Phylogeny II Parsimony ML SEMPHY Phylogenetic Tree branch

Phylogeny II : Parsimony, ML, SEMPHY .
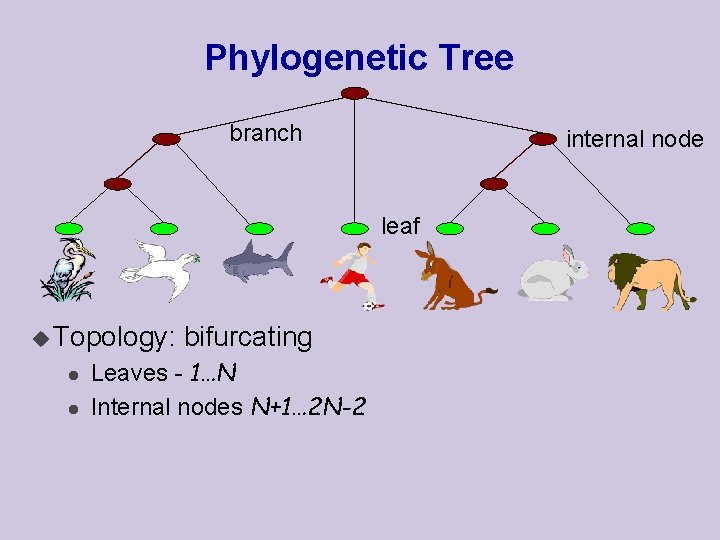
Phylogenetic Tree branch internal node leaf u Topology: l l bifurcating Leaves - 1…N Internal nodes N+1… 2 N-2

Character Based Methods u We start with a multiple alignments u Assumptions: l All sequences are homologous l Each position in alignment is homologous l Positions evolve independently l No gaps u Seek to explain the evolution of each position in the alignment

Parsimony u Character-based method Assumptions: u Independence of characters (no interactions) u Best tree is one where minimal changes take place
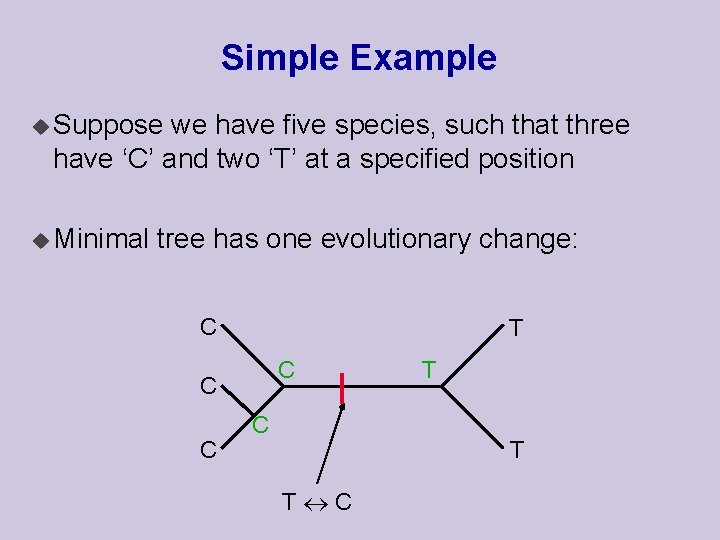
Simple Example u Suppose we have five species, such that three have ‘C’ and two ‘T’ at a specified position u Minimal tree has one evolutionary change: C T C C T T T C
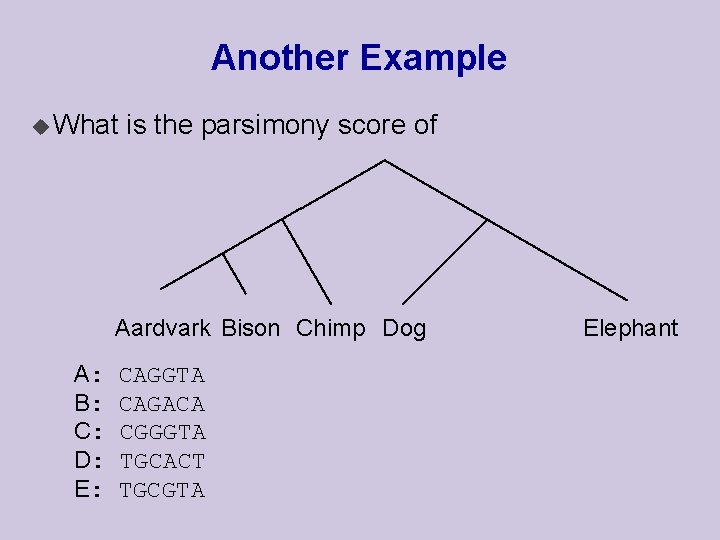
Another Example u What is the parsimony score of Aardvark Bison Chimp Dog A: B: C: D: E: CAGGTA CAGACA CGGGTA TGCACT TGCGTA Elephant

