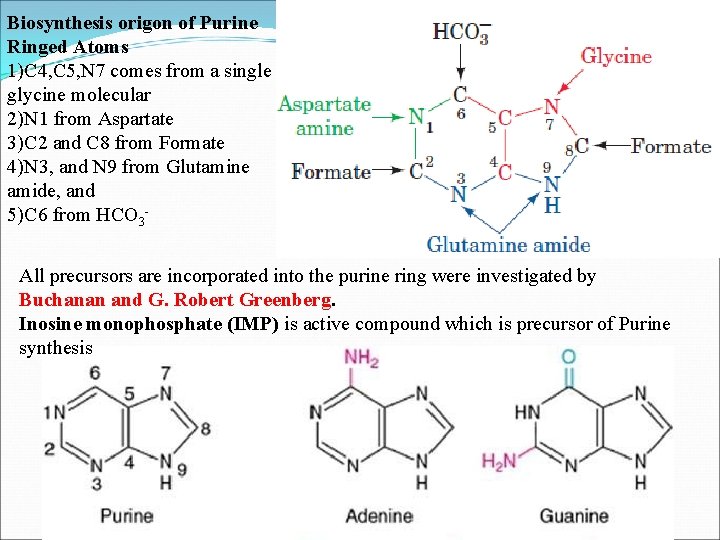
Biosynthesis origon of Purine Ringed Atoms 1)C 4, C 5, N 7 comes from a single glycine molecular 2)N 1 from Aspartate 3)C 2 and C 8 from Formate 4)N 3, and N 9 from Glutamine amide, and 5)C 6 from HCO 3 All precursors are incorporated into the purine ring were investigated by Buchanan and G. Robert Greenberg. Inosine monophosphate (IMP) is active compound which is precursor of Purine synthesis
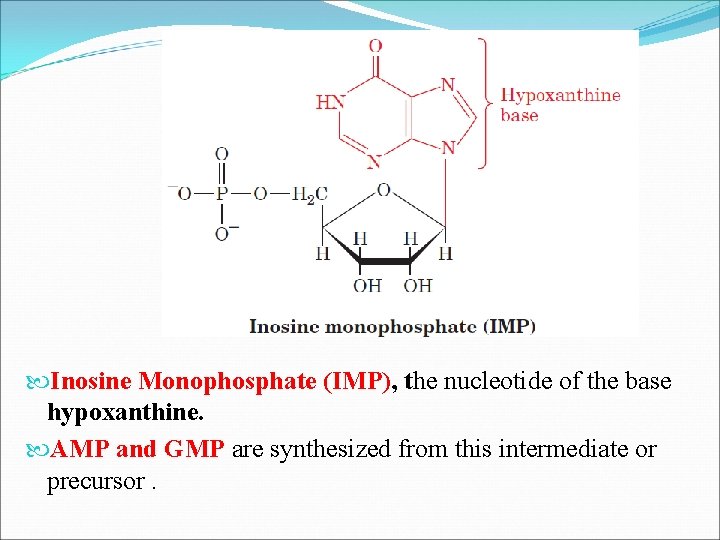
Inosine Monophosphate (IMP), the nucleotide of the base hypoxanthine. AMP and GMP are synthesized from this intermediate or precursor.
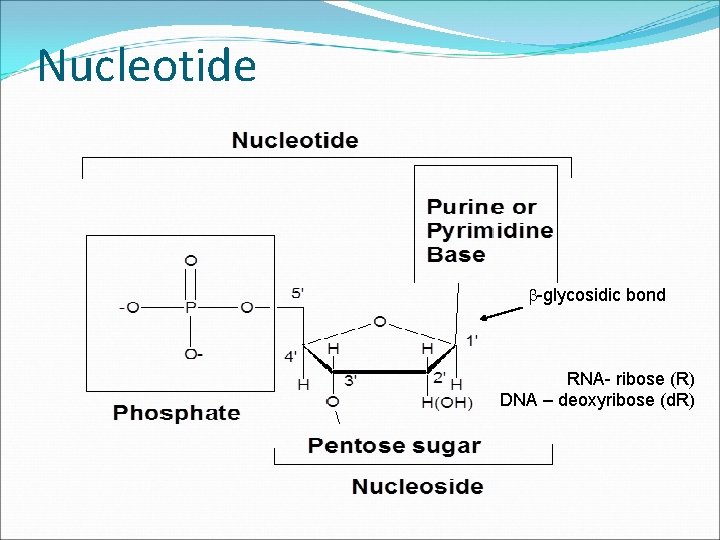
Nucleotide b-glycosidic bond RNA- ribose (R) DNA – deoxyribose (d. R)
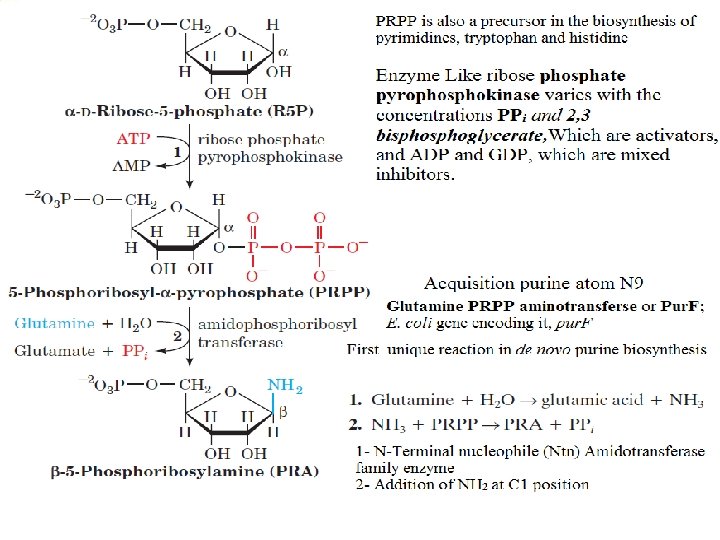
Biosynthesis of Inosine Monophospahte



