Chapter 26 Phylogeny and the Tree of Life








- Slides: 27

Chapter 26 Phylogeny and the Tree of Life

What you need to know: • The taxonomic categories and how they indicate relatedness. • How systematics is used to develop phylogenetic trees. • The three domains of life including their similarities and their differences.

Systematics: classifying organisms and determining their evolutionary relationships Taxonomy (classification) Systematics Phylogenetics (evolutionary history)

Tools used to determine evolutionary relationships: 1. Fossils 2. Morphology (homologous structures) 3. Molecular evidence (DNA, amino acids) Who is more closely related? Animals and fungi are more closely related than either is to plants.

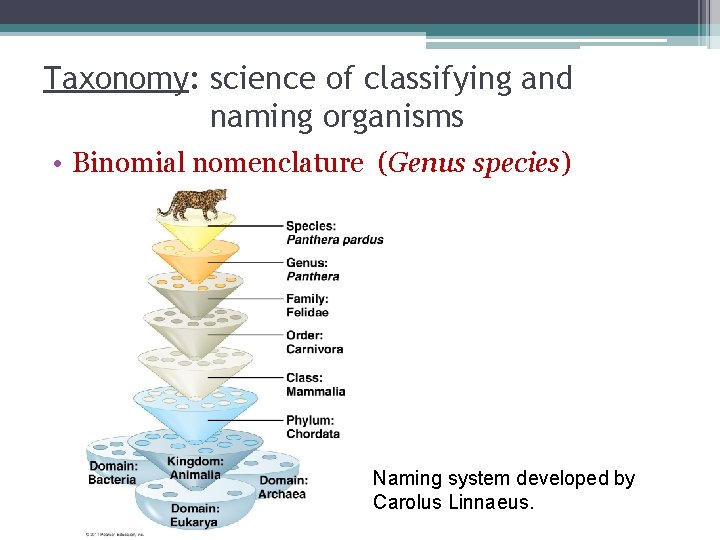
Taxonomy: science of classifying and naming organisms • Binomial nomenclature (Genus species) Naming system developed by Carolus Linnaeus.

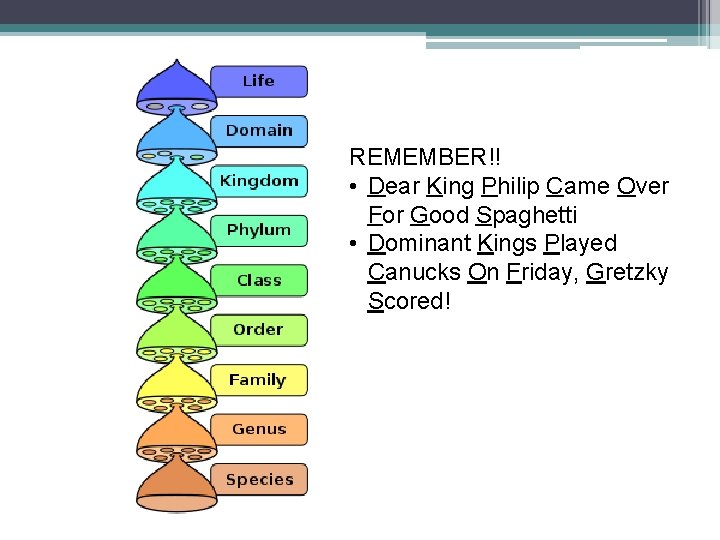
REMEMBER!! • Dear King Philip Came Over For Good Spaghetti • Dominant Kings Played Canucks On Friday, Gretzky Scored!