Evaluating Parsimony Scores u How do we compute the Parsimony score for a given tree? u Traditional Parsimony l Each base change has a cost of 1 u Weighted l Parsimony Each change is weighted by the score c(a, b)
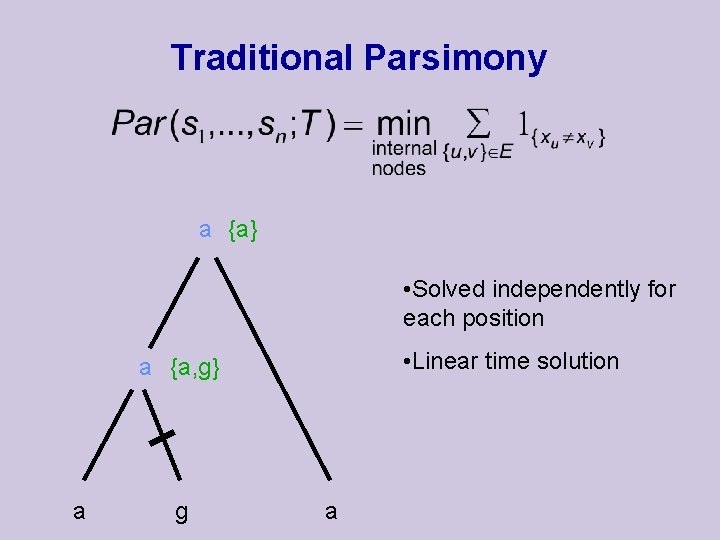
Traditional Parsimony a {a} • Solved independently for each position • Linear time solution a {a, g} a g a

Evaluating Weighted Parsimony Dynamic programming on the tree Initialization: u For each leaf i set S(i, a) = 0 if i is labeled by a, otherwise S(i, a) = Iteration: u if k is node with children i and j, then S(k, a) = minb(S(i, b)+c(a, b)) + minb(S(j, b)+c(a, b)) Termination: u cost of tree is mina. S(r, a) where r is the root

