METABOLISM OF PURINES PURINES NH 2 O N

METABOLISM OF PURINES
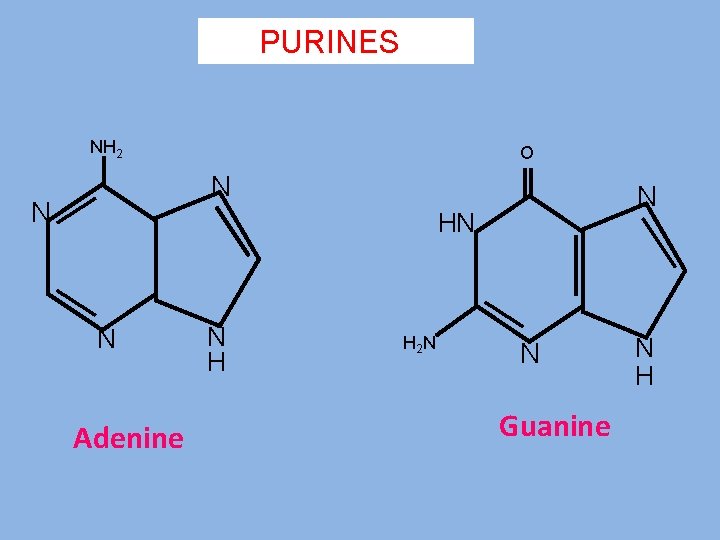
PURINES NH 2 O N N N HN N Adenine N H H 2 N N Guanine N H
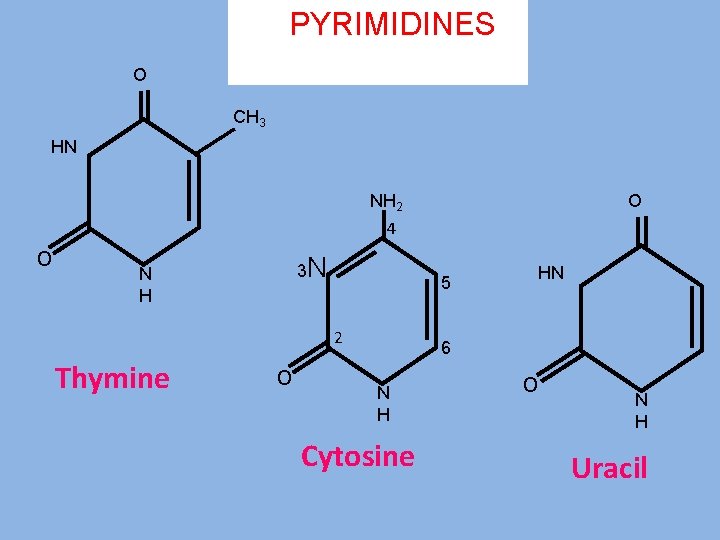
PYRIMIDINES O CH 3 HN NH 2 O 4 o N H 3 N 5 2 Thymine o HN 6 N H Cytosine o N H Uracil
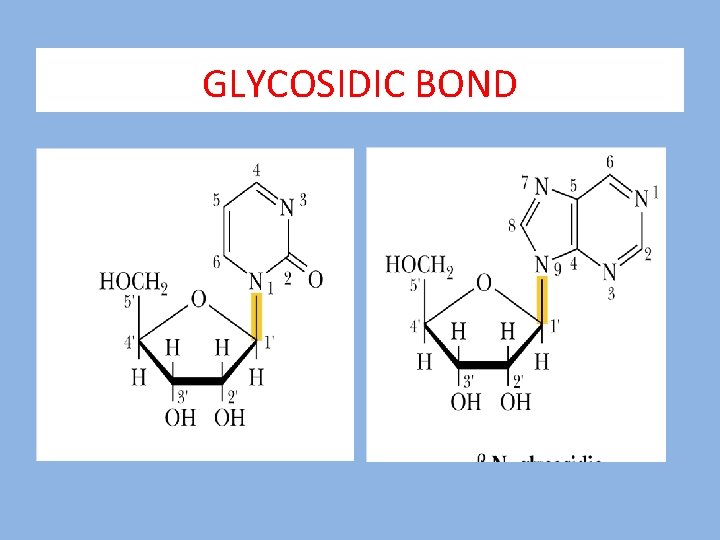
GLYCOSIDIC BOND
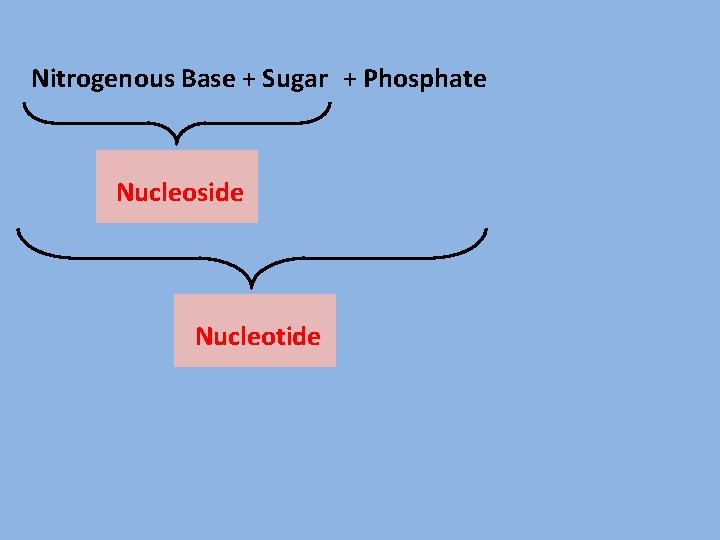
Nitrogenous Base + Sugar + Phosphate Nucleoside Nucleotide
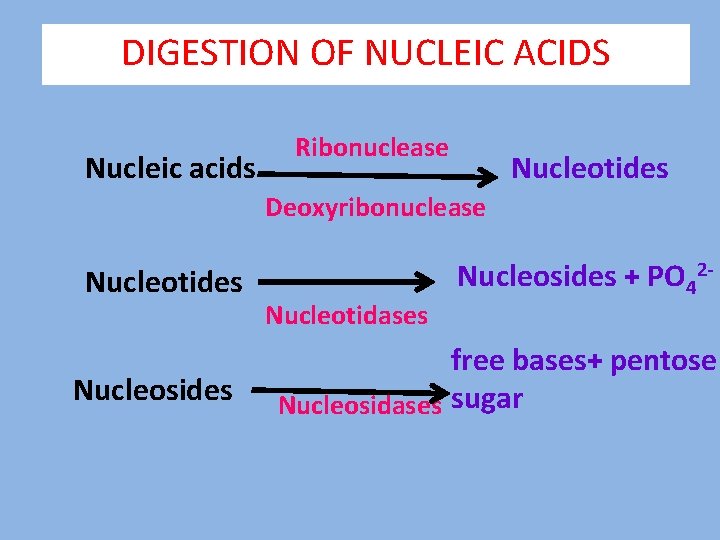
DIGESTION OF NUCLEIC ACIDS Nucleic acids Ribonuclease Nucleotides Deoxyribonuclease Nucleotides Nucleosides Nucleotidases Nucleosides + PO 42 - free bases+ pentose Nucleosidases sugar

BIOSYNTHESIS OF PURINE NUCLEOTIDES • Liver is the major site. • Two pathways: 1. De novo synthesis 2. Salvage pathway • Operates in the Cytoplasm

De Novo Synthesis Of Purines Purine ring synthesis from different small components (amphibolic intermediates)
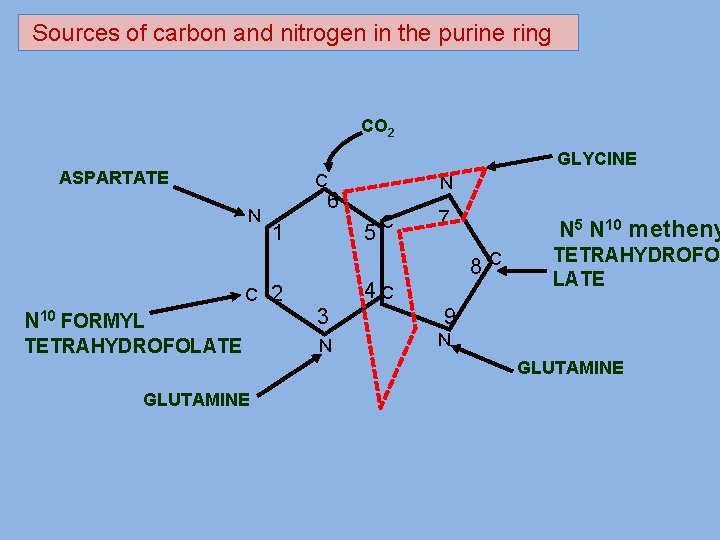
Sources of carbon and nitrogen in the purine ring CO 2 GLYCINE ASPARTATE C N N 6 5 1 C 2 C 7 N 5 N 10 metheny 8 C 4 C N 10 FORMYL 3 9 TETRAHYDROFOLATE N N TETRAHYDROFO LATE GLUTAMINE

De Novo Synthesis Of Purines • IMP formation • Conversion of IMP to AMP and GMP • Formation of nucleoside diphosphates and triphosphates
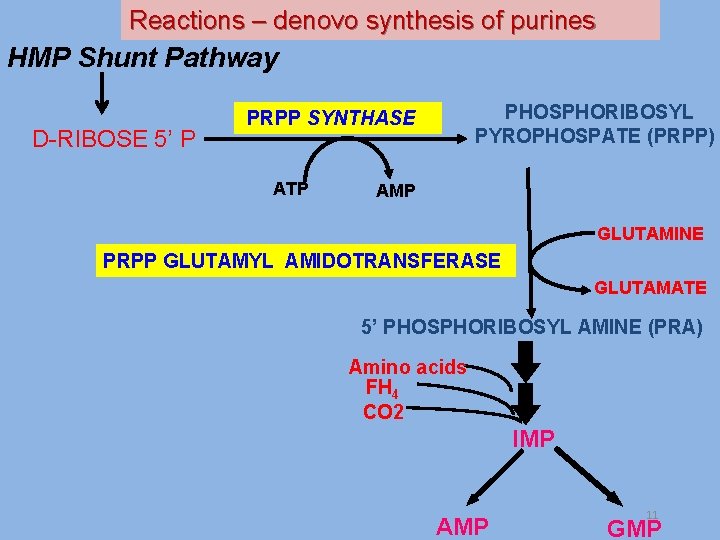
Reactions – denovo synthesis of purines HMP Shunt Pathway D-RIBOSE 5’ P PHOSPHORIBOSYL PYROPHOSPATE (PRPP) PRPP SYNTHASE ATP AMP GLUTAMINE PRPP GLUTAMYL AMIDOTRANSFERASE GLUTAMATE 5’ PHOSPHORIBOSYL AMINE (PRA) Amino acids FH 4 CO 2 IMP AMP 11 GMP
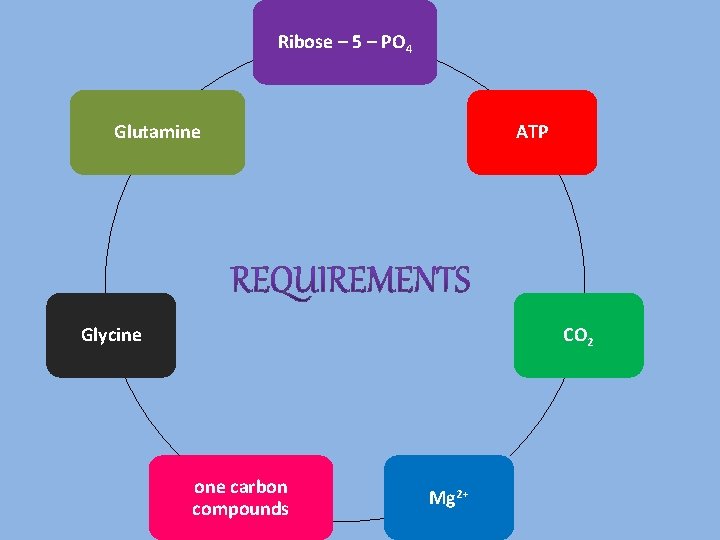
Ribose – 5 – PO 4 Glutamine ATP Glycine CO 2 one carbon compounds Mg 2+
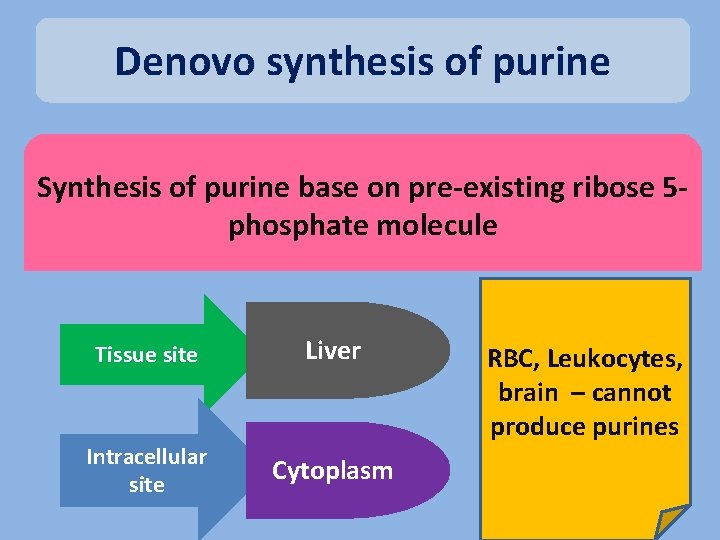
Denovo synthesis of purine Synthesis of purine base on pre-existing ribose 5 phosphate molecule Tissue site Liver Intracellular site Cytoplasm RBC, Leukocytes, brain – cannot produce purines

