TTT TTC TTA TTG CTT CTC CTA CTG



































- Slides: 81






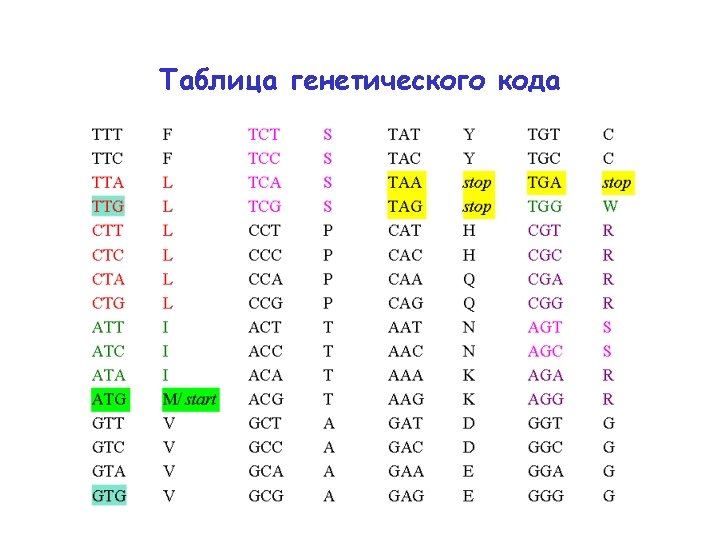
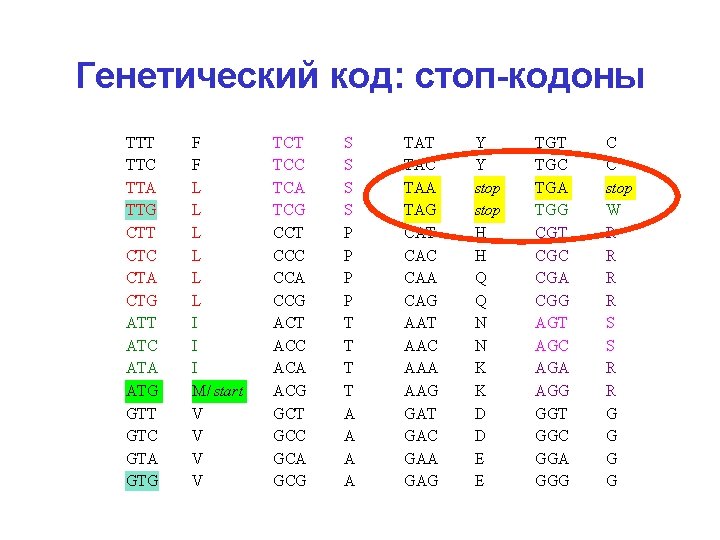
Генетический код: стоп-кодоны TTT TTC TTA TTG CTT CTC CTA CTG ATT ATC ATA ATG GTT GTC GTA GTG F F L L L I I I M/ start V V TCT TCC TCA TCG CCT CCC CCA CCG ACT ACC ACA ACG GCT GCC GCA GCG S S P P T T A A TAT TAC TAA TAG CAT CAC CAA CAG AAT AAC AAA AAG GАT GАC GАA GАG Y Y stop H H Q Q N N K K D D E E TGT TGC TGA TGG CGT CGC CGA CGG AGT AGC AGA AGG GGT GGC GGA GGG C C stop W R R S S R R G G



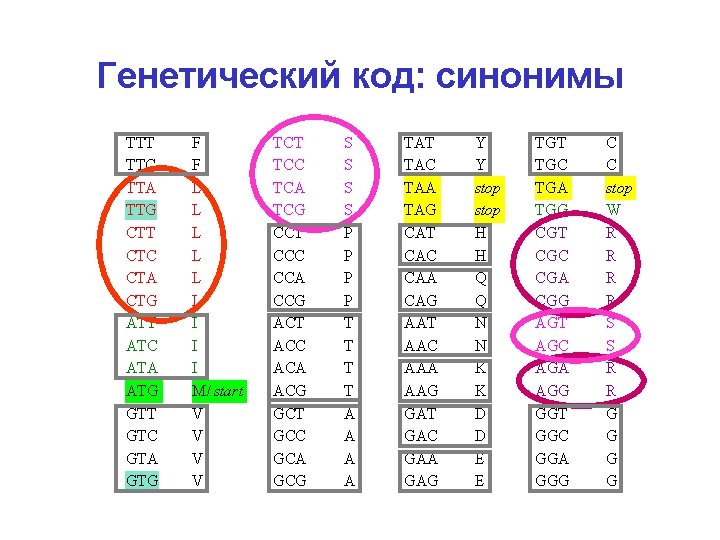
Генетический код: синонимы TTT TTC TTA TTG CTT CTC CTA CTG ATT ATC ATA ATG GTT GTC GTA GTG F F L L L I I I M/ start V V TCT TCC TCA TCG CCT CCC CCA CCG ACT ACC ACA ACG GCT GCC GCA GCG S S P P T T A A TAT TAC TAA TAG CAT CAC CAA CAG AAT AAC AAA AAG GАT GАC GАA GАG Y Y stop H H Q Q N N K K D D E E TGT TGC TGA TGG CGT CGC CGA CGG AGT AGC AGA AGG GGT GGC GGA GGG C C stop W R R S S R R G G


Gen. Mark, окно 96 нт

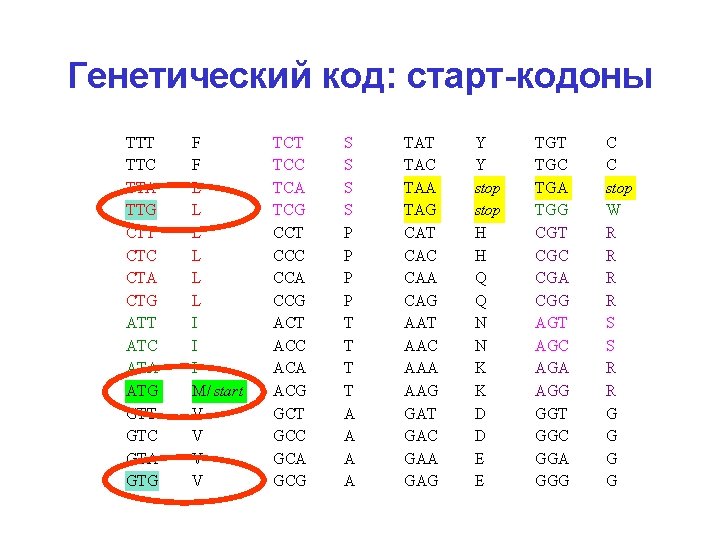
Генетический код: старт-кодоны TTT TTC TTA TTG CTT CTC CTA CTG ATT ATC ATA ATG GTT GTC GTA GTG F F L L L I I I M/ start V V TCT TCC TCA TCG CCT CCC CCA CCG ACT ACC ACA ACG GCT GCC GCA GCG S S P P T T A A TAT TAC TAA TAG CAT CAC CAA CAG AAT AAC AAA AAG GАT GАC GАA GАG Y Y stop H H Q Q N N K K D D E E TGT TGC TGA TGG CGT CGC CGA CGG AGT AGC AGA AGG GGT GGC GGA GGG C C stop W R R S S R R G G

Начала генов Bacillus subtilis dna. N ACATTATCCGTTAGGAGGATAAAAATG gyr. A GTGATACTTCAGGGAGGTTTTTTAATG ser. S TCAATAAAAAAAGGAGTGTTTCGCATG bof. A CAAGCGAAGGAGATGAGAAGATTCATG csf. B GCTAACTGTACGGAGGTGGAGAAGATG xpa. C ATAGACACAGGAGTCGATTATCTCATG met. S ACATTCTGATTAGGAGGTTTCAAGATG gca. D AAAAGGGATATTGGAGGCCAATAAATG spo. VC TATGTGACTAAGGGAGGATTCGCCATG fts. H GCTTACTGTGGGAGGAGGTAAGGAATG pab. B AAAGAAAATAGAGGAATGATACAAATG rpl. J CAAGAATCTACAGGAGGTGTAACCATG tuf. A AAAGCTCTTAAGGAGGATTTTAGAATG rps. J TGTAGGCGAAAAGGAGGGAAAATAATG rpo. A CGTTTTGAAGGAGGGTTTTAAGTAATG rpl. M AGATCATTTAGGAGGGGAAATTCAATG