Cost of Evaluating Parsimony u Score is evaluated on each position independetly. Scores are then summed over all positions. u If there are n nodes, m characters, and k possible values for each character, then complexity is O(nmk) u By keeping traceback information, we can reconstruct most parsimonious values at each ancestor node
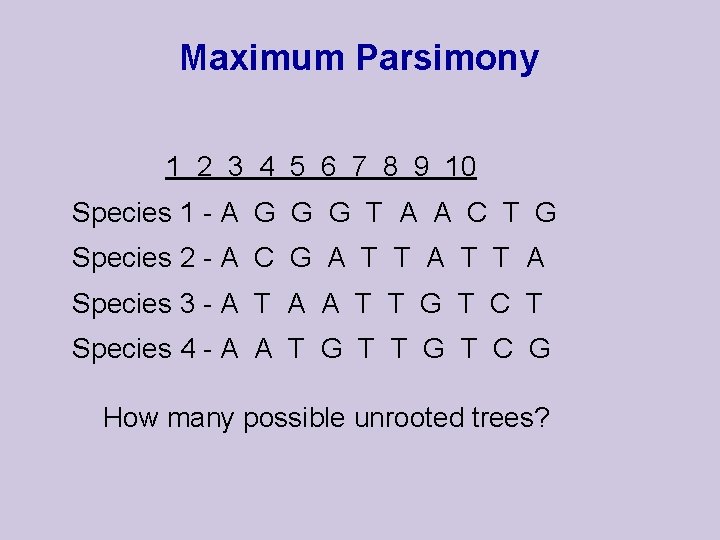
Maximum Parsimony 1 2 3 4 5 6 7 8 9 10 Species 1 - A G G G T A A C T G Species 2 - A C G A T T A Species 3 - A T A A T T G T C T Species 4 - A A T G T C G How many possible unrooted trees?
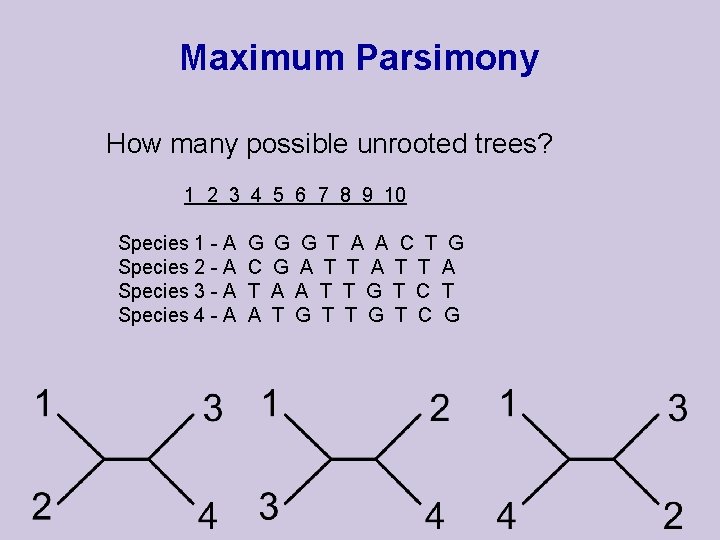
Maximum Parsimony How many possible unrooted trees? 1 2 3 4 5 6 7 8 9 10 Species 1 - A Species 2 - A Species 3 - A Species 4 - A G C T A G G T A A C T G G A T T A A A T T G T C G
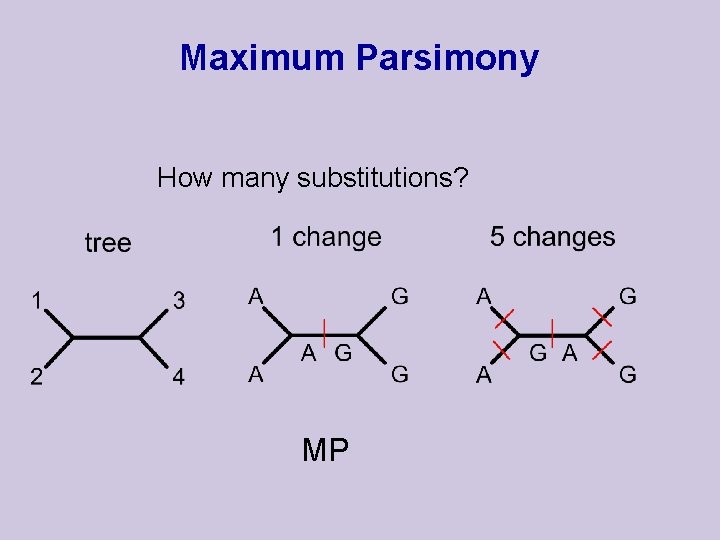
Maximum Parsimony How many substitutions? MP
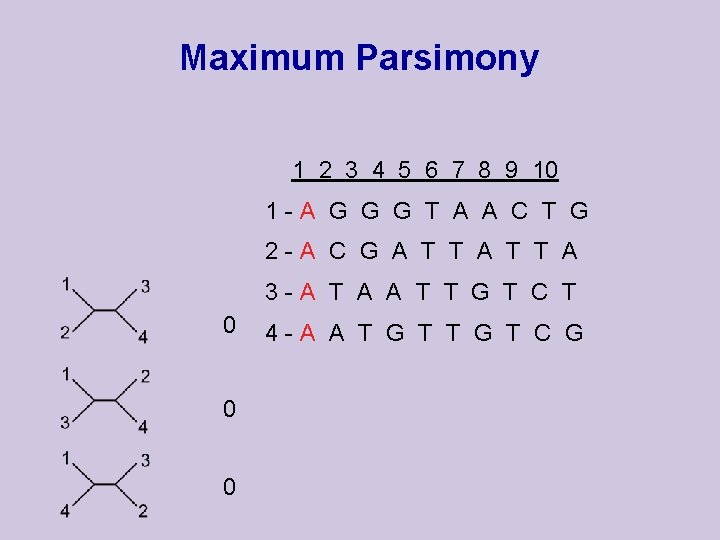
Maximum Parsimony 1 2 3 4 5 6 7 8 9 10 1 -A G G G T A A C T G 2 -A C G A T T A 3 -A T A A T T G T C T 0 0 0 4 -A A T G T C G
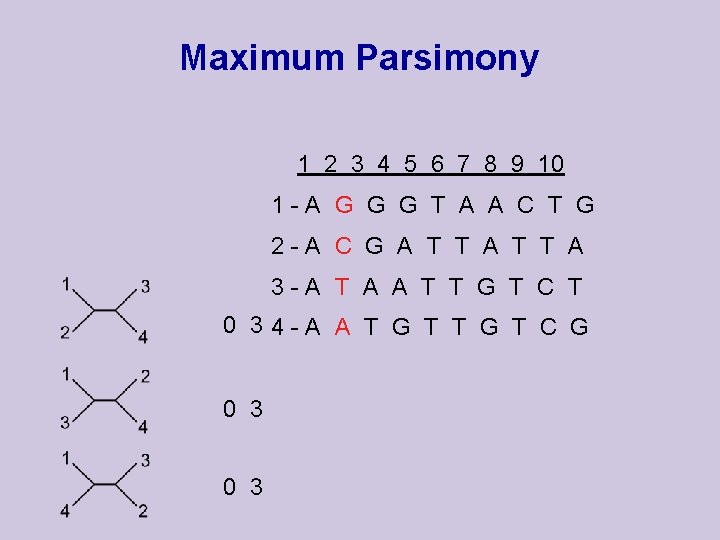
