DOE Genomics GTL Annual Retreat 2005 SequenceBased Discovery

DOE Genomics: GTL Annual Retreat 2005 Sequence-Based Discovery Carl Abulencia Diversa Corporation San Diego, California

Sequence-Based Discovery Goals • Gain a better understanding of how organisms live in contaminated environments, and how they have adapted to external stressors. • Obtain sequence data by accessing the genomes from the organisms living in contaminated environments. • Analyze the genes, operons, and stress response pathways from the natural diversity to study evolutionary changes, selections, expansions and gene transfers.
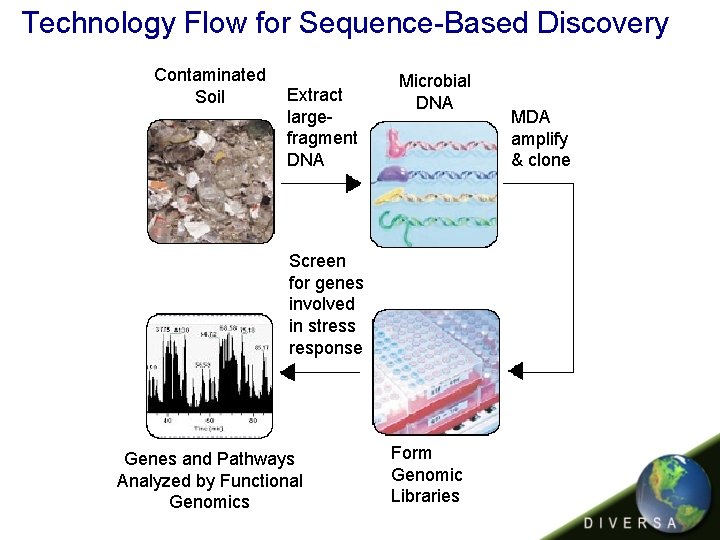
Technology Flow for Sequence-Based Discovery Contaminated Soil Extract largefragment DNA Microbial DNA Screen for genes involved in stress response Pathways Genes andanalyzed Pathways Analyzed by computational by Functional core Genomics Form Genomic Libraries MDA amplify & clone
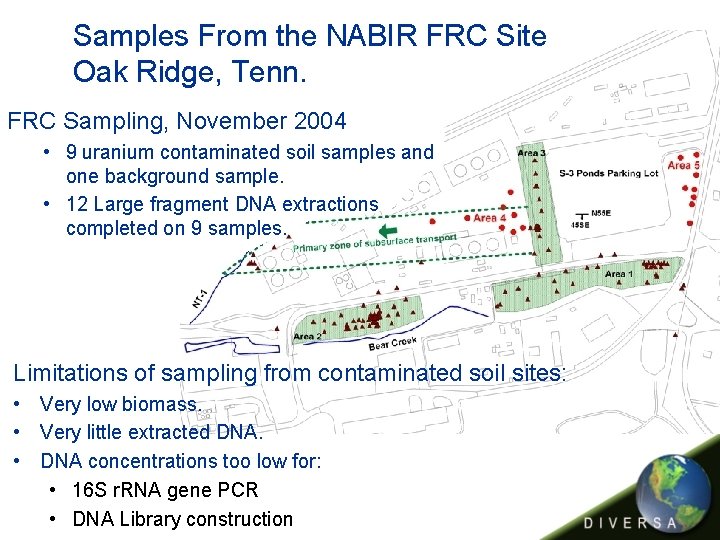
Samples From the NABIR FRC Site Oak Ridge, Tenn. FRC Sampling, November 2004 • 9 uranium contaminated soil samples and one background sample. • 12 Large fragment DNA extractions completed on 9 samples. Limitations of sampling from contaminated soil sites: • Very low biomass. • Very little extracted DNA. • DNA concentrations too low for: • 16 S r. RNA gene PCR • DNA Library construction

Amplification of Sample DNA Whole Genome Amplification Multiple Displacement Amplification (MDA) 97 kb 49 kb 12 kb • Isothermal amplification method. • phi 29 polymerase. • Processive (up to 70 kb). • Strand-displacing. • 3’-5’ exonuclease activity (proofreading activity). • Random hexamer primers. • 3 hour – 16 hour reaction time.