q Purines & Pyrimidines are dietarily nonessential. q Humans synthesize ample amounts of purines & pyrimidines nucleotides from amphibolic intermediates (denovo). q Ingested nucleic acids are degraded to purine & pyrimidines. q Little or no dietary purine & pyrimidines is incorporated into nucleic acids, parenterally administered compounds are incorporated. q However, injected purine & pyrimidines analogs may be incorporated into DNA.
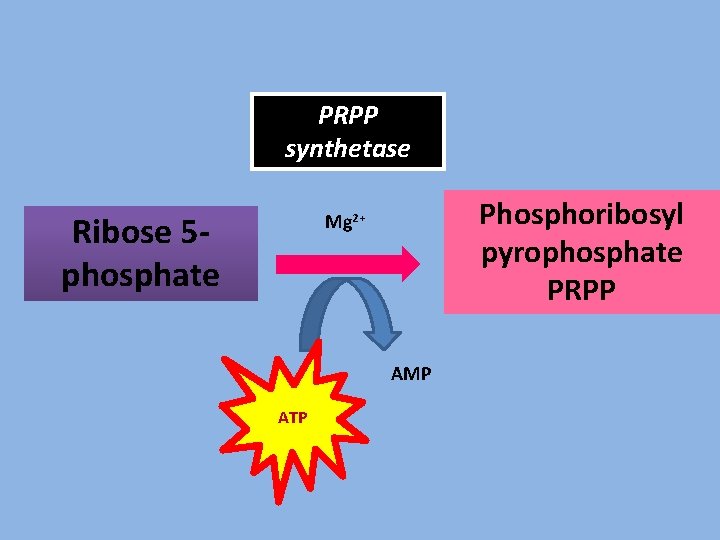
PRPP synthetase Ribose 5 phosphate Phosphoribosyl pyrophosphate PRPP Mg 2+ AMP ATP
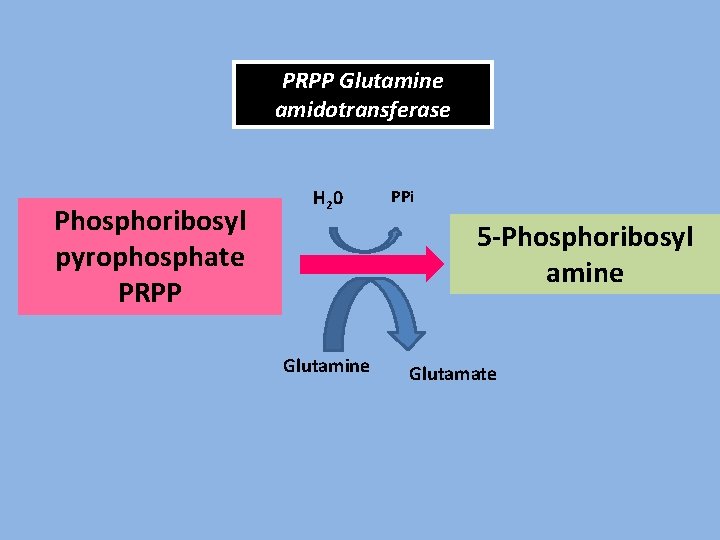
PRPP Glutamine amidotransferase Phosphoribosyl pyrophosphate PRPP H 2 0 PPi 5 -Phosphoribosyl amine Glutamate
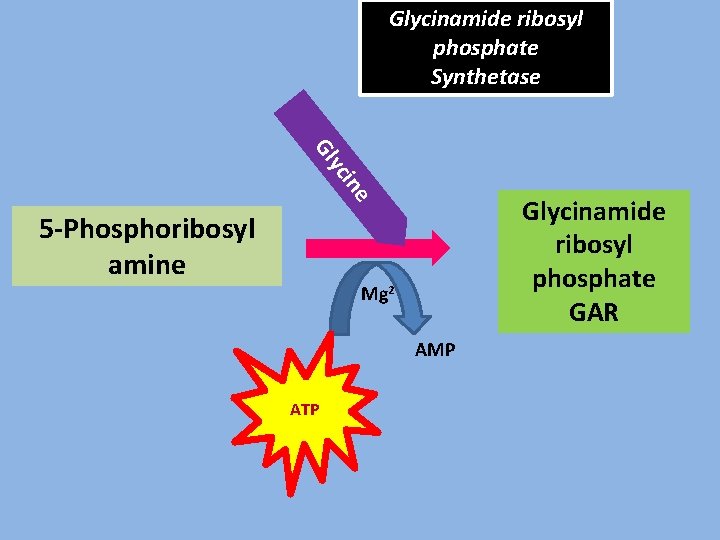
Glycinamide ribosyl phosphate Synthetase ine yc Gl 5 -Phosphoribosyl amine Glycinamide ribosyl phosphate GAR Mg 2+ AMP ATP
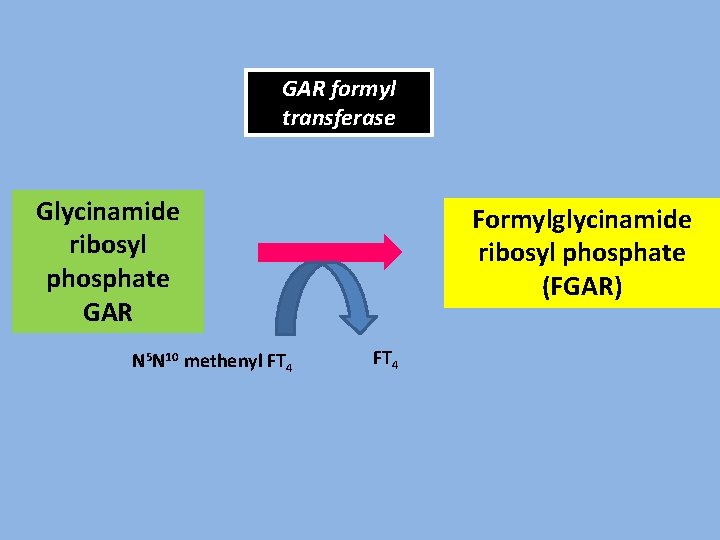
GAR formyl transferase Glycinamide ribosyl phosphate GAR N 5 N 10 methenyl FT 4 Formylglycinamide ribosyl phosphate (FGAR) FT 4
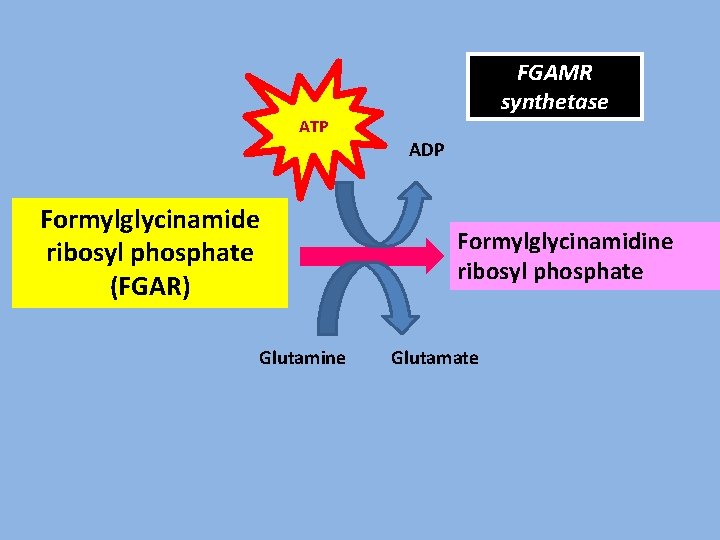
FGAMR synthetase ATP ADP Formylglycinamide ribosyl phosphate (FGAR) Glutamine Formylglycinamidine ribosyl phosphate Glutamate
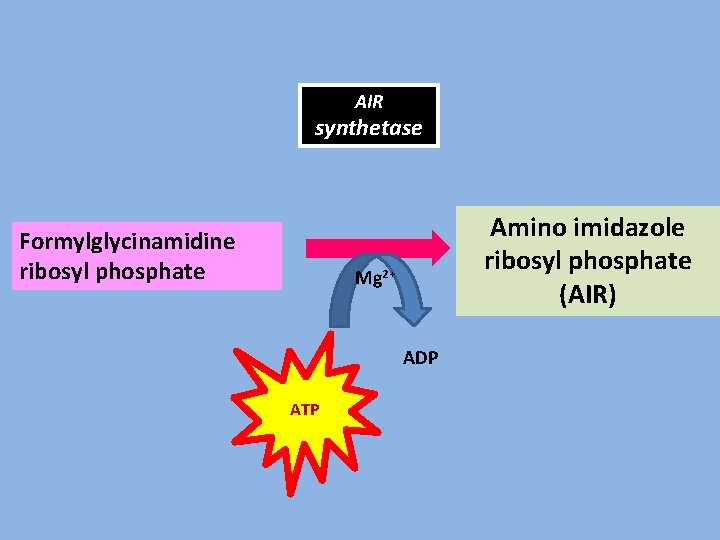
AIR synthetase Formylglycinamidine ribosyl phosphate Amino imidazole ribosyl phosphate (AIR) Mg 2+ ADP ATP
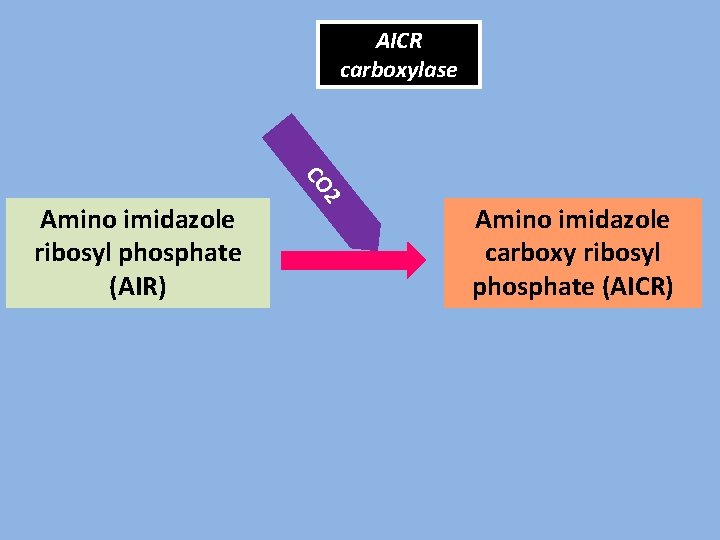
AICR carboxylase 2 CO Amino imidazole ribosyl phosphate (AIR) Amino imidazole carboxy ribosyl phosphate (AICR)
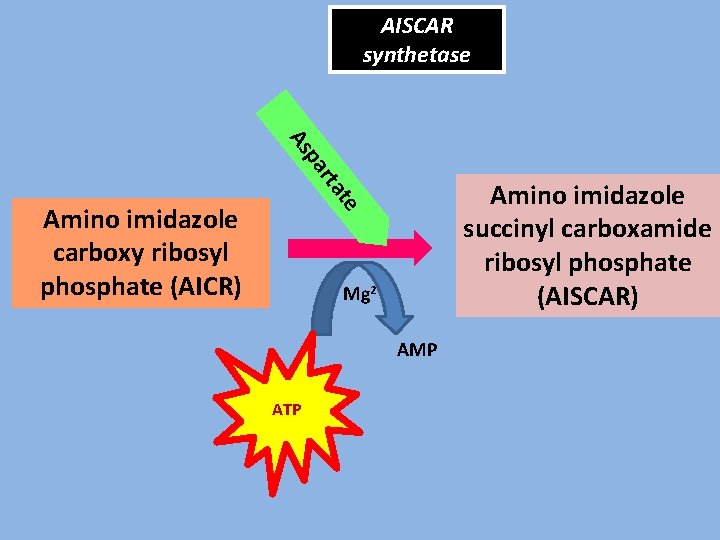