Maximum Parsimony 1 2 3 4 5 6 7 8 9 10 1 -A G G G T A A C T G 2 -A C G A T T A 3 -A T A A T T G T C T 0 34 -A A T G T C G 0 3
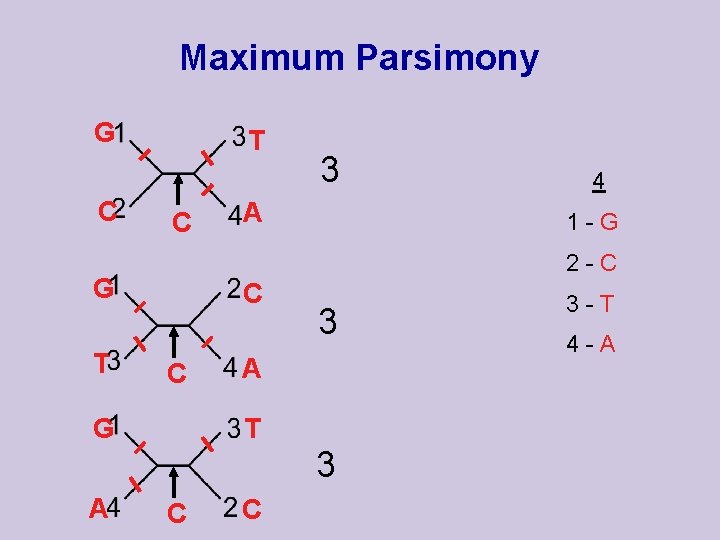
Maximum Parsimony G C T C A C C G 1 -G 3 A T 3 A 4 2 -C G T 3 C C 3 -T 4 -A
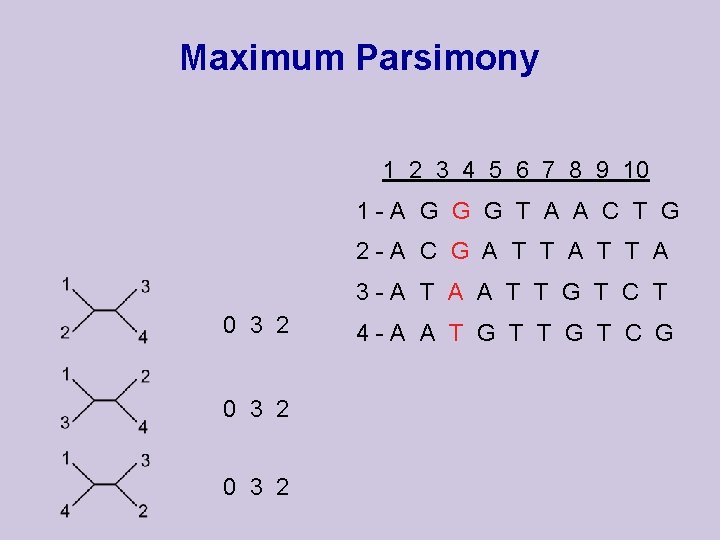
Maximum Parsimony 1 2 3 4 5 6 7 8 9 10 1 -A G G G T A A C T G 2 -A C G A T T A 3 -A T A A T T G T C T 0 3 2 4 -A A T G T C G
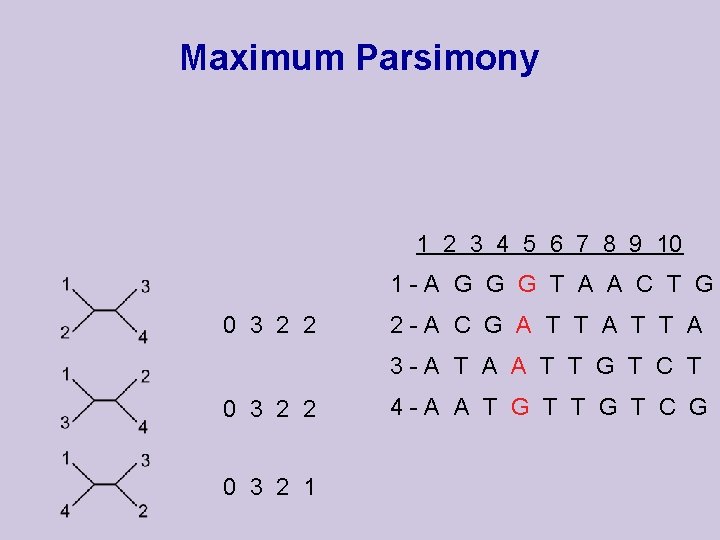
Maximum Parsimony 1 2 3 4 5 6 7 8 9 10 1 -A G G G T A A C T G 0 3 2 2 2 -A C G A T T A 3 -A T A A T T G T C T 0 3 2 2 0 3 2 1 4 -A A T G T C G
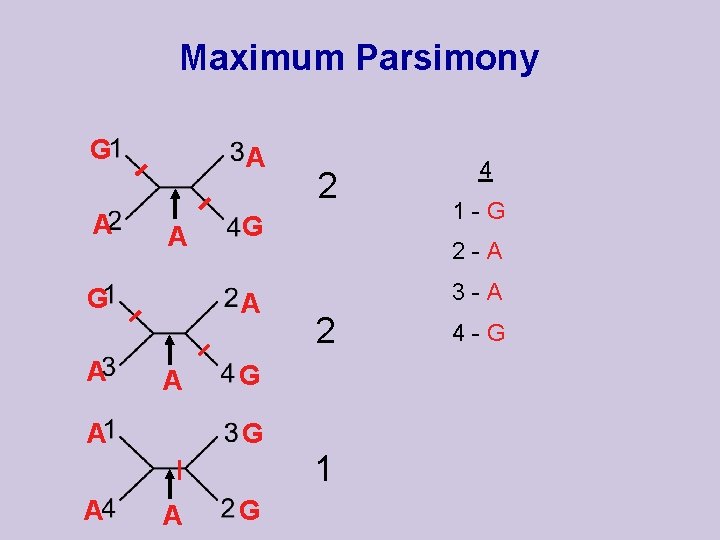
Maximum Parsimony G A A A G 1 -G 2 -A 3 -A 2 G G A 2 4 1 4 -G
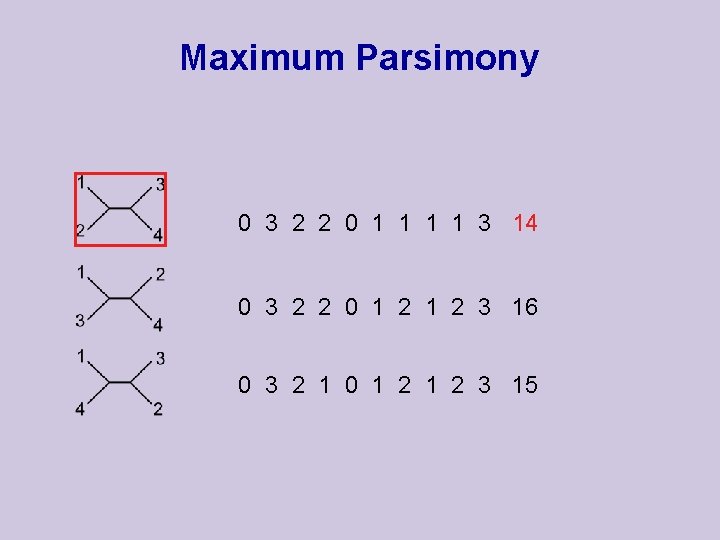
Maximum Parsimony 0 3 2 2 0 1 1 3 14 0 3 2 2 0 1 2 3 16 0 3 2 1 0 1 2 3 15
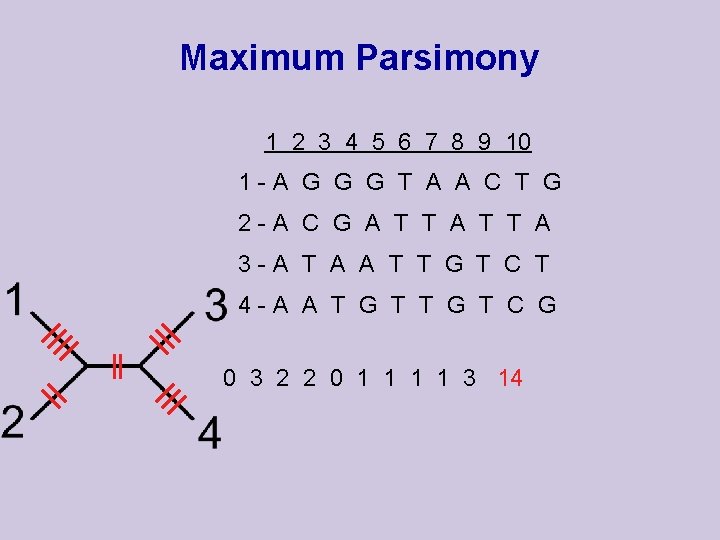
Maximum Parsimony 1 2 3 4 5 6 7 8 9 10 1 -A G G G T A A C T G 2 -A C G A T T A 3 -A T A A T T G T C T 4 -A A T G T C G 0 3 2 2 0 1 1 3 14
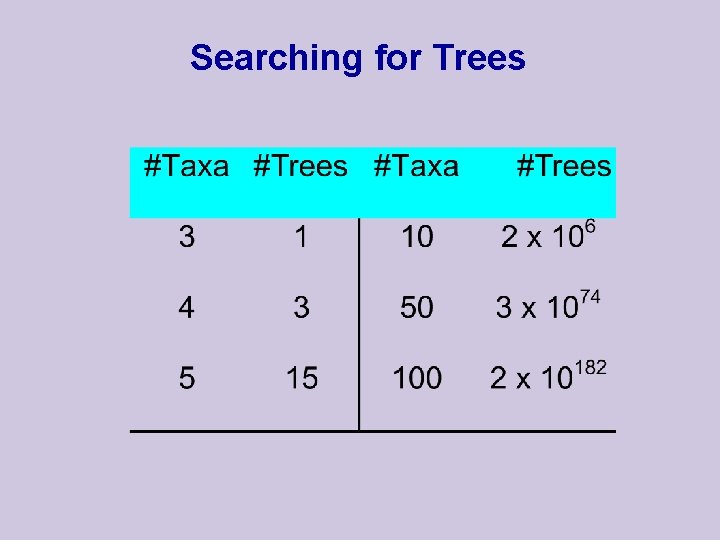
Searching for Trees

