Protein degradation Ubiquitinproteasome system Autophagylysosome system Degradation signals

Protein degradation • Ubiquitin-proteasome system • Autophagy-lysosome system • Degradation signals in proteolysis • Protein N-termini vs. proteolysis Yong Tae Kwon Protein Metabolism Medical Research Center Department of Biomedical Sciences Seoul National University School of Medicine

Proteins are degraded • Proteins of the body are in a dynamic state of synthesis and degradation. • Approximately 6% of our cellular proteins are degraded and resynthesized daily. § Quality control § Proteins being synthesized are folded incorrectly § Proteins become denature/misfolded/damaged § Regulation of biological pathways § Cell cycle, apoptosis, differentiation, development § Receptor mediated endocytosis § Signaling pathways § Generating energy upon starvation § Autophagy
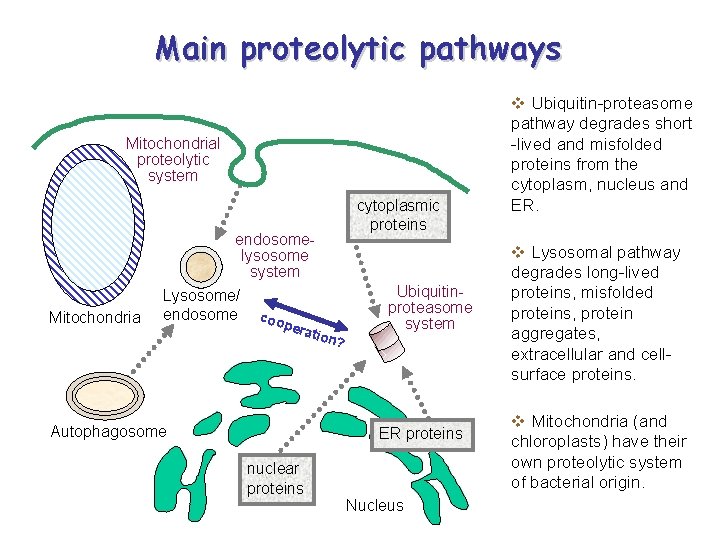
Main proteolytic pathways Mitochondrial proteolytic system endosomelysosome system Mitochondria Lysosome/ endosome coo pera Autophagosome tion ? cytoplasmic proteins Ubiquitinproteasome system ER proteins nuclear proteins Nucleus v Ubiquitin-proteasome pathway degrades short -lived and misfolded proteins from the cytoplasm, nucleus and ER. v Lysosomal pathway degrades long-lived proteins, misfolded proteins, protein aggregates, extracellular and cellsurface proteins. v Mitochondria (and chloroplasts) have their own proteolytic system of bacterial origin.
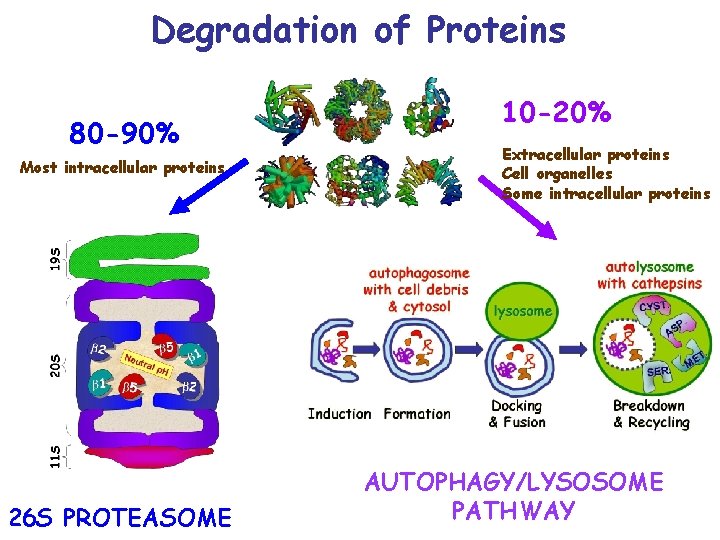
Degradation of Proteins 80 -90% Most intracellular proteins 26 S PROTEASOME 10 -20% Extracellular proteins Cell organelles Some intracellular proteins AUTOPHAGY/LYSOSOME PATHWAY

Content 1. Protein degradation: overview 2. Ubiquitin-proteasome system (UPS) 3. UPS in cell cycle and cancer 4. Drugs targeting UPS 5. Autophagy 6. Protein quality control: UPS and autophagy 7. Protein degradation in neurodegenerative diseases
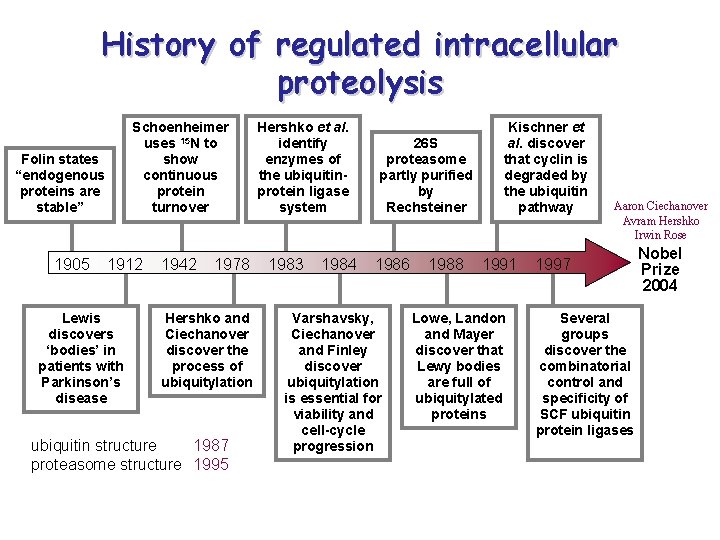
History of regulated intracellular proteolysis Schoenheimer uses 15 N to show continuous protein turnover Folin states “endogenous proteins are stable” 1905 1912 Lewis discovers ‘bodies’ in patients with Parkinson’s disease 1942 1978 Hershko and Ciechanover discover the process of ubiquitylation ubiquitin structure 1987 proteasome structure 1995 Hershko et al. identify enzymes of the ubiquitinprotein ligase system 1983 1984 26 S proteasome partly purified by Rechsteiner 1986 Varshavsky, Ciechanover and Finley discover ubiquitylation is essential for viability and cell-cycle progression 1988 Kischner et al. discover that cyclin is degraded by the ubiquitin pathway 1991 Lowe, Landon and Mayer discover that Lewy bodies are full of ubiquitylated proteins Aaron Ciechanover Avram Hershko Irwin Rose 1997 Several groups discover the combinatorial control and specificity of SCF ubiquitin protein ligases Nobel Prize 2004

The Nobel Prize in Chemistry 2004 "for the discovery of ubiquitin-mediated protein degradation” Aaron Ciechanover 1/3 of the prize Israel Avram Hershko 1/3 of the prize Israel Professor Protein Metabolism Medical Research Center Seoul National University School of Medicine Irwin Rose 1/3 of the prize USA
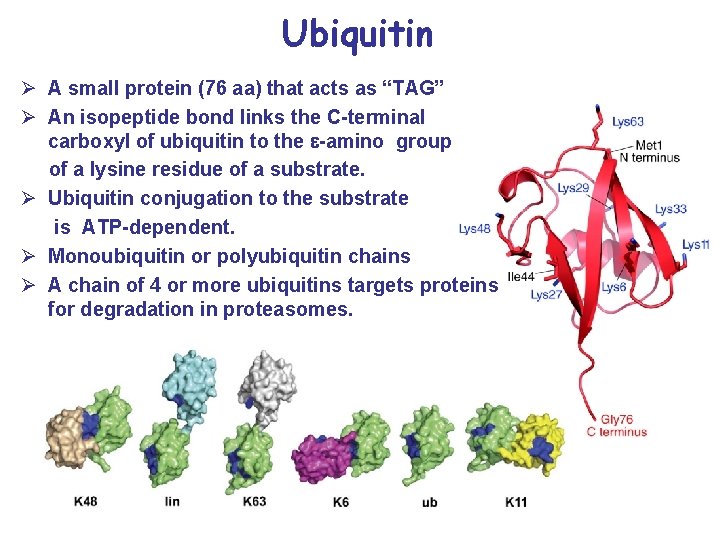
Ubiquitin Ø A small protein (76 aa) that acts as “TAG” Ø An isopeptide bond links the C-terminal carboxyl of ubiquitin to the e-amino group of a lysine residue of a substrate. Ø Ubiquitin conjugation to the substrate is ATP-dependent. Ø Monoubiquitin or polyubiquitin chains Ø A chain of 4 or more ubiquitins targets proteins for degradation in proteasomes.