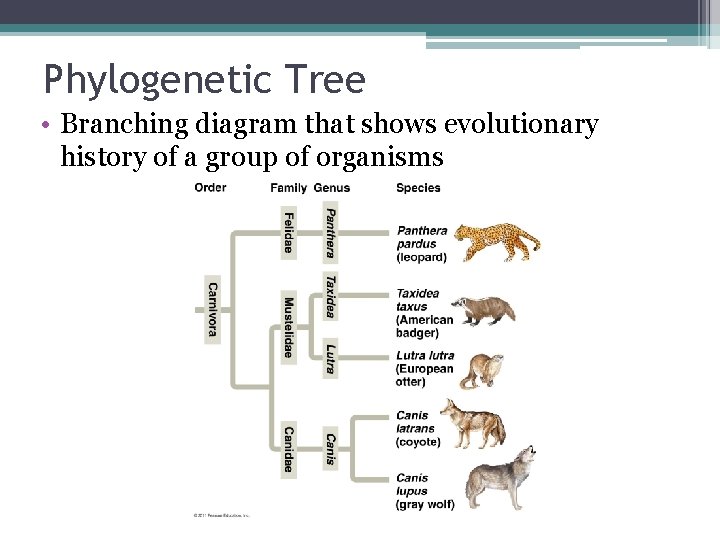
Phylogenetic Tree • Branching diagram that shows evolutionary history of a group of organisms

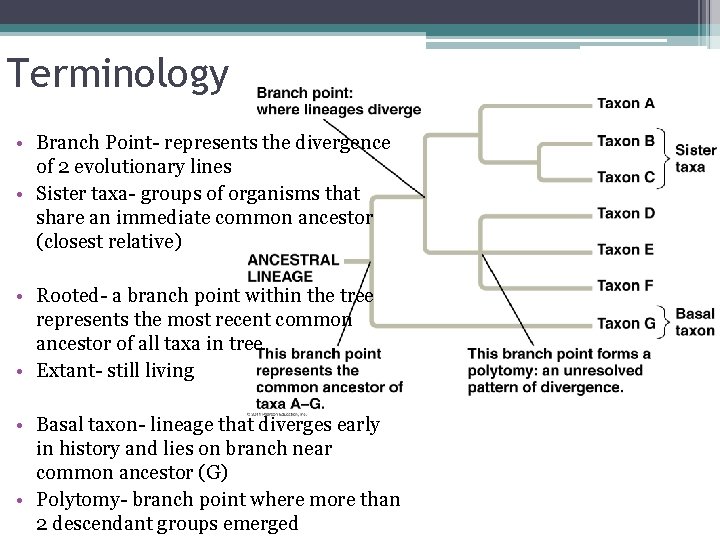
Terminology • Branch Point- represents the divergence of 2 evolutionary lines • Sister taxa- groups of organisms that share an immediate common ancestor (closest relative) • Rooted- a branch point within the tree represents the most recent common ancestor of all taxa in tree • Extant- still living • Basal taxon- lineage that diverges early in history and lies on branch near common ancestor (G) • Polytomy- branch point where more than 2 descendant groups emerged

3 points about Phylogenetic Trees 1) Intended to show patterns of descent not phenotypic similarities Ex) Crocodiles most closely related to birds than lizards but look more like lizards

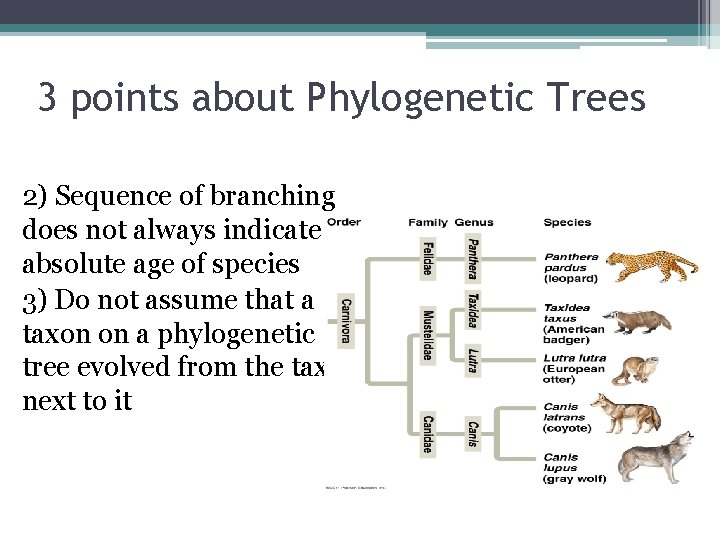
3 points about Phylogenetic Trees 2) Sequence of branching does not always indicate absolute age of species 3) Do not assume that a taxon on a phylogenetic tree evolved from the taxon next to it