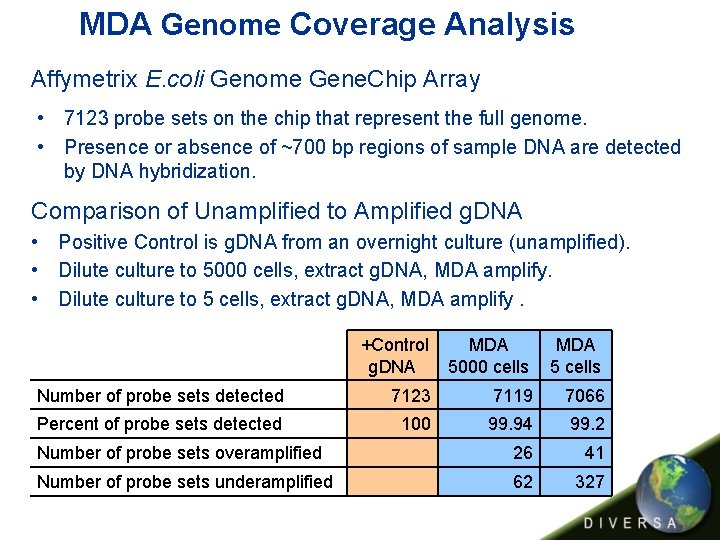
MDA Genome Coverage Analysis Affymetrix E. coli Genome Gene. Chip Array • 7123 probe sets on the chip that represent the full genome. • Presence or absence of ~700 bp regions of sample DNA are detected by DNA hybridization. Comparison of Unamplified to Amplified g. DNA • Positive Control is g. DNA from an overnight culture (unamplified). • Dilute culture to 5000 cells, extract g. DNA, MDA amplify. • Dilute culture to 5 cells, extract g. DNA, MDA amplify. +Control g. DNA MDA 5000 cells MDA 5 cells Number of probe sets detected 7123 7119 7066 Percent of probe sets detected 100 99. 94 99. 2 Number of probe sets overamplified 26 41 Number of probe sets underamplified 62 327
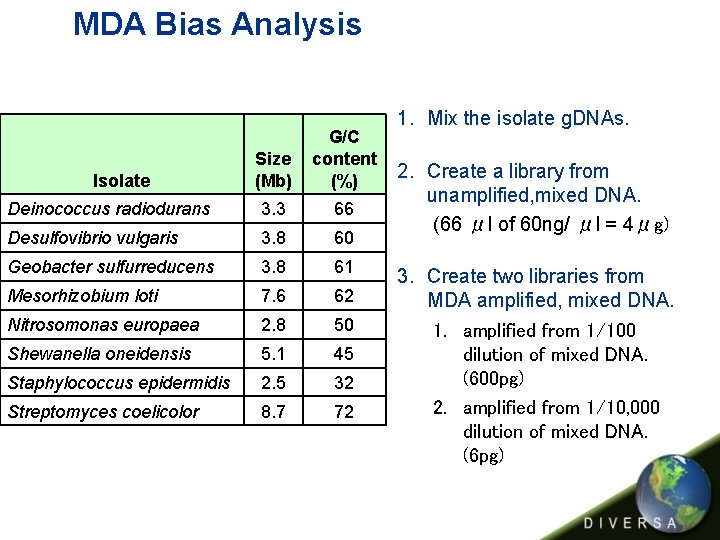
MDA Bias Analysis Size (Mb) G/C content (%) Deinococcus radiodurans 3. 3 66 Desulfovibrio vulgaris 3. 8 60 Geobacter sulfurreducens 3. 8 61 Mesorhizobium loti 7. 6 62 Nitrosomonas europaea 2. 8 50 Shewanella oneidensis 5. 1 45 Staphylococcus epidermidis 2. 5 32 Streptomyces coelicolor 8. 7 72 Isolate 1. Mix the isolate g. DNAs. 2. Create a library from unamplified, mixed DNA. (66 μl of 60 ng/ μl = 4μg) 3. Create two libraries from MDA amplified, mixed DNA. 1. amplified from 1/100 dilution of mixed DNA. (600 pg) 2. amplified from 1/10, 000 dilution of mixed DNA. (6 pg)
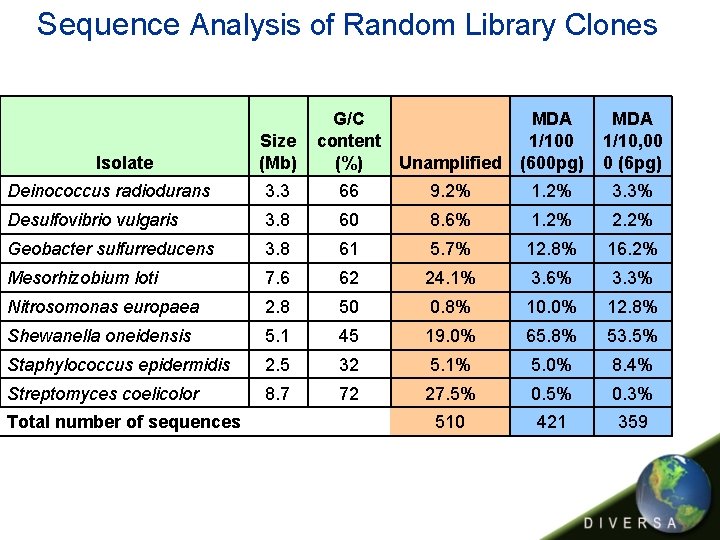