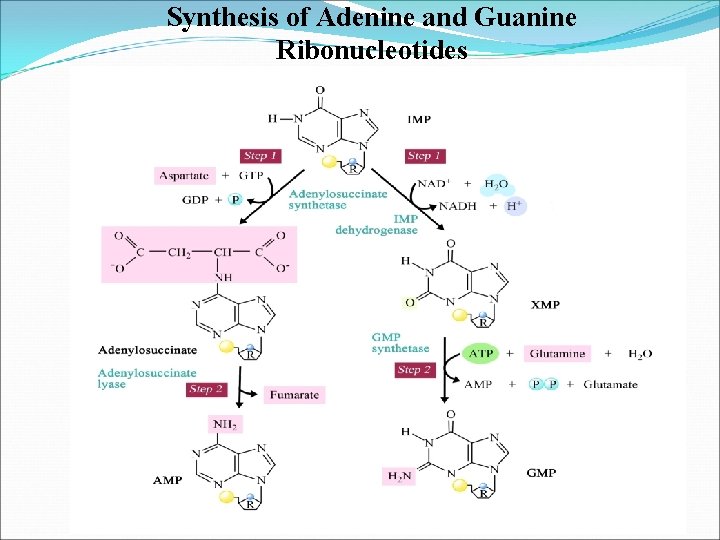
Synthesis of Adenine and Guanine Ribonucleotides
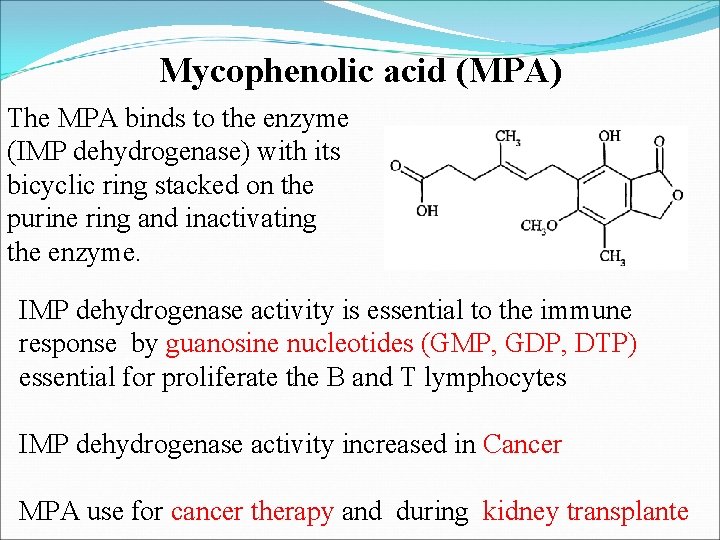
Mycophenolic acid (MPA) The MPA binds to the enzyme (IMP dehydrogenase) with its bicyclic ring stacked on the purine ring and inactivating the enzyme. IMP dehydrogenase activity is essential to the immune response by guanosine nucleotides (GMP, GDP, DTP) essential for proliferate the B and T lymphocytes IMP dehydrogenase activity increased in Cancer MPA use for cancer therapy and during kidney transplante

Synthesis of ADP, ATP, GDP and GTP Corresponding nucleoside triphosphates required for the synthesis of Nucleic acid In phosphorylation reactions respective nucleoside monophosphate synthesize in presence of base specific nucleoside monophosphate kinases These enzymes do not discriminate between ribose and deoxyribose in the substrate. Adenylate kinase is particularly active in the liver and in muscle, where the turnover of energy from ATP is high
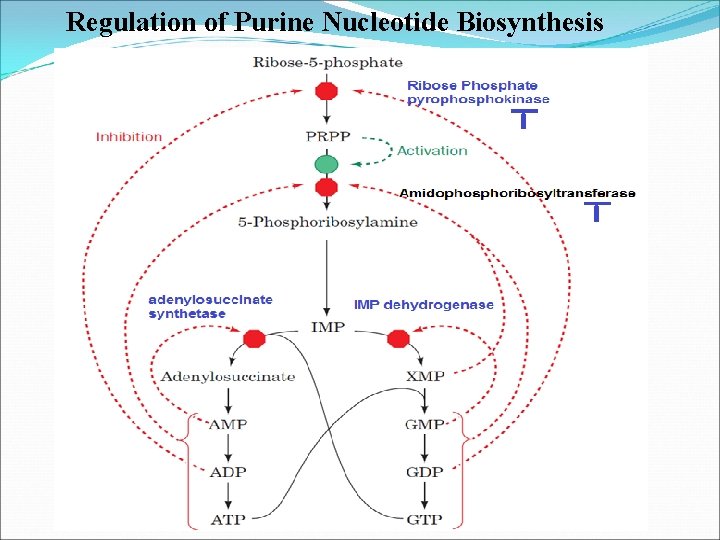
Regulation of Purine Nucleotide Biosynthesis