AISCAR synthetase rta pa As te Amino imidazole succinyl carboxamide ribosyl phosphate (AISCAR) Amino imidazole carboxy ribosyl phosphate (AICR) Mg 2+ AMP ATP
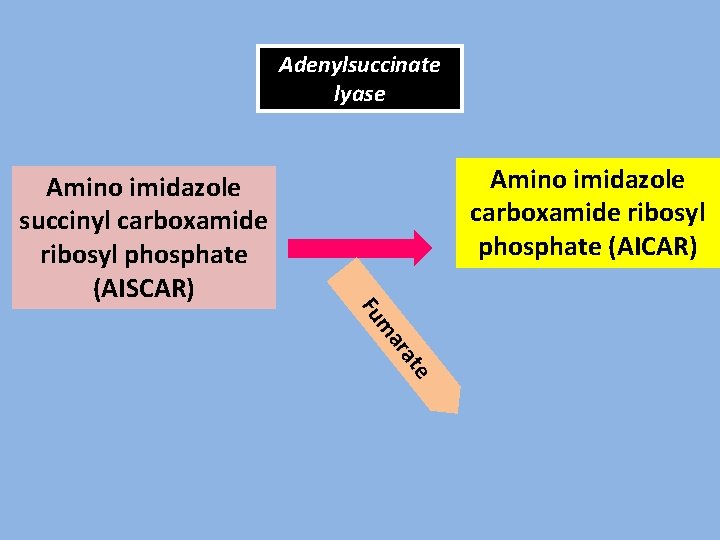
Adenylsuccinate lyase Amino imidazole carboxamide ribosyl phosphate (AICAR) Amino imidazole succinyl carboxamide ribosyl phosphate (AISCAR) e at ar m Fu
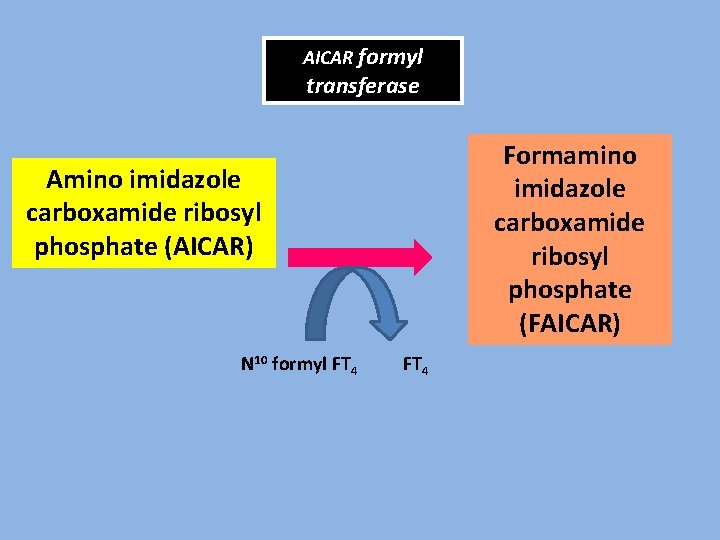
AICAR formyl transferase Formamino imidazole carboxamide ribosyl phosphate (FAICAR) Amino imidazole carboxamide ribosyl phosphate (AICAR) N 10 formyl FT 4
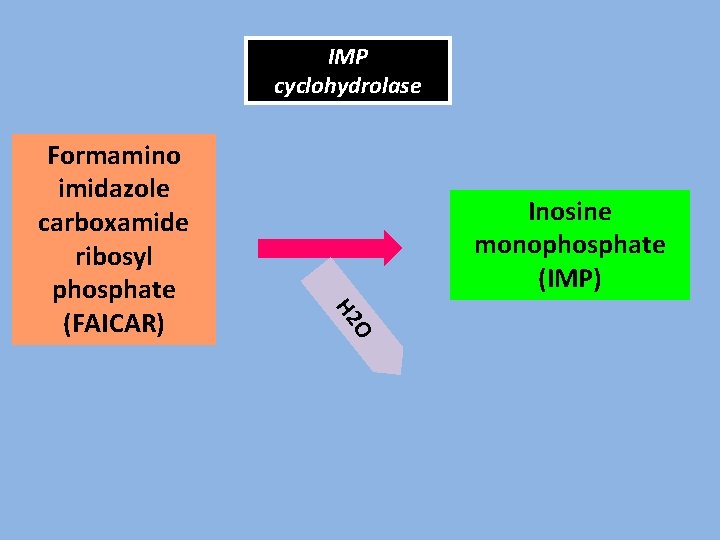
IMP cyclohydrolase Inosine monophosphate (IMP) O H 2 Formamino imidazole carboxamide ribosyl phosphate (FAICAR)
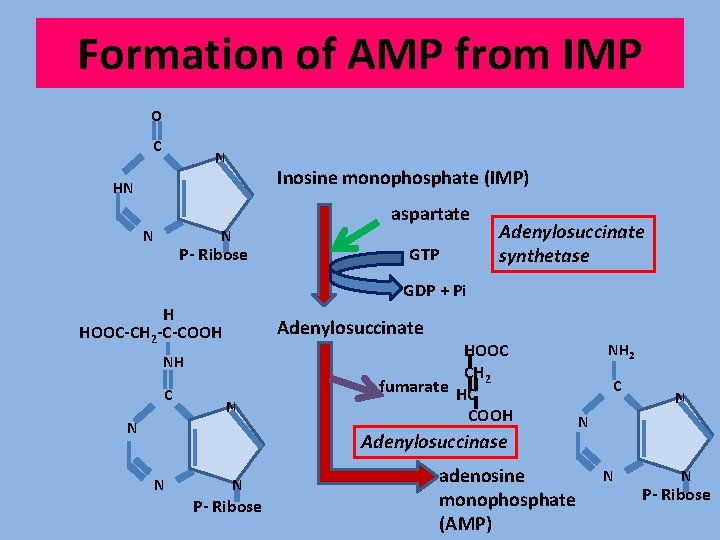
Formation of AMP from IMP O C N Inosine monophosphate (IMP) HN N aspartate N P- Ribose GTP Adenylosuccinate synthetase GDP + Pi H HOOC-CH 2 -C-COOH Adenylosuccinate NH C N N HOOC CH 2 fumarate HC COOH Adenylosuccinase N N P- Ribose adenosine monophosphate (AMP) NH 2 C N N P- Ribose
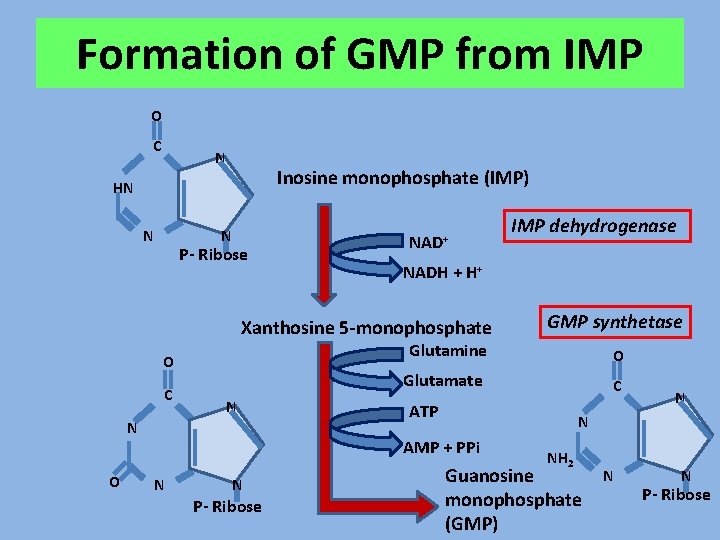
Formation of GMP from IMP O C N Inosine monophosphate (IMP) HN N N P- Ribose NAD+ NADH + H+ Xanthosine 5 -monophosphate N N P- Ribose C ATP N N AMP + PPi N O Glutamate N O GMP synthetase Glutamine O C IMP dehydrogenase NH 2 Guanosine monophosphate (GMP) N N P- Ribose
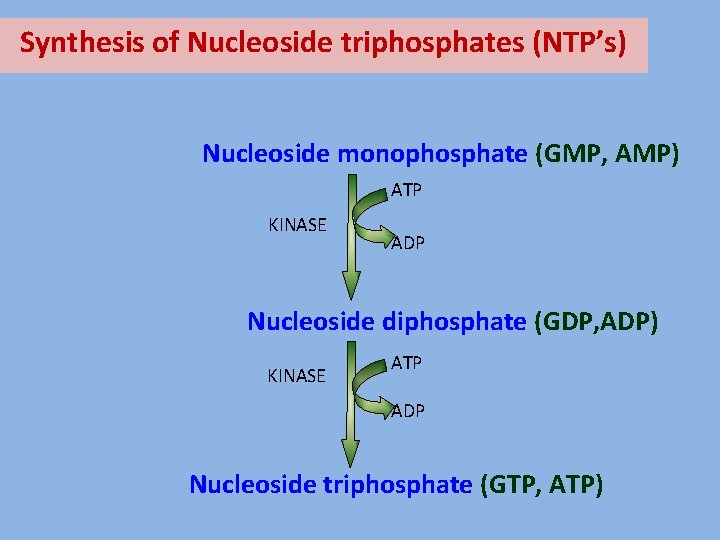
Synthesis of Nucleoside triphosphates (NTP’s) Nucleoside monophosphate (GMP, AMP) ATP KINASE ADP Nucleoside diphosphate (GDP, ADP) KINASE ATP ADP Nucleoside triphosphate (GTP, ATP)