Sequence Analysis of Random Library Clones Isolate Size (Mb) G/C content (%) Unamplified MDA 1/100 (600 pg) MDA 1/10, 00 0 (6 pg) Deinococcus radiodurans 3. 3 66 9. 2% 1. 2% 3. 3% Desulfovibrio vulgaris 3. 8 60 8. 6% 1. 2% 2. 2% Geobacter sulfurreducens 3. 8 61 5. 7% 12. 8% 16. 2% Mesorhizobium loti 7. 6 62 24. 1% 3. 6% 3. 3% Nitrosomonas europaea 2. 8 50 0. 8% 10. 0% 12. 8% Shewanella oneidensis 5. 1 45 19. 0% 65. 8% 53. 5% Staphylococcus epidermidis 2. 5 32 5. 1% 5. 0% 8. 4% Streptomyces coelicolor 8. 7 72 27. 5% 0. 3% 510 421 359 Total number of sequences
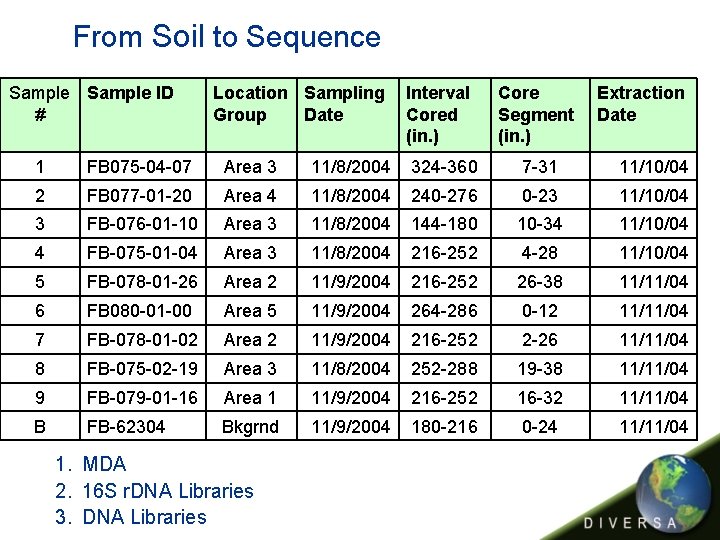
From Soil to Sequence Sample ID # Location Sampling Group Date Interval Cored (in. ) Core Segment (in. ) Extraction Date 1 FB 075 -04 -07 Area 3 11/8/2004 324 -360 7 -31 11/10/04 2 FB 077 -01 -20 Area 4 11/8/2004 240 -276 0 -23 11/10/04 3 FB-076 -01 -10 Area 3 11/8/2004 144 -180 10 -34 11/10/04 4 FB-075 -01 -04 Area 3 11/8/2004 216 -252 4 -28 11/10/04 5 FB-078 -01 -26 Area 2 11/9/2004 216 -252 26 -38 11/11/04 6 FB 080 -01 -00 Area 5 11/9/2004 264 -286 0 -12 11/11/04 7 FB-078 -01 -02 Area 2 11/9/2004 216 -252 2 -26 11/11/04 8 FB-075 -02 -19 Area 3 11/8/2004 252 -288 19 -38 11/11/04 9 FB-079 -01 -16 Area 1 11/9/2004 216 -252 16 -32 11/11/04 B FB-62304 Bkgrnd 11/9/2004 180 -216 0 -24 11/11/04 1. MDA 2. 16 S r. DNA Libraries 3. DNA Libraries
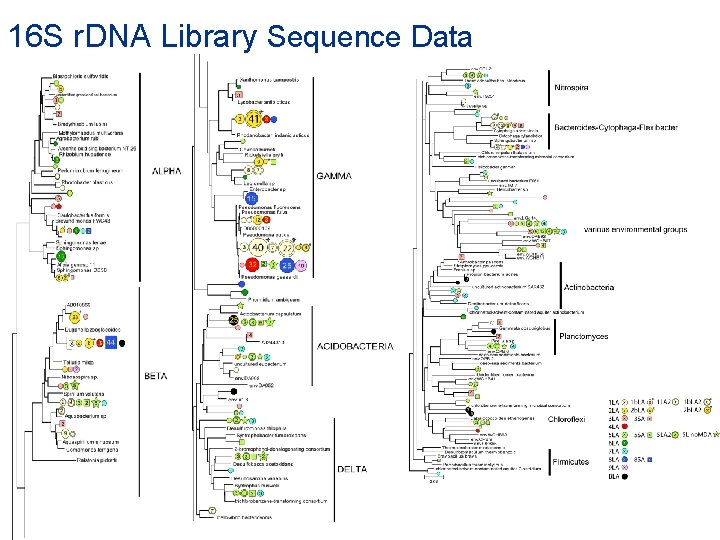
16 S r. DNA Library Sequence Data
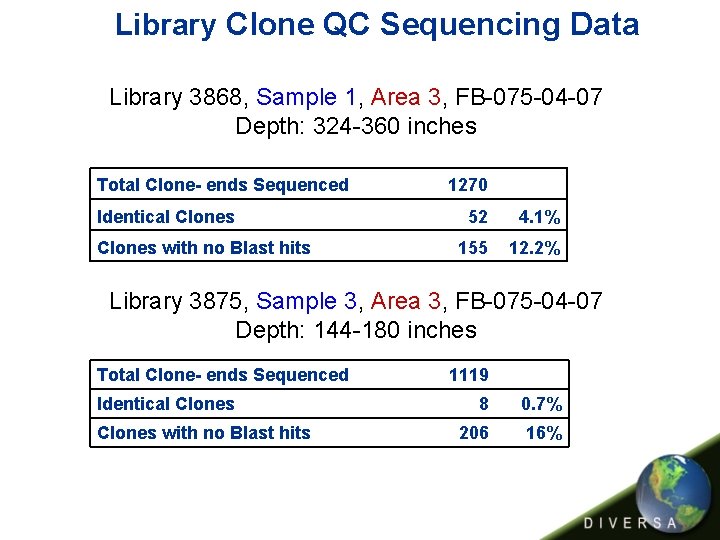
Library Clone QC Sequencing Data Library 3868, Sample 1, Area 3, FB-075 -04 -07 Depth: 324 -360 inches Total Clone- ends Sequenced Identical Clones with no Blast hits 1270 52 4. 1% 155 12. 2% Library 3875, Sample 3, Area 3, FB-075 -04 -07 Depth: 144 -180 inches Total Clone- ends Sequenced Identical Clones with no Blast hits 1119 8 0. 7% 206 16%