Purine synthesis pathways involved in nucleic acid metabolism and enhanced during cell proliferation o. IMP, ATP, and GTP synthesized in most of the cells o Enzyme binds ATP, ADP, and AMP at one inhibitory site and GTP, GDP, and GMP at another o. The rate of IMP production is consequently independently but synergistically controlled by the levels of adenine nucleotides and guanine nucleotides. o. A second level of regulation occurs immediately below the branch point leading from IMP to AMP and GMP

Lecture 3

Salvage pathway ØPurine bases produced by degradation of RNA or DNA and intermediate of purine synthesis can be directly converted to the corresponding nucleotides. ØBreak down of Purine nucleotides in Phosphate, ribose (deoxyribose) and the bases adenine, guanine, and/or hypoxanthine during the digestion and cellular metabolism ØHypoxanthine is the purine base present on Inosine-5´monophosphate (IMP)

v. Two phosphoribosyl transferases are involved in salvage pathway Ø APRT (adenine phosphoribosyl transferase) for adenine to form AMP Ø HGPRT (hypoxanthine guanine phosphoribosyl transferase) for guanine or hypoxanthine v. The significance of salvage pathway Ø Save the fuel. Ø Some tissues and organs such as brain and bone marrow are only capable of synthesizing nucleotides by salvage pathway

Purine Salvage Pathway Absence of activity of HGPRT leads to Lesch-Nyhan syndrome

v. Adenine phosphoribosyl transferase (APRT) Ø Adenine + PRPP AMP + PP i v. Hypoxanthine-Guanine phosphoribosyl transferase (HGPRT) Ø Hypoxanthine + PRPP Ø Guanine + PRPP IMP + PPi GMP + PP i These are all reversible reactions AMP, IMP, GMP do not need to be resynthesized de novo !

Lesch-Nyhan syndrome ØFirstly described in 1964 by Michael Lesch and William L. Nyhan. ØThere is a defect or lack in the Hypoxanthine-Guanine phosphoribosyl transferase (HGPRT) enzyme encoded by HPRT 1 gen ØA defect may be in the production or activity of HGPRT ØCauses increased level of Hypoxanthine and Guanine (in degradation to uric acid) ØSex-linked (X-linked) metabolic disorder: only males ØThe prevalence is approximately 1 in 380, 000 ØMutations in the HPRT 1 gen

ØThe rate of purine synthesis is increased about 200 fold ØLoss of HGPRT leads to elevated PRPP levels and stimulation of de novo purine synthesis. ØUric acid level rises and there is gout disease ØIn addition there are mental aberrations or changes ØPatients will self-mutilate by biting lips and fingers off
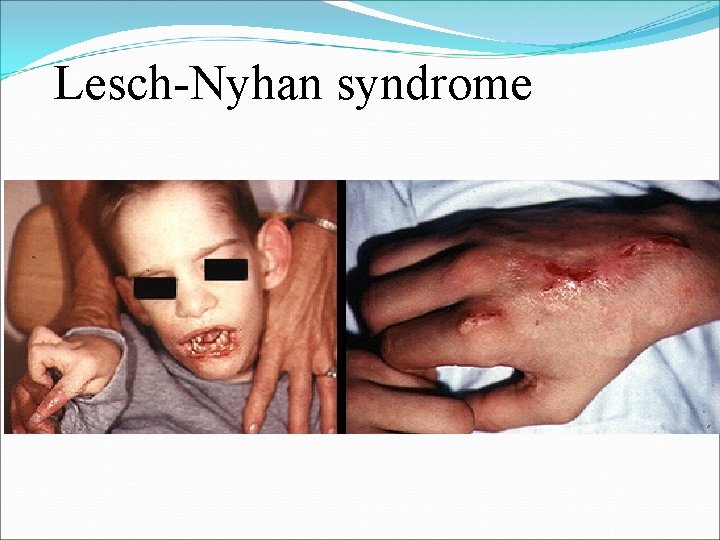
Lesch-Nyhan syndrome

Autism v. About 25% of autistic persons may purines at high level v. So purine test is a diagnostic test for autistic patients ØBiochemical findings from this test disappear adolescence ( between 13 -19 age) ØMust obtain urine specimen in infancy, but it’s may difficult to do! o Pink urine due to uric acid crystals may be seen in diapers

Degradation of Purine Nucleotide

Purine Catabolism and Salvage All purine degradation leads to uric acid v. Ribonucleases, Deoxyribonucleases, Polynucleotidases degraded nucleic acids v. Ingested nucleic acids are degraded to nucleotides in intestine Ø Pancreatic nucleases Ø Intestinal phosphodiesterases v. Group-specific nucleotidases and non-specific phosphatases degrade nucleotides into nucleoside Ø Direct absorption of nucleosides Ø Further degradation

Nucleosidase (Hydrolysis reactions) The is intracellular reaction Nucleoside + H 2 O base + ribose n. Phosphorylase Nucleoside + Pi base + r-1 -phosphate Importantly, Most of the ingested nucleic acids are degraded and excreted