Участок связывания рибосом dna. N ACATTATCCGTTAGGAGGATAAAAATG gyr. A GTGATACTTCAGGGAGGTTTTTTAATG ser. S TCAATAAAAAAAGGAGTGTTTCGCATG bof. A CAAGCGAAGGAGATGAGAAGATTCATG csf. B GCTAACTGTACGGAGGTGGAGAAGATG xpa. C ATAGACACAGGAGTCGATTATCTCATG met. S ACATTCTGATTAGGAGGTTTCAAGATG gca. D AAAAGGGATATTGGAGGCCAATAAATG spo. VC TATGTGACTAAGGGAGGATTCGCCATG fts. H GCTTACTGTGGGAGGAGGTAAGGAATG pab. B AAAGAAAATAGAGGAATGATACAAATG rpl. J CAAGAATCTACAGGAGGTGTAACCATG tuf. A AAAGCTCTTAAGGAGGATTTTAGAATG rps. J TGTAGGCGAAAAGGAGGGAAAATAATG rpo. A CGTTTTGAAGGAGGGTTTTAAGTAATG rpl. M AGATCATTTAGGAGGGGAAATTCAATG

Сравнительный анализ (один и тот же ген в нескольких геномах) Гены консервативнее, чем межгенные области (точнее, особенности эволюции другие) Sty TCGCTCG--CAGCGGAAAGAGGATTACGCCCTTCGCCTGGAGGCTGTGCAGGGGC---GCCGGAGATGGGATGCATAATT Stm TCGCTCG--CAGCGGAAAGAGGATTACGCCCTTCGCCTGGAGGCTGTGCAGGGGC---GCCGGAGATGGGATGCATAATT Sen TCGCTCG--CAGCGGAAAGAGGATTACGCCCTTCGCCTGGAGGCTGTGCAGGGGC---GCCGGAGATGGGATGCATAATT Eco TTGCCCG--TGCCAGACGGCAGATTATCTCCCTGACCTGGTGGTTGCCCAGGAGGAGGGCCGGAAATAGGTTGTATCATT Kpn ----CGG--TGGCGCAGTGCCTGATGGG-CCTCGCCCTGGAGGACGGTCTGGCAT---ATCAGCAAGGGGGTGCGTCATG Ype TTGTTAGAACAGGGGAAAACGGTAAACAGTGTGGCATTAGATGTCGGTTATAGCT-----CCGCCTCTGCTTTTATCGCC * * * * * * * Sty AATTATCCTTTAAC-----CATAAATCTGAGCAATA-TATGCTTGGCGGCCAGATTATGGC--ACACTTGTCCGG Stm AATTATCCTTTAAC-----CATAAATCTGAGCAATA-TATGCCTGGCGGCCAGATTATGGC--ACACTTGTCCGG Sen AATTATCCTTTAAC-----CATAAATCTGAGCAATA-TATGCCTGGCGGCCAGATTATGGC--ACACTTGTCCGG Eco ACGTATCCTTATAC-----CTGAAATCTTCGCAAG--TATGCCTGGCCGCGAGATTATGGC--ACACTTGTCCGG Kpn ATTCATCCTTTCGATATCGCGGTGCTGGAACCAGGTGATGAGTATGCCTGGCGGCCAGATTATGGC--ACACTTCCCCAG Ype ATGTTTCAGCAAATAT----CGGGTACCA-CGCCTGAGCGTTTCCGGCGGGGCAATAGTGGCTTATACTAAGCCCC * ** * * *** * ** **** * *** Sty TTAACTCTCGTT-CTCAAACAG------GTACGACAGTC--GTGAAAATTCTCGTTGATGAAAATATGCCTTACGCCCGC Stm TTAACTCTCGTT-CTCAAACAG------GTACGACAGTC--GTGAAAATTCTCGTTGATGAAAATATGCCTTACGCCCGC Sen TTAACTCTCGTT-CTCAAACAG------GTACGACAGTC--GTGAAAATTCTCGTTGATGAAAATATGCCTTACGCCCGC Eco TTAACTCTCGT--CTCATACAG------GTAACACAAAC--GTGAAAATCCTTGTTGATGAAAATATGCCTTATGCCCGC Kpn TTAACTCTCGTT-CTCAGACAG------GTACTGAACT---GTGAAAATCCTCGTTGATGAAAATATGCCCGT Ype CTGTTTTTCATCTGTATGGCAGTTCGCTGTCGGAGAGTAAAGTGAAAATTCTGGTTGATGAAAATATGCCGTACGCTGAG * * ** * * *** **** ** ********* ** ** 123123123123123123123

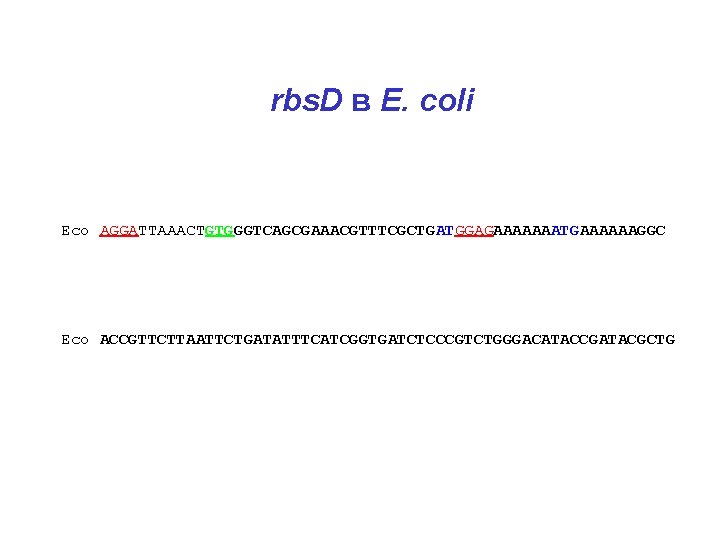
rbs. D в E. coli Eco AGGATTAAACTGTGGGTCAGCGAAACGTTTCGCTGATGGAGAAAAAAATGAAAAAAGGC Eco ACCGTTCTTAATTCTGATATTTCATCGGTGATCTCCCGTCTGGGACATACCGATACGCTG

rbs. D в энтеробактериях Sty AGGGTTACACTGCGGC-CAGCGAAACGTTTCGCTAGTGGAGCAGAAAAATGAAGAAAGGC Sen AGGGTTACACTGCGGC-CAGCGAAACGTTTCGCTAGTGGAGCAGAAAAATGAAGAAAGGC Stm GGGGTTACACTGCGGC-CAGCGAAACGTTTCGCTAGTGGAGCAGAAAAATGAAGAAAGGC Eco AGGATTAAACTGTGGGTCAGCGAAACGTTTCGCTGATGGAGAA-AAAAATGAAAAAAGGC Ype TTTTCTAAACTCCTTGTTAGCGAAACGTTTCGCTCTTGGAGTA-GATCATGAAAAAAGGT ** ******** * * ***** Sty ACCGTACTCAACTCTGAAATCTCGTCGGTCATTTCCCGTCTGGGGCATACTGATACTCTG Sen ACCGTACTCAACTCTGAAATCTCGTCGGTCATTTCCCGTCTGGGGCATACTGATACTCTG Stm ACCGTACTCAACTCTGAAATCTCGTCGGTCATTTCCCGTCTGGGGCATACTGATACTCTG Eco ACCGTTCTTAATTCTGATATTTCATCGGTGATCTCCCGTCTGGGACATACCGATACGCTG Ype GTATTACTGAACGCTGATATTTCCGCGGTTATCTCCCGTCTGGGCCATACCGATCAGATT * ** ** ****** ***** *

rbs. D в энтеробактериях: ответ Sty AGGGTTACACTGCGGC-CAGCGAAACGTTTCGCTAGTGGAGCAGAAAAATGAAGAAAGGC Sen AGGGTTACACTGCGGC-CAGCGAAACGTTTCGCTAGTGGAGCAGAAAAATGAAGAAAGGC Stm GGGGTTACACTGCGGC-CAGCGAAACGTTTCGCTAGTGGAGCAGAAAAATGAAGAAAGGC Eco AGGATTAAACTGTGGGTCAGCGAAACGTTTCGCTGATGGAGAA-AAAAATGAAAAAAGGC Ype TTTTCTAAACTCCTTGTTAGCGAAACGTTTCGCTCTTGGAGTA-GATCATGAAAAAAGGT ** ******** * * ***** Sty ACCGTACTCAACTCTGAAATCTCGTCGGTCATTTCCCGTCTGGGGCATACTGATACTCTG Sen ACCGTACTCAACTCTGAAATCTCGTCGGTCATTTCCCGTCTGGGGCATACTGATACTCTG Stm ACCGTACTCAACTCTGAAATCTCGTCGGTCATTTCCCGTCTGGGGCATACTGATACTCTG Eco ACCGTTCTTAATTCTGATATTTCATCGGTGATCTCCCGTCTGGGACATACCGATACGCTG Ype GTATTACTGAACGCTGATATTTCCGCGGTTATCTCCCGTCTGGGCCATACCGATCAGATT * ** ** ****** ***** *



