Genomics I High throughput sequencing Jim Noonan Department




















- Slides: 38

Genomics I: High throughput sequencing Jim Noonan Department of Genetics

A working definition of genomics The global study of how biological information is encoded in genome sequence • Genes • Regulatory sequences • Genetic variation How this information is read out to produce distinct biological outcomes • Gene expression and regulation • Cellular identity, differentiation and development • Phenotypic variation among individuals and species

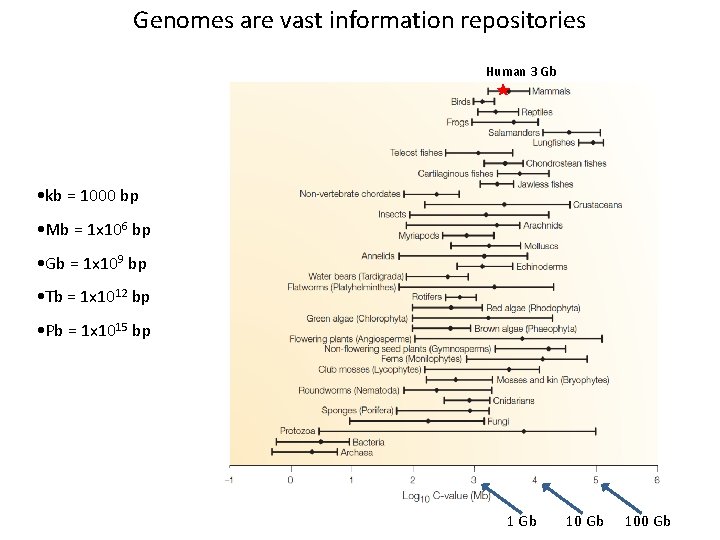
Genomes are vast information repositories Human 3 Gb • kb = 1000 bp • Mb = 1 x 106 bp • Gb = 1 x 109 bp • Tb = 1 x 1012 bp • Pb = 1 x 1015 bp 1 Gb 100 Gb

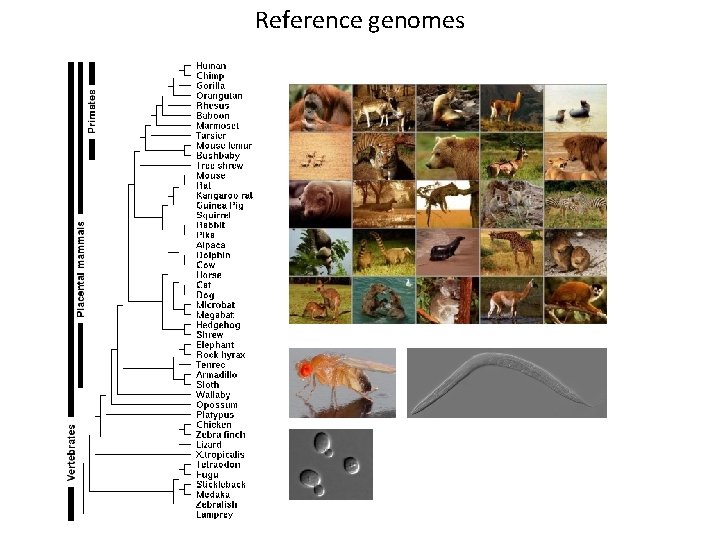
Reference genomes

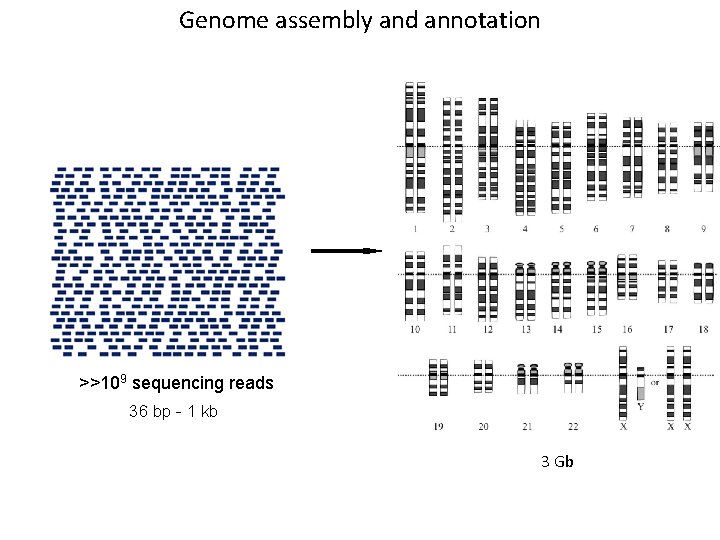
Genome assembly and annotation >>109 sequencing reads 36 bp - 1 kb 3 Gb

