Wellcome Trust graduate course Computational Methods series Sequencebased

Wellcome Trust graduate course. - Computational Methods series. --- Sequence-based bioinformatics. Dr. Hyunji Kim Department of Biochemistry, University of Oxford, South Parks Road, Oxford, OX 1 3 QU, UK Email: hyunji. kim@bioch. ox. ac. uk

Basic Tools 1) BLAST/WUBLAST A search engine to find sequences of your interest. BLAST can sophisticate its search, by varying substitution matrices/filtering options on a specified database. http: //www. ncbi. nlm. nih. gov/BLAST/, http: //www. ebi. ac. uk/blast 2/, 2) Clustal. W/T-Coffee/Muscle Helps us make sense of a bunch of unaligned sequences, via generating multiple or pairwise sequence alignments. Uses a progressive-alignment method. http: //www. ebi. ac. uk/clustalw/ 3) HMMer/PSI-BLAST Builds a profile Hidden Markov Model from a set of sequences aligned. Aligns sequences using a p. HMM, searches from a sequence database, and can assign functions to a given sequence. http: //hmmer. wustl. edu/ 4) Phylip/Tree. Dyn Calculates a distance matrix from a set of sequences. Derives phylogenetic trees, by taking such matrix as input, based upon theories of minimum evolution, parsimony and more. http: //evolution. genetics. washington. edu/phylip. html

5) Databases • • Nucleotide databases; EMBL, Genbank &DDBJ Protein databases; fully annotated, e. g. Swiss-Prot v 52. 3, as of 17 th of Apr. , 2007. (264, 492 entries) a computer-annotated, e. g. Tr. EMBL v 35. 3 • Genomics databases; Ensembl & Eukaryota, Bacteria and Archaea genomes 20+14; (v 44), 51, 445, 40, as of 20 th of Apr. , 2007. http: //www. ebi. ac. uk/uniprot/index. html, http: //www. ensembl. org/, http: //www. ebi. ac. uk/genomes/index. html 6) Major Bioinformatics Centres, around the globe. http: //www. ebi. ac. uk/, http: //www. ncbi. nlm. nih. gov/, http: //www. ddbj. nig. ac. jp/, http: //us. expasy. org/, http: //www. sanger. ac. uk/, http: //geneontology. org/