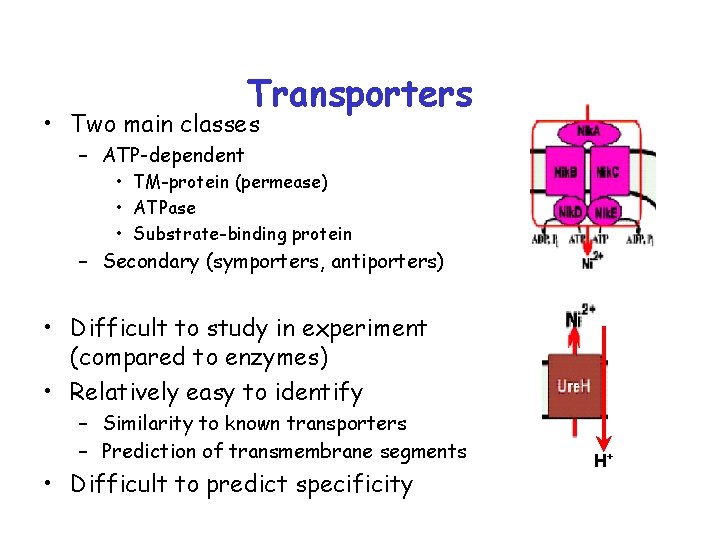
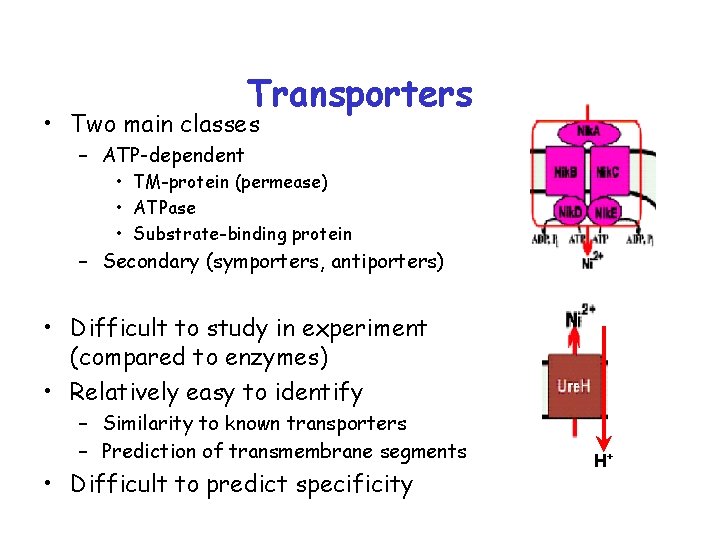
Transporters • Two main classes – ATP-dependent • TM-protein (permease) • ATPase • Substrate-binding protein – Secondary (symporters, antiporters) • Difficult to study in experiment (compared to enzymes) • Relatively easy to identify – Similarity to known transporters – Prediction of transmembrane segments • Difficult to predict specificity H+


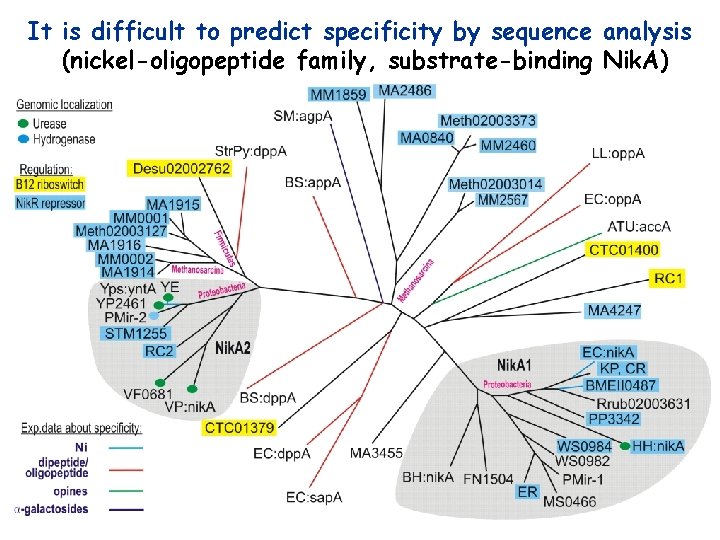
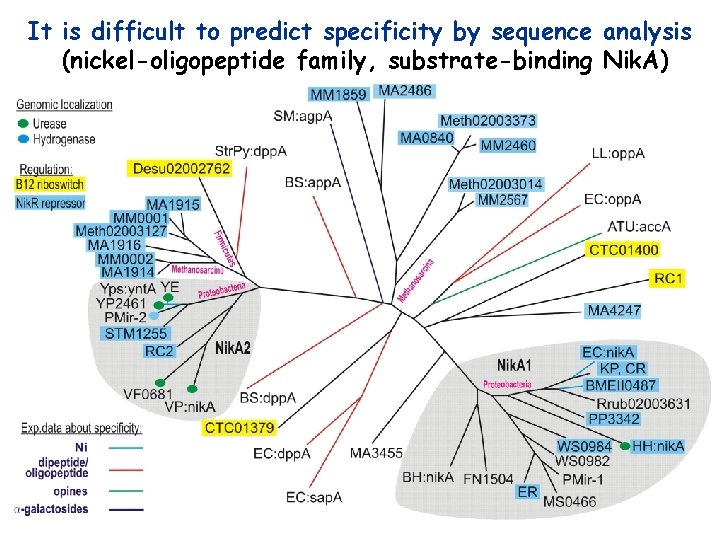
It is difficult to predict specificity by sequence analysis (nickel-oligopeptide family, substrate-binding Nik. A)

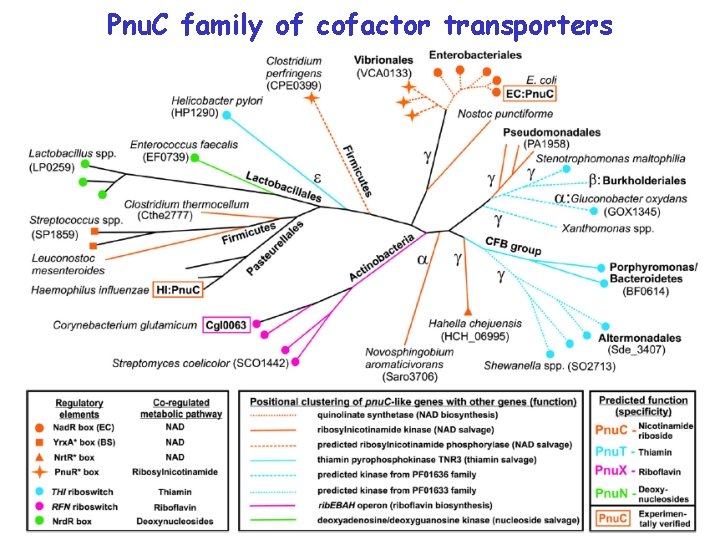
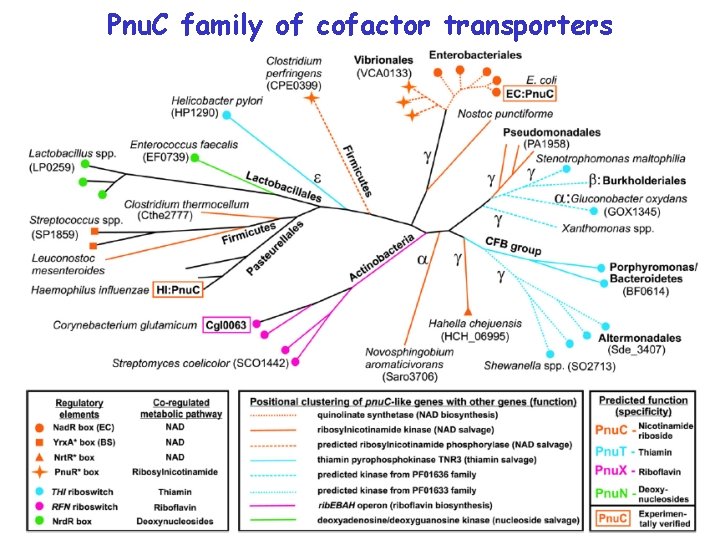
Pnu. C family of cofactor transporters

Riboflavin biosynthesis pathway

5’ UTR regions of riboflavin genes from various bacteria

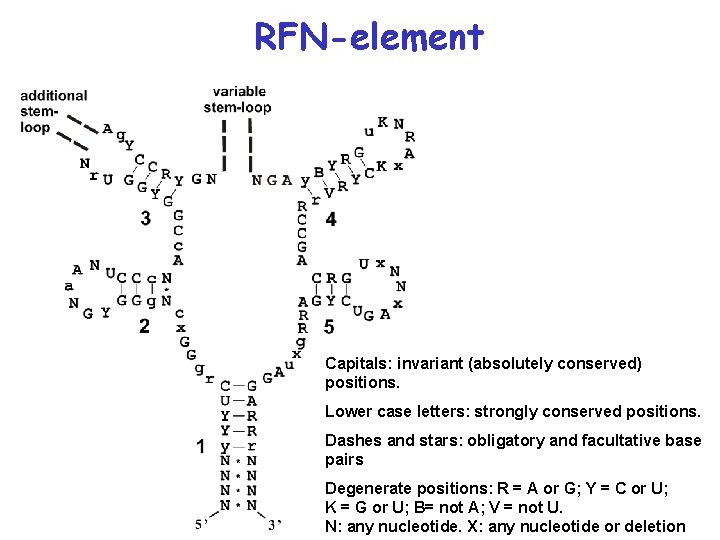
Conserved secondary structure of the RFN-element Capitals: invariant (absolutely conserved) positions. Lower case letters: strongly conserved positions. Dashes and stars: obligatory and facultative base pairs N: any nucleotide. X: any nucleotide or deletion