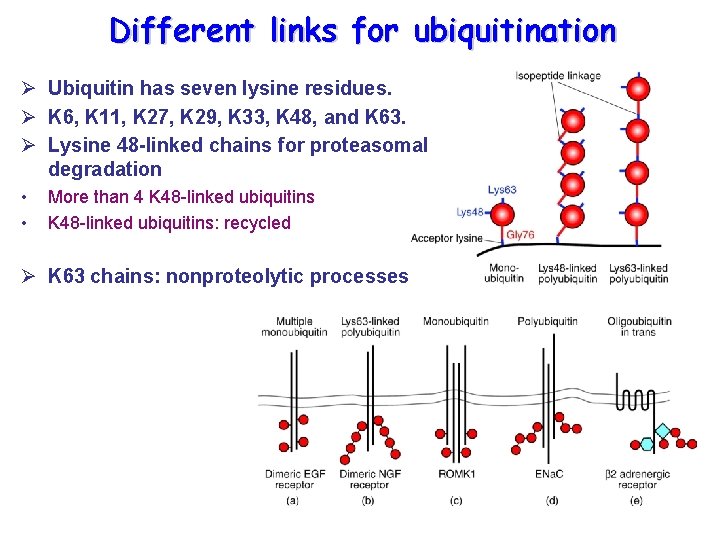
Different links for ubiquitination Ø Ubiquitin has seven lysine residues. Ø K 6, K 11, K 27, K 29, K 33, K 48, and K 63. Ø Lysine 48 -linked chains for proteasomal degradation • • More than 4 K 48 -linked ubiquitins: recycled Ø K 63 chains: nonproteolytic processes
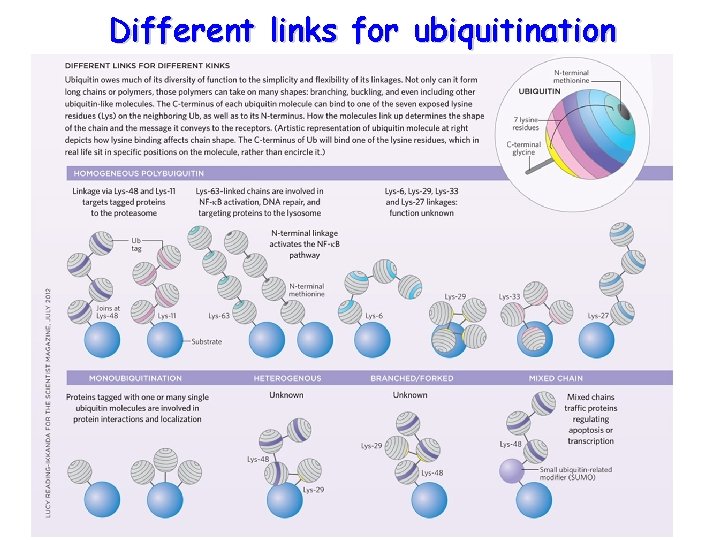
Different links for ubiquitination

Functions of Ubiquitination • Monoubiquitination Ø Receptor internalization Ø Endocytosis – lysosome Ø Transcription regulation • Polyubiquitination Ø Targets proteins from cytoplasm, nucleus & ER for degradation by the proteasome Ø DNA repair, signaling, autophagy

Four Step in Ubiquitin-Proteasome System § § Recognition • Degrons of substrates recognized by E 3 Ubiquitination • E 1, E 2, E 3 Deubiquitination • Deubiquitination enzymes Degradation • Proteasome
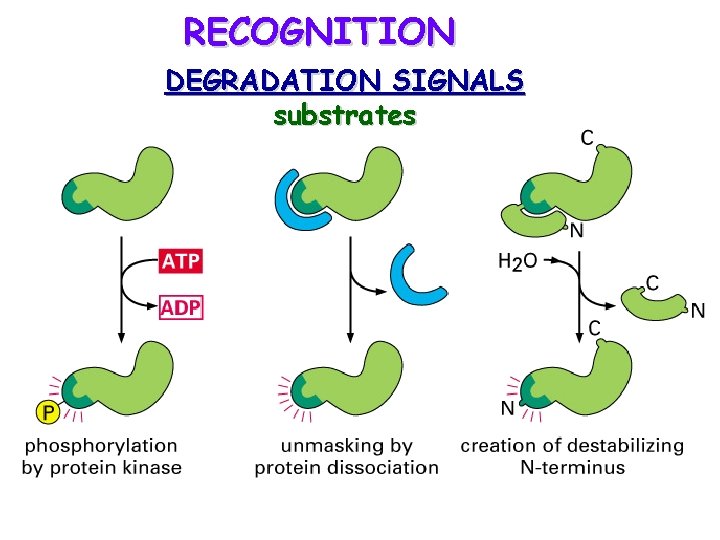
RECOGNITION DEGRADATION SIGNALS substrates
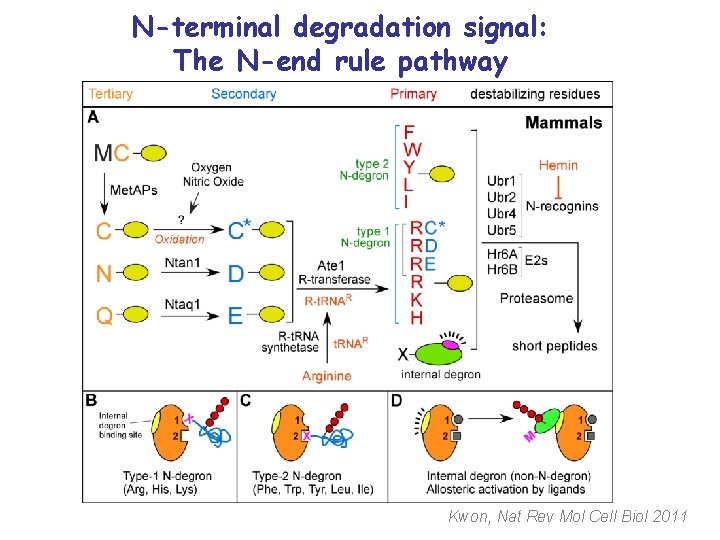
N-terminal degradation signal: The N-end rule pathway Kwon, Nat Rev Mol Cell Biol 2011
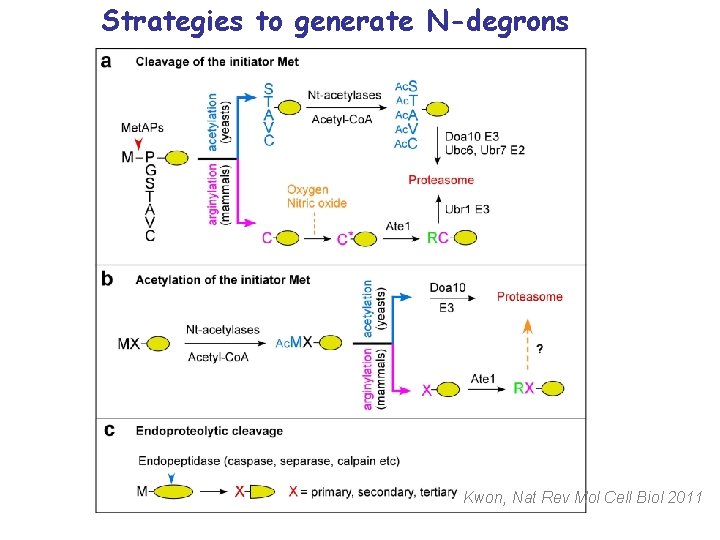
Strategies to generate N-degrons Kwon, Nat Rev Mol Cell Biol 2011