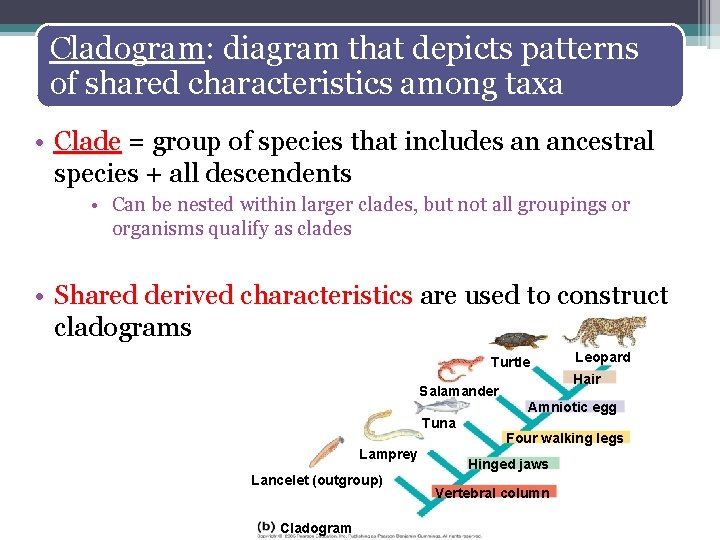
Cladogram: diagram that depicts patterns of shared characteristics among taxa • Clade = group of species that includes an ancestral species + all descendents • Can be nested within larger clades, but not all groupings or organisms qualify as clades • Shared derived characteristics are used to construct cladograms Turtle Leopard Hair Salamander Amniotic egg Tuna Lamprey Lancelet (outgroup) Cladogram Four walking legs Hinged jaws Vertebral column

Taxon is equivalent to a clade if it is monophyletic • Consists of an ancestral species and ALL its descendants

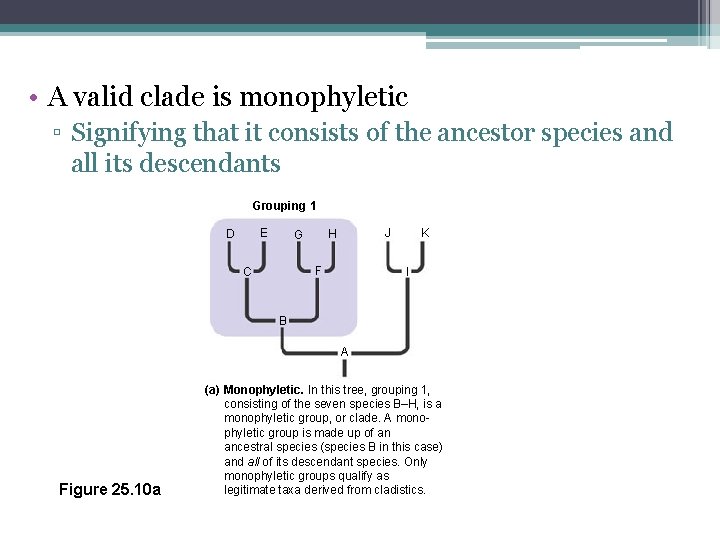
• A valid clade is monophyletic ▫ Signifying that it consists of the ancestor species and all its descendants Grouping 1 E D J H G F C K I B A Figure 25. 10 a (a) Monophyletic. In this tree, grouping 1, consisting of the seven species B–H, is a monophyletic group, or clade. A monophyletic group is made up of an ancestral species (species B in this case) and all of its descendant species. Only monophyletic groups qualify as legitimate taxa derived from cladistics.