Intracellular Purine Catabolism Nucleotides broken into nucleosides by action of nucleotidase (Hydrolysis reactions) Purine nucleoside phosphorylase (PNP) Inosine to Hypoxanthine Xanthosine to Xanthine Guanosine to Guanine

Intracellular Purine Catabolism v. Xanthine is the point of convergence for the metabolism of the purine bases v. Xanthine converted to Uric acid ØXanthine oxidase catalyzes two reactions v. Purine ribonucleotide (In case of RNA) degradation pathway is same for purine deoxyribonucleotides (In case of DNA)
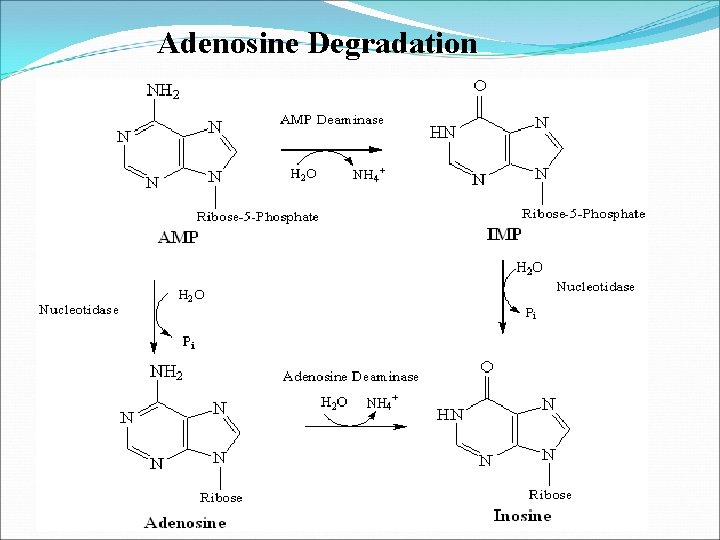
Adenosine Degradation
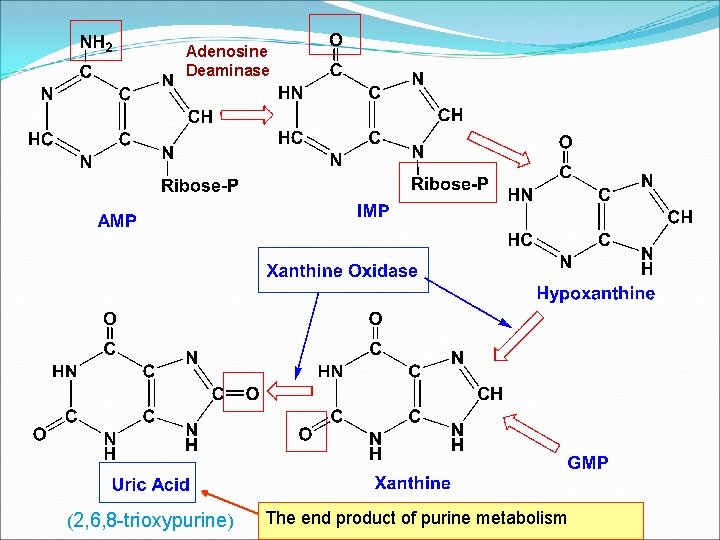
Adenosine Deaminase (2, 6, 8 -trioxypurine) The end product of purine metabolism

Uric acid is the excreted end product of purine catabolism in primates, birds, and some other animals. The rate of uric acid excretion by the normal adult human is about 0. 6 g/24 h, arising in part from ingested purines and in part from the turnover of the purine nucleotides of nucleic acids. The normal concentration of uric acid in the serum of adults is in the range of 3 -7 mg/dl.