Searching for sequences by homology - BLAST

x y i j

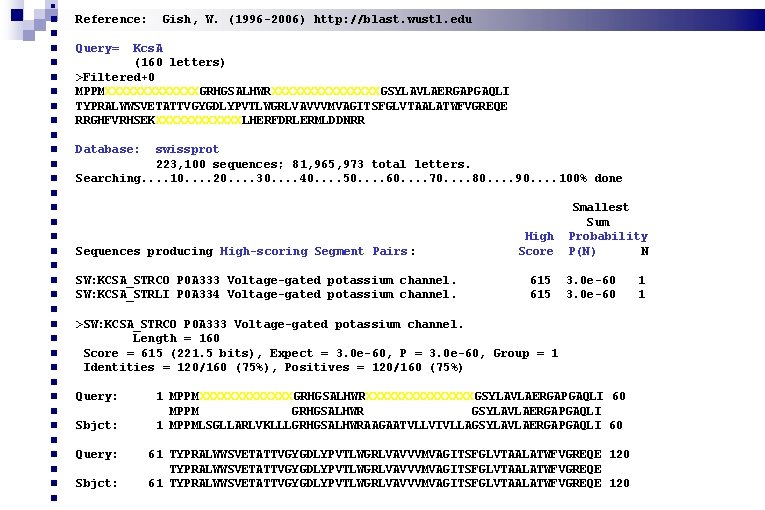
Reference: Gish, W. (1996 -2006) http: //blast. wustl. edu Query= Kcs. A (160 letters) >Filtered+0 MPPMXXXXXXXGRHGSALHWRXXXXXXXXGSYLAVLAERGAPGAQLI TYPRALWWSVETATTVGYGDLYPVTLWGRLVAVVVMVAGITSFGLVTAALATWFVGREQE RRGHFVRHSEKXXXXXXLHERFDRLERMLDDNRR Database: swissprot 223, 100 sequences; 81, 965, 973 total letters. Searching. . 10. . 20. . 30. . 40. . 50. . 60. . 70. . 80. . 90. . 100% done Sequences producing High-scoring Segment Pairs : High Score Smallest Sum Probability P(N) N SW: KCSA_STRCO P 0 A 333 Voltage-gated potassium channel. SW: KCSA_STRLI P 0 A 334 Voltage-gated potassium channel. 615 3. 0 e-60 >SW: KCSA_STRCO P 0 A 333 Voltage-gated potassium channel. Length = 160 Score = 615 (221. 5 bits), Expect = 3. 0 e-60, P = 3. 0 e-60, Group = 1 Identities = 120/160 (75%), Positives = 120/160 (75%) Query: Sbjct: 1 MPPMXXXXXXXGRHGSALHWRXXXXXXXXGSYLAVLAERGAPGAQLI 60 MPPM GRHGSALHWR GSYLAVLAERGAPGAQLI 1 MPPMLSGLLARLVKLLLGRHGSALHWRAAGAATVLLVIVLLAGSYLAVLAERGAPGAQLI 60 Query: Sbjct: 61 TYPRALWWSVETATTVGYGDLYPVTLWGRLVAVVVMVAGITSFGLVTAALATWFVGREQE 120 TYPRALWWSVETATTVGYGDLYPVTLWGRLVAVVVMVAGITSFGLVTAALATWFVGREQE 61 TYPRALWWSVETATTVGYGDLYPVTLWGRLVAVVVMVAGITSFGLVTAALATWFVGREQE 120 1 1

Multiple sequence alignment – Clustal. W

*************************** CLUSTAL W (1. 83) Multiple Sequence Alignments *************************** 1. Sequence Input From Disc 2. Multiple Alignments 3. Profile / Structure Alignments 4. Phylogenetic trees S. Execute a system command H. HELP X. EXIT (leave program) Your choice: 2 1. Do complete multiple alignment now (Slow/Accurate) 2. Produce guide tree file only 3. Do alignment using old guide tree file 4. Toggle Slow/Fast pairwise alignments = SLOW 5. Pairwise alignment parameters 6. Multiple alignment parameters ****** MULTIPLE ALIGNMENT MENU ****** 7. Reset gaps before alignment? = OFF 8. Toggle screen display = ON 9. Output format options S. Execute a system command H. HELP or press [RETURN] to go back to main menu Your choice:

AVFPMILW Red Small (small+ hydrophobic (incl. aromatic -Y)) DE Blue Acidic RHK Magenta Basic STYHCNGQ Green Hydroxyl, Amine Others Gray CLUSTAL W (1. 82) multiple sequence alignment KVAP_AERPE MVP_METJA O 28600 Q 8 TXQ 4 Q 6 L 2 S 2 Q 979 Z 2 O 26605 Q 9 HIA 8 Q 97 CK 5 FDALW-WAVVTATTVGYGDVVP-ATPIGKVIGIAVMLTGISALTLLIGTVSNMF-----FDAFY-FTTISITTVGYGDITP-KTDAGKLI---IIFS---VLFFISGLITS-------FDSLY-MTVITITTTGYGEVKP-MGPGGRVISMLLMFVGVGTF--------LTCLY-FTAATITTVGYGDVVP-TTEAGRLLSVIVMFSGIGVASYAL------FTSLW-WTMQTITTVGYGDTPV-YGFYGRINGMLIMVFGIGTIGYVTASLAT-------FTAIW-FTMETVTTVGYGDVVP-VSNLGRVVAMLIMVSGIGLLGTLTATISAYLF----Q EDSLW-YVLQTITTVGYGDIVP-VTSLGRFTGMVIMFSAIASTSLITASATSTLLERGEQ GNAFY-YTGEVITTLGFGDILP-VTMDAKIFTISLAFLGVAIFFSSITALILPSVERRLG GTALY-YTGETVTTLGFGDILP-VDLESRLFTISLAFLGVAIFFSAMTALITPTIERRVG 79 70 64 73 79 80 114 94 84