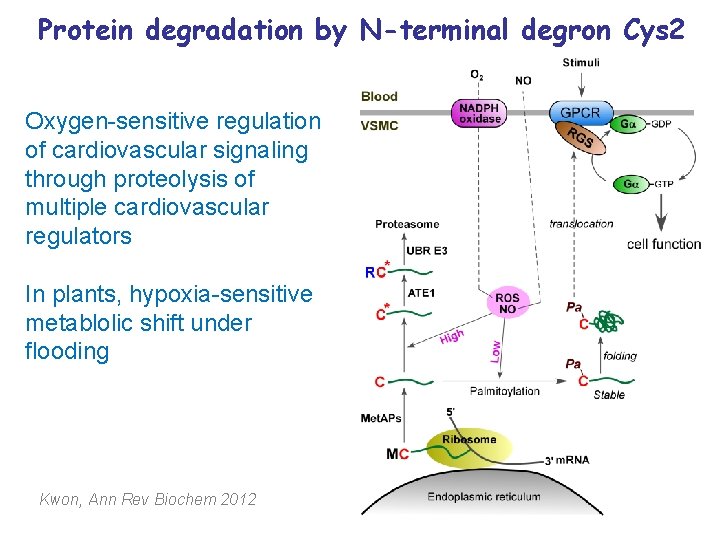
Protein degradation by N-terminal degron Cys 2 Oxygen-sensitive regulation of cardiovascular signaling through proteolysis of multiple cardiovascular regulators In plants, hypoxia-sensitive metablolic shift under flooding Kwon, Ann Rev Biochem 2012

Creation of N-degrons by endopeptidase cleavage N-terminal Asn, Gln, Asp, Glu of C-terminal fragments Caspases alone cleave more than 1, 000 proteins. Neurodegeneration-associated proteins: A , Tau etc Cell cycle regulators: BRCA 1, CDC 6 etc
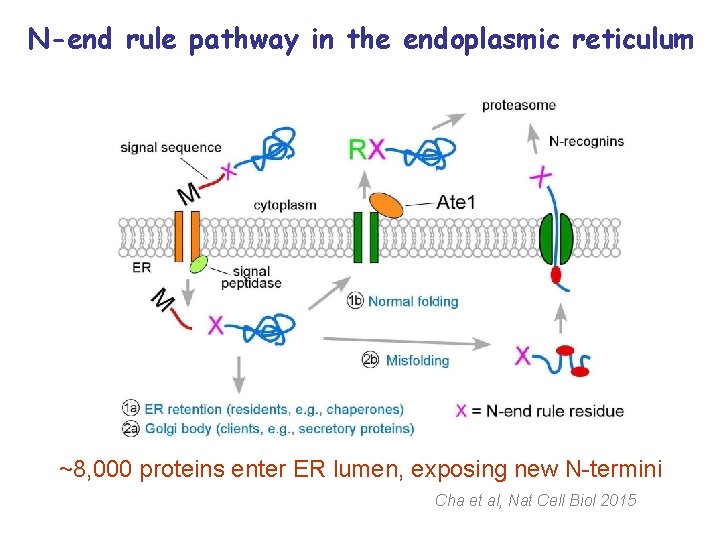
N-end rule pathway in the endoplasmic reticulum ~8, 000 proteins enter ER lumen, exposing new N-termini Cha et al, Nat Cell Biol 2015
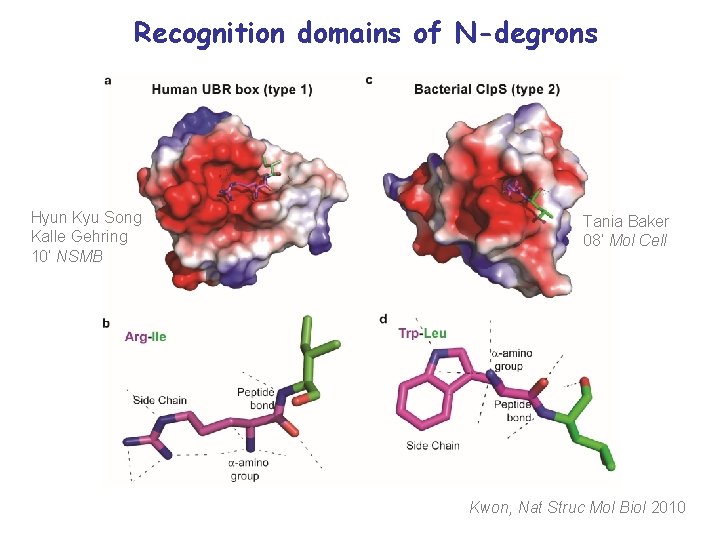
Recognition domains of N-degrons Hyun Kyu Song Kalle Gehring 10’ NSMB Tania Baker 08’ Mol Cell Kwon, Nat Struc Mol Biol 2010
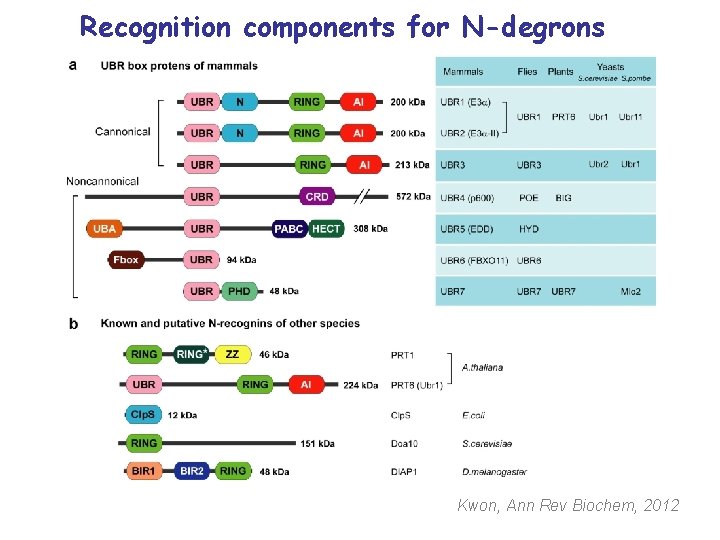
Recognition components for N-degrons Kwon, Ann Rev Biochem, 2012

The Ubiquitin-Proteasome System Regulates cellular concentrations of short-lived proteins Selective recognition by ubiquitin ligases (E 3) Via ubiquitination Proteasomal degradation into small peptides
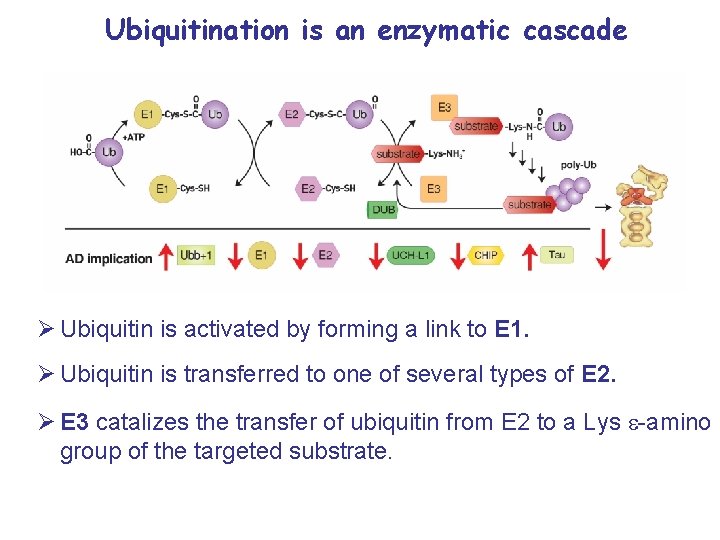