GOUT Gout is a disease of the joints, usually in males, caused by an elevated concentration of uric acid in the blood and tissues The joints become inflamed, painful, and arthritic, owing to the abnormal deposition of crystals of sodium urate The kidneys are also affected, because excess uric acid is deposited in the kidney tubules
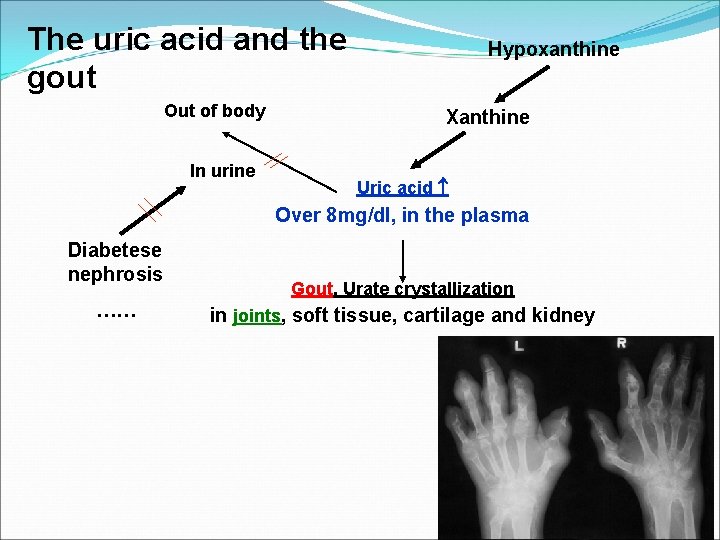
The uric acid and the gout Out of body In urine Hypoxanthine Xanthine Uric acid Over 8 mg/dl, in the plasma Diabetese nephrosis …… Gout, Urate crystallization in joints, soft tissue, cartilage and kidney
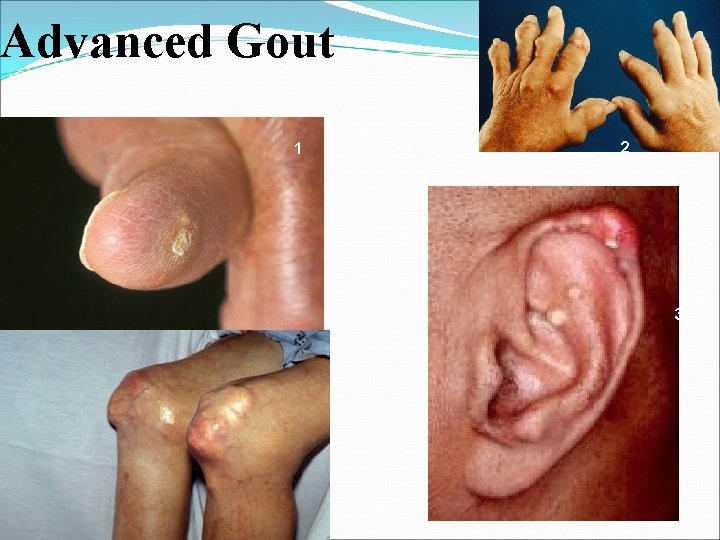
Advanced Gout 2 1 1 3
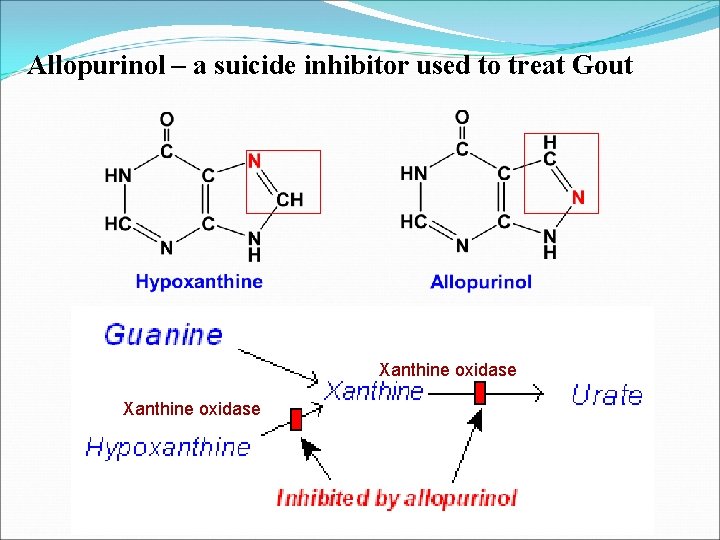
Allopurinol – a suicide inhibitor used to treat Gout Xanthine oxidase

Pyrimidine Synthesis
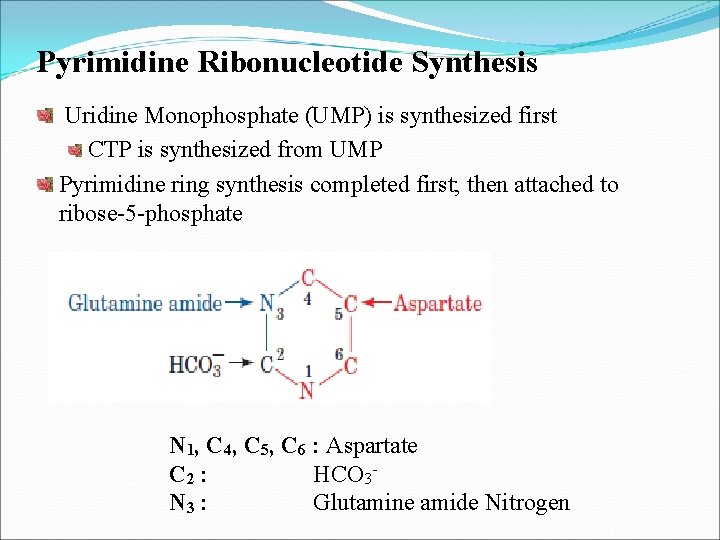
Pyrimidine Ribonucleotide Synthesis Uridine Monophosphate (UMP) is synthesized first CTP is synthesized from UMP Pyrimidine ring synthesis completed first; then attached to ribose-5 -phosphate N 1, C 4, C 5, C 6 : Aspartate C 2 : HCO 3 N 3 : Glutamine amide Nitrogen

Neurospora crassa are unable to synthesize pyrimidines therefore require both cytosine and uracil in their growth medium for proper growth In case of pyrimidine synthesis, pyrimidine ring is coupled to the ribose-5 -phosphate moiety after the ring has been synthesized