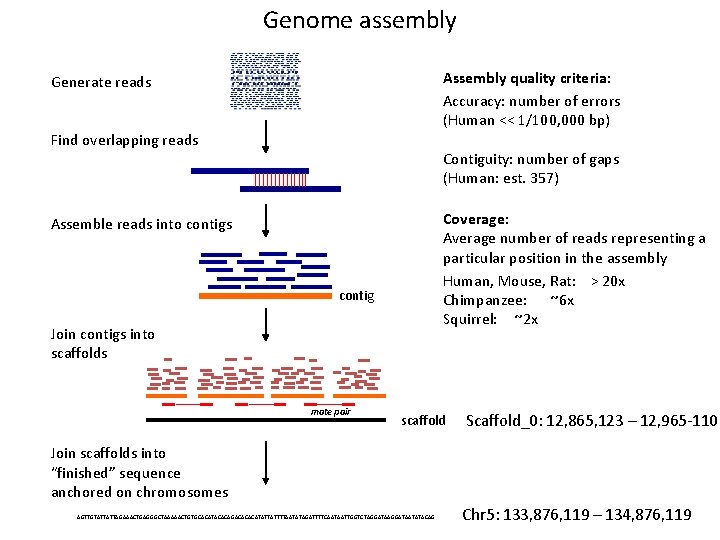
Genome assembly Assembly quality criteria: Accuracy: number of errors (Human << 1/100, 000 bp) Generate reads Find overlapping reads Contiguity: number of gaps (Human: est. 357) Coverage: Average number of reads representing a particular position in the assembly Human, Mouse, Rat: > 20 x Chimpanzee: ~6 x Squirrel: ~2 x Assemble reads into contigs contig Join contigs into scaffolds mate pair scaffold Scaffold_0: 12, 865, 123 – 12, 965 -110 Join scaffolds into “finished” sequence anchored on chromosomes AGTTGTATTATTAGAAACTGAGGGCTAAAAACTGTGCACATACACAGACACACATATTATTTTAATATAGATTTTCAATAATTGGTCTAGGATAATATACAG Chr 5: 133, 876, 119 – 134, 876, 119