Interesting Library QC Sequences E value Top Blast hit 3. 00 E-59 gi|47059343| heavy metal transporting ATPase [Ornithobacterium rhinotracheale] 3. 00 E-46 gi|39996434| heavy metal efflux pump, Czc. A family [Geobacter sulfurreducens PCA] 1. 00 E-17 gi|48846102| COG 1235: Metal-dependent hydrolases of the beta-lactamase superfamily I [Geobacter metallireducens GS-15] 1. 00 E-11 gi|18977112|ref|NP_578469. 1| heavy-metal transporting cpx-type atpase [Pyrococcus furiosus DSM 3638] 2. 00 E-87 gi|56460133|ref|YP_155414. 1| Predicted extracellular metal-dependent peptidase [Idiomarina loihiensis L 2 TR] 1. 00 E-45 gi|53760196| COG 0347: Nitrogen regulatory protein PII [Methylobacillus flagellatus KT] 3. 00 E-24 gi|46131534| COG 0715: ABC-type nitrate/sulfonate/bicarbonate transport systems, periplasmic components [Ralstonia eutropha JMP 134] 1. 00 E-10 gi|62484831| predicted cytotoxic translational repressor of toxic-antitoxic stability system [uncultured bacterium] 2. 00 E-56 gi|46579304| aminotransferase, classes I and II [Desulfovibrio vulgaris subsp. vulgaris str. Hildenborough]

Library Screening for Histidine Kinase Genes • Histidine kinase superfamily chosen for sequence-based discovery from environmental libraries. • Cells sense and respond to extracellular stimuli through phosphotransfermediated signaling pathways controlled by HKs and response regulators. • Library clones containing HKs retrieved by DNA hybridization using degenerate probes designed from a subfamily of HKs from D. vulgaris. Sense Primer 5’-GGCSCAYGARATSAACAACCC-3’ 5’-GGTSGTGAAGAASGGYTCGAA-3 ’ Antisense Primer