Profile alignment & Pattern recognition: HMMer More sensitive homology-search: PSI-BLAST & HMMer
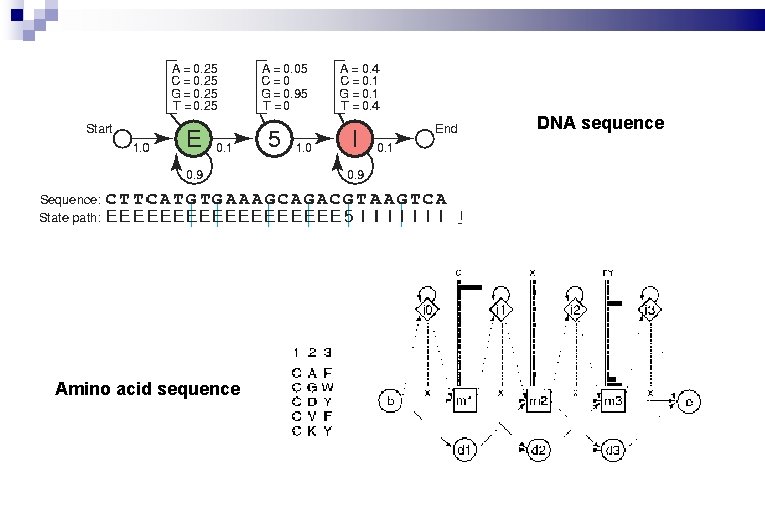
DNA sequence Amino acid sequence

PSI-BLAST

Phylogeny: Phylip & Treedyn
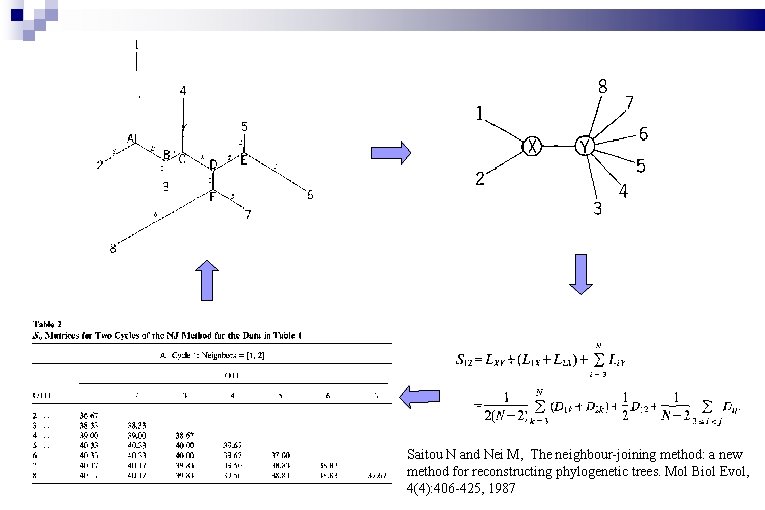
Saitou N and Nei M, The neighbour-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol, 4(4): 406 -425, 1987