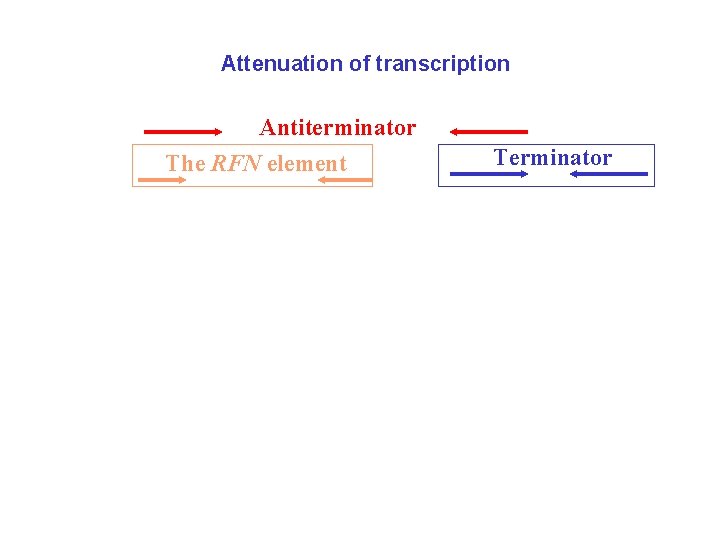
Attenuation of transcription Antiterminator The RFN element Antiterminator Terminator

Attenuation of translation Antisequestor The RFN element SD-sequestor

Рибопереключатель RFN: регуляторный механизм Transcription attenuation Translation attenuation



Метаболическая реконструкция тиаминового биосинтеза = thi. N (confirmed) Transport of HMP Transport of HET (Gram-positive bacteria) (Gram-negative bacteria)







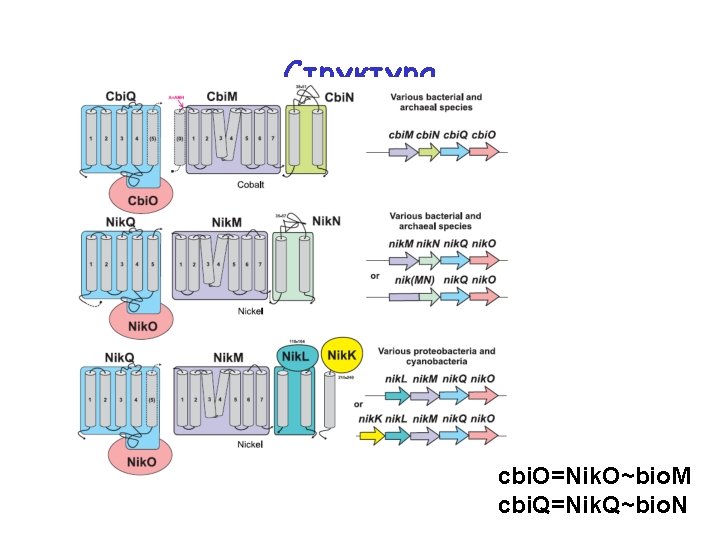
Структура cbi. O=Nik. O~bio. M cbi. Q=Nik. Q~bio. N









Transporters • Two main classes – ATP-dependent • TM-protein (permease) • ATPase • Substrate-binding protein – Secondary (symporters, antiporters) • Difficult to study in experiment (compared to enzymes) • Relatively easy to identify – Similarity to known transporters – Prediction of transmembrane segments • Difficult to predict specificity H+

It is difficult to predict specificity by sequence analysis (nickel-oligopeptide family, substrate-binding Nik. A)

Pnu. C family of cofactor transporters

Riboflavin biosynthesis pathway

5’ UTR regions of riboflavin genes

RFN-element Capitals: invariant (absolutely conserved) positions. Lower case letters: strongly conserved positions. Dashes and stars: obligatory and facultative base pairs Degenerate positions: R = A or G; Y = C or U; K = G or U; B= not A; V = not U. N: any nucleotide. X: any nucleotide or deletion

RFN: the mechanism of regulation • Transcription attenuation • Translation attenuation

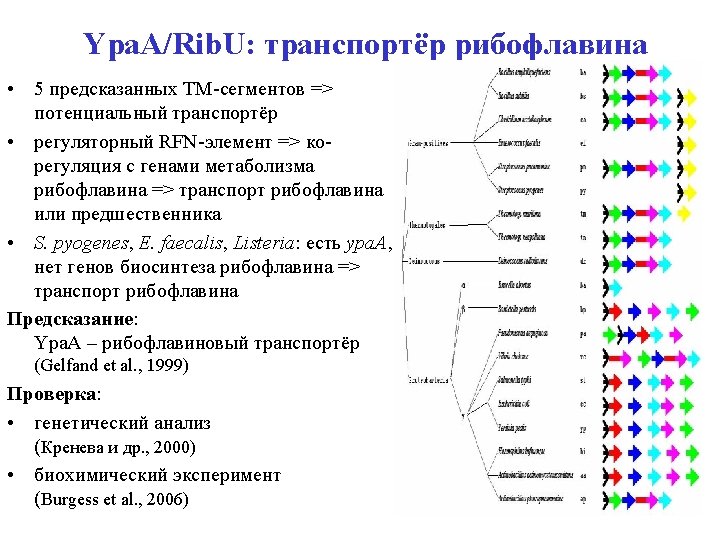
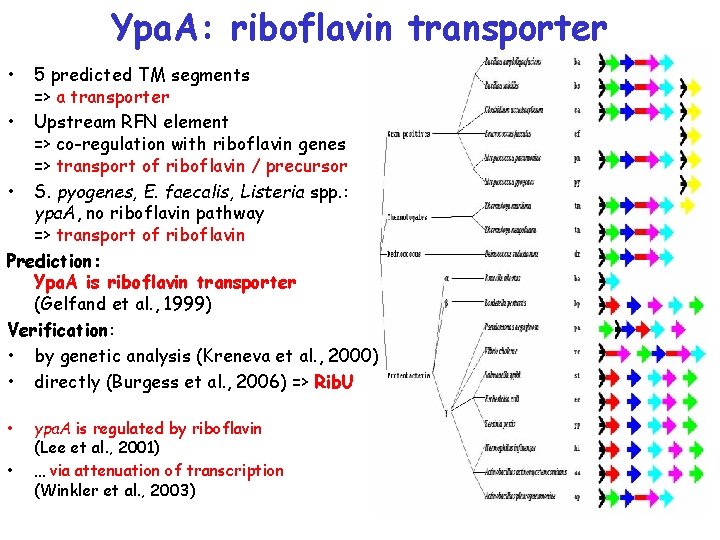
Ypa. A: riboflavin transporter • 5 predicted TM segments => a transporter • Upstream RFN element => co-regulation with riboflavin genes => transport of riboflavin / precursor • S. pyogenes, E. faecalis, Listeria spp. : ypa. A, no riboflavin pathway => transport of riboflavin Prediction: Ypa. A is riboflavin transporter (Gelfand et al. , 1999) Verification: • by genetic analysis (Kreneva et al. , 2000) • directly (Burgess et al. , 2006) => Rib. U • • ypa. A is regulated by riboflavin (Lee et al. , 2001) … via attenuation of transcription (Winkler et al. , 2003)