Paraphyletic Group • Consists of an ancestral species and some, but not all, of its descendants

• A paraphyletic clade ▫ Is a grouping that consists of an ancestral species and some, but not all, of the descendants Grouping 2 G E D C J H K I F B A Figure 25. 10 b (b) Paraphyletic. Grouping 2 does not meet the cladistic criterion: It is paraphyletic, which means that it consists of an ancestor (A in this case) and some, but not all, of that ancestor’s descendants. (Grouping 2 includes the descendants I, J, and K, but excludes B–H, which also descended from A. )

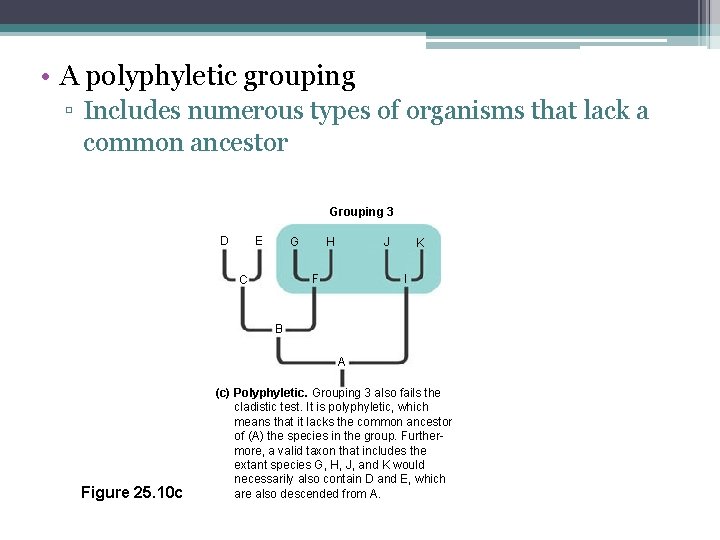
Polyphyletic Group • Includes distantly related species but does not include their most recent common ancestor

• A polyphyletic grouping ▫ Includes numerous types of organisms that lack a common ancestor Grouping 3 D E G J H I F C K B A Figure 25. 10 c (c) Polyphyletic. Grouping 3 also fails the cladistic test. It is polyphyletic, which means that it lacks the common ancestor of (A) the species in the group. Furthermore, a valid taxon that includes the extant species G, H, J, and K would necessarily also contain D and E, which are also descended from A.

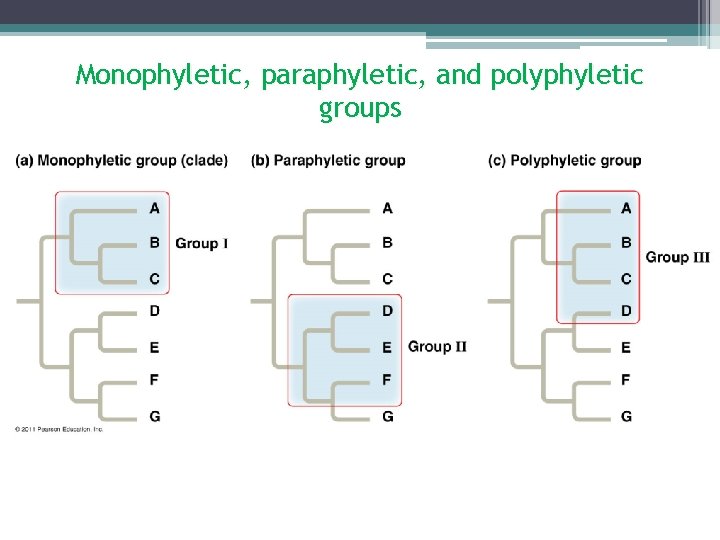
Monophyletic, paraphyletic, and polyphyletic groups