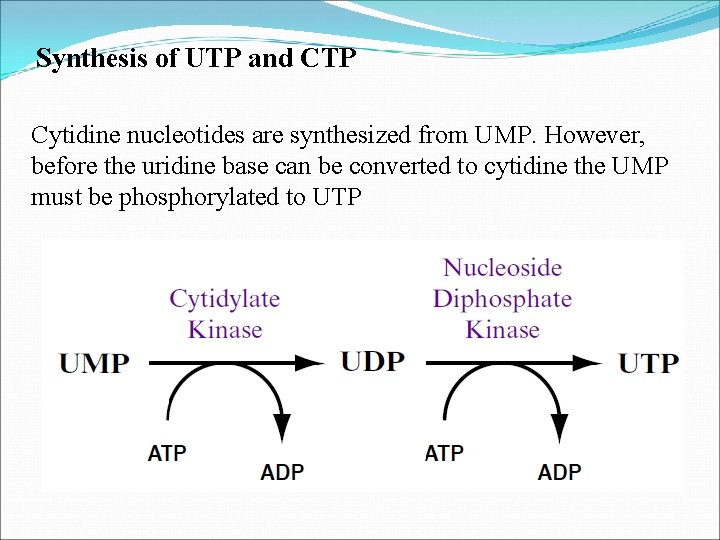
Synthesis of UTP and CTP Cytidine nucleotides are synthesized from UMP. However, before the uridine base can be converted to cytidine the UMP must be phosphorylated to UTP
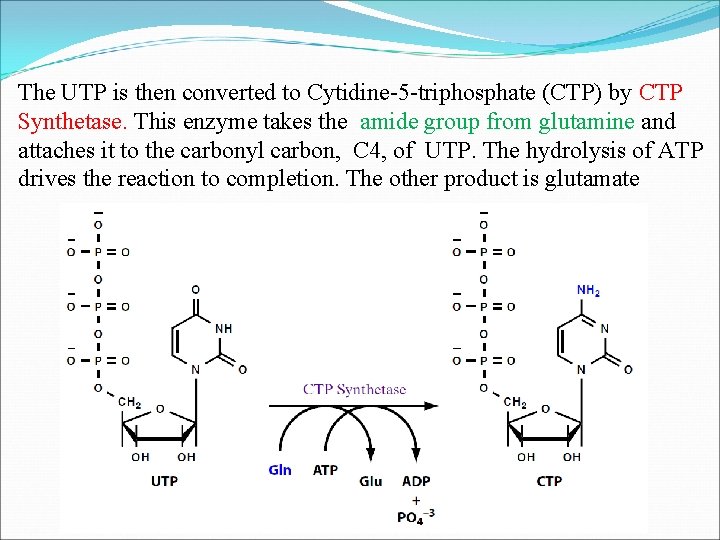
The UTP is then converted to Cytidine-5 -triphosphate (CTP) by CTP Synthetase. This enzyme takes the amide group from glutamine and attaches it to the carbonyl carbon, C 4, of UTP. The hydrolysis of ATP drives the reaction to completion. The other product is glutamate

Pyrimidine nucleotide biosynthesis is controlled at the step catalyzed by Carbamoyl phosphate Synthetase II This is an allosteric enzyme. PRPP and AT P activate the enzyme and UDP and UTP are allosteric inhibitors of its activity
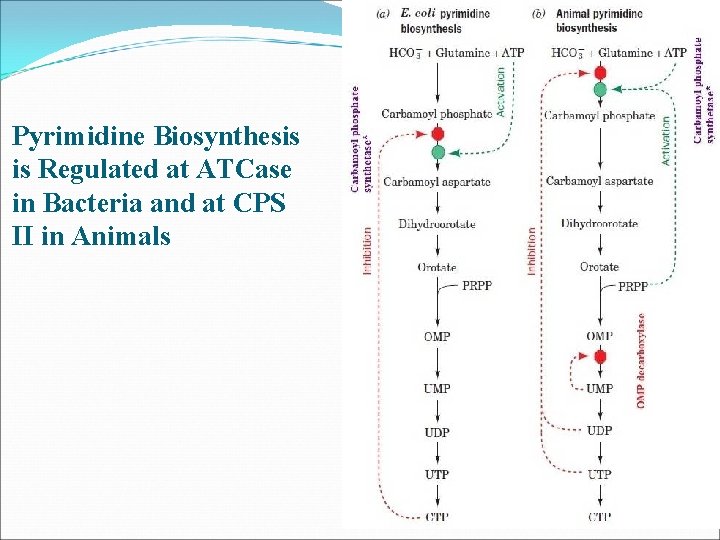
Pyrimidine Biosynthesis is Regulated at ATCase in Bacteria and at CPS II in Animals

UMP Synthesis Overview Need 2 ATPs : Both of them used in first step One transfers phosphate, the other is hydrolyzed to ADP and Pi Two condensation Reactions: Form carbamoyl aspartate and dihydroorotate (intramolecular) Dihydroorotate dehydrogenase is an key intra-mitochondrial enzyme; oxidizing power comes from quinone reduction Attachment of base to ribose ring is catalyzed by orotate phospho ribo syl transferase (OPRT); Phosphoribosyl pyrophosphate (PRPP) provides ribose-5 -P PPi splits off PRPP – irreversible