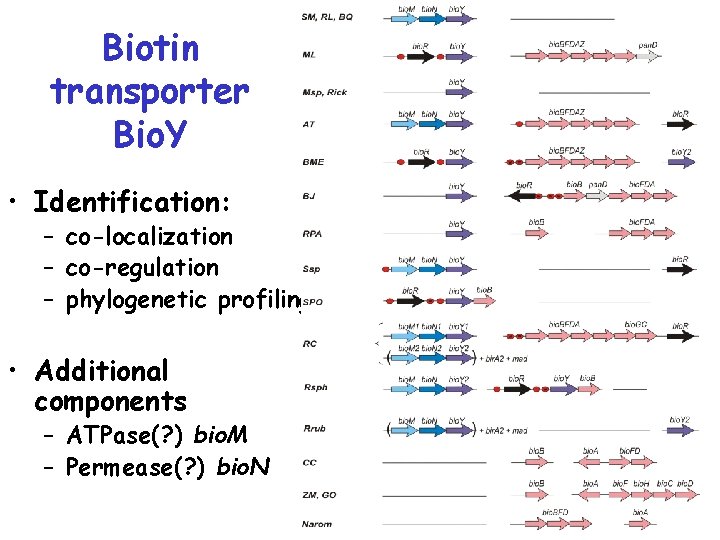
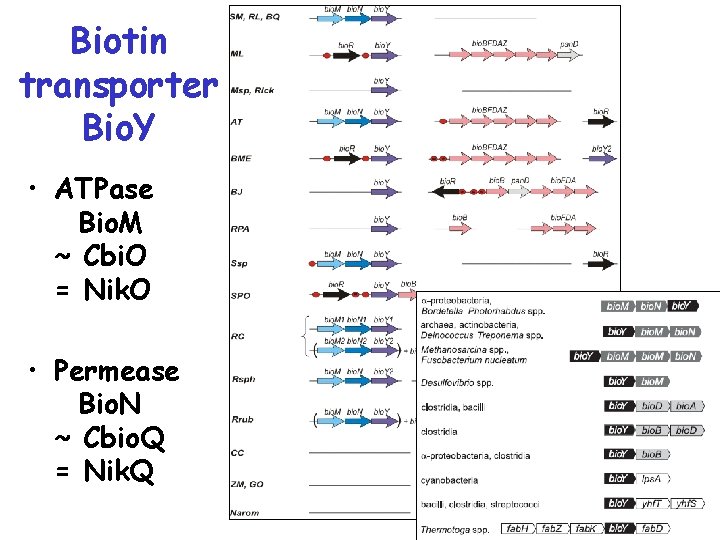
Biotin transporter Bio. Y • Identification: – co-localization – co-regulation – phylogenetic profiling • Additional components – ATPase(? ) bio. M – Permease(? ) bio. N

Thiamin biosynthesis = thi. N (confirmed) Transport of HMP Transport of HET (Gram-positive bacteria) (Gram-negative bacteria)

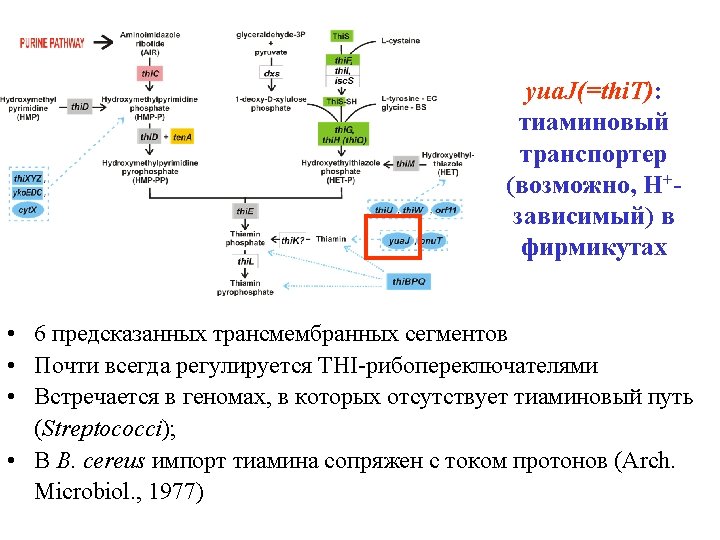
yua. J(=thi. T): thiamine transporter • 6 predicted TM-segments • Regulated by THI riboswitches • Streptococci: Thi. T, no thiamine pathway

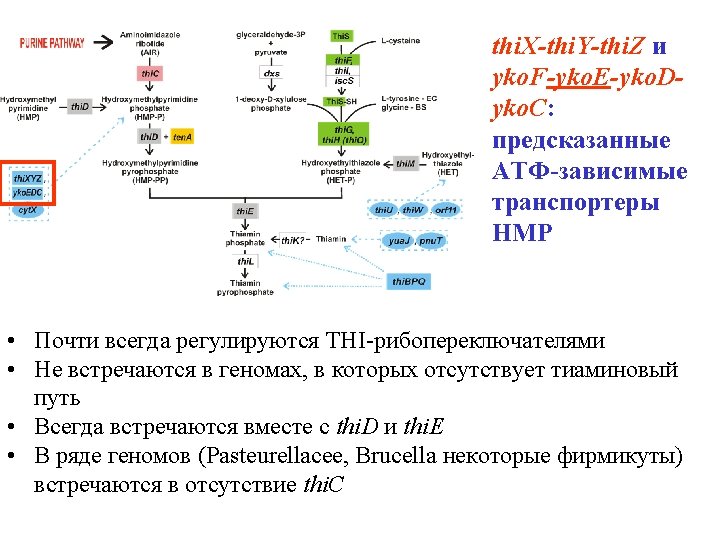
yko. FEDC: ATP-dependent HMP transporter • • Regulated by THI riboswitches Newer occurs in genomes lacking thiamine pathway Always co-occurs with thi. D and thi. E Sometimes occurs without thi. C

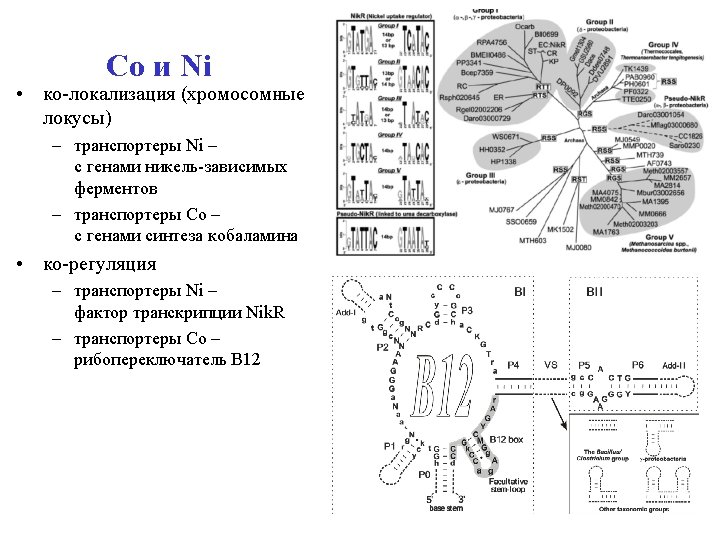
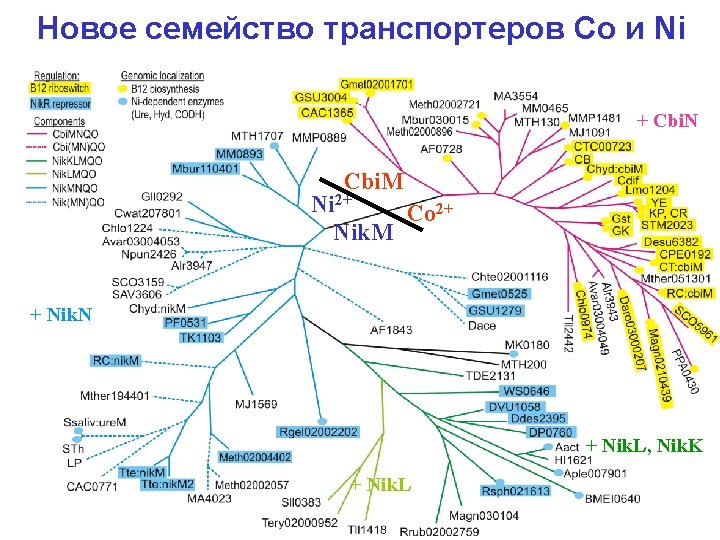
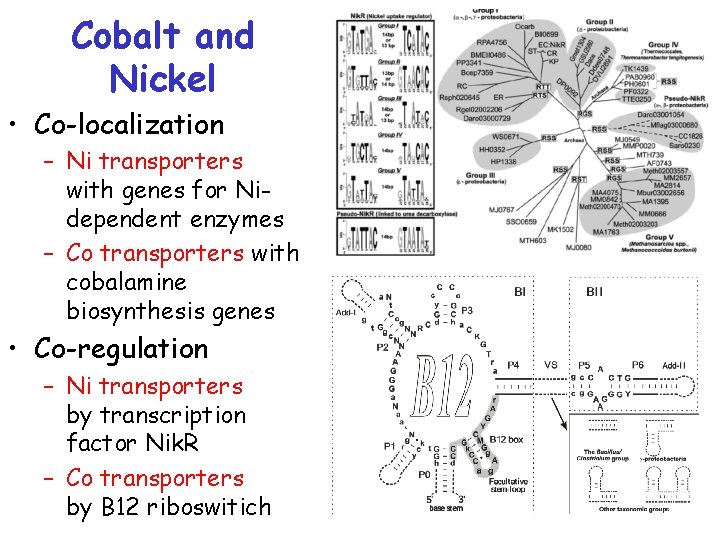
Cobalt and Nickel • Co-localization – Ni transporters with genes for Nidependent enzymes – Co transporters with cobalamine biosynthesis genes • Co-regulation – Ni transporters by transcription factor Nik. R – Co transporters by В 12 riboswitich