Tree. Dyn

Protein secondary structure prediction: two consensus methods

http: //sbcb. bioch. ox. ac. uk/TM_noj. html

Example Output ALOM 2 DAS HMMTOP 2 MEMSAT 1. 5 PHD SPLIT 4 TMAP TMFINDER TMHMM 2 TMPRED TOPPRED 2 Consensus 640 650 660 670 680 690 700 | | | | MFAKGYGKNNEPLRGYILTFLIALGFILIAELNVIAPIISNFFLASYALINFSVFHASLAKSPGWRPAFK ***************************** *************************** ************************* ***************** ------? ? ? hhhh. HHHHHHHHHh. HHhhhhh? ? ? ------ Dr. Jonathan Cuthbertson developed Transmembrane Prediction Server.
Pongo http: //pongo. biocomp. unibo. it/pongo
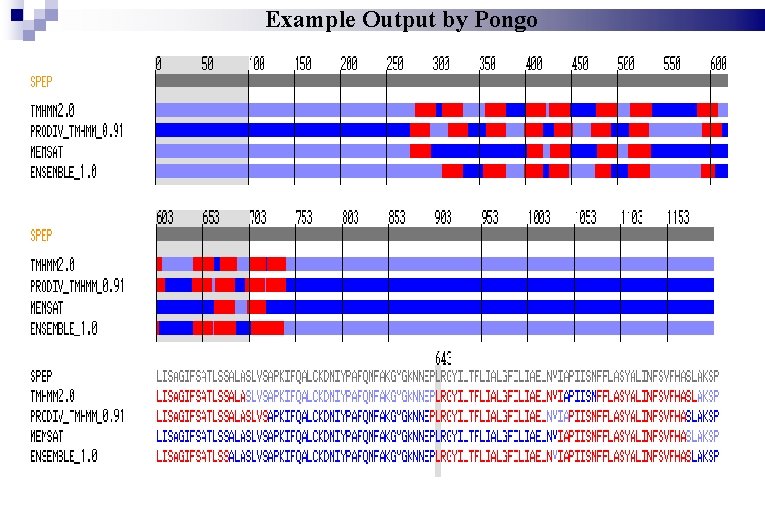
Example Output by Pongo

Background for practical sessions
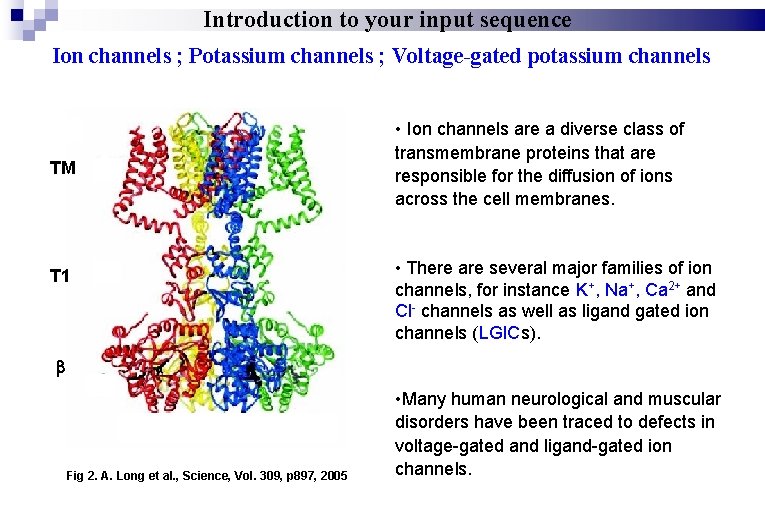
Introduction to your input sequence Ion channels ; Potassium channels ; Voltage-gated potassium channels TM T 1 • Ion channels are a diverse class of transmembrane proteins that are responsible for the diffusion of ions across the cell membranes. • There are several major families of ion channels, for instance K+, Na+, Ca 2+ and Cl- channels as well as ligand gated ion channels (LGICs). Fig 2. A. Long et al. , Science, Vol. 309, p 897, 2005 • Many human neurological and muscular disorders have been traced to defects in voltage-gated and ligand-gated ion channels.