Regulation • Availability of PRPP • PRPP synthetase – Feedback inhibited by purine 5 -nucleotides – Inhibited by AMP, GMP & to smaller extent by ADP, GTP • Glutamine PRPP amidotransferase – Rate limiting step – Allosteric inhibition by AMP at one inhibitory site and GMP at another inhibitory site – Allosteric activation by PRPP

Regulation • Adenylosuccinate synthetase – Stimulated by GTP – Excess GTP leads to increased production of AMP – Feedback inhibited by AMP • GMP synthetase – Stimulated by ATP – Excess ATP leads to production of GMP • IMP dehydrogenase – Feedback inhibited by GMP
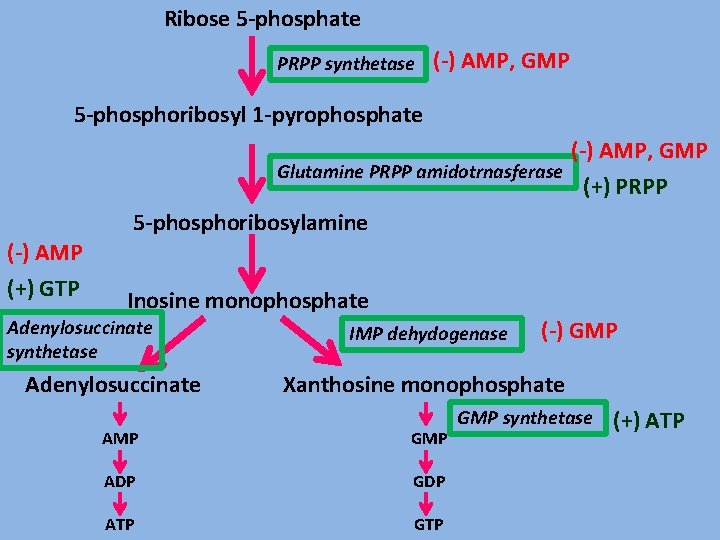
Ribose 5 -phosphate PRPP synthetase (-) AMP, GMP 5 -phosphoribosyl 1 -pyrophosphate (-) AMP, GMP Glutamine PRPP amidotrnasferase (+) PRPP 5 -phosphoribosylamine (-) AMP (+) GTP Inosine monophosphate Adenylosuccinate synthetase Adenylosuccinate IMP dehydogenase (-) GMP Xanthosine monophosphate AMP GMP ADP GDP ATP GMP synthetase (+) ATP

Denovo synthesis of purines

SALVAGE PATHWAY: . Phosphoribosylation of purines and Phosphorylation of purine nucleosides • Recycling of purines formed by degradation of nucleotides. • Enzymes: APRTase, HGPRTase and kinases • Important in RBCs and brain • Less energy consumption

SALVAGE PATHWAY: . 1. Salvage Reactions by phosphoribosylation of purine bases 2. Salvage Reactions by phosphorylation of purine nucleosides

1. Salvage Reactions by phosphoribosylation of purine bases Adenine -Adenine phosphoribosyl transferase Hypoxanthine –guanine phosphoribosyl transferase Guanine
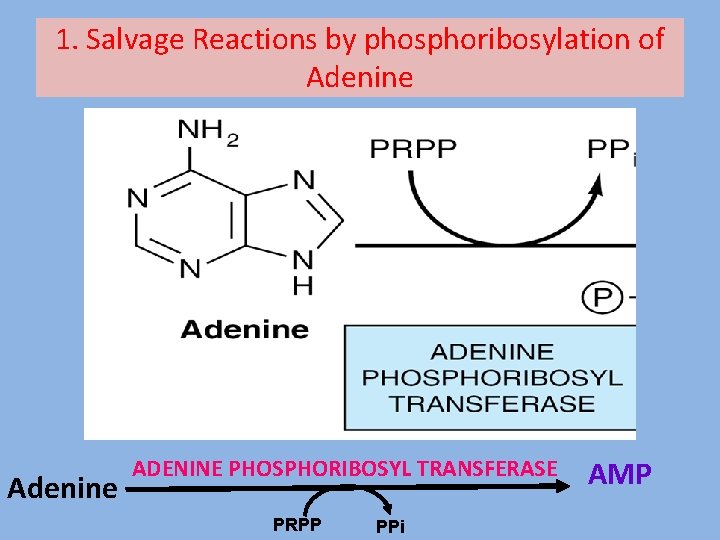
1. Salvage Reactions by phosphoribosylation of Adenine ADENINE PHOSPHORIBOSYL TRANSFERASE PRPP PPi AMP
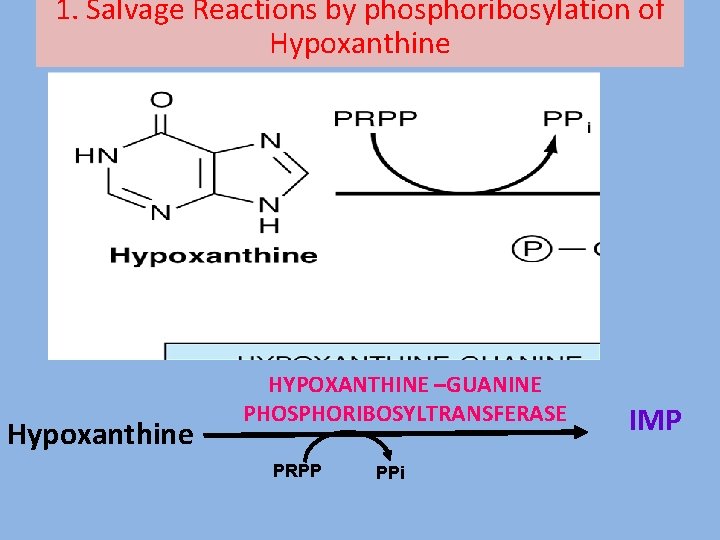
1. Salvage Reactions by phosphoribosylation of Hypoxanthine HYPOXANTHINE –GUANINE PHOSPHORIBOSYLTRANSFERASE PRPP PPi IMP
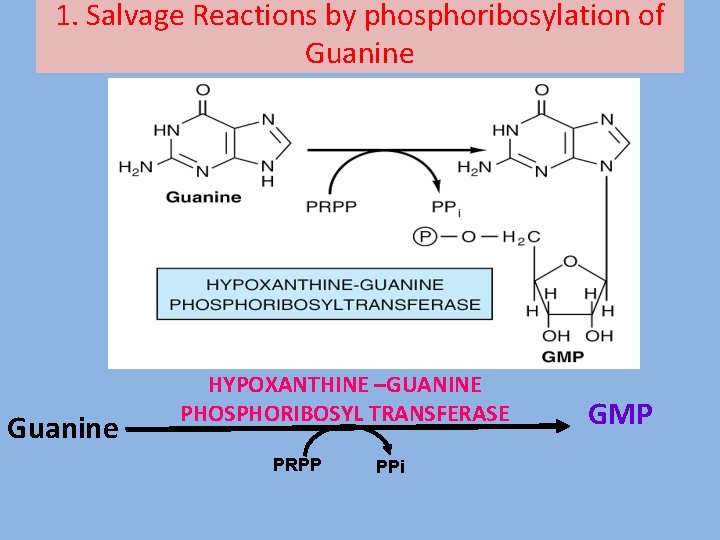
1. Salvage Reactions by phosphoribosylation of Guanine HYPOXANTHINE –GUANINE PHOSPHORIBOSYL TRANSFERASE PRPP PPi GMP
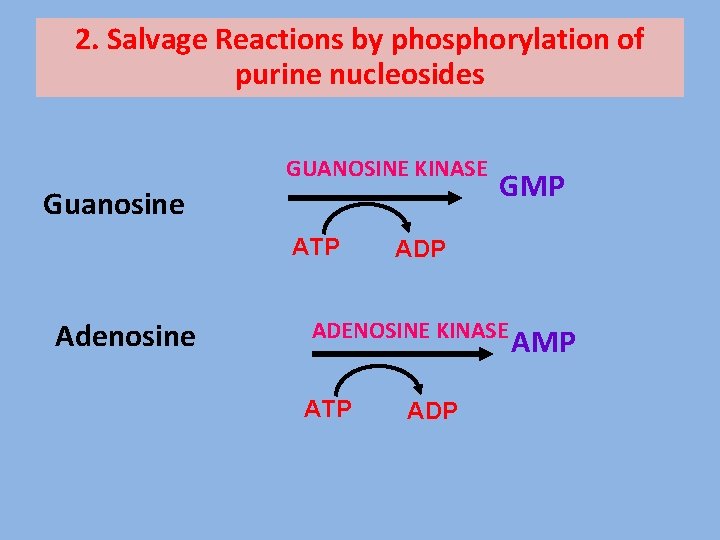
2. Salvage Reactions by phosphorylation of purine nucleosides GUANOSINE KINASE Guanosine ATP Adenosine GMP ADENOSINE KINASE ATP ADP AMP
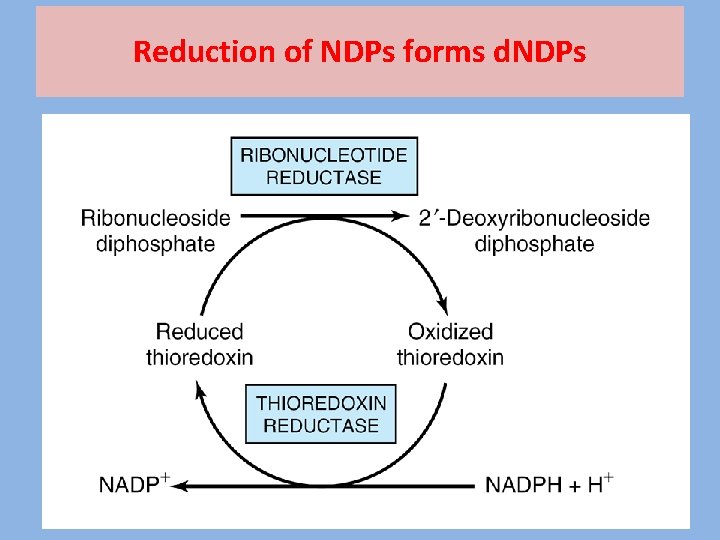
Reduction of NDPs forms d. NDPs

Regulation of Purine nucleotide synthesis PRPP pool size - ribose 5 -phsophate and PRPP synthase. PRPP glutamyl amidotransferse Hypoxanthine guanine phosphoribosyl transferase
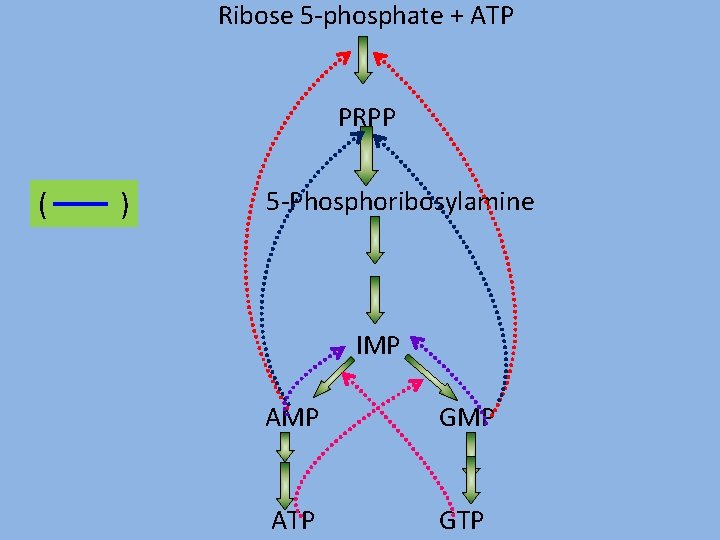
Ribose 5 -phosphate + ATP PRPP ( ) 5 -Phosphoribosylamine IMP AMP GMP ATP GTP
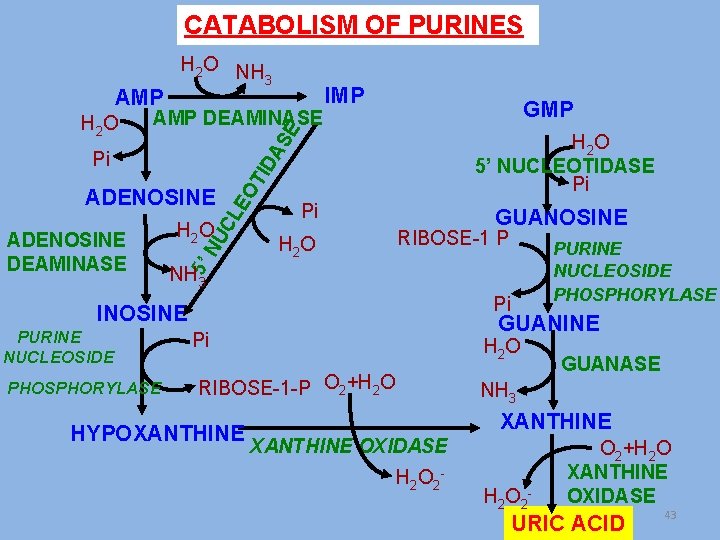
CATABOLISM OF PURINES AMP DEAMINASE GMP H 2 O 5’ NUCLEOTIDASE Pi ADENOSINE Pi H 2 O GUANOSINE RIBOSE-1 P NH 3 Pi INOSINE PURINE NUCLEOSIDE PHOSPHORYLASE 5’ NU C H 2 O LE OT ID A Pi ADENOSINE DEAMINASE IMP SE H 2 O H 2 O NH 3 GUANINE Pi H 2 O RIBOSE-1 -P O 2+H 2 O HYPOXANTHINE GUANASE NH 3 XANTHINE OXIDASE H 2 O 2 - O 2+H 2 O XANTHINE OXIDASE URIC ACID 43