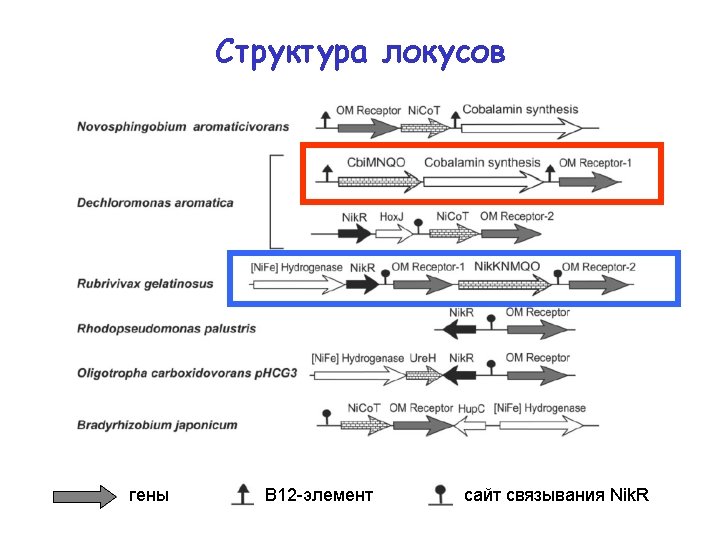
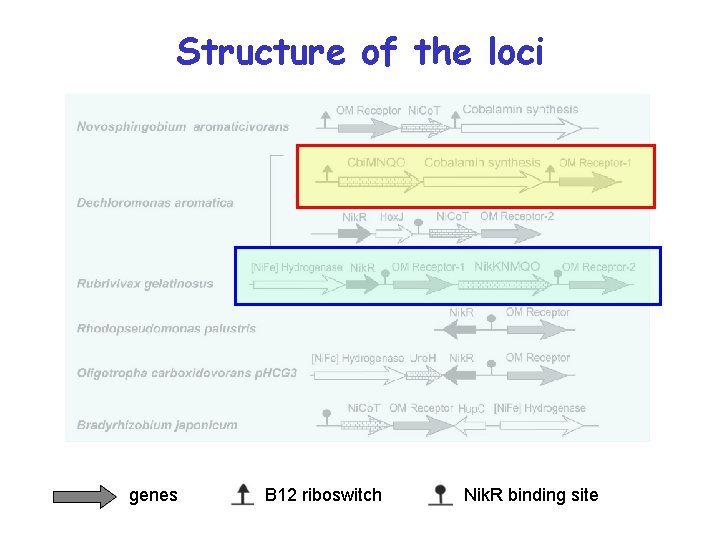
Structure of the loci genes B 12 riboswitch Nik. R binding site

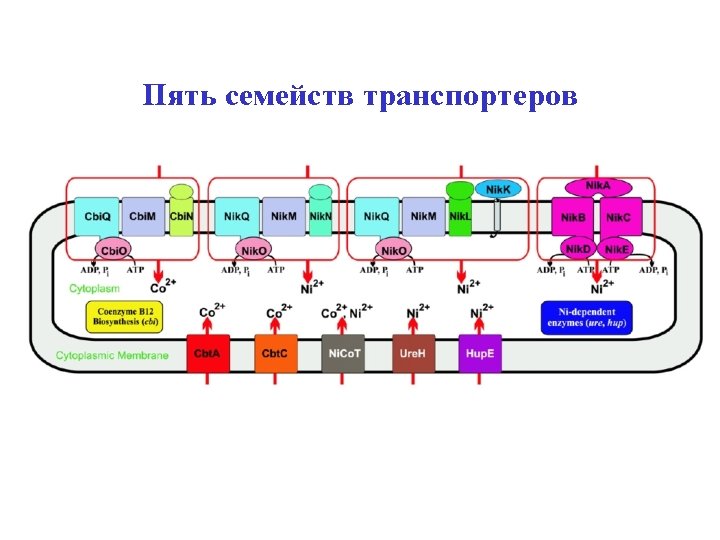
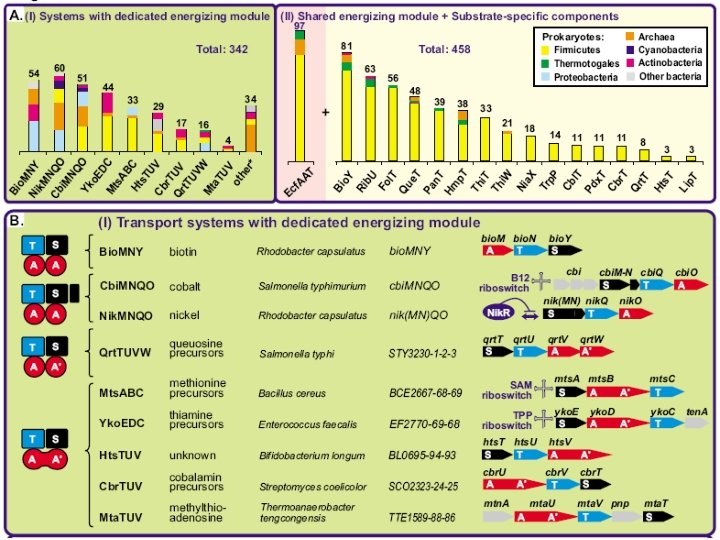
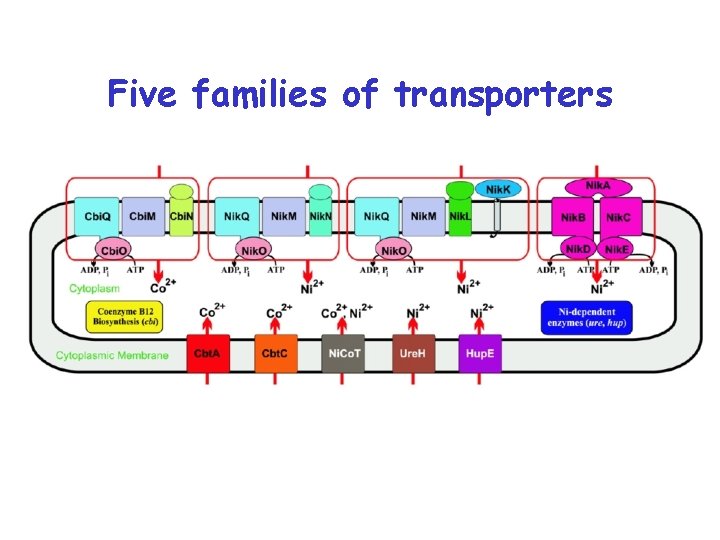
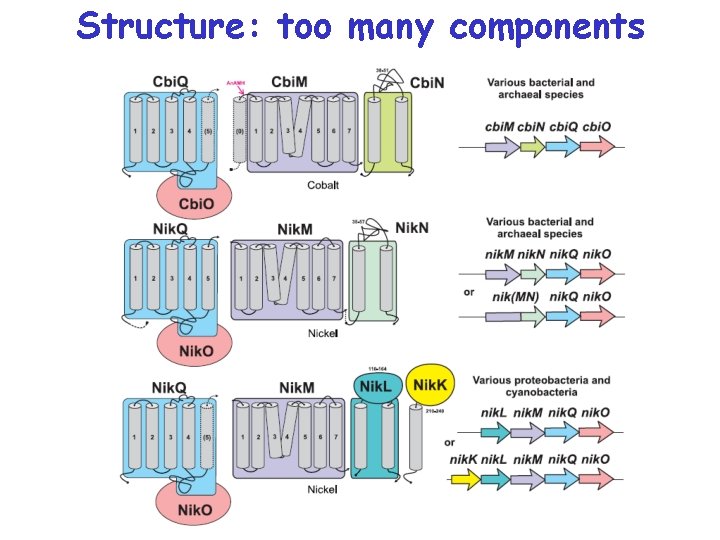
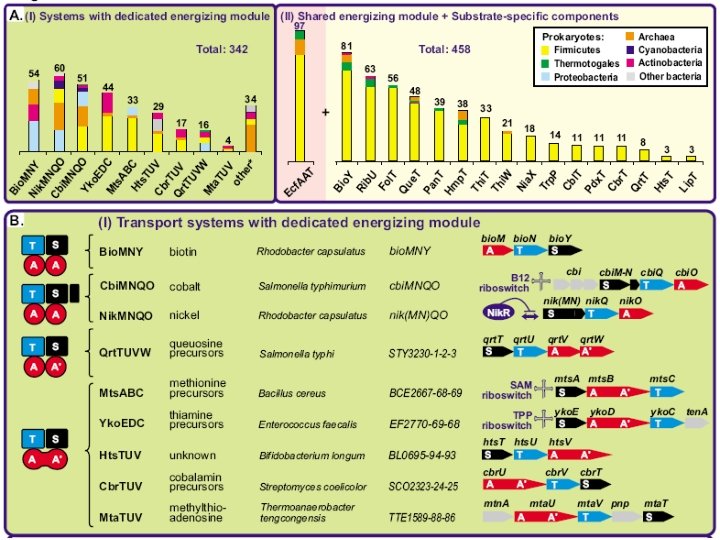
Five families of transporters