Pyrimidines Degradation Catabolism of cytosine and uracil yields alanine, ammonium ion, and CO 2 Catabolism of thymine yields amino iso butyric acid, ammonium ion, and CO 2 In some organisms, free pyrimidines are salvaged and recycled to form nucleotides via phosphoribosyltransferase reactions. In humans, pyrimidines are recycled from nucleosides, but free pyrimidine bases are not salvaged.
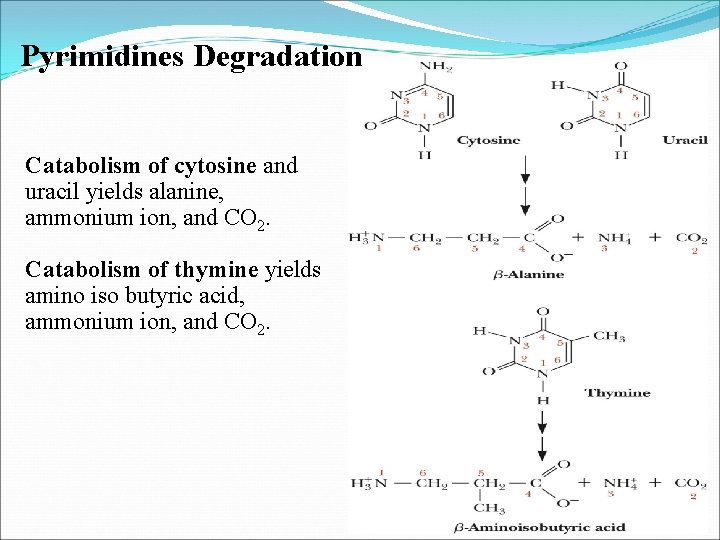
Pyrimidines Degradation Catabolism of cytosine and uracil yields alanine, ammonium ion, and CO 2. Catabolism of thymine yields amino iso butyric acid, ammonium ion, and CO 2.

Biosynthesis of Nucleotides The four ribonucleotides obtained from the biosynthesis pathways of AMP, GMP, UMP and CTP which are reduced to the deoxyribonucleotides needed for DNA synthesis Before their reduction to deoxyribonucleotides, they must all be converted to the diphosphate forms The addition or removal of phosphate is accomplished by enzymes like Nucleoside Mono-phosphate Kinases Nucleoside Di-phosphate Kinase

Ribonucleotide reductases (RNRs) is an enzymes that catalyze the formation of deoxyribonucleotides by the reduction of the corresponding ribonucleotides Class I and II RNRs are widely distributed among prokaryotes depending upon species Class III RNRs occur in prokaryotes that can grow anaerobically Class I RNRs present in all eukaryotes except a few unicellular species

Thioredoxin and Glutaredoxin They are Class I Ribonucleotide Reductase’s (RNRs) This enzyme has two other proteins one is an enzyme other is a small disulfide containing protein In Thioredoxin Reductase, Thioredoxin make up the electron transport chain and reduction of the ribose of the nucleotides NADPH passes a pair of electrons to a FAD covalently linked to Thioredoxin Reductase to form NADP and FADH 2 The FADH 2 then reduces a disulfide (–S–S–) group on Thioredoxin Reductase to a pair of thiol (–SH) groups FADH 2 is oxidized to FAD. Thioredoxin Reductase passes the electrons to a disulfide (–S–S–) on the protein Thioredoxin. oxidized to a disulfide (–S–S–)

The disulfide (–S–S–) on Thioredoxin is reduced to a pair of thiol (–SH) groups and the thiols (–SH) on Thioredoxin Reductase are oxidized to a disulfide (–S–S–) Reductase picks up the elections from Thioredoxin, oxidizing the thiol (–SH) groups on Thioredoxin to a disulfide (–S–S–) and reducing the disulfide (–S–S–) group on Ribonucleotide Reductase to a pair of thiol (–SH) groups Finally, the electrons are used to reduce the ribose sugar of the ribonucleotides to 2 - deoxyribose. During this reduction reaction the thiol (–SH) groups on Ribonucleotide Reductase are oxidized to a disulfide (–S–S–)


In the case of Glutaredoxin Reductase, Glutaredoxin make up the electron transport chain and reduce the ribose to deoxyribose Transfer of electrons to reduce the ribose is identical to Thioredoxin Reductase , except the first step in which NADPH reduces a molecule of oxidized glutathione in the first step of the electron pathway rather than reducing a molecule of FAD
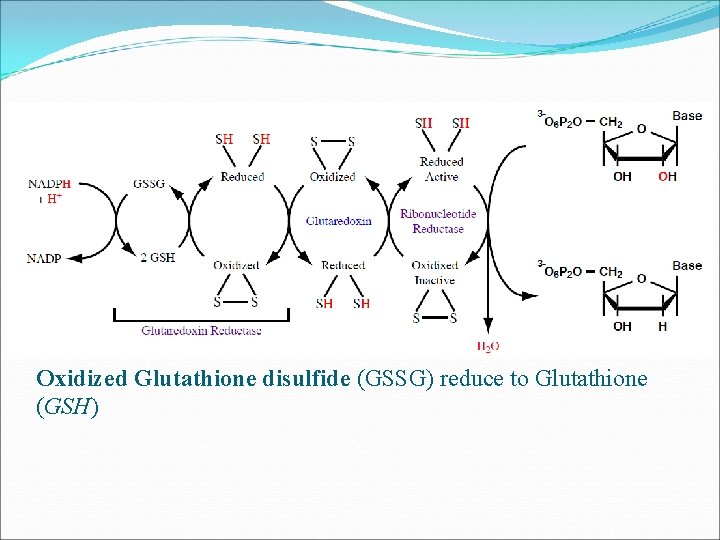
Oxidized Glutathione disulfide (GSSG) reduce to Glutathione (GSH)