URIC ACID • Total body urate pool : 800 to 1600 mg in ♂ 500 to 700 mg in ♀ • Turnover rate : 600 mg / 24 hrs. • Normal uric acid level : 2 - 5 mg / d. L in ♀ 3 - 7 mg / d. L in ♂

DISORDERS OF PURINE METABOLISM Hyperuricemia • Gout • Lesch – Nyhan syndrome Hypouricemia • Immunodeficiency(Adenosine deaminase deficiency)

Hyperuricemia Uric acid level > 7 mg / dl in males and > 6 mg / dl in females GOUT Metabolic disease associated with hyperuricemia

GOUT Types v. Primary gout v. Secondary gout

PRIMARY GOUT v Glucose-6 -phosphatase deficiency. v. Increased activity of Glutathione reductase v. Abnormality in PRPP synthase. v Abnormality in PRPP Glutamyl amidotransferase v Deficiency of enzymes of salvage pathway
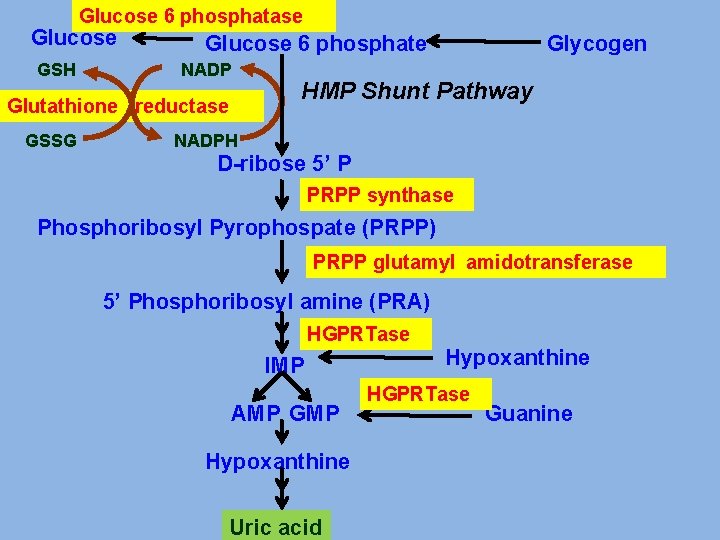
Glucose 6 phosphatase Glucose GSH Glucose 6 phosphate NADP Glutathione reductase GSSG Glycogen HMP Shunt Pathway NADPH D-ribose 5’ P PRPP synthase Phosphoribosyl Pyrophospate (PRPP) PRPP glutamyl amidotransferase 5’ Phosphoribosyl amine (PRA) HGPRTase IMP AMP GMP Hypoxanthine Uric acid Hypoxanthine HGPRTase Guanine
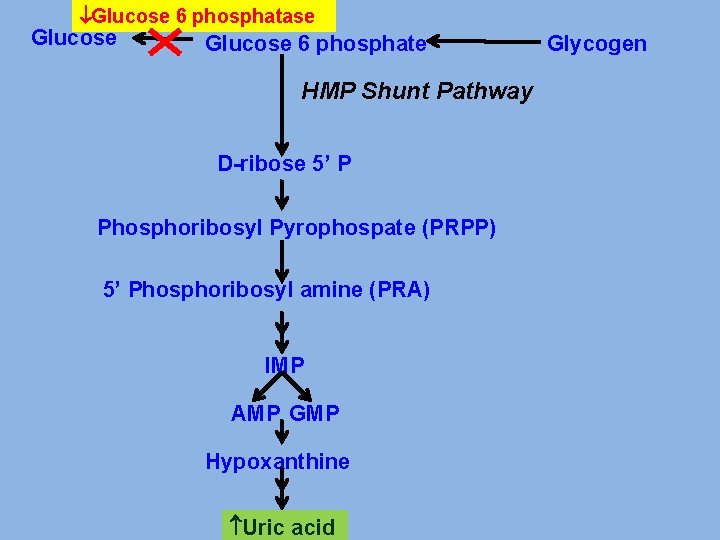
Glucose 6 phosphatase Glucose 6 phosphate HMP Shunt Pathway D-ribose 5’ P Phosphoribosyl Pyrophospate (PRPP) 5’ Phosphoribosyl amine (PRA) IMP AMP GMP Hypoxanthine Uric acid Glycogen
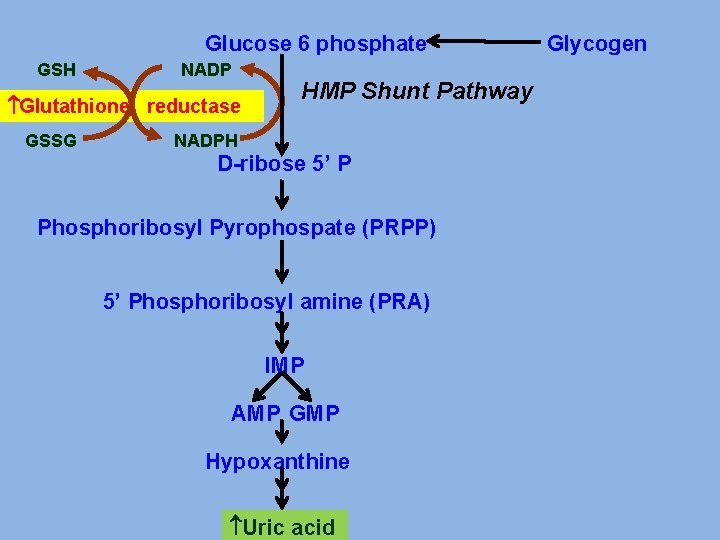
Glucose 6 phosphate GSH NADP Glutathione reductase GSSG HMP Shunt Pathway NADPH D-ribose 5’ P Phosphoribosyl Pyrophospate (PRPP) 5’ Phosphoribosyl amine (PRA) IMP AMP GMP Hypoxanthine Uric acid Glycogen
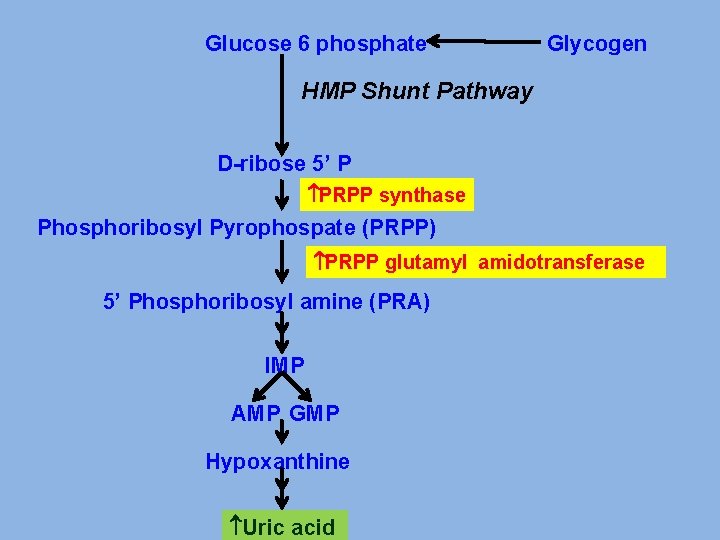
Glucose 6 phosphate Glycogen HMP Shunt Pathway D-ribose 5’ P PRPP synthase Phosphoribosyl Pyrophospate (PRPP) PRPP glutamyl amidotransferase 5’ Phosphoribosyl amine (PRA) IMP AMP GMP Hypoxanthine Uric acid
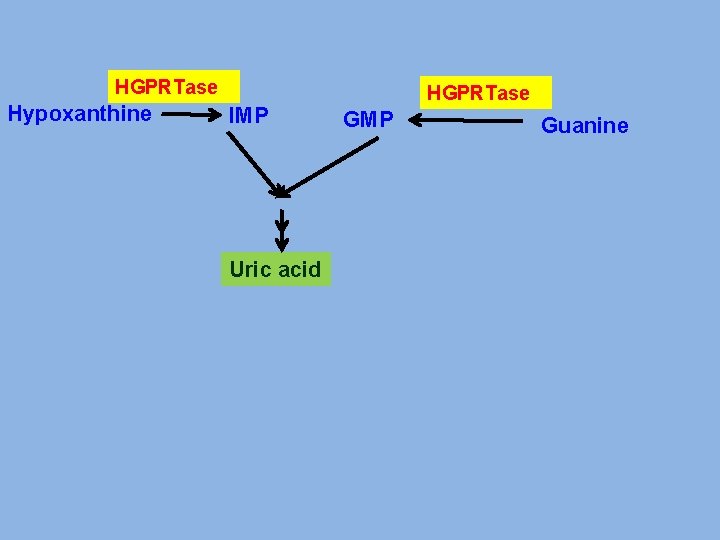
HGPRTase Hypoxanthine IMP Uric acid HGPRTase GMP Guanine
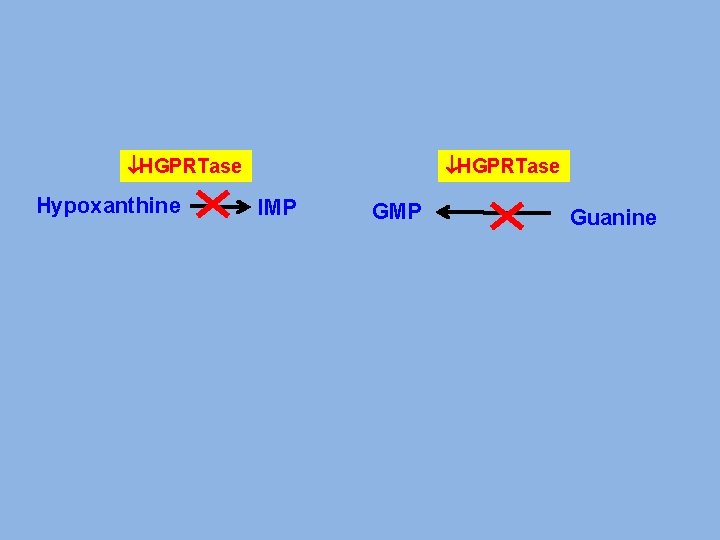
HGPRTase Hypoxanthine HGPRTase IMP Guanine
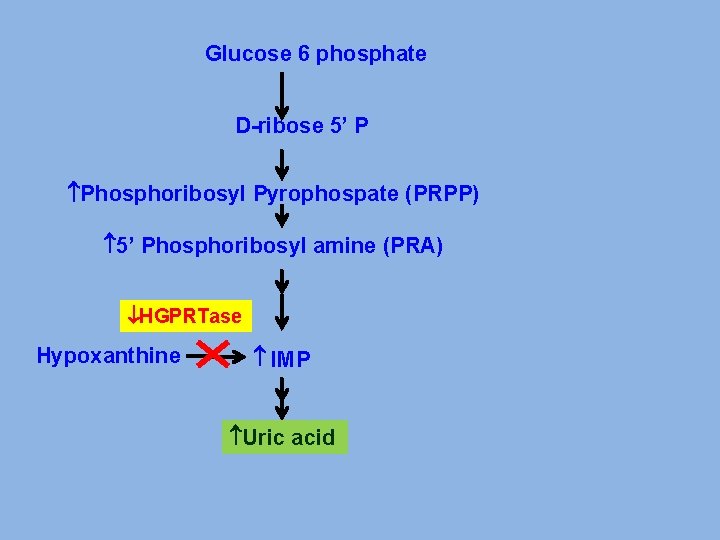
Glucose 6 phosphate D-ribose 5’ P Phosphoribosyl Pyrophospate (PRPP) 5’ Phosphoribosyl amine (PRA) HGPRTase Hypoxanthine IMP Uric acid
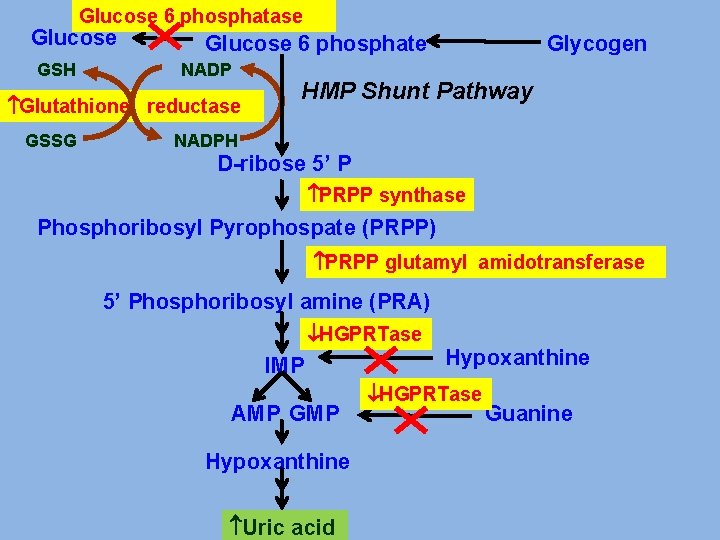
Glucose 6 phosphatase Glucose GSH Glucose 6 phosphate NADP Glutathione reductase GSSG Glycogen HMP Shunt Pathway NADPH D-ribose 5’ P PRPP synthase Phosphoribosyl Pyrophospate (PRPP) PRPP glutamyl amidotransferase 5’ Phosphoribosyl amine (PRA) HGPRTase IMP AMP GMP Hypoxanthine Uric acid Hypoxanthine HGPRTase Guanine