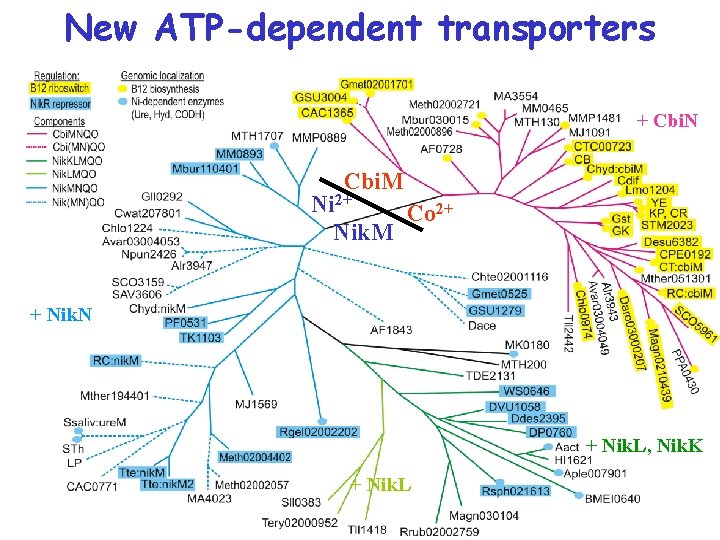
New ATP-dependent transporters + Cbi. N Cbi. M Ni 2+ Co 2+ Nik. M + Nik. N + Nik. L, Nik. K + Nik. L

Dmitry Rodionov Thomas Eitinger

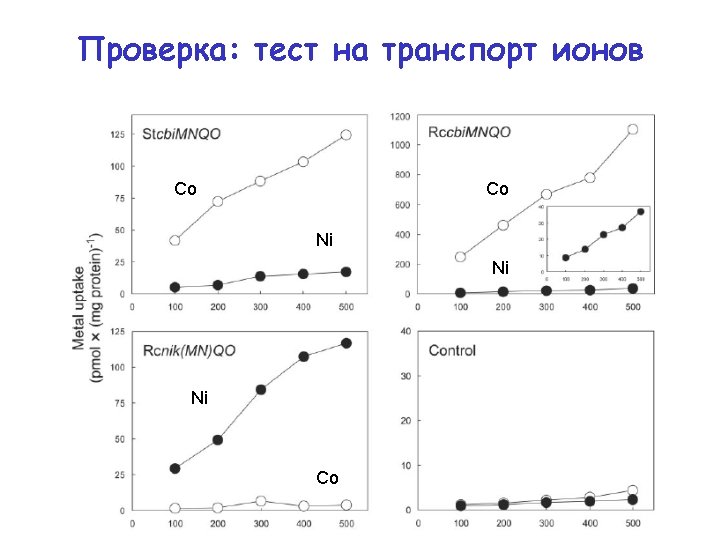
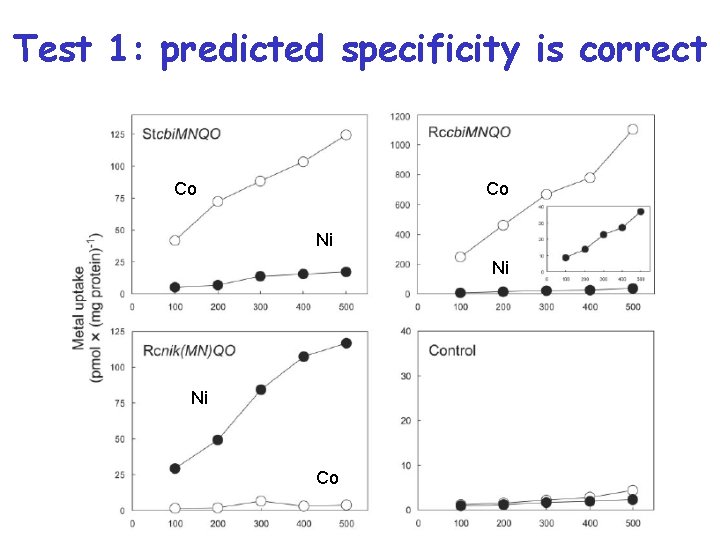
Test 1: predicted specificity is correct Co Co Ni Ni Ni Co

Structure: too many components

Biotin transporter Bio. Y • ATPase Bio. M ~ Cbi. O = Nik. O • Permease Bio. N ~ Cbio. Q = Nik. Q

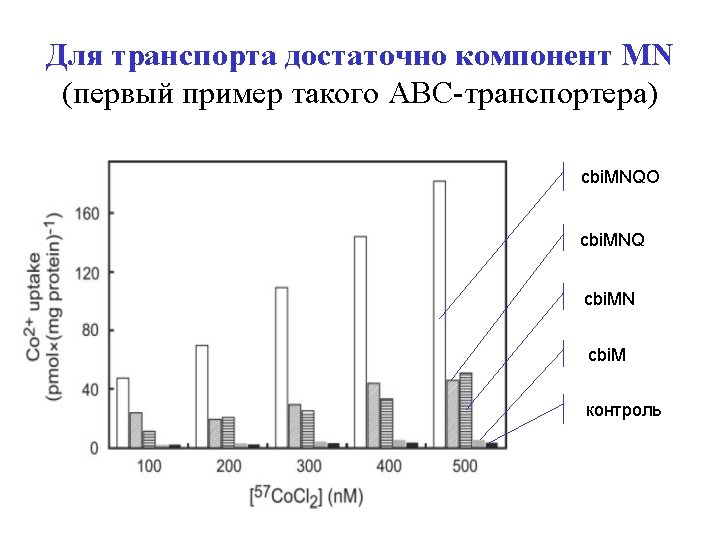
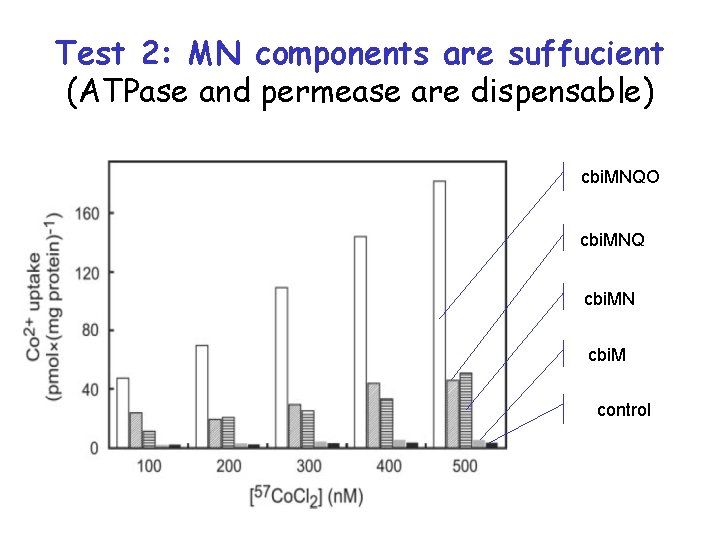
Test 2: MN components are suffucient (ATPase and permease are dispensable) cbi. MNQO cbi. MNQ cbi. MN cbi. M control

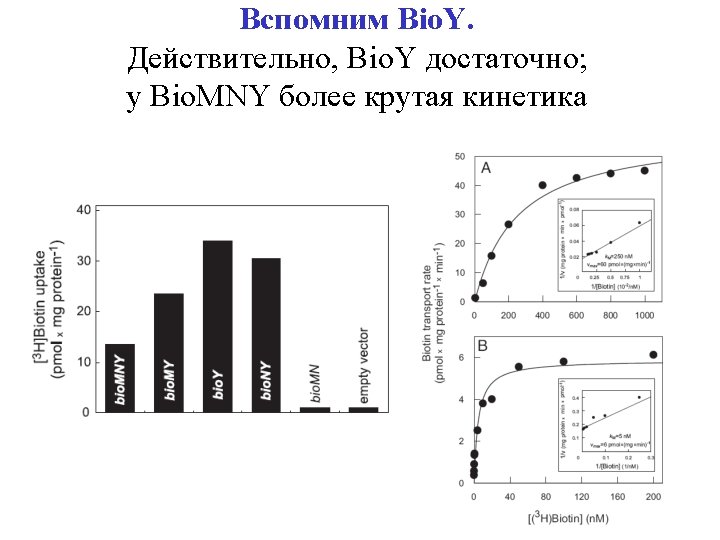
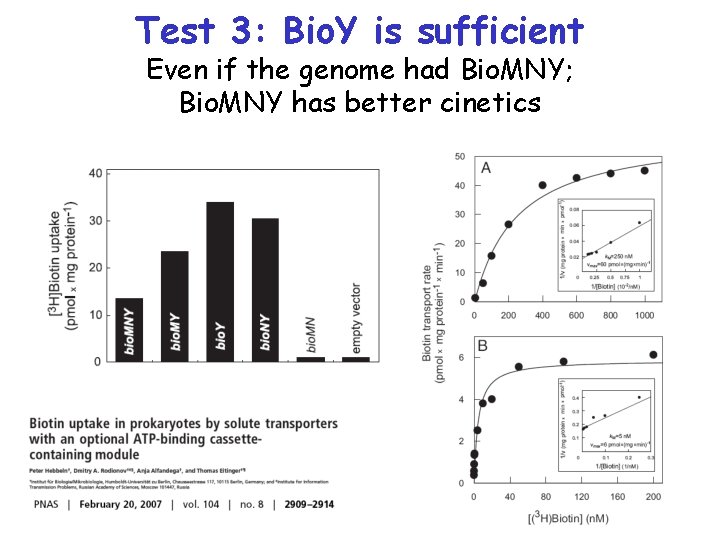
Test 3: Bio. Y is sufficient Even if the genome had Bio. MNY; Bio. MNY has better cinetics