TATCATGCTTGTCATTCGTTTATCAGAGGCCAAATGTTTTTCTTTGTAAACGTGTGTAAAACATTCTCAGAATTTTAAACAATAACAAATCAGGGCTGAATGTGGCC CTGTCCAAATCAAACAGTTGTATTATTAGAAACTGAGGGCTAAAAACTGTGCACATACACAGACACACATATTATTTTAATATAGATTTTCAATAATTGGTCTAGGAT AAGTTTAAGCAAGAAGAAAACAAAGACTGTTACTATGGAAAAATGAAAATAGATTTTAAAACATGTTAATTCACGTTACTTTTTGTTAAATTTACTTTTCTTCTTTCAC TTTAGGAACAATAAATCACATTAATTCCTTATCTCATGTGAAATTTCATATTTATGATACCTTTAAATGTCATTTGTTGAAGATTATTCATTTTTTCA CCAGGCACAAGACCAGTATTATGTTCTAGGCATTGGGGATACCATGTTCACAAGACTATGATTTACAGGATCAGATGTGGACTCTCAAATTCGACTGAGAAT GTTAATTTCAAGTTGTAATTGATGCTAGAAAGACAATGAAACAGAGCCATGTGACCAATGAGATGAGGGTGGCAGCAGCCTGTTTTAGATAAGGTACCTGA GATTAGTGTCTTCAGATATGCCTTAATGATATGAAAGAACCATTCATGGGAAGGCCTAGCATTAAAAACCGTCTAGGCAGAATGAGCAGCAAGTGCAAGGGTCCT AGGAAGAGAAACTATGGAAAAATGAAAATAGATTTTAAAACATGTTAATTCACGTTACTTTTTGTTAAATTTACTTTTCTTCTTTCACTTCTTACCTGTCAATGT CACATTAATTCCTTATCTCATGTGAAATTTCATATTTATGATACCTTTAAATGTCATTTGTTGAAGATTATTCATTTTTTCAATAAATATTTTTTAGAA ATTATGTTCTAGGCATTGGGGATACCATGTTCACAAGACTATGATTTACAGGATCAGATGTGGACTCTCAAATTCGACTGAGAATAAAACAGACACTAAACA ATTGATGCTACTATGGAAAAATGAAAATAGATTTTAAAACATGTTAATTCACGTTACTTTTTGTTAAATTTACTTTTCTTCTTTCACTTCTTACCTGTCAATGTTATTAA ATTCCTTATCTCATGTGAAATTTCATATTTATGATACCTTTAAATGTCATTTGTTGAAGATTATTCATTTTTTCAATAAATATTTTTTAGAATAATAA GTTCTAGGCATTGGGGATACCATGTTCACAAGACTATGATTTACAGGATCAGATGTGGACTCTCAAATTCGACTGAGAATAAAACAGACACAAGTAAA GCTATCCCAGGCACAAGACCAGTATTATGTTCTAGGCATTGGGGATACCATTACCTGTCAATGTTATTAATATTTTTAGGAACAATAAATCACATTAATTCCAACATG TGTCATTCGTTTATCAGAGGCCAAATGTTTTTCTTTGTAAACGTGTGTAAAACATTCTCAGAATTTTAAACAATAACAAATCAGGGCTGAATGTGGCCAACATGCAA CAAACAGTTGTATTATTAGAAACTGAGGGCTAAAAACTGTGCACATACACAGACACACATATTATTTTAATATAGATTTTCAATAATTGGTCTAGGATAATA GCAAGAAGAAAACAAAGACTGTTACTATGGAAAAATGAAAATAGATTTTAAAACATGTTAATTCACGTTACTTTTTGTTAAATTTACTTTTCTTCTTTCACTTCTTACCT CAATAAATCACATTAATTCCTTATCTCATGTGAAATTTCATATTTATGATACCTTTAAATGTCATTTGTTGAAGATTATTCATTTTTTCAATAAATATT GACCAGTATTATGTTCTAGGCATTGGGGATACCATGTTCACAAGACTATGATTTACAGGATCAGATGTGGACTCTCAAATTCGACTGAGAATAAAACAGACA AGTTGTAATTGATGCTAGAAAGACAATGAAACAGAGCCATGTGACCAATGAGATGAGGGTGGCAGCAGCCTGTTTTAGATAAGGTACCTGATTGGTGGGA CTTCAGATATGCCTTAATGATATGAAAGAACCATTCATGGGAAGGCCTAGCATTAAAAACCGTCTAGGCAGAATGAGCAGCAAGTGCAAGGGTCCTGGATAGGAA GAGAAACTATGGAAAAATGAAAATAGATTTTAAAACATGTTAATTCACGTTACTTTTTGTTAAATTTACTTTTCTTCTTTCACTTCTTACCTGTCAATGTTATTAATATTT CTTATCTCATGTGAAATTTCATATTTATGATACCTTTAAATGTCATTTGTTGAAGATTATTCATTTTTTCAATAAATATTTTTTAGAATAATAAGTCCC GGCATTGGGGATACCATGTTCACAAGACTATGATTTACAGGATCAGATGTGGACTCTCAAATTCGACTGAGAATAAAACAGACACTAAACAAGTAAAG CTATGGAAAAATGAAAATAGATTTTAAAACATGTTAATTCACGTTACTTTTTGTTAAATTTACTTTTCTTCTTTCACTTCTTACCTGTCAATGTTATTAATATTTTTAGGA CATGTGAAATTTCATATTTATGATACCTTTAAATGTCATTTGTTGAAGATTATTCATTTTTTCAATAAATATTTTTTAGAATAATAAGTCCCAGGCA GGGGATACCATGTTCACAAGACTATGATTTACAGGATCAGATGTGGACTCTCAAATTCGACTGAGAATAAAACAGACACAAGTAAAGTTAATTT CACAAGACCAGTATTATGTTCTAGGCATTGGGGATACCATTACCTGTCAATGTTATTAATATTTTTAGGAACAATAAATCACATTAATTCCAACATGCAAAGAGGAAA ATCAGAGGCCAAATGTTTTTCTTTGTAAACGTGTGTAAAACATTCTCAGAATTTTAAACAATAACAAATCAGGGCTGAATGTGGCCAACATGCAAAGAGGAAATCT TTATTAGAAACTGAGGGCTAAAAACTGTGCACATACACAGACACACATATTATTTTAATATAGATTTTCAATAATTGGTCTAGGATAATATACAGAGAACA CAAAGACTGTTACTATGGAAAAATGAAAATAGATTTTAAAACATGTTAATTCACGTTACTTTTTGTTAAATTTACTTTTCTTCTTTCACTTCTTACCTGTCAATGTTATTA AATTCCTTATCTCATGTGAAATTTCATATTTATGATACCTTTAAATGTCATTTGTTGAAGATTATTCATTTTTTCAATAAATATTTTTTAGAATAATA GTTCTAGGCATTGGGGATACCATGTTCACAAGACTATGATTTACAGGATCAGATGTGGACTCTCAAATTCGACTGAGAATAAAACAGACACTAAACAAGTAA TGCTAGAAAGACAATGAAACAGAGCCATGTGACCAATGAGATGAGGGTGGCAGCAGCCTGTTTTAGATAAGGTACCTGATTGGTGGGATTGGAAGACCTC CCTTAATGATATGAAAGAACCATTCATGGGAAGGCCTAGCATTAAAAACCGTCTAGGCAGAATGAGCAGCAAGTGCAAGGGTCCTGGATAGGAATGAGCTGGATA GGAAAAATGAAAATAGATTTTAAAACATGTTAATTCACGTTACTTTTTGTTAAATTTACTTTTCTTCTTTCACTTCTTACCTGTCAATGTTATTAATATTTTTAGGAACAA GTGAAATTTCATATTTATGATACCTTTAAATGTCATTTGTTGAAGATTATTCATTTTTTCAATAAATATTTTTTAGAATAATAAGTCCCAGGCACAAG GGATACCATGTTCACAAGACTATGATTTACAGGATCAGATGTGGACTCTCAAATTCGACTGAGAATAAAACAGACACTAAACAAGTAAAGTTAATTTCA ATGAAAATAGATTTTAAAACATGTTAATTCACGTTACTTTTTGTTAAATTTACTTTTCTTCTTTCACTTCTTACCTGTCAATGTTATTAATATTTTTAGGAACAATAAAT TTTCATATTTATGATACCTTTAAATGTCATTTGTTGAAGATTATTCATTTTTTCAATAAATATTTTTTAGAATAATAAGTCCCAGGCACAAGACCAG CCATGTTCACAAGACTATGATTTACAGGATCAGATGTGGACTCTCAAATTCGACTGAGAATAAAACAGACACAAGTAAAGTTAATTTCAAGTTGT CAGTATTATGTTCTAGGCATTGGGGATACCATTACCTGTCAATGTTATTAATATTTTTAGGAACAATAAATCACATTAATTCCAACATGCAAAGAGGAAATCTCCATA GCCAAATGTTTTTCTTTGTAAACGTGTGTAAAACATTCTCAGAATTTTAAACAATAACAAATCAGGGCTGAATGTGGCCAACATGCAAAGAGGAAATCTCCCATCTG AACTGAGGGCTAAAAACTGTGCACATACACAGACACACATATTATTTTAATATAGATTTTCAATAATTGGTCTAGGATAATATACAGAGAACATGCCAAAA GTTACTATGGAAAAATGAAAATAGATTTTAAAACATGTTAATTCACGTTACTTTTTGTTAAATTTACTTTTCTTCTTTCACTTCTTACCTGTCAATGTTATTAATATTTT