Searching for the Optimal Tree u Exhaustive Search l Very intensive u Branch and Bound l A compromise u Heuristic l Fast l Usually starts with NJ
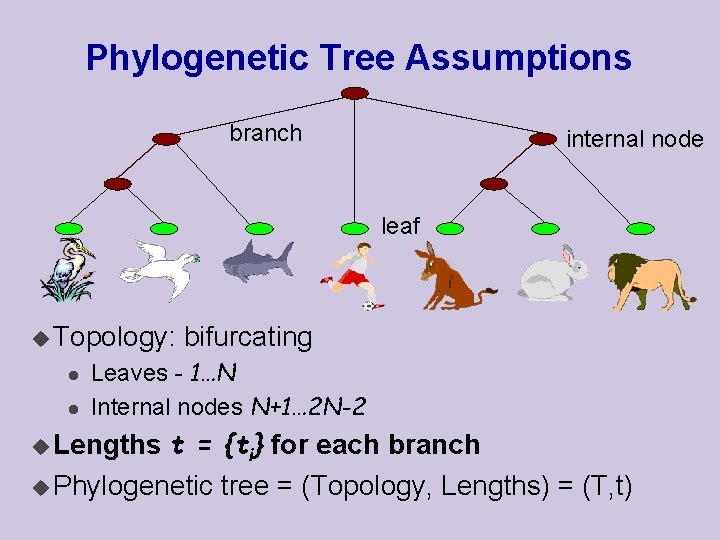
Phylogenetic Tree Assumptions branch internal node leaf u Topology: l l bifurcating Leaves - 1…N Internal nodes N+1… 2 N-2 t = {ti} for each branch u Phylogenetic tree = (Topology, Lengths) = (T, t) u Lengths