Ubiquitination is an enzymatic cascade Ø Ubiquitin is activated by forming a link to E 1. Ø Ubiquitin is transferred to one of several types of E 2. Ø E 3 catalizes the transfer of ubiquitin from E 2 to a Lys e-amino group of the targeted substrate.
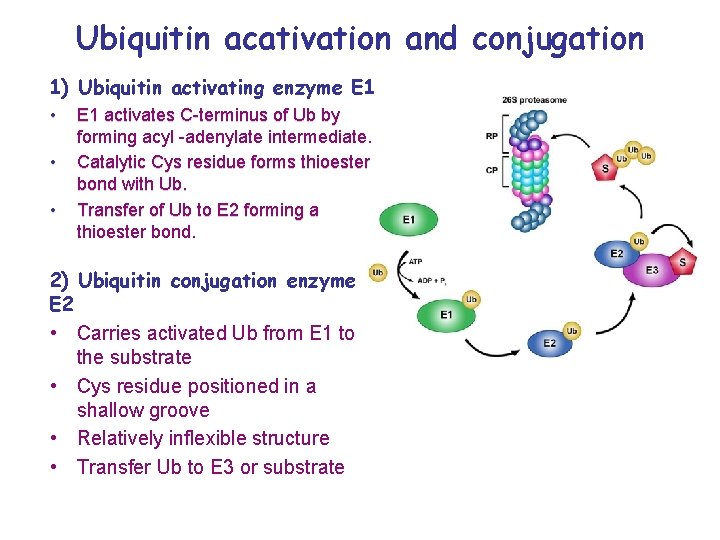
Ubiquitin acativation and conjugation 1) Ubiquitin activating enzyme E 1 • • • E 1 activates C-terminus of Ub by forming acyl -adenylate intermediate. Catalytic Cys residue forms thioester bond with Ub. Transfer of Ub to E 2 forming a thioester bond. 2) Ubiquitin conjugation enzyme E 2 • Carries activated Ub from E 1 to the substrate • Cys residue positioned in a shallow groove • Relatively inflexible structure • Transfer Ub to E 3 or substrate
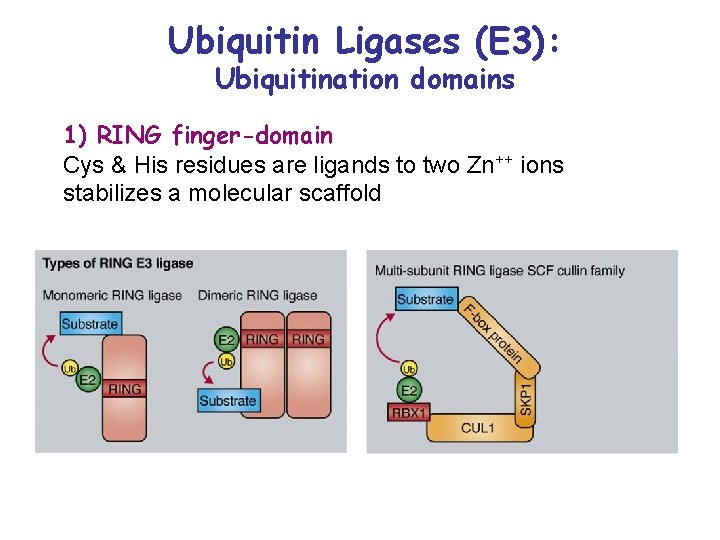
Ubiquitin Ligases (E 3): Ubiquitination domains 1) RING finger-domain Cys & His residues are ligands to two Zn++ ions stabilizes a molecular scaffold
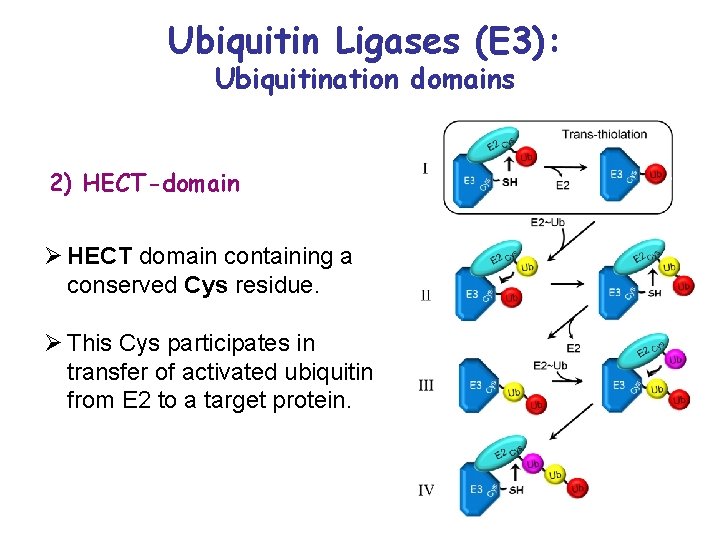
Ubiquitin Ligases (E 3): Ubiquitination domains 2) HECT-domain Ø HECT domain containing a conserved Cys residue. Ø This Cys participates in transfer of activated ubiquitin from E 2 to a target protein.