SECONDARY GOUT • Increased turnover rate of nucleic acids. - Leukemias, polycythemia. - Treatment of Malignant tumors. - Psoriasis. • Reduced excretion rate. - Renal failure. - Thiazide diuretics. - Lactic acidosis and Ketoacidosis.
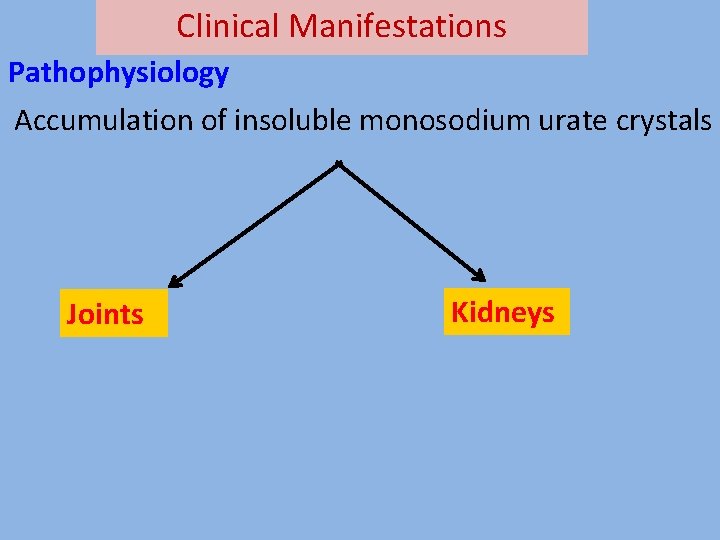
Clinical Manifestations Pathophysiology Accumulation of insoluble monosodium urate crystals Joints Kidneys
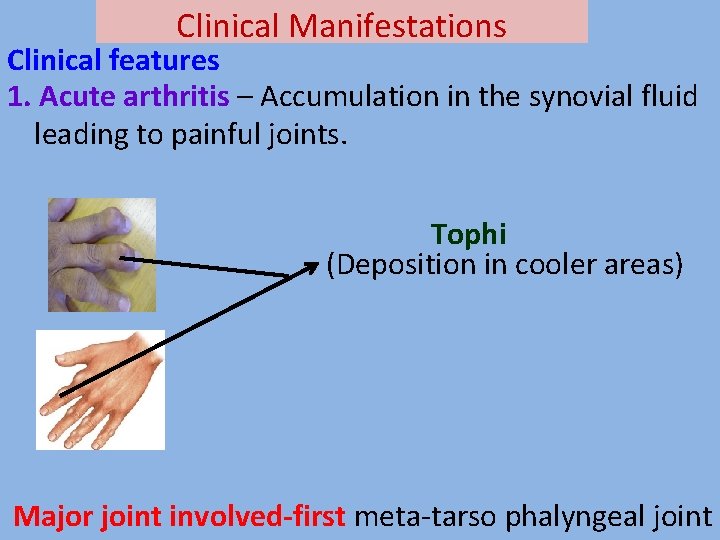
Clinical Manifestations Clinical features 1. Acute arthritis – Accumulation in the synovial fluid leading to painful joints. Tophi (Deposition in cooler areas) Major joint involved-first meta-tarso phalyngeal joint

Clinical Manifestations Clinical features 1. Acute arthritis Precipitating factors- Alcohol Lactic acidosis Decreases renal excretion of uric acid Increased serum uric acid levels 2. Renal lithiasis– Precipitation of insoluble sodium urate crystals in the kidneys and ureter leading to stone formation.

Treatment 1. Allopurinol 2. Probenecid 3. Restrict alcohol
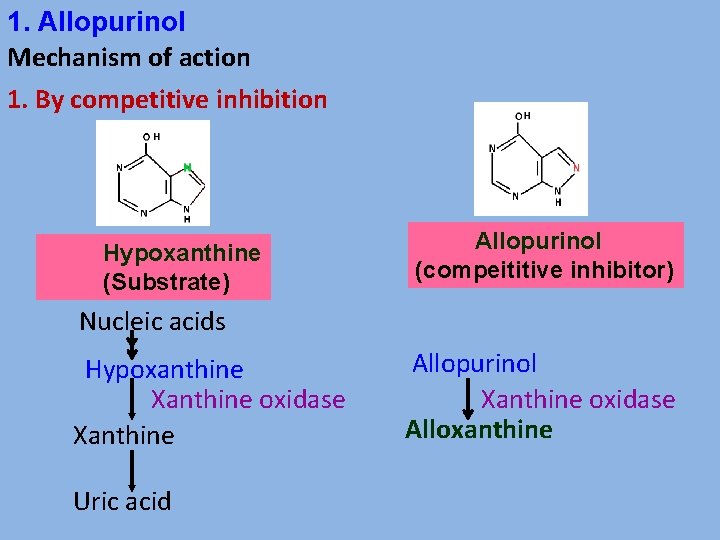
1. Allopurinol Mechanism of action 1. By competitive inhibition Hypoxanthine (Substrate) Allopurinol (compeititive inhibitor) Nucleic acids Hypoxanthine Xanthine oxidase Xanthine Uric acid Allopurinol Xanthine oxidase Alloxanthine
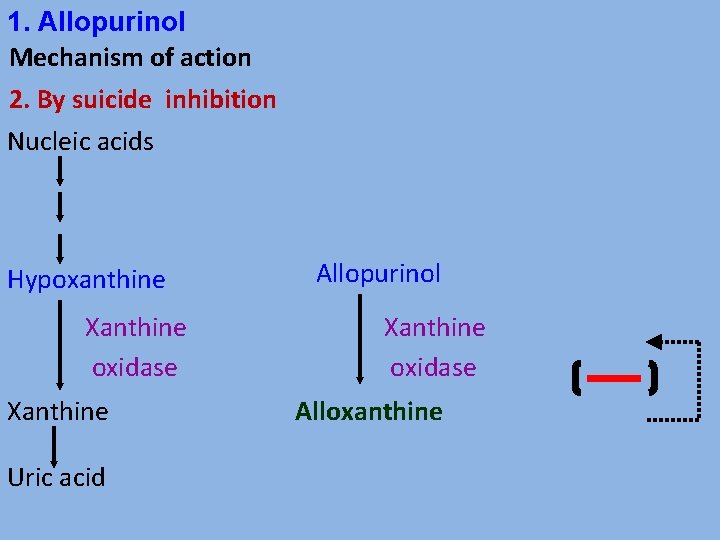
1. Allopurinol Mechanism of action 2. By suicide inhibition Nucleic acids Hypoxanthine Xanthine oxidase Xanthine Uric acid Allopurinol Xanthine oxidase Alloxanthine

2. Probenecid Mechanism of action Probenecid Decreases the reabsorption of uric acid from renal tubules Increased renal excretion of uric acid Decreased serum uric acid levels 3. Restrict alcohol

LESCH NYHAN SYNDROME • ‘ X linked ’ disorder of Purine metabolism. • Total or partial deficiency of HGPRTase • Features: - Self mutilation, mental retardation and ↑ uric acid. - Nephrolithiasis, gout.

HYPOURICEMIA • Adenosine deaminase deficiency. • Congenital xanthine oxidase deficiency • Purine nucleoside phosphorylase deficiency.
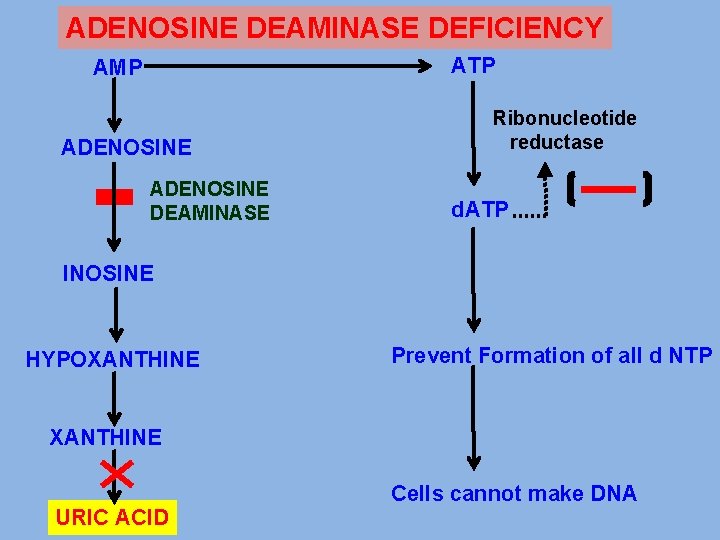
ADENOSINE DEAMINASE DEFICIENCY ATP AMP ADENOSINE DEAMINASE Ribonucleotide reductase d. ATP INOSINE HYPOXANTHINE Prevent Formation of all d NTP XANTHINE URIC ACID Cells cannot make DNA

METABOLISM OF PYRIMIDINES
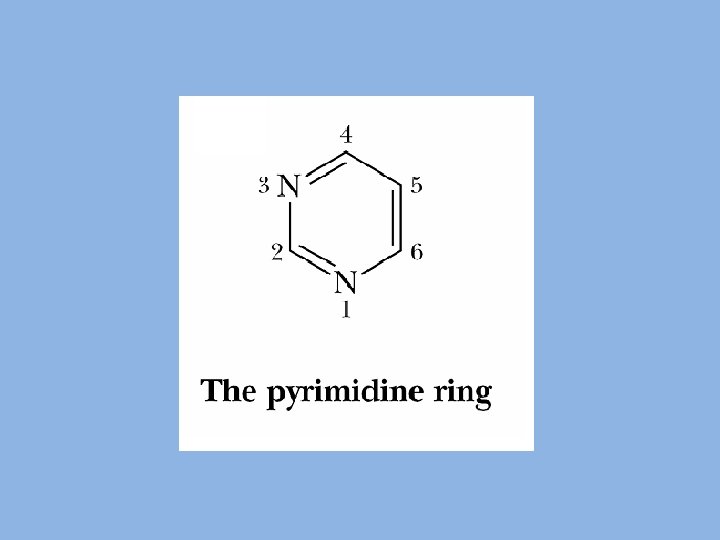

BIOSYNTHESIS OF PYRIMIDINE NUCLEOTIDES • Two pathways: 1. De novo synthesis 2. Salvage pathway • Subcellular site: Cytoplasm