Genome annotation Genes: ~3 billion bp ACAATAAATCACATTAATTCCTTATCTCATGTGAAATTTCATATTTATGATACCTTTAAATGTCATTTGTTGAAGATTATTCATTTTTTCATTC AATATTTTTTAGAATAATAAGTCCCAGGCACAAGACCAGTATTATG TTCTAGGCATTGGGGATACCATGTTCACAAGACTATGATTTACAG GATCAGATGTGGACTCTCAAATTCGACTGAGAATAAAACAGACACTAAA CAAGTAAAGTTAATTTCAAGTTGTAATTGATGCTAGAAAGACAATG AAACAGAGCCATGTGACCAATGAGATGAGGGTGGCAGCAGCCT GTTTTAGATAAGGTACCTGATTGGTGGGATTGGAAGACCTCTCTGAGAT TAGTGTCTTCAGATATGCCTTAATGATATGAAAGAACCATTCATGGGAA GGCCTAGCATTAAAAACCGTCTAGGCAGAATGAGCAGCAAGTGCAAGG GTCCTGGATAGGAATGAGCTGGATATACTCAAGGAAGAGAAACT ATGGAAAAATGAAAATAGATTTTAAAACATGTTAATTCACGTTACTTTTTG TTAAATTTACTTTTCTTCTTTCACTTCTTACCTGTCAATGTTATTAATATTT TTAGGAACAATAAATCACATTAATTCCTTATCTCATGTGAAATTTCATATT TATGATACCTTTAAATGTCATTTGTTGAAGATTATTCATTTTT TCATTCAATATTTTTTAGAATAATAAGTCCCAGGCACAAGACCAGT ATTATGTTCTAGGCATTGGGGATACCATGTTCACAAGACTATGAT TTACAGGATCAGATGTGGACTCTCAAATTCGACTGAGAATAAAACAGAC ACTAAACAAGTAAAGTTAATTTCAAGTTGTAATTGATGCTACTATG GAAAAATGAAAATAGATTTTAAAACATGTTAATTCACGTTACTTTTTGTTA AATTTACTTTTCTTCTTTCACTTCTTACCTGTCAATGTTATTAATATTTTTA GGAACAATAAATCACATTAATTCCTTATCTCATGTGAAATTTCATATTTAT GATTGATACCTTTAAATGTCATTTGTTGAAGATTATTCATTTTTTCAATATTTTTTAGAATAATAAGTCCCAGGCACAAGACCAGTATT ATGTTCTAGGCATTGGGGATACCATGTTCACAAGACTATGATTTA CAGGATCAGATGTGGACTCTCAAATTCGACTGAGAATAAAACAGACACA AACAAGTAAAGTTAATTTCAAGTTGTAATTGATGCTATCCCAGGCA CAAGACCA…. - Coding, noncoding, mi. RNA, etc. Isoforms Expression Genetic variation: - SNPs and CNVs Sequence conservation Regulatory sequences: - Promoters Enhancers Insulators Epigenetics: - DNA methylation Chromatin

Portals to access and interpret genomes UCSC Genome Browser (genome. ucsc. edu): Visualization, data recovery, simple analysis (also http: //genome-preview. ucsc. edu/) ENSEMBL (ensembl. org): Visualization, data recovery, simple analysis Integrative Genomics Viewer (broadinstitute. orgsoftware/igv/): Local genome viewer (visualize local and remote data) Galaxy (main. g 2. bx. psu. edu): Complex data analysis and workflows

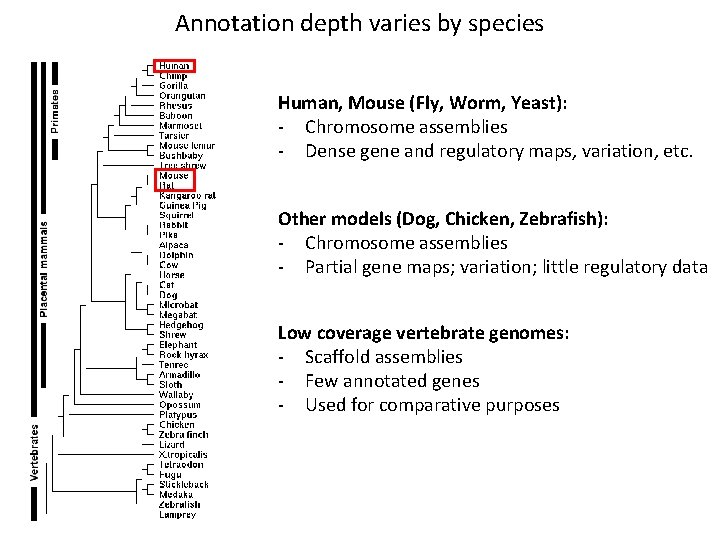
Annotation depth varies by species Human, Mouse (Fly, Worm, Yeast): - Chromosome assemblies - Dense gene and regulatory maps, variation, etc. Other models (Dog, Chicken, Zebrafish): - Chromosome assemblies - Partial gene maps; variation; little regulatory data Low coverage vertebrate genomes: - Scaffold assemblies - Few annotated genes - Used for comparative purposes