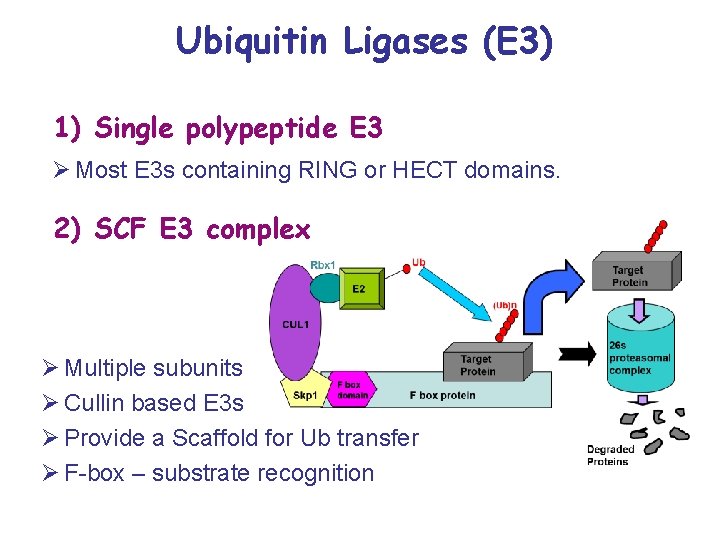
Ubiquitin Ligases (E 3) 1) Single polypeptide E 3 Ø Most E 3 s containing RING or HECT domains. 2) SCF E 3 complex Ø Multiple subunits Ø Cullin based E 3 s Ø Provide a Scaffold for Ub transfer Ø F-box – substrate recognition
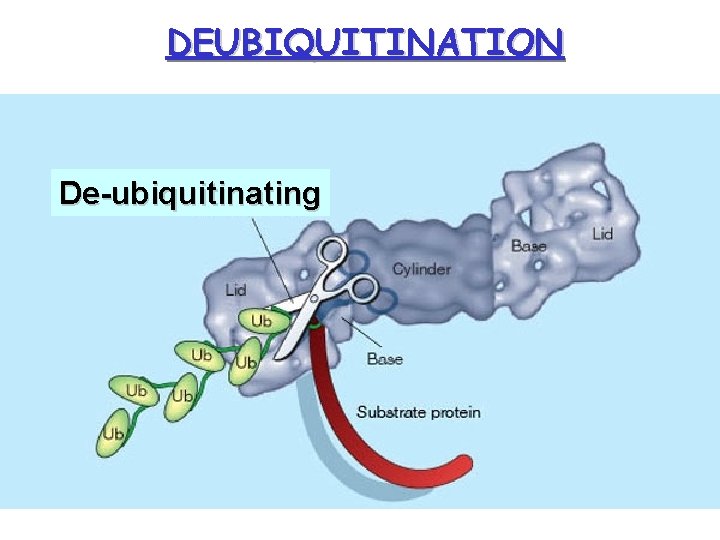
DEUBIQUITINATION De-ubiquitinating
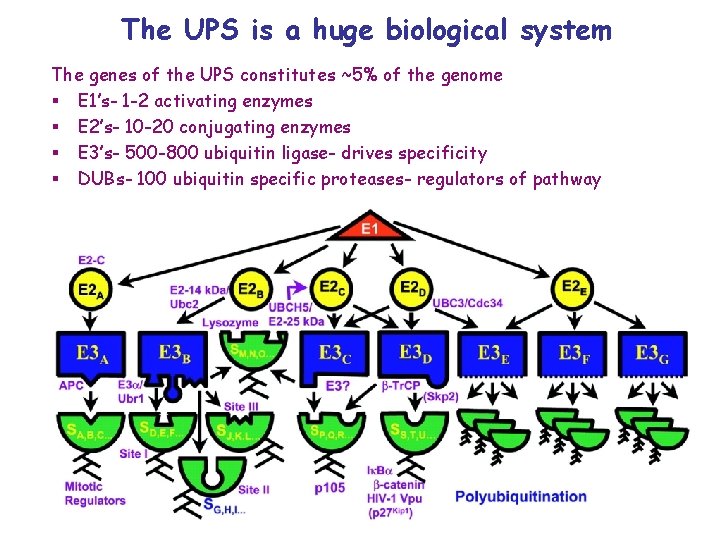
The UPS is a huge biological system The genes of the UPS constitutes ~5% of the genome § E 1’s- 1 -2 activating enzymes § E 2’s- 10 -20 conjugating enzymes § E 3’s- 500 -800 ubiquitin ligase- drives specificity § DUBs- 100 ubiquitin specific proteases- regulators of pathway
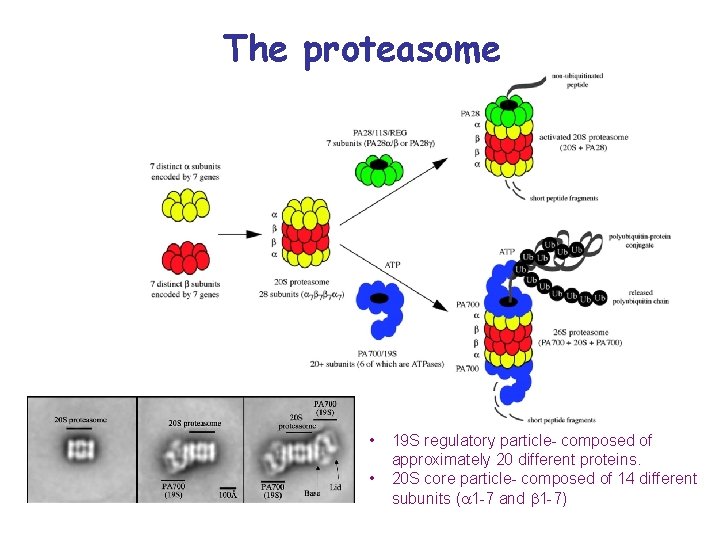
The proteasome • • 19 S regulatory particle- composed of approximately 20 different proteins. 20 S core particle- composed of 14 different subunits ( 1 -7 and 1 -7)
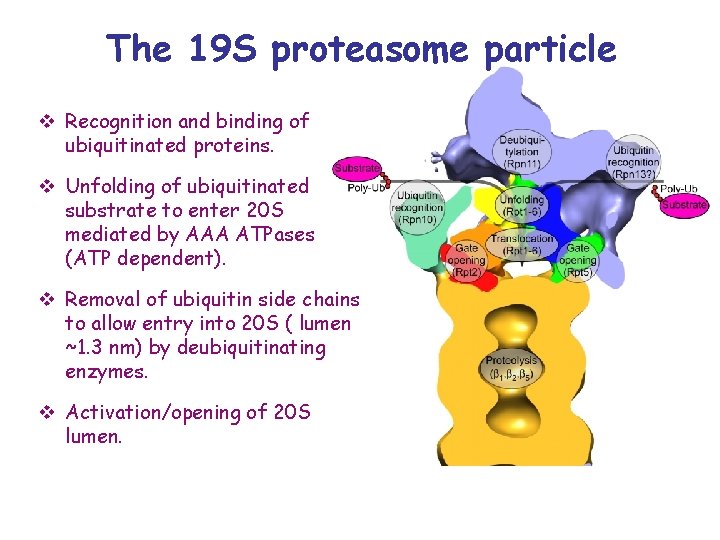
The 19 S proteasome particle v Recognition and binding of ubiquitinated proteins. v Unfolding of ubiquitinated substrate to enter 20 S mediated by AAA ATPases (ATP dependent). v Removal of ubiquitin side chains to allow entry into 20 S ( lumen ~1. 3 nm) by deubiquitinating enzymes. v Activation/opening of 20 S lumen.