Probabilistic Methods u The phylogenetic tree represents a generative probabilistic model (like HMMs) for the observed sequences. u Background probabilities: q(a) u Mutation probabilities: P(a|b, t) u Models for evolutionary mutations l Jukes Cantor l Kimura 2 -parameter model u Such models are used to derive the probabilities

Jukes Cantor model u. A model for mutation rates • Mutation occurs at a constant rate • Each nucleotide is equally likely to mutate into any other nucleotide with rate a.
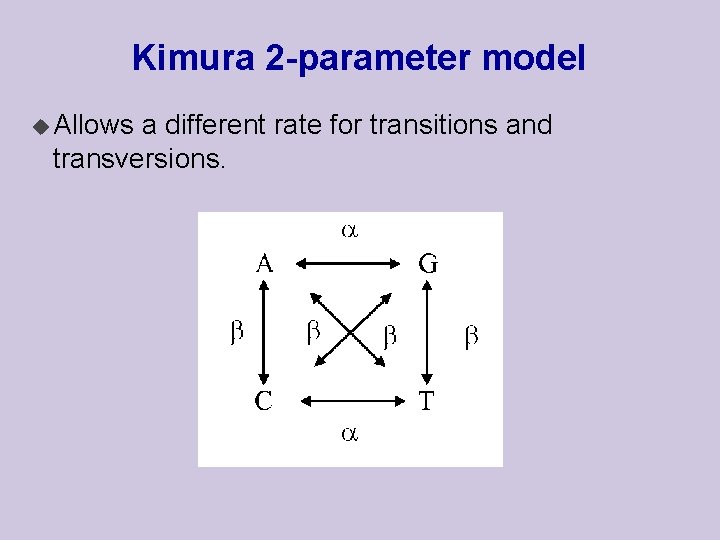
Kimura 2 -parameter model u Allows a different rate for transitions and transversions.

Mutation Probabilities u The rate matrix R is used to derive the mutation probability matrix S: u S is obtained by integration. For Jukes Cantor: uq can be obtained by setting t to infinity
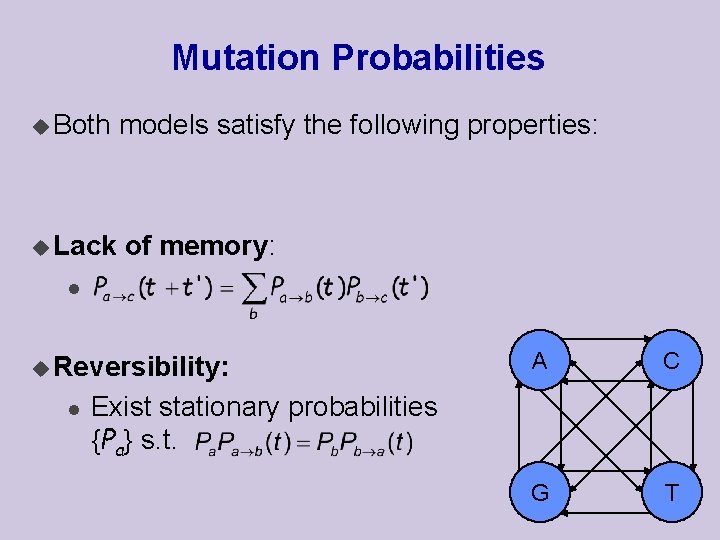
Mutation Probabilities u Both u Lack models satisfy the following properties: of memory: l u Reversibility: l A C G T Exist stationary probabilities {Pa} s. t.
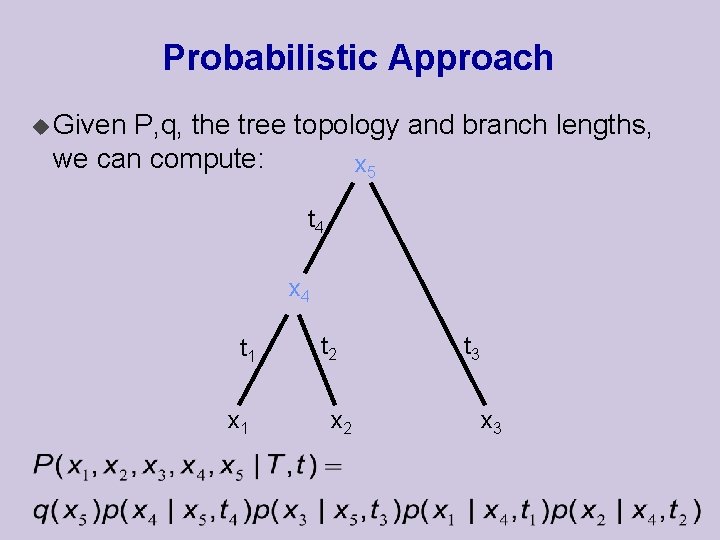
Probabilistic Approach u Given P, q, the tree topology and branch lengths, we can compute: x 5 t 4 x 4 t 1 x 1 t 2 x 2 t 3 x 3

Computing the Tree Likelihood u We are interested in the probability of observed data given tree and branch “lengths”: u Computed by summing over internal nodes u This can be done efficiently using a tree upward traversal pass.