DE NOVO SYNTHESIS Pyrimidine ring synthesis from different small components (amphibolic intermediates) PYRIMIDINE RING IS SYNTHESIZED BEFORE BEING ATTACHED TO RIBOSE 5 -PHOSPHATE ( DONATED BY PRPP)
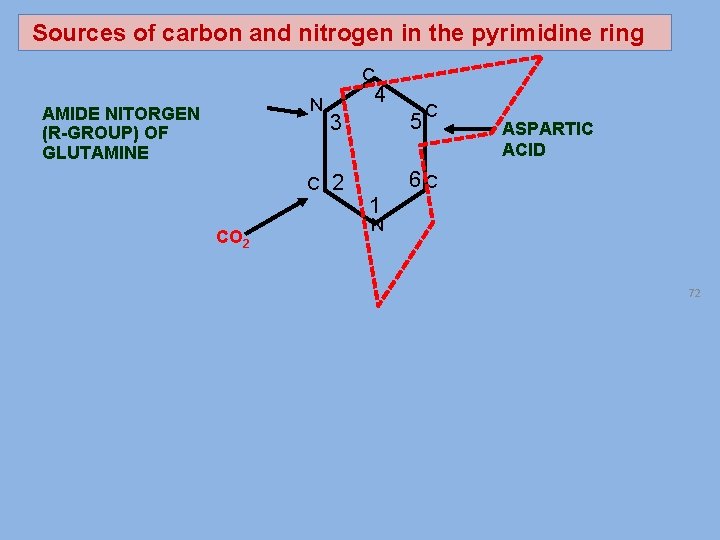
Sources of carbon and nitrogen in the pyrimidine ring C N AMIDE NITORGEN (R-GROUP) OF GLUTAMINE 4 C 2 CO 2 5 3 C ASPARTIC ACID 6 C 1 N 72
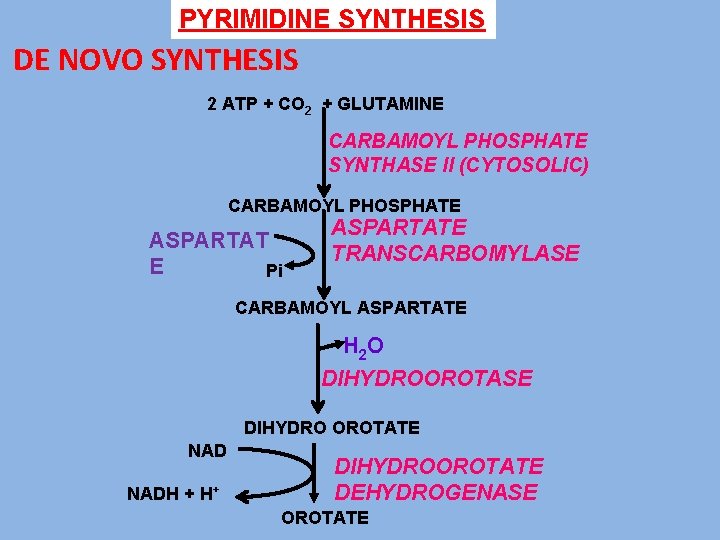
PYRIMIDINE SYNTHESIS DE NOVO SYNTHESIS 2 ATP + CO 2 + GLUTAMINE CARBAMOYL PHOSPHATE SYNTHASE II (CYTOSOLIC) CARBAMOYL PHOSPHATE ASPARTAT E Pi ASPARTATE TRANSCARBOMYLASE CARBAMOYL ASPARTATE H 2 O DIHYDROOROTASE DIHYDRO OROTATE NADH + H+ DIHYDROOROTATE DEHYDROGENASE OROTATE
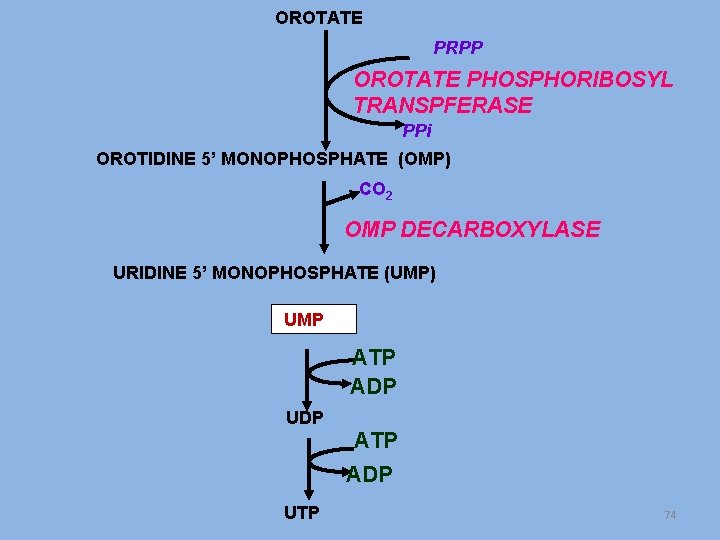
OROTATE PRPP OROTATE PHOSPHORIBOSYL TRANSPFERASE PPi OROTIDINE 5’ MONOPHOSPHATE (OMP) CO 2 OMP DECARBOXYLASE URIDINE 5’ MONOPHOSPHATE (UMP) UMP ATP ADP UDP ATP ADP UTP 74
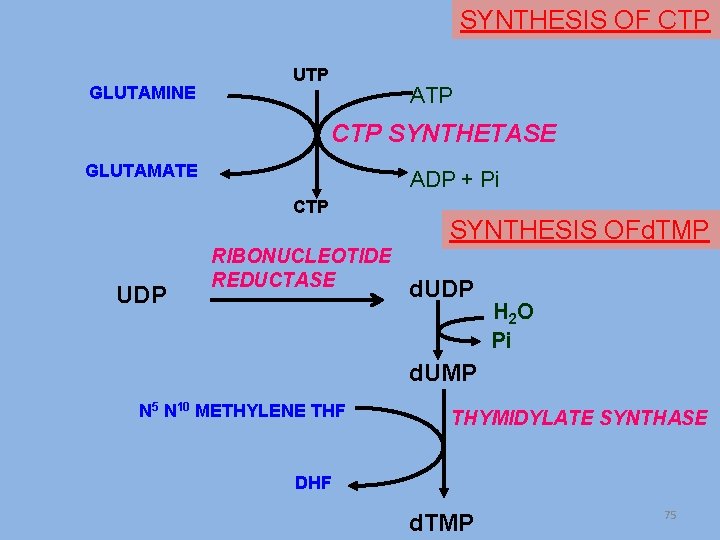
SYNTHESIS OF CTP GLUTAMINE UTP ATP CTP SYNTHETASE GLUTAMATE ADP + Pi CTP UDP RIBONUCLEOTIDE REDUCTASE SYNTHESIS OFd. TMP d. UDP H 2 O Pi d. UMP N 5 N 10 METHYLENE THF THYMIDYLATE SYNTHASE DHF d. TMP 75
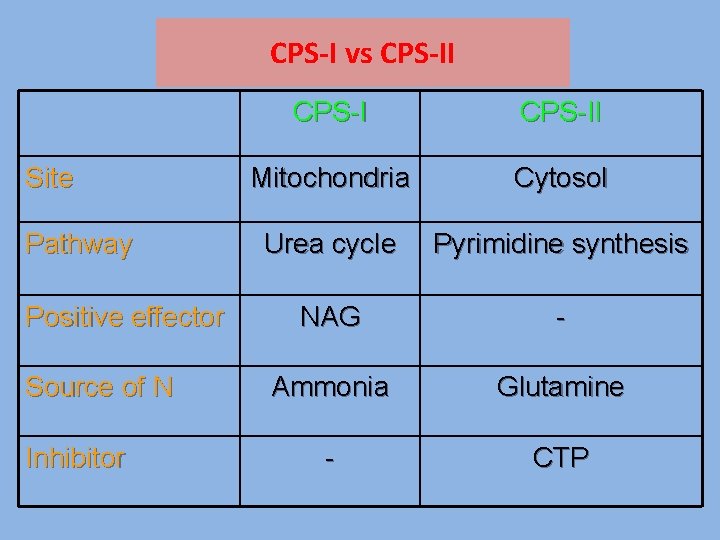
CPS-I vs CPS-II Site Pathway Positive effector Source of N Inhibitor CPS-II Mitochondria Cytosol Urea cycle Pyrimidine synthesis NAG - Ammonia Glutamine - CTP
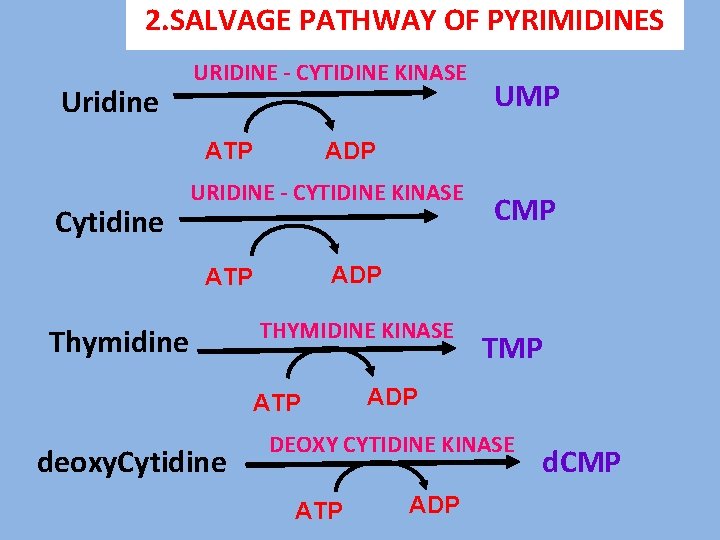
2. SALVAGE PATHWAY OF PYRIMIDINES Uridine URIDINE - CYTIDINE KINASE ATP Cytidine ADP URIDINE - CYTIDINE KINASE THYMIDINE KINASE ATP deoxy. Cytidine CMP ADP ATP Thymidine UMP TMP ADP DEOXY CYTIDINE KINASE ATP ADP d. CMP

REGULATION OF PYRIMIDINE SYNTHESIS • CPS II, ATC and DHOase • OPRTase and OMP decarboxylase • ATC is inhibited by CTP • CPS II is inhibited by UTP • OMP decarboxylase is inhibited by UMP. • CTP synthase requires ATP.

Catabolism OF PYRIMIDINE Catabolism of cytosine and uracil yields alanine, co 2, NH 3 Catabolism of Thymine yields aminoisobutyrate, co 2, NH 3 79

DISORDERS OROTIC ACIDURIA Hereditary disorder due to defect in enzymes of pyrimidine synthesis , resulting in excretion of orotic acid in urine Types Type 1 Type 2
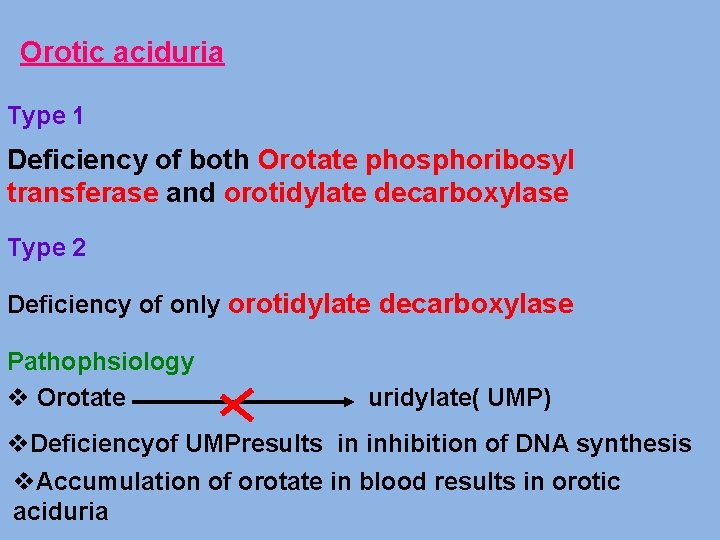
Orotic aciduria Type 1 Deficiency of both Orotate phosphoribosyl transferase and orotidylate decarboxylase Type 2 Deficiency of only orotidylate decarboxylase Pathophsiology v Orotate uridylate( UMP) v. Deficiencyof UMPresults in inhibition of DNA synthesis v. Accumulation of orotate in blood results in orotic aciduria

Clinical Features Growth retardation Megaloblastic anaemia Treatment Diet rich in uridine or cytidine. They may be converted to. UTP which can act as feed back inhibitor.