The 20 S Proteasome particle v Contains the endopeptidase activity v The alpha subunits function is to control the opening and closing of the 20 S gate (interacts with 19 S) v The beta subunits 1, 2 and 5 contain the endopeptidase activity of the proteasome. v The proteasome degrades the proteins to ~8 amino-acid peptides.
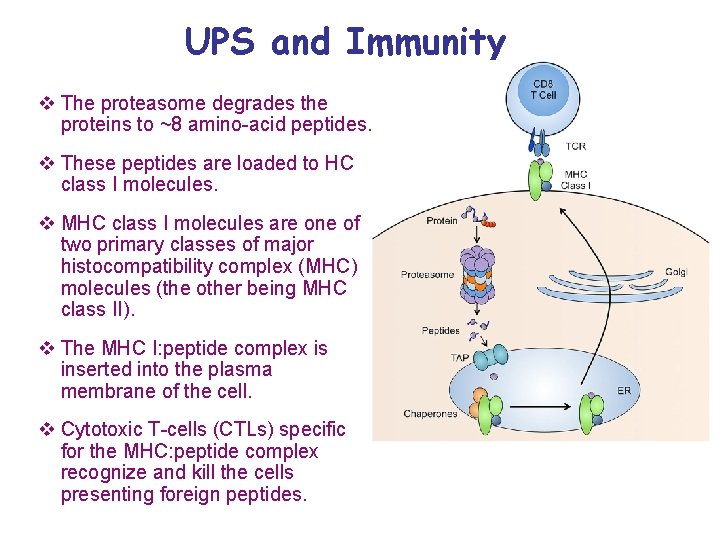
UPS and Immunity v The proteasome degrades the proteins to ~8 amino-acid peptides. v These peptides are loaded to HC class I molecules. v MHC class I molecules are one of two primary classes of major histocompatibility complex (MHC) molecules (the other being MHC class II). v The MHC I: peptide complex is inserted into the plasma membrane of the cell. v Cytotoxic T-cells (CTLs) specific for the MHC: peptide complex recognize and kill the cells presenting foreign peptides.
UPS, Cell Cycle and Cancer Why is cell cycle important? ~1. 5 M new cancer patients in US ~0. 5 M deaths in US ~13% of all deaths ~7. 6 M deaths worldwide in 2007 Cell cycle checkpoint DNA repair Apoptosis Cancer All Connected!!!
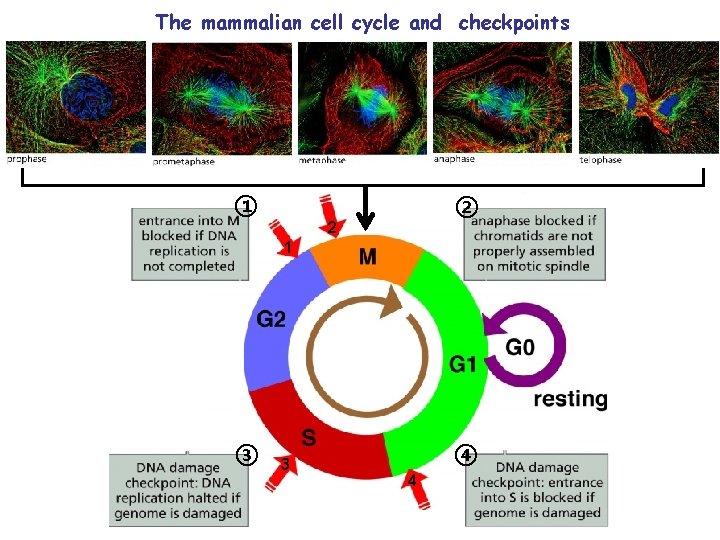
The mammalian cell cycle and checkpoints ① ② 2 1 ③ 3 ④ 4
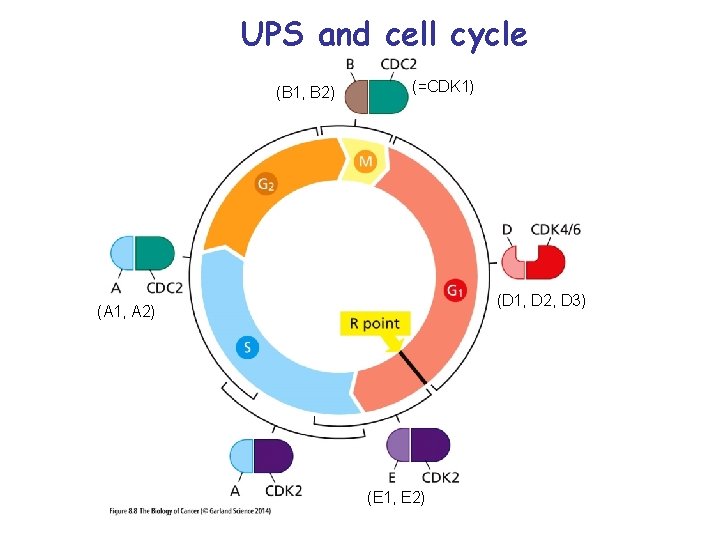
UPS and cell cycle (B 1, B 2) (=CDK 1) (D 1, D 2, D 3) (A 1, A 2) (E 1, E 2)
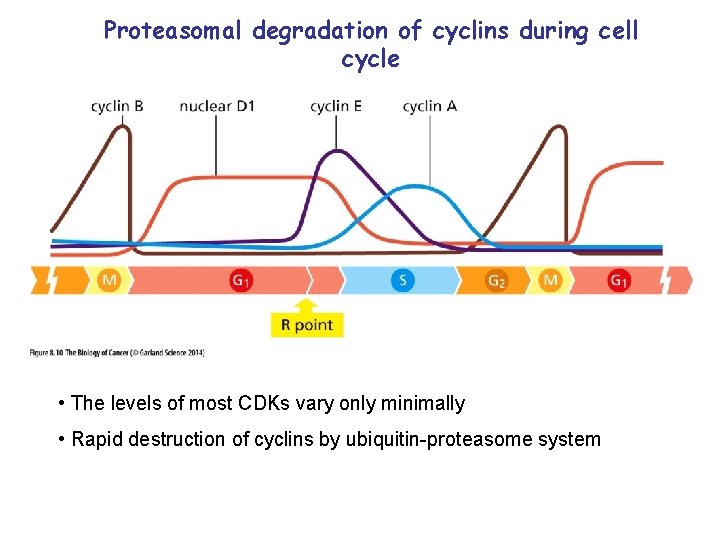
Proteasomal degradation of cyclins during cell cycle • The levels of most CDKs vary only minimally • Rapid destruction of cyclins by ubiquitin-proteasome system
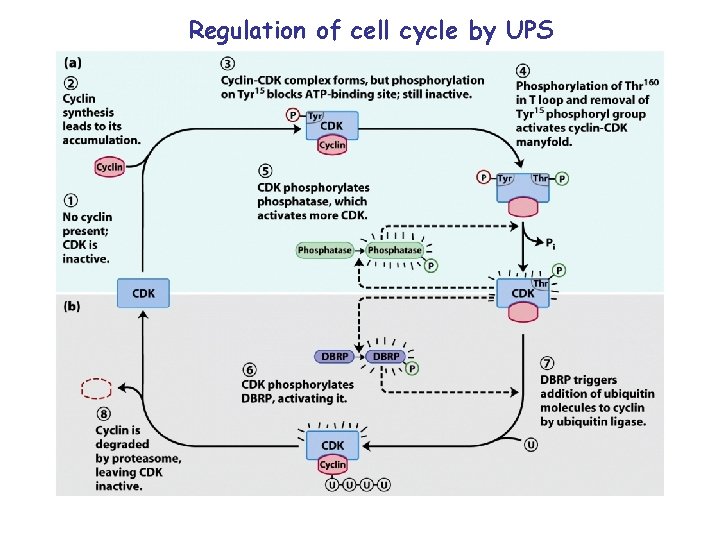