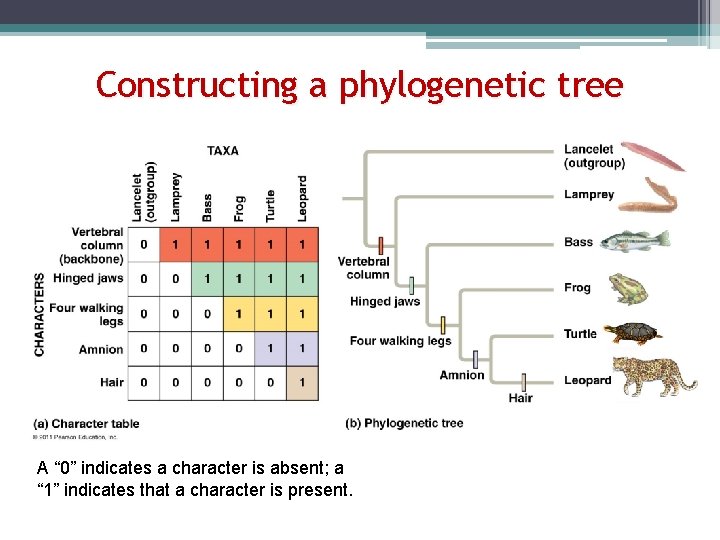
Constructing a phylogenetic tree A “ 0” indicates a character is absent; a “ 1” indicates that a character is present.

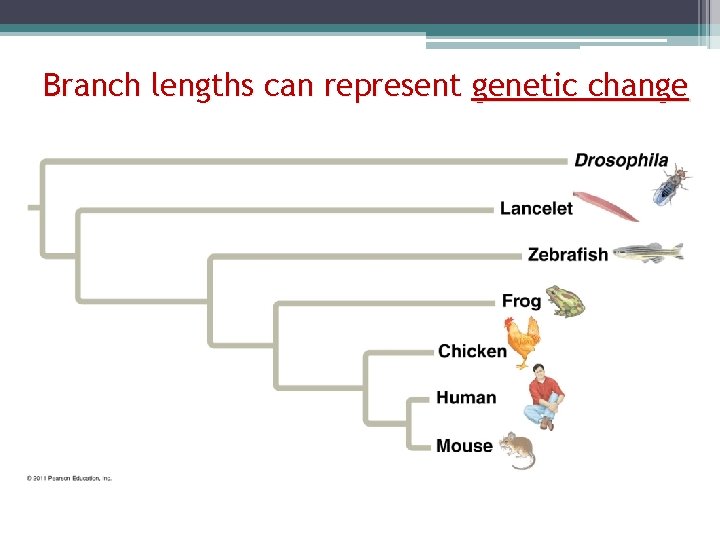
Branch lengths can represent genetic change