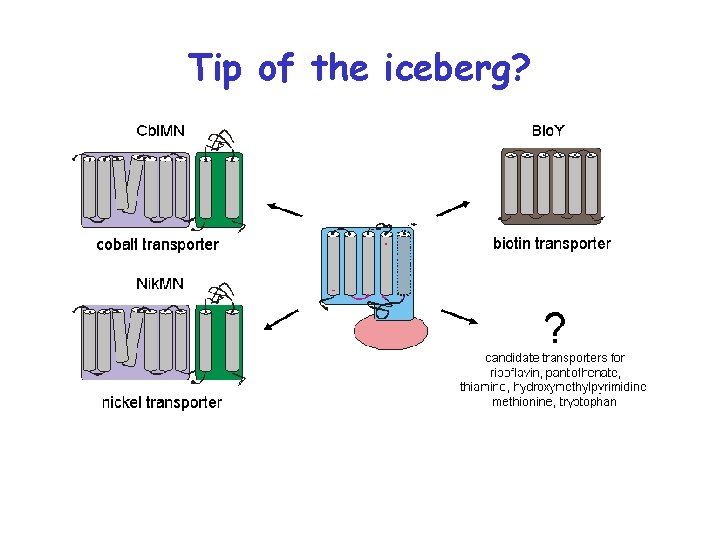
Tip of the iceberg?




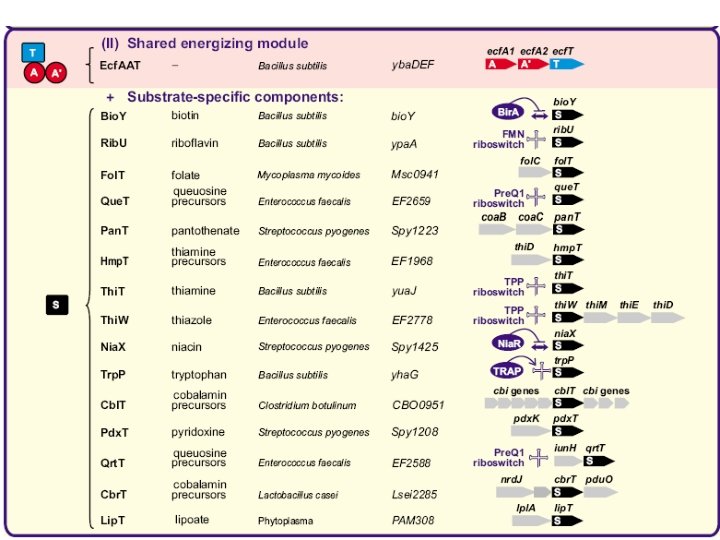
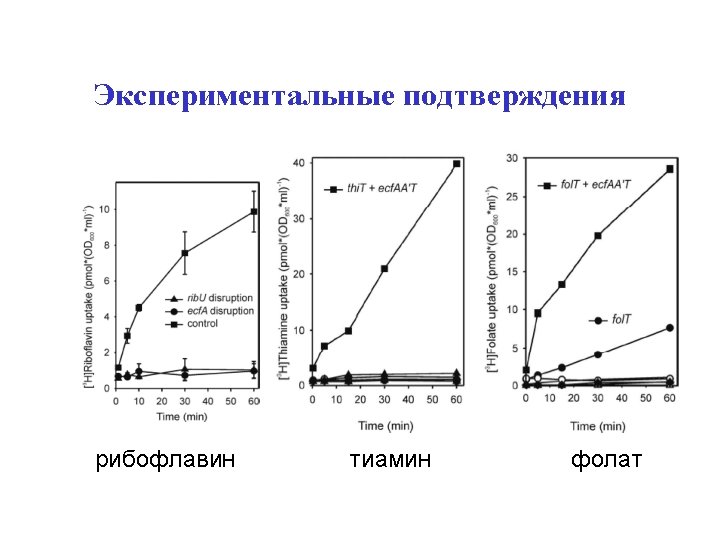
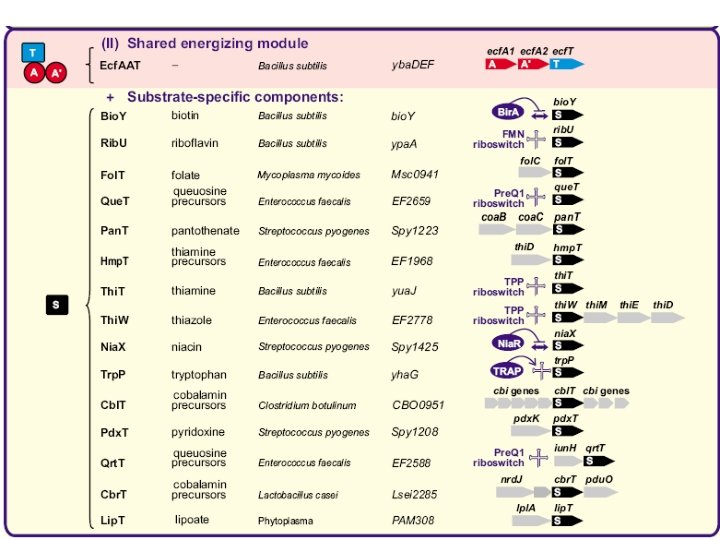
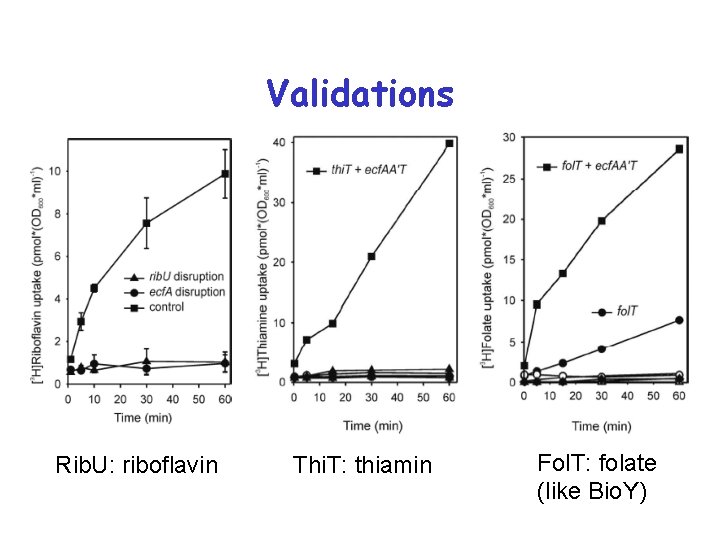
Validations Rib. U: riboflavin Thi. T: thiamin Fol. T: folate (like Bio. Y)

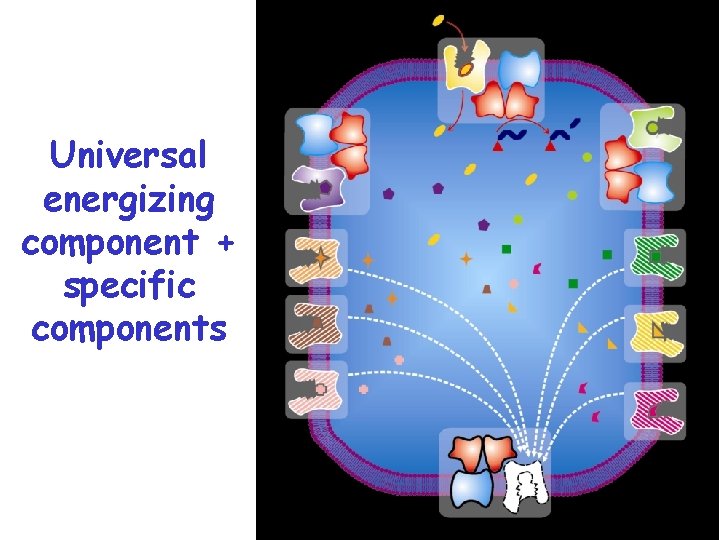
Universal energizing component + specific components