Regulation of cell cycle by UPS
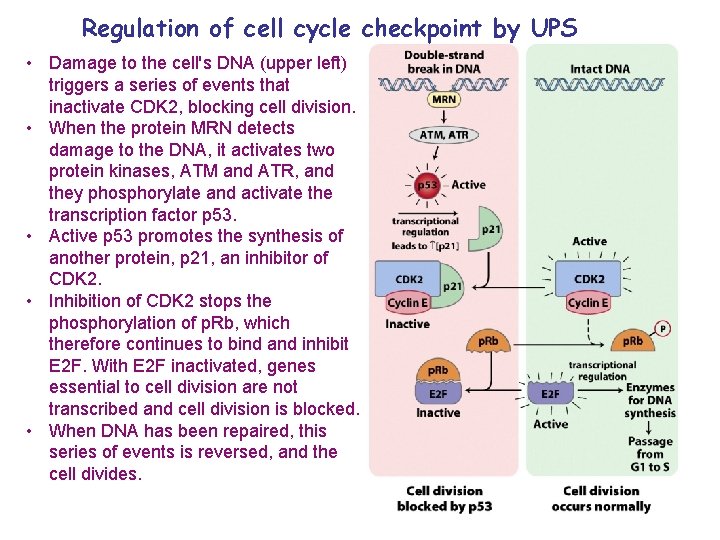
Regulation of cell cycle checkpoint by UPS • Damage to the cell's DNA (upper left) triggers a series of events that inactivate CDK 2, blocking cell division. • When the protein MRN detects damage to the DNA, it activates two protein kinases, ATM and ATR, and they phosphorylate and activate the transcription factor p 53. • Active p 53 promotes the synthesis of another protein, p 21, an inhibitor of CDK 2. • Inhibition of CDK 2 stops the phosphorylation of p. Rb, which therefore continues to bind and inhibit E 2 F. With E 2 F inactivated, genes essential to cell division are not transcribed and cell division is blocked. • When DNA has been repaired, this series of events is reversed, and the cell divides.
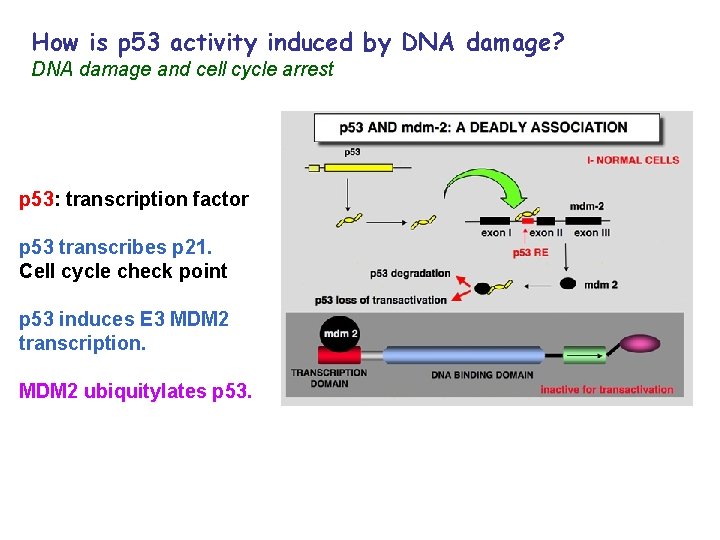
How is p 53 activity induced by DNA damage? DNA damage and cell cycle arrest p 53: transcription factor p 53 transcribes p 21. Cell cycle check point p 53 induces E 3 MDM 2 transcription. MDM 2 ubiquitylates p 53.
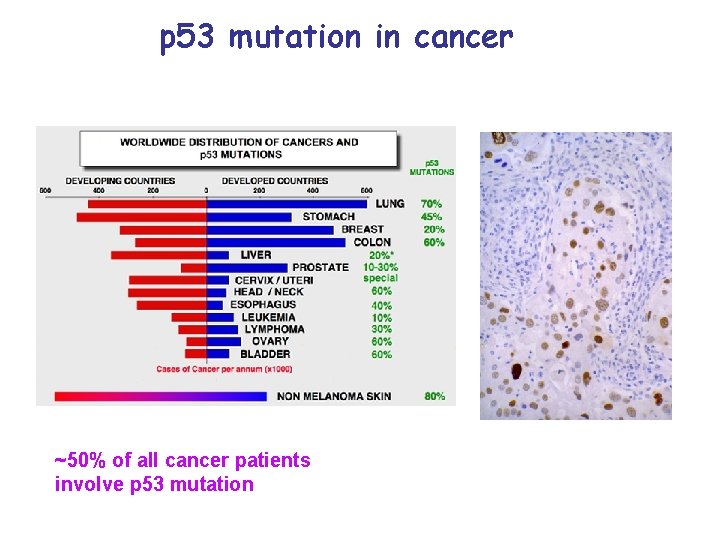
p 53 mutation in cancer ~50% of all cancer patients involve p 53 mutation
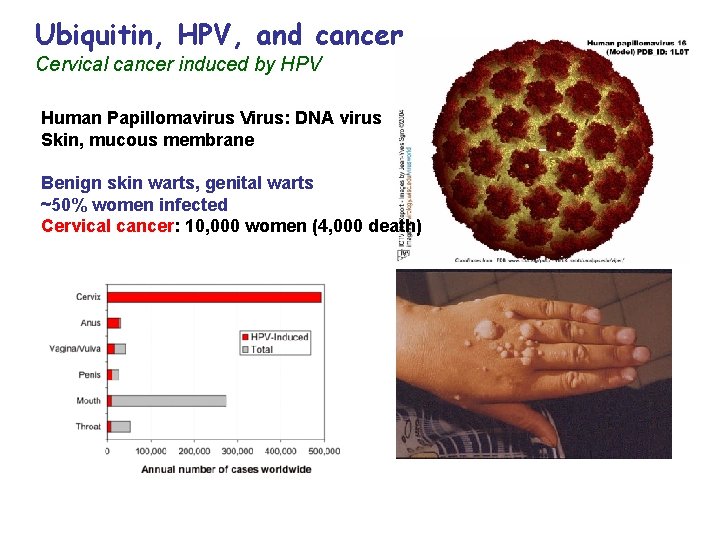
Ubiquitin, HPV, and cancer Cervical cancer induced by HPV Human Papillomavirus Virus: DNA virus Skin, mucous membrane Benign skin warts, genital warts ~50% women infected Cervical cancer: 10, 000 women (4, 000 death)
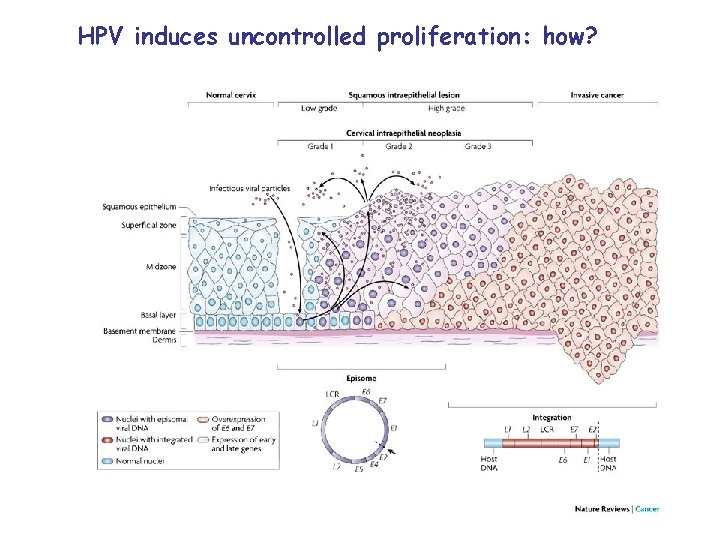
HPV induces uncontrolled proliferation: how?