Inhibitors of Purine and Pyrimidine Nucleotide Synthesis 6 -Mercaptopurine Thioguanine used in the treatment of leukaemia Methotrexate used as anticancer drug Sulfonamides used as antibiotic Azaserine and diazonorleucine are glutamine analog and inhibit the enzymes that utilize glutamine as amino group donor in the biosynthesis of purine nucleotides. 5 -Fluorouracil is an anti cancer drug which inhibits thymidylate synthase enzyme

Thank you

• Adenylosuccinate synthase - AMP (-) - GTP (+) • IMP dehydrogenase - GMP (-) • HGPRTase : AMP and GMP (-)
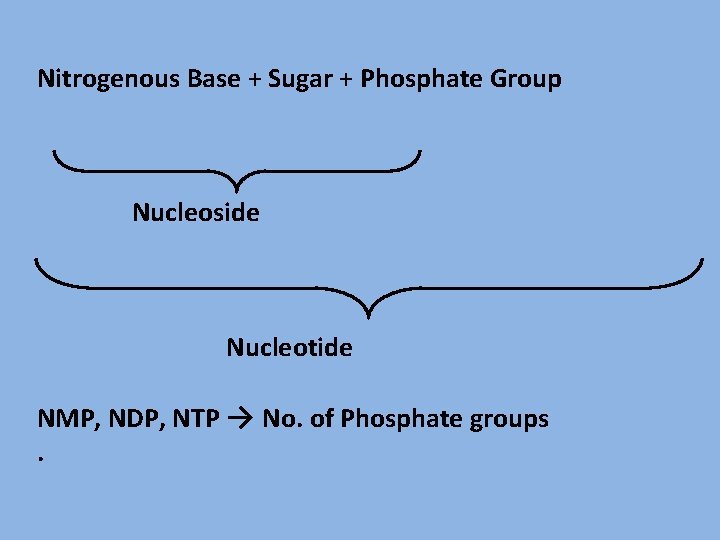
Nitrogenous Base + Sugar + Phosphate Group Nucleoside Nucleotide NMP, NDP, NTP → No. of Phosphate groups.
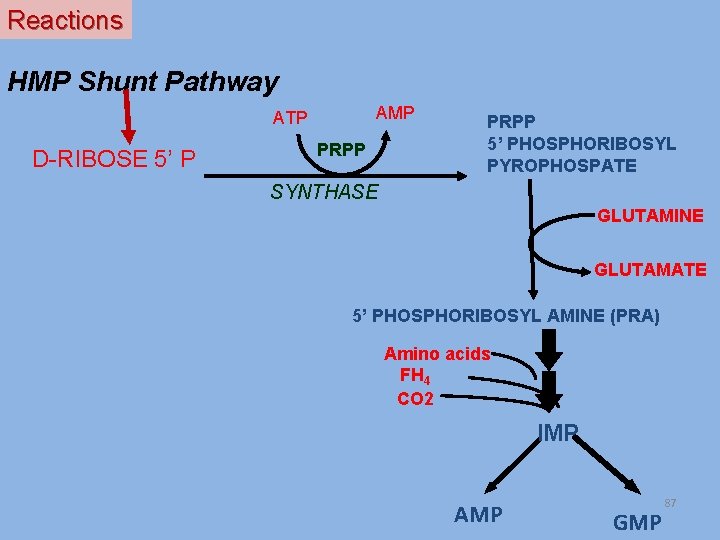
Reactions HMP Shunt Pathway AMP ATP D-RIBOSE 5’ P PRPP 5’ PHOSPHORIBOSYL PYROPHOSPATE SYNTHASE GLUTAMINE GLUTAMATE 5’ PHOSPHORIBOSYL AMINE (PRA) Amino acids FH 4 CO 2 IMP AMP GMP 87
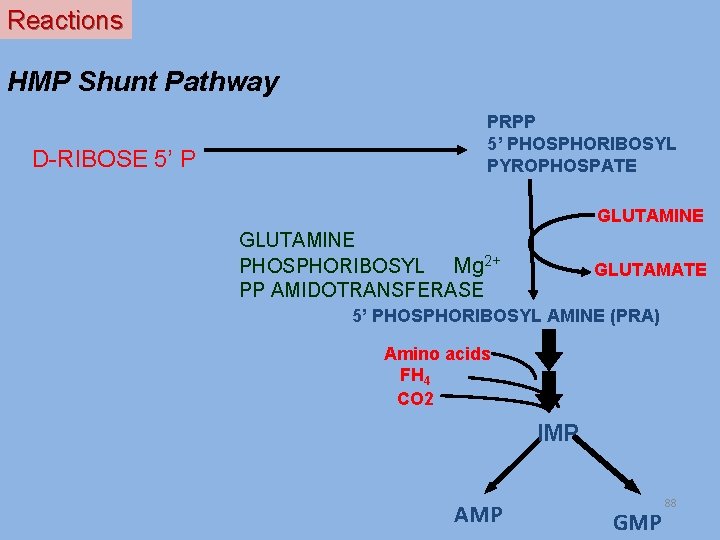
Reactions HMP Shunt Pathway D-RIBOSE 5’ P PRPP 5’ PHOSPHORIBOSYL PYROPHOSPATE GLUTAMINE PHOSPHORIBOSYL Mg 2+ PP AMIDOTRANSFERASE GLUTAMATE 5’ PHOSPHORIBOSYL AMINE (PRA) Amino acids FH 4 CO 2 IMP AMP GMP 88
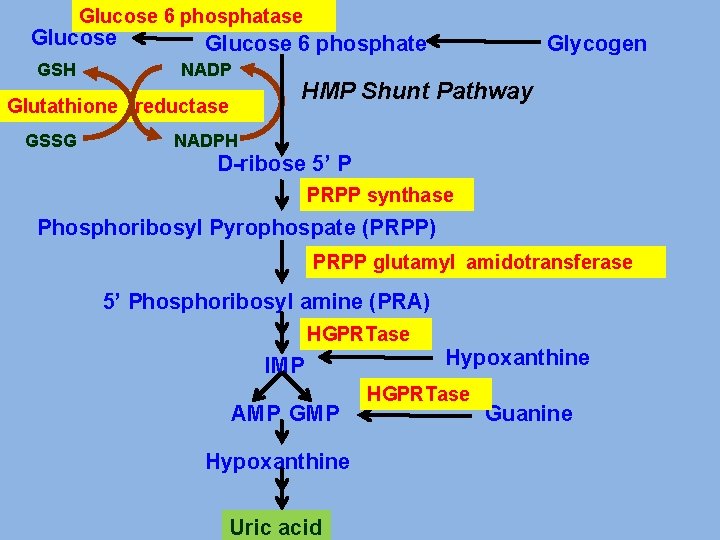
Glucose 6 phosphatase Glucose GSH Glucose 6 phosphate NADP Glutathione reductase GSSG Glycogen HMP Shunt Pathway NADPH D-ribose 5’ P PRPP synthase Phosphoribosyl Pyrophospate (PRPP) PRPP glutamyl amidotransferase 5’ Phosphoribosyl amine (PRA) HGPRTase IMP AMP GMP Hypoxanthine Uric acid Hypoxanthine HGPRTase Guanine
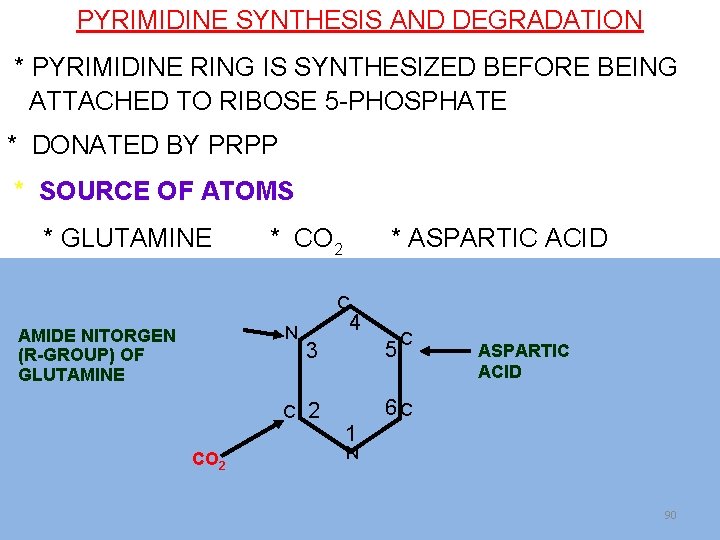
PYRIMIDINE SYNTHESIS AND DEGRADATION * PYRIMIDINE RING IS SYNTHESIZED BEFORE BEING ATTACHED TO RIBOSE 5 -PHOSPHATE * DONATED BY PRPP * SOURCE OF ATOMS * GLUTAMINE * CO 2 * ASPARTIC ACID C N AMIDE NITORGEN (R-GROUP) OF GLUTAMINE 4 C 2 CO 2 5 3 C ASPARTIC ACID 6 C 1 N 90

ADENOSINE DEAMINASE DEFICIENCY • EXPRESSED IN CYTOSOL OF CELLS • HUMAN LYMPHOCYTES HAVE HIGHEST ACTIVITY • DEFICIENCY OF ADA CUASES -ACCUMULATION OF adenosine -CONVERTED TO NTP AND d NTP -AS d ATP RIBONUCLEOTIDE REDUCTASE INHIBITED -PREVENT FORMATION OF ALL d NTP -CELLS CANNOT MAKE DNA
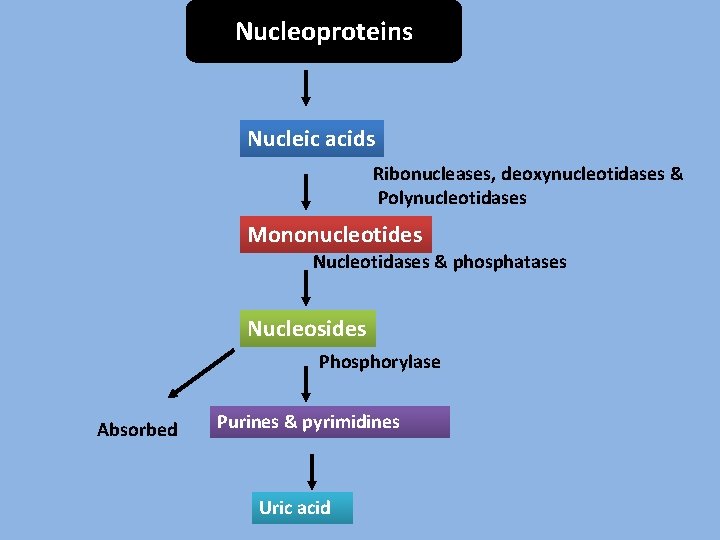
Nucleoproteins Nucleic acids Ribonucleases, deoxynucleotidases & Polynucleotidases Mononucleotides Nucleotidases & phosphatases Nucleosides Phosphorylase Absorbed Purines & pyrimidines Uric acid
- Slides: 92