Tree Likelihood Computation u Define P(Lk|a)= prob. of leaves below node k given that xk=a u Init: for leaves: P(Lk|a)=1 if xk=a ; 0 otherwise u Iteration: if k is node with children i and j, then u Termination: Likelihood is

Maximum Likelihood (ML) u Score each tree by l Assumption of independent positions u Branch lengths t can be optimized l Gradient ascent l EM u We look for the highest scoring tree l Exhaustive l Sampling methods (Metropolis)
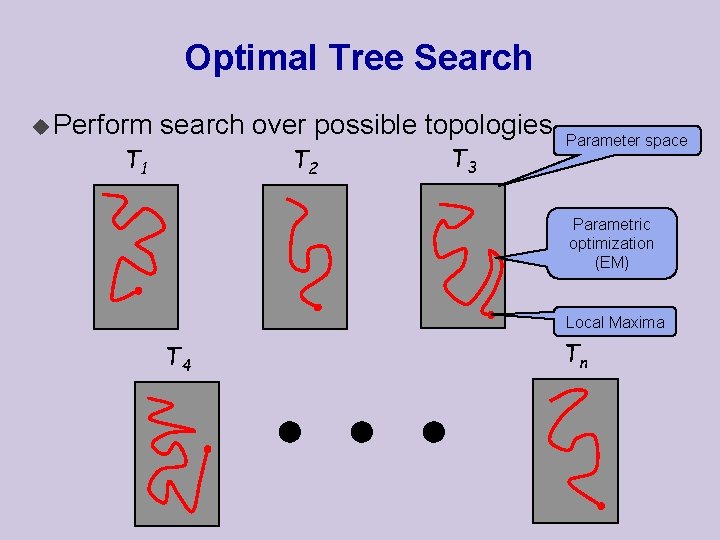
Optimal Tree Search u Perform search over possible topologies T 1 T 2 T 3 Parameter space Parametric optimization (EM) Local Maxima T 4 Tn

Computational Problem u Such procedures are computationally expensive! u Computation of optimal parameters, per candidate, requires non-trivial optimization step. u Spend non-negligible computation on a candidate, even if it is a low scoring one. u In practice, such learning procedures can only consider small sets of candidate structures

Structural EM Idea: Use parameters found for current topology to help evaluate new topologies. Outline: u Perform search in (T, t) space. u Use EM-like iterations: l E-step: use current solution to compute expected sufficient statistics for all topologies l M-step: select new topology based on these expected sufficient statistics

The Complete-Data Scenario Suppose we observe H, the ancestral sequences. Define: F is a linear function of Si, j is a matrix of # of co-occurrences for each pair (a, b) in the taxa i, j Find: topology T that maximizes