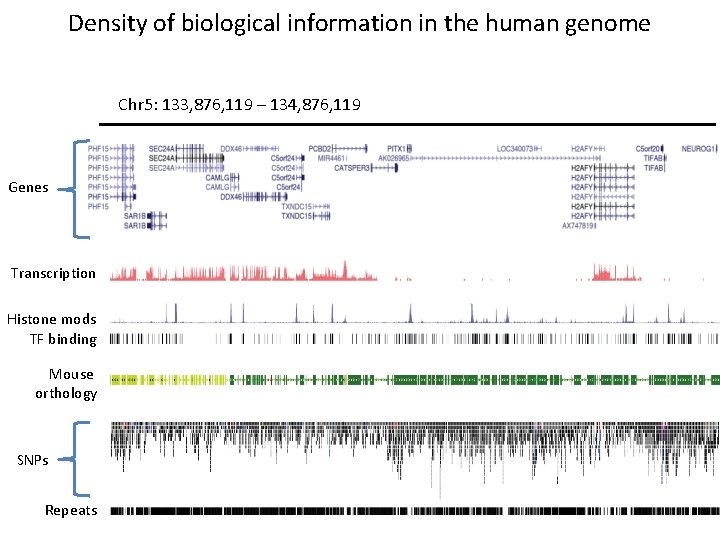
Density of biological information in the human genome Chr 5: 133, 876, 119 – 134, 876, 119 Genes Transcription Histone mods TF binding Mouse orthology SNPs Repeats

Sequence as the readout for biological processes Determining the biological state of cells, tissues and organisms requires the quantification of sequence information • Gene expression • Protein-DNA interactions (Ch. IP) • DNA-DNA interactions (3 C/4 C/5 C) • Chromatin state • DNA methylation • Genetic variation (SNPs/CNVs) Indirect measures Direct measures CTATGATCAGTC. . . TCAATCTG. . . GGACTTCGAGATC. . . AAGTCGCTGACGT. . . microarrays, PCR, etc. Sequencing

Outline First-generation sequencing technology • Sanger sequencing • Parallelization in human genome project Current massively parallel sequencing strategies “Second Generation” • 454 • Illumina • SOLi. D • Ion Torrent & Ion Proton • Sequencing Services (Complete Genomics) “Third Generation” • Pacific Biosciences • Oxford Nanopore

Metrics for evaluating sequencing methods Throughput • Number of high quality bases per unit time • Number of independent samples run in parallel - multiplexing • Difficulty of sample prep Yield • Number of useful/mappable reads per sample • Read length Cost • Per run and per base • Equipment • Reagents • Labor • Analysis The goal is to increase throughput and yield while reducing cost

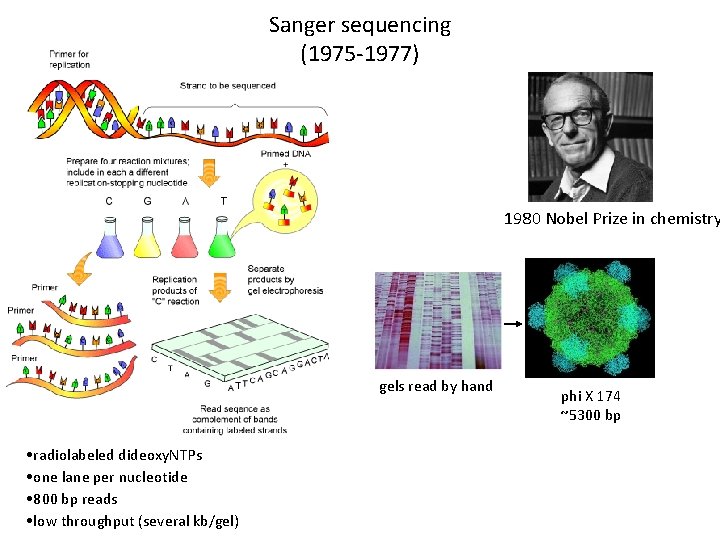
Sanger sequencing (1975 -1977) 1980 Nobel Prize in chemistry gels read by hand • radiolabeled dideoxy. NTPs • one lane per nucleotide • 800 bp reads • low throughput (several kb/gel) phi X 174 ~5300 bp

Sequencing the reference human genome (1990 -present; ‘finished’ 2003) • Industrialization of Sanger sequencing, library construction, sample preparation, analysis, etc. • $3 billion total cost • 1 Gb/month at largest centers (2005) • YCGA = 9. 6 Tb per month (2011)

Second-generation sequencing “Democratizing” sequencing production • Massive parallelization • Reduction in per-base cost • Eliminate need for huge infrastructure • Millions of reads - >1 Gb sequence per run Novel sequencing applications • RNA-seq • Ch. IP-seq Counting applications • Methyl-seq • Whole-genome and targeted resequencing Challenges • Read length • Quality • Data analysis and storage