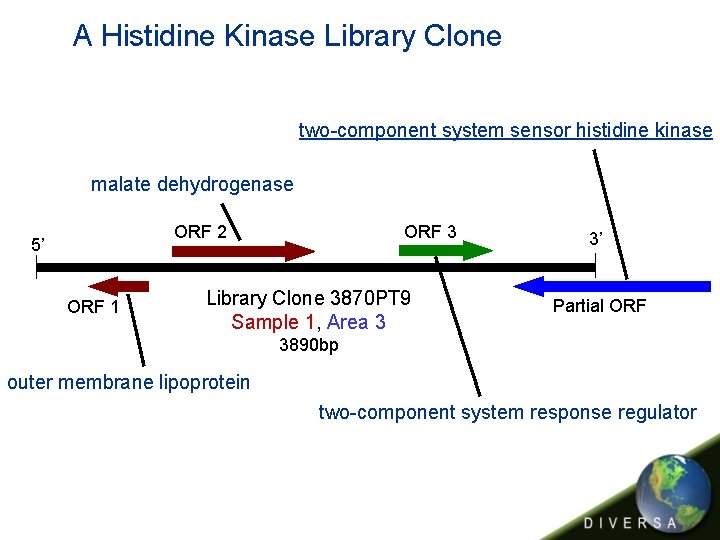
A Histidine Kinase Library Clone two-component system sensor histidine kinase malate dehydrogenase ORF 3 ORF 2 5’ ORF 1 Library Clone 3870 PT 9 Sample 1, Area 3 3’ Partial ORF 3890 bp outer membrane lipoprotein two-component system response regulator

Histidine Kinase Discovery Progress Sequence-Based Screening • 12 Libraries Screened, 50, 000 clones/ library • 26 HK clones in Sequencing Library QC sequencing • 34 HK clones fully sequenced

Summary • g. DNA extracted from 9 contaminated soil samples • MDA used to amplify g. DNA from low-yield samples • 16 S r. DNA libraries constructed from each sample • Library diversity verified by random QC sequencing • Histidine kinases discovered in libraries by random sequencing • Desulfovibrio-like histidine kinases discovered in libraries by sequence-based hybridization • 16 S r. DNA sequences, library QC sequences, and full length HK clone sequences all uploaded to the VIMSS database • A manuscript for publication is near completion detailing analysis of DNA libraries constructed from contaminated environmental soil samples

Acknowledgments • LBNL – Dominique Joyner, Sharon Borglin, Eoin Brodie, Terry Hazen • LBNL – Eric Alm, Adam Arkin • LBNL – Tamas Torok • ORNL – David Watson • Diversa – Denise Wyborski, Mircea Podar, Joe Garcia, Cathy Chang, Sequencing Group, Keiko Obokata, Toby Richardson, Sherman Chang, Karsten Zengler, Martin Keller
- Slides: 17