Your expected blastp-output K+ channels, blastp Homologues are visualised in BLIXEM.
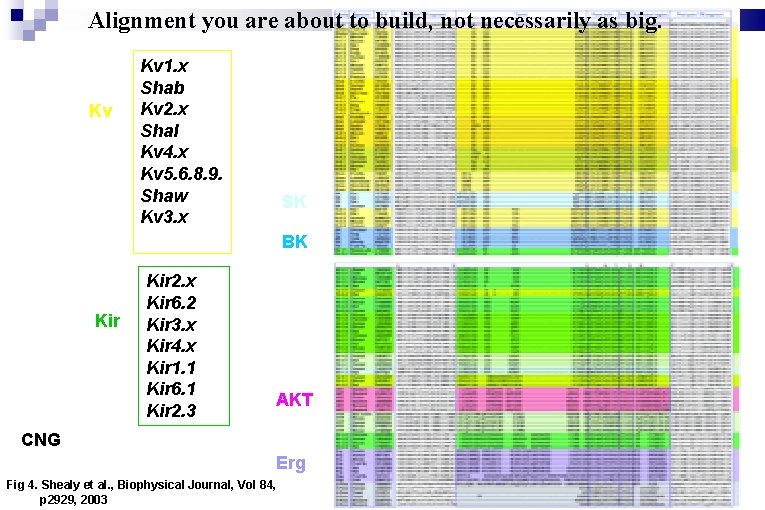
Alignment you are about to build, not necessarily as big. Kv Kv 1. x Shab Kv 2. x Shal Kv 4. x Kv 5. 6. 8. 9. Shaw Kv 3. x SK BK Kir 2. x Kir 6. 2 Kir 3. x Kir 4. x Kir 1. 1 Kir 6. 1 Kir 2. 3 AKT CNG Erg Fig 4. Shealy et al. , Biophysical Journal, Vol 84, p 2929, 2003

Example of p. HMM-related output hmmsearch - search a sequence database with a profile HMM - - - - HMM file: Sequence database: - - - - - - - - - Kv. hmm [Kv_homologues] infile_comb - - - - - Query HMM: Kv_homologues HMM has been calibrated; E-values are empirical estimates] Scores for complete sequences (score includes all domains): Sequence -------CIKS_DROME Q 9 VX 00_DROME CIKB_DROME O 62350_Celegans Q 9 VLC 6_DROME CIKW_DROME Q 8 SYL 2_DROME Q 22012_Celegans Filtered_5 DROME Filtered_6 DROME Q 9 XXD 1_Celegans Description ------ Score ----241. 2 234. 3 159. 3 156. 7 156. 6 156. 5 155. 3 140. 5 125. 0 E-value N ------- --3. 2 e-71 1 3. 9 e-69 1 1. 5 e-46 1 8. 8 e-46 1 9. 6 e-46 1 1 e-45 1 2. 4 e-45 1 6. 6 e-41 1 3. 1 e-36 1
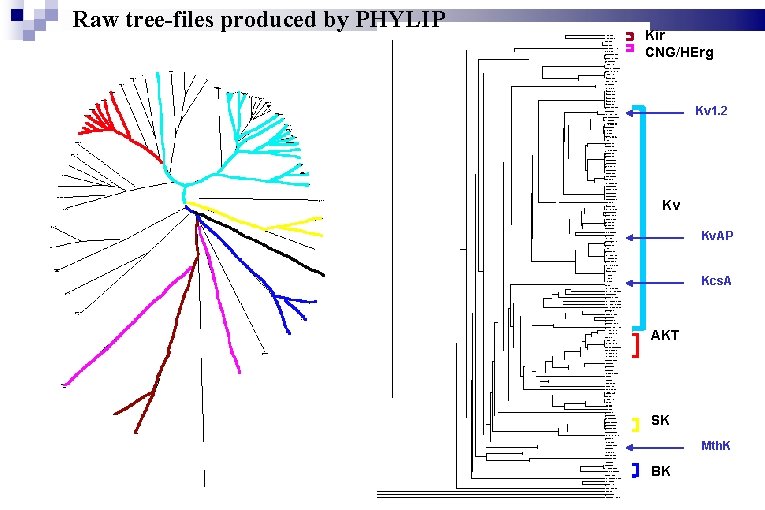
Raw tree-files produced by PHYLIP Kir CNG/HErg Kv 1. 2 Kv Kv. AP Kcs. A AKT SK Mth. K BK

Phylogenetic trees modified in Tree. Dyn
- Slides: 33