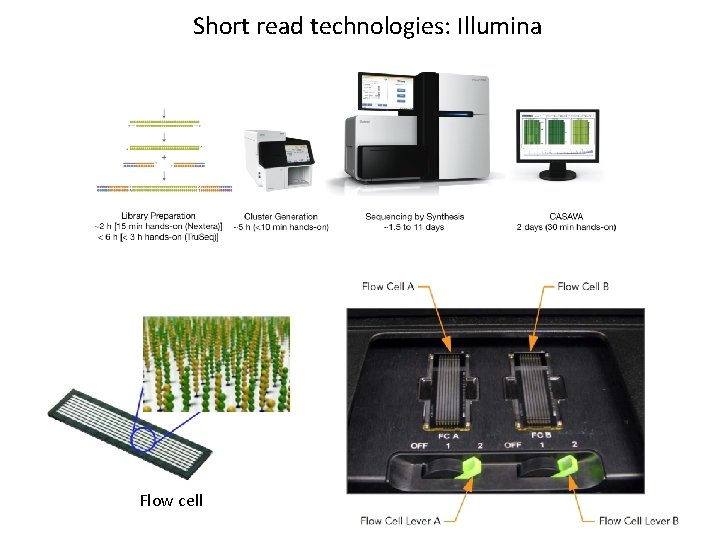
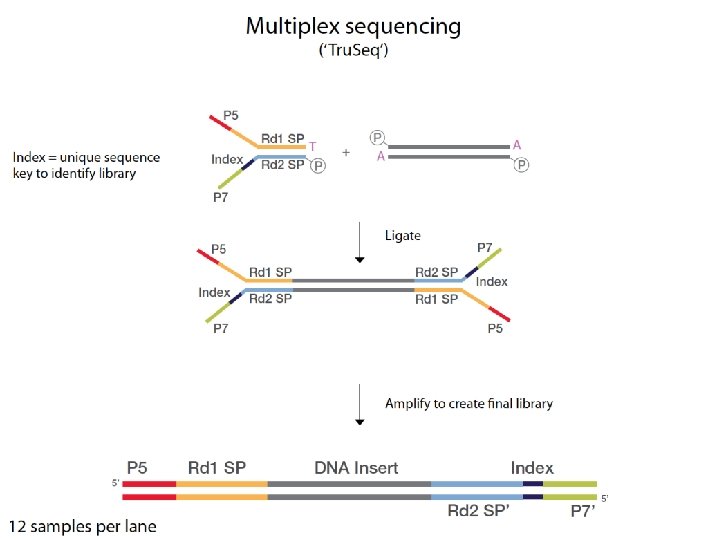
Short read technologies: Illumina Flow cell

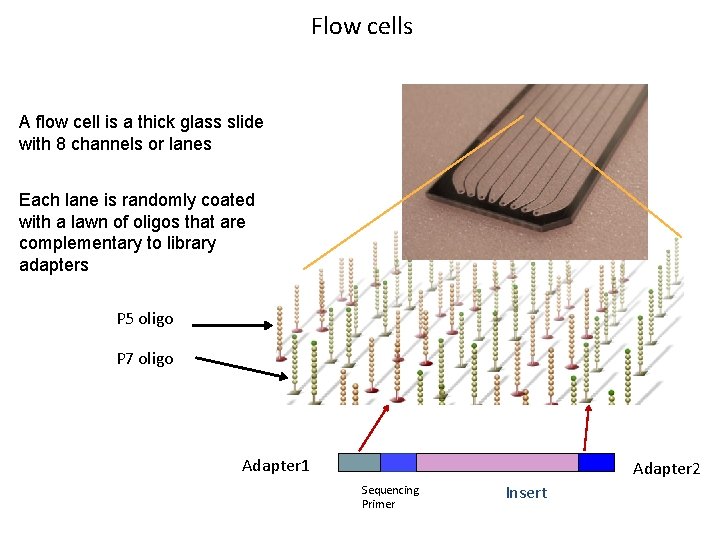
Flow cells A flow cell is a thick glass slide with 8 channels or lanes Each lane is randomly coated with a lawn of oligos that are complementary to library adapters P 5 oligo P 7 oligo Adapter 1 Adapter 2 Sequencing Primer Insert

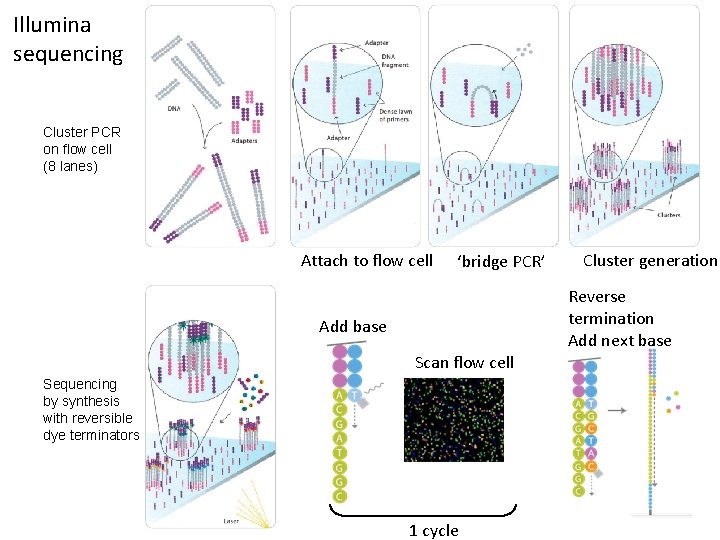
Illumina sequencing Cluster PCR on flow cell (8 lanes) Attach to flow cell ‘bridge PCR’ Cluster generation Reverse termination Add next base Add base Scan flow cell Sequencing by synthesis with reversible dye terminators 1 cycle