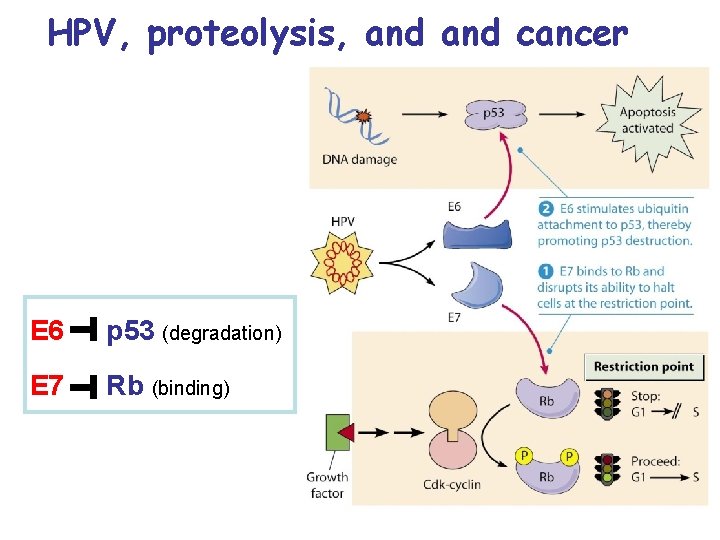
HPV, proteolysis, and cancer E 6 p 53 (degradation) E 7 Rb (binding)
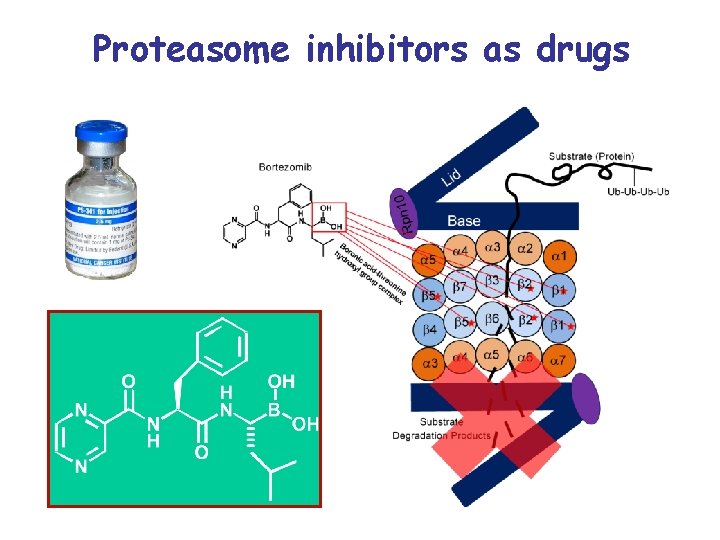
Proteasome inhibitors as drugs

Valcade: anticancer drug • Cytotoxicity involves multiple mechanisms of action – – – Stabilization of cell-cycle regulatory proteins Inhibition of NF- B activation Anti-angiogenic activity Induction of apoptosis Override of Bcl-2 resistance Hypoxic cells are hypersensitive
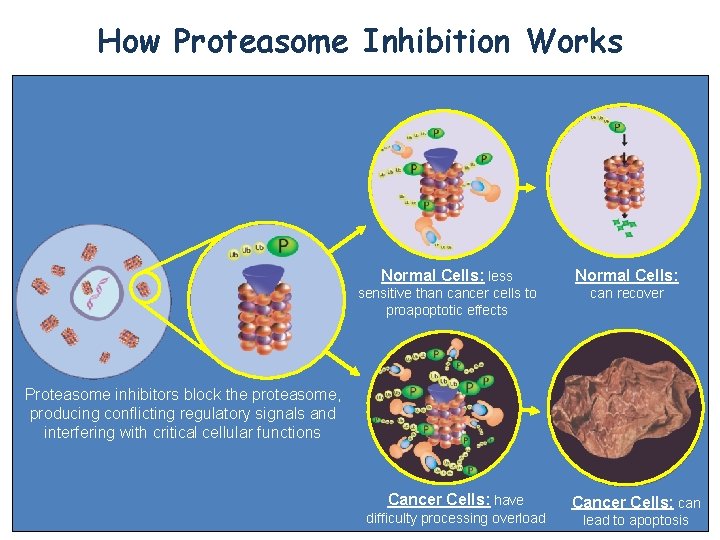
How Proteasome Inhibition Works Normal Cells: less Normal Cells: sensitive than cancer cells to proapoptotic effects can recover Proteasome inhibitors block the proteasome, producing conflicting regulatory signals and interfering with critical cellular functions Cancer Cells: have difficulty processing overload Cancer Cells: can lead to apoptosis
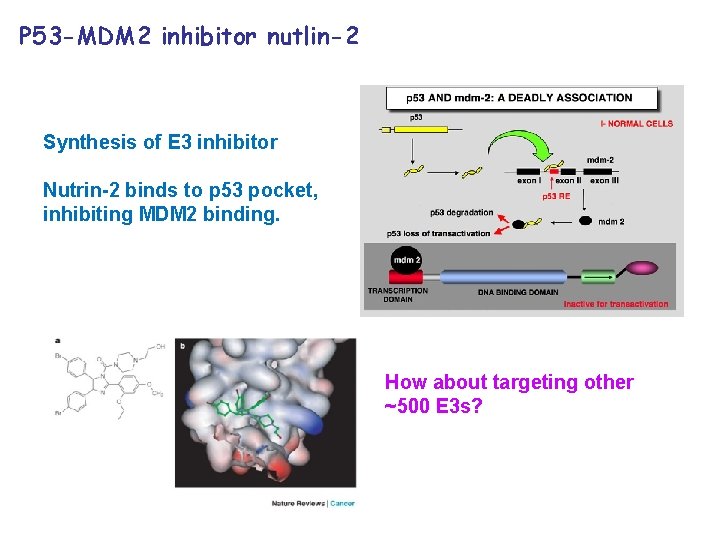
P 53 -MDM 2 inhibitor nutlin-2 Synthesis of E 3 inhibitor Nutrin-2 binds to p 53 pocket, inhibiting MDM 2 binding. How about targeting other ~500 E 3 s?
- Slides: 47