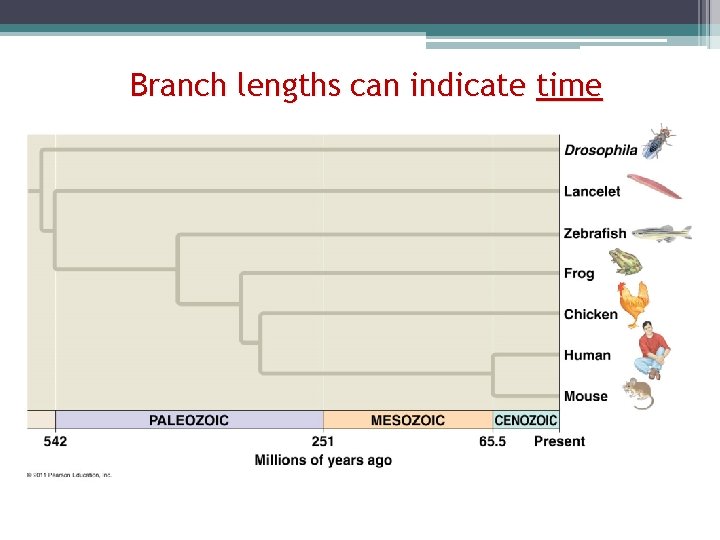
Branch lengths can indicate time

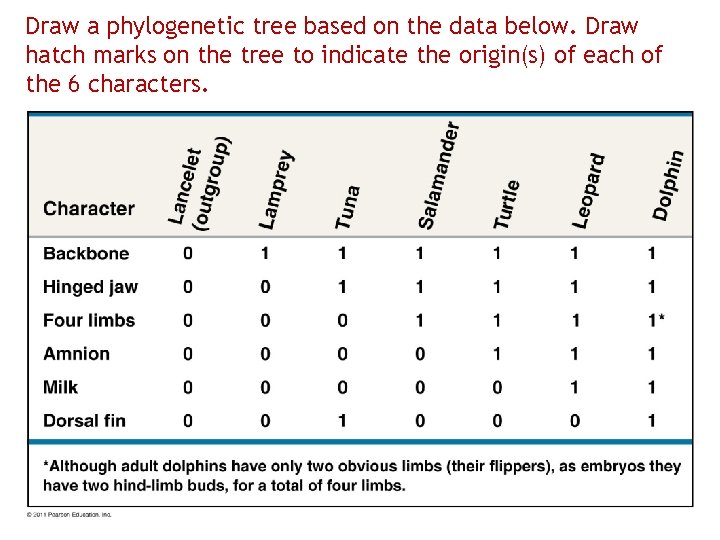
Draw a phylogenetic tree based on the data below. Draw hatch marks on the tree to indicate the origin(s) of each of the 6 characters.

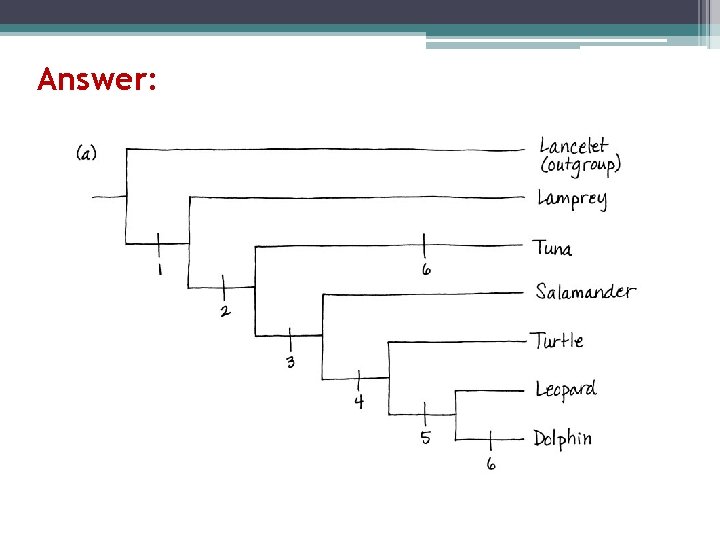
Answer:

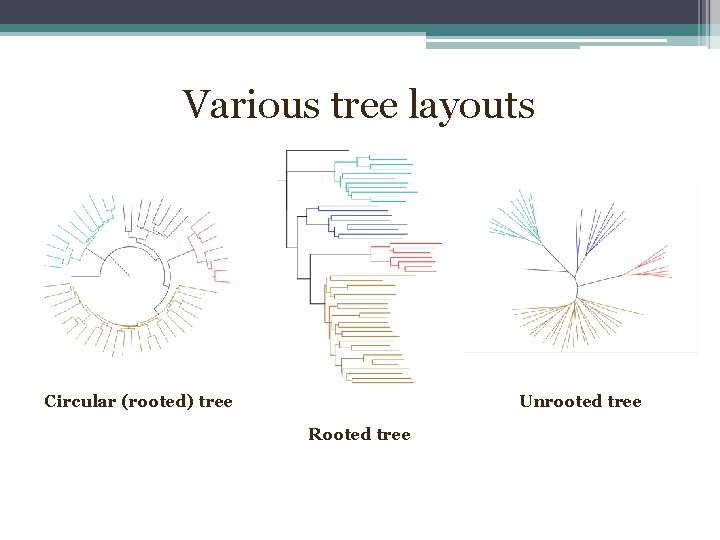
Various tree layouts Circular (rooted) tree Unrooted tree Rooted tree