Hi. Seq 2500 1 Instrument – 2 Run Modes High Output Mode Rapid Run Mode 600 Gb in ~10. 5 days Current v 3 flow cell Current v 3 reagents c. Bot required 120 Gb in ~1 day New 2 -lane flow cell New reagents No c. Bot required User configurable 6 human genomes in 10. 5 days Highest Output 1 human genome in a day Fastest turnaround

Mi. Seq • Run-times – 50 cycle – 4 hours – 300 cycle – 27 hours • Two sequencing options – 50 cycles – 300 cycles (2 x 150 bp) • One lane – 6 -7 million clusters – Up to 8 billion bases (300 cycles) Ideal for: R&D, CLIA, small genomes and projects where longer reads are important

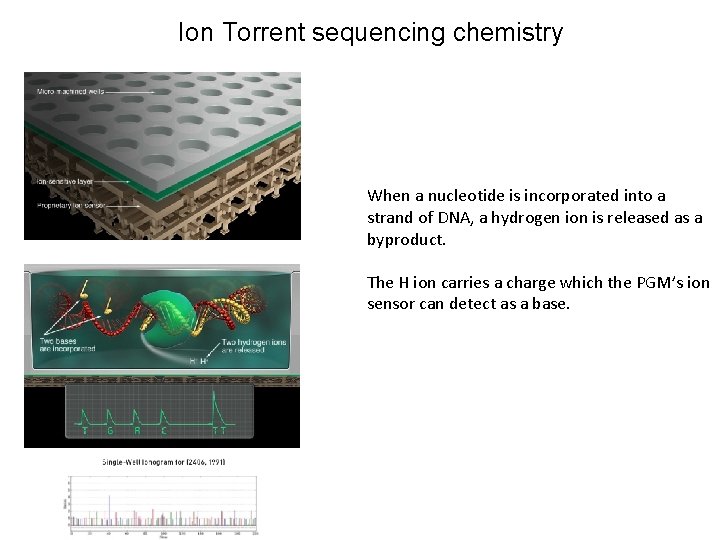
Ion Torrent and Ion Proton Sequencing on semiconductor chip

Ion Torrent sequencing chemistry When a nucleotide is incorporated into a strand of DNA, a hydrogen ion is released as a byproduct. The H ion carries a charge which the PGM’s ion sensor can detect as a base.

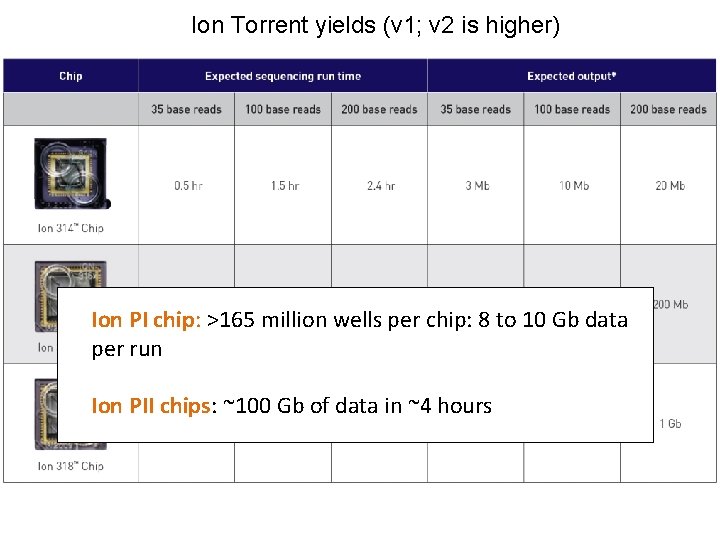
Ion Torrent yields (v 1; v 2 is higher) Ion PI chip: >165 million wells per chip: 8 to 10 Gb data per run Ion PII chips: ~100 Gb of data in ~4 hours

Advantages and limitations Advantages • Low equipment cost • Rapid run times: 3 to 4 hours • Simple Chemistry Limitations • • • Homopolymers detection Error rates Slow on introducing newer chips: Overpromise PGM and Proton: two separate systems Library prep: Emulsion PCR

Third-generation sequencing High-throughput single molecule sequencing in real time at low cost Pacific Biosciences • Sequence in real time with fluorescent NTPs • Rate limited by processivity of polymerase • Very long reads possible (6 kb) • Not well parallelized (few reads)