Activity of enzyme Ribonucleotide Reductase has a unique control mechanism to assure that the DNA are synthesized in adequate and balanced amounts This enzyme contains Activity Site, Specificity Site, and the catalytic site The activity site turns the enzyme “ON” by ATP or “OFF” by deoxy-ATP Specificity Site controls which nucleotide will be reduced Catalytic site performs the reduction

When the Specificity Site is occupied by ATP or deoxy ATP (d. ATP) CDP and UDP are reduced respectively Specificity Site is occupied by deoxy. TTP (d. TTP) and deoxy. GTP (d. GTP) GDP and ADP are reduced respectively The Specificity Site assures balance and adequate synthesis of deoxyribonucleotides


When the concentration of ATP is high, the cell is energy rich synthesize DNA and divide. ATP binds to the activity site to turn the enzyme “ON”. ATP also binds to the specificity site to stimulate the reduction of the pyrimidines, UDP and CDP Deoxy. UDP is the precursor of deoxythymidine (d. TTP), the base pair partner of d. ATP in DNA High ATP concentrations stimulate the synthesis of its partner in DNA and the partner of deoxyguanosine As deoxythymidine (d. TTP) concentrations build up it signals that the deoxy pyrimidines are present in adequate amounts for DNA replication

d. TTP binds to the specificity site and stimulates the reduction of one of the purines, GDP to d. GDP When d. GTP concentrations enhance, it binds to the specificity site and stimulates the reduction of the other purine like ADP is reduced to d. ADP As d. ATP concentrations increase they signal that all four deoxy nucleotide triphosphates are present in adequate amounts for DNA replication. d. ATP replaces ATP in the activity site and the enzyme is turned “OFF”.

To be noted Deoxyuridylate nucleotides do not incorporated into DNA Two mechanisms to assure it 1. Enzyme Deoxyuridine Triphosphate Diphosphohydrolase or rapidly converts any deoxy UTP to UMP 2. The deoxy UMP is rapidly and quantitatively converted to deoxy TMP

Conversion of d. UMP to d. TMP Deoxy UMP (d. UMP) converts to deoxy TMP (d. TMP) by help of Enzyme Thymidylate Synthase or Deoxyuridine Triphosphate Diphosphohydrolase N 5, N 10 -Methylene-Tetrahydrofolate (N 5, N 10 -Methylene. TH 4) N 5, N 10 -Methylene-TH 4 having two functions during this conversion Donates a one carbon fragment to the d. UMP nucleotide On other side, tetrahydrofolate molecule acts as a reducing agent and donates a pair of hydrogen atoms to reduce the one carbon fragment from the methylene oxidation state to the methyl oxidation state
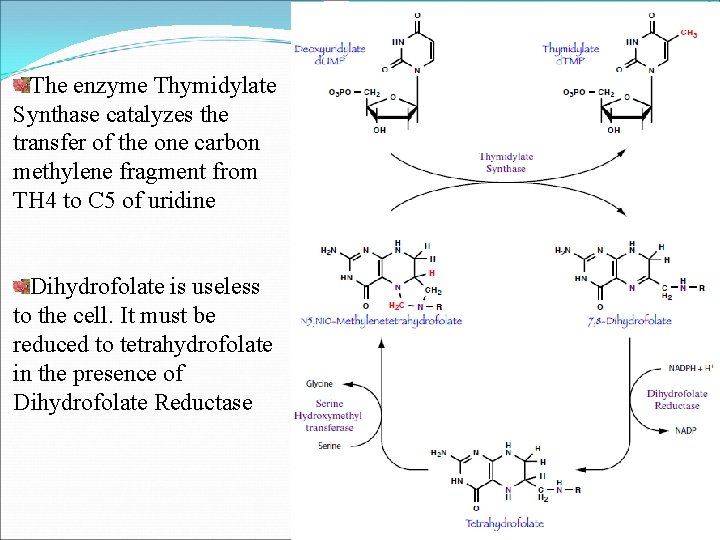
The enzyme Thymidylate Synthase catalyzes the transfer of the one carbon methylene fragment from TH 4 to C 5 of uridine Dihydrofolate is useless to the cell. It must be reduced to tetrahydrofolate in the presence of Dihydrofolate Reductase

NADPH and a hydrogen ion (H+) donates the hydrogens and electrons necessary for the reaction. Then TH 4 is reformed, it accepts a one carbon fragment from serine and it is ready for the next cycle of reactions Anticancer drug The enzyme Dihydrofolate Reductase was the first target for cancer chemotherapeutic agents d. TMP and (d. TTP) is essential for DNA replication, inhibiting the formation of d. TTP would inhibit DNA replication. With DNA synthesis inhibited, cancer cells would cease to divide, and the tumor would stop growing. The drug Methotrexate is a specific competitive inhibitor of the enzyme Dihydrofolate Reductase

The Mycophenolic acid (MPA) binds to the enzyme (IMP dehydrogenase) and block the formation of GMP and ultimately reduced the cell proliferation
- Slides: 73