Expected Likelihood with a tree (T 0, t 0) u Compute u Start Formal justification: u Define: Theorem: Consequence: improvement in expected score improvement in likelihood
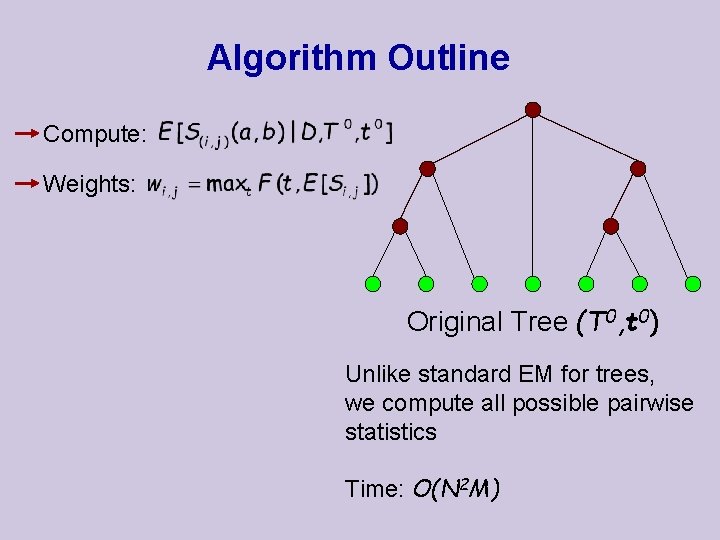
Algorithm Outline Compute: Weights: Original Tree (T 0, t 0) Unlike standard EM for trees, we compute all possible pairwise statistics Time: O(N 2 M)
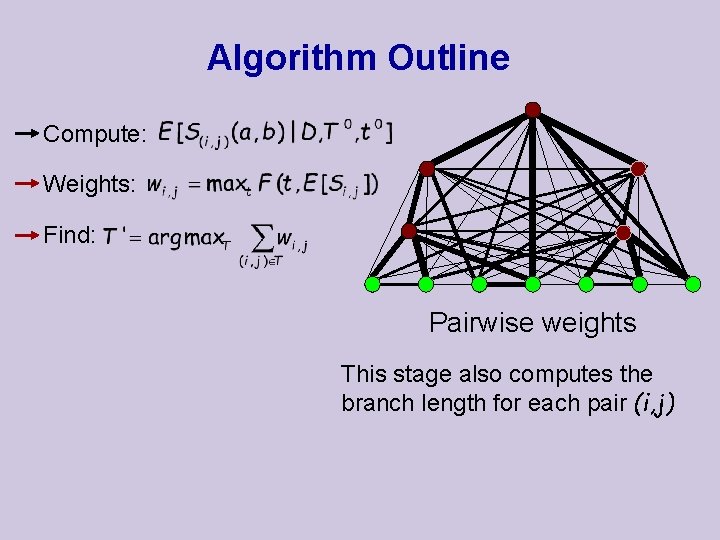
Algorithm Outline Compute: Weights: Find: Pairwise weights This stage also computes the branch length for each pair (i, j)
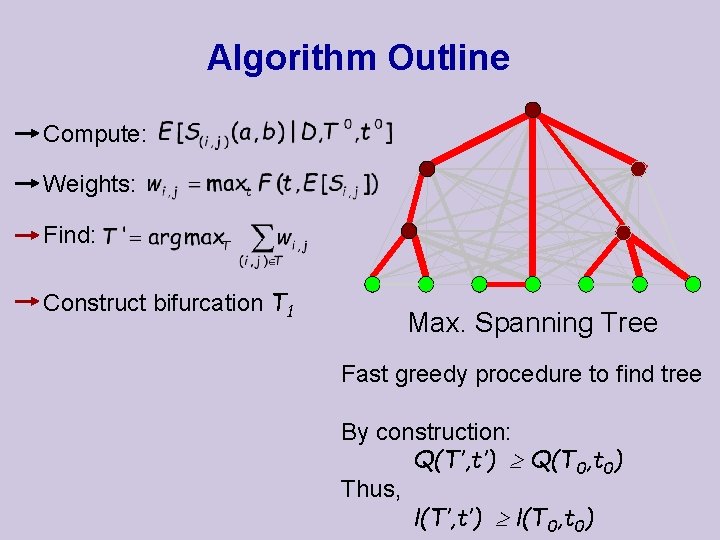
Algorithm Outline Compute: Weights: Find: Construct bifurcation T 1 Max. Spanning Tree Fast greedy procedure to find tree By construction: Q(T’, t’) Q(T 0, t 0) Thus, l(T’, t’) l(T 0, t 0)
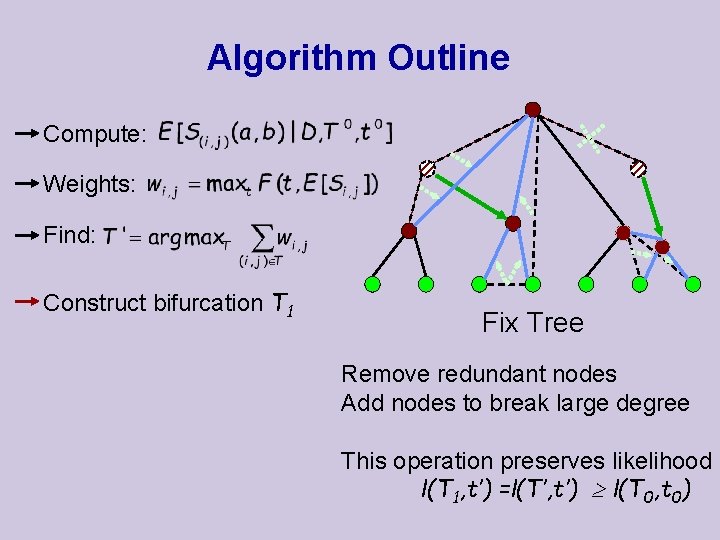
Algorithm Outline Compute: Weights: Find: Construct bifurcation T 1 Fix Tree Remove redundant nodes Add nodes to break large degree This operation preserves likelihood l(T 1, t’) =l(T’, t’) l(T 0, t 0)

Assessing trees: the Bootstrap u Often we don’t trust the tree found as the “correct” one. u Bootstrapping: l Sample (with replacement) n positions from the alignment l Learn the best tree for each sample l Look for tree features which are frequent in all trees. u For some models this procedure approximates the tree posterior P(T| X 1, …, Xn)
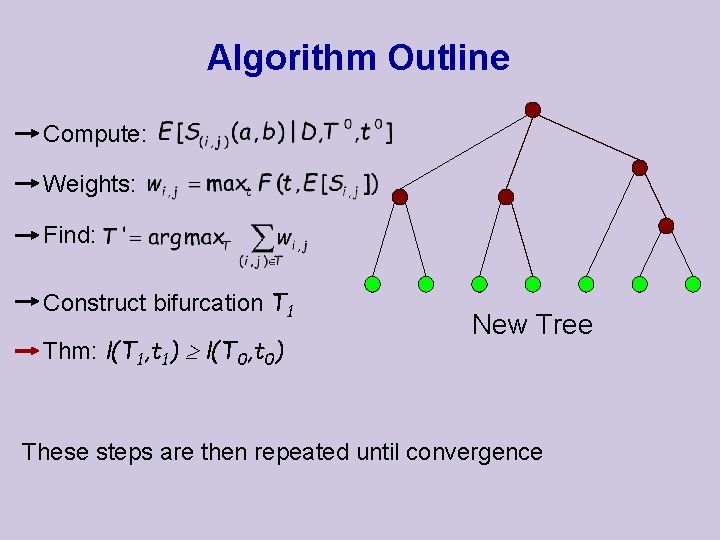
Algorithm Outline Compute: Weights: Find: Construct bifurcation T 1 Thm: l(T 1, t 1) l(T 0, t 0) New Tree These steps are then repeated until convergence
- Slides: 44