Sequencing in real time: Pacific Biosciences SMRT cells Zero Mode Waveguides

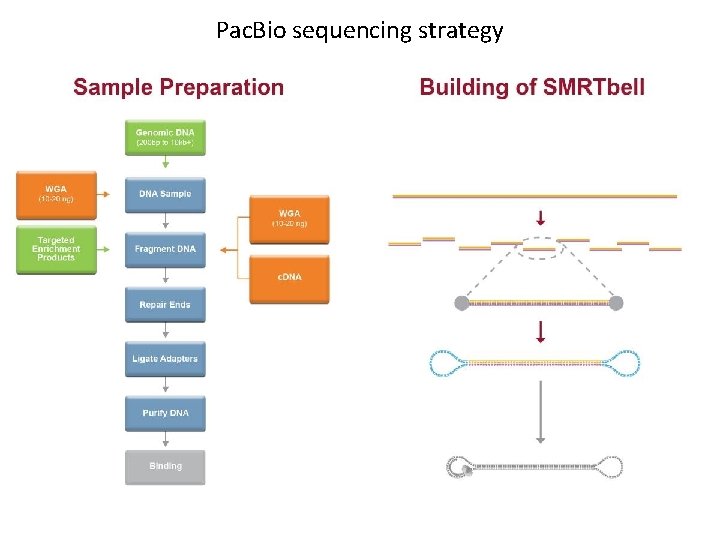
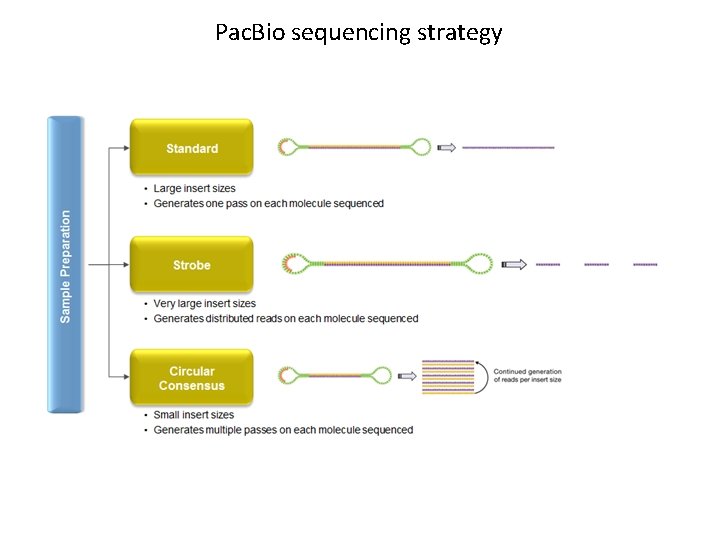
Pac. Bio sequencing strategy

Pac. Bio sequencing strategy

Applications q Targeted sequencing SNP and structure variants detection q Repetitive regions q Full length transcript profiling De novo assembly and genome finishing q Bacterial genomes q Fungal genomes q Gap-captured sequencing q Targeted captured sequencing Base modifications detection q Methylation q DNA damage q q q YCGA Pac. Bio RS **Projects at YCGA Shrikant Mane

Pac. Bio vs Illumina Sequencing Chemistry Pac. Bio RS (Third generation) Illumina Hi. Seq (Second generation) Sequencing by synthesis (SBS) Single Molecule Real Time (SMRT) Sequencing by synthesis (SBS) Sequencing Smart Cell made up of Flow cell has made of 150, 000 ZMWs 8 separate lanes substrate Data output per 60 billion/day at a cost of $. 06 per 1 to 2 billion/ day. $1. 5/ Mb Mb day Read Length Average up to 5 Kb 50 bp to 150 bp Raw: 10 -15 %. With 30 x coverage: Error rates 0. 5 to 1 % Q 50 (< 0. 01) Sample Library SMRT Bell template ds. DNA with adaptors (175 bp to (Single-strand circular DNA) 250 bp to 1 Kb) 10 Kb insert Shrikant Mane

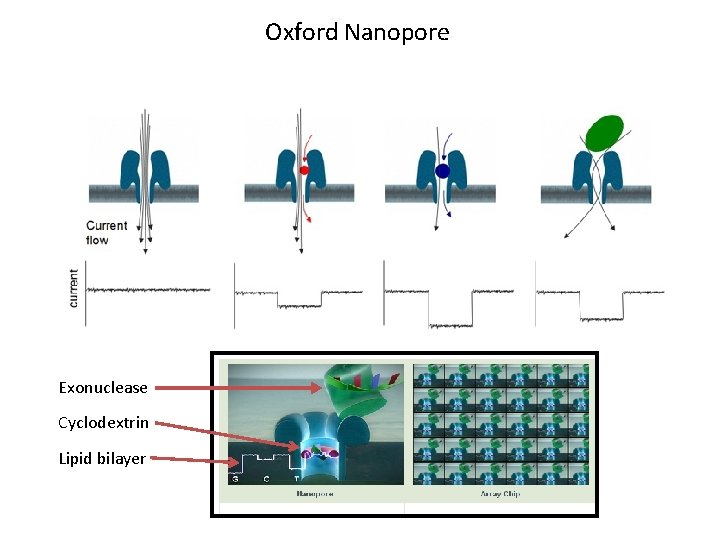
Oxford Nanopore Exonuclease Cyclodextrin Lipid bilayer

Oxford Nanopore Grid. ION: enables scaled-up measurements of multiple nanopores and analysis of data in real time Mini. ION

Advantages and limitations • Nanopores offer a label-free, electrical, single-molecule DNA sequencing method • No costly fluorescent labeling reagents • No need for expensive optical hardware and sophisticated instrumentation to detect DNA bases • Runs as long as needed • High error rates ~5% • Not available yet

Conclusions • High-throughput sequencing has become democratized moved out of industrial-scale genome centers • Sequence is no longer limiting - next generation of sequencers will make sequencing very inexpensive • Earlier methods for counting / resequencing applications are largely obsolete • Scale of data production outstripping our ability to store and analyze it • Next: Applications of the technology