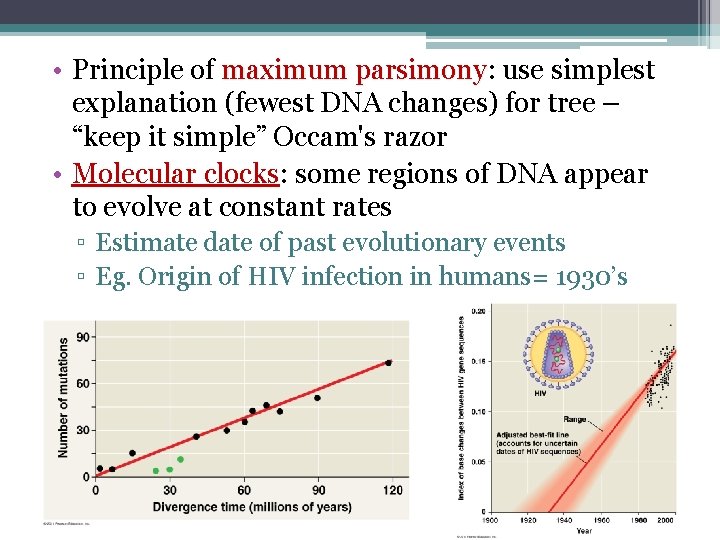
• Principle of maximum parsimony: parsimony use simplest explanation (fewest DNA changes) for tree – “keep it simple” Occam's razor • Molecular clocks: some regions of DNA appear to evolve at constant rates ▫ Estimate date of past evolutionary events ▫ Eg. Origin of HIV infection in humans= 1930’s

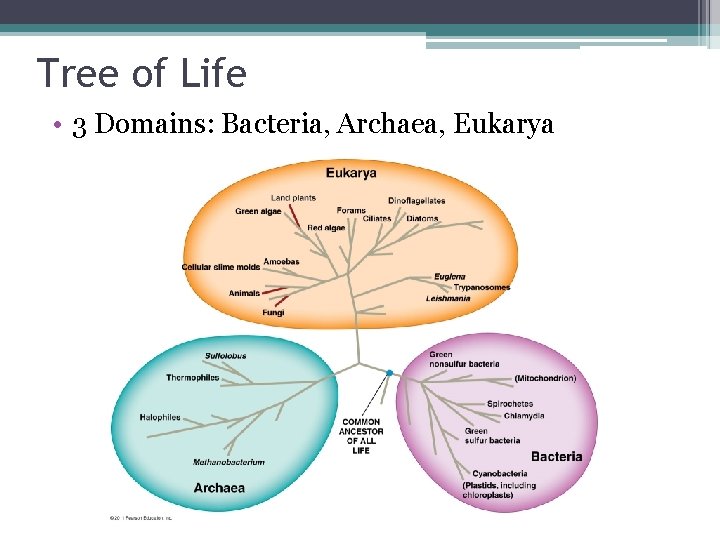
Tree of Life • 3 Domains: Bacteria, Archaea, Eukarya

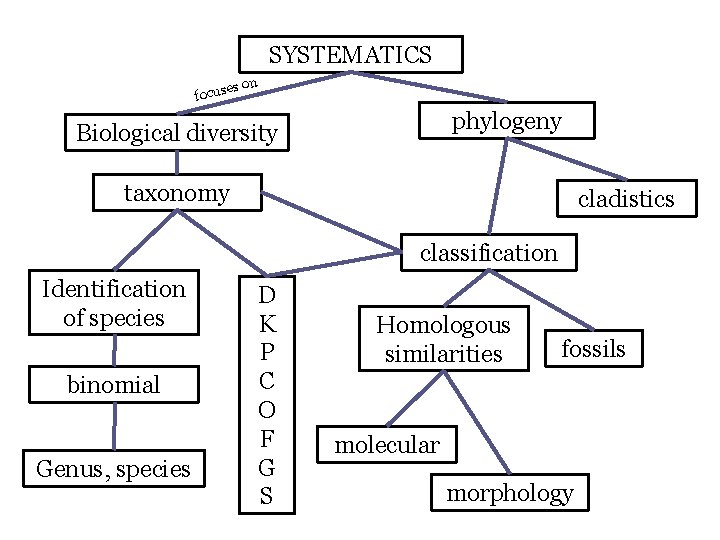
SYSTEMATICS es focus on phylogeny Biological diversity taxonomy cladistics classification Identification of species binomial Genus, species D K P C O F G S Homologous similarities fossils molecular morphology