UALR Biosciences Core Facility High Throughput DNA Sequencing

UALR Biosciences Core Facility High Throughput DNA Sequencing n Fragment Analyses n Real Time PCR n In situ Hybridization n Proteomics n

High Throughput DNA Sequencing Beckman CEQ 2000 XL DNA Analysis System

Beckman CEQ 2000 XL DNA Analysis System This is an automated unit capable of reading and analyzing DNA sequencing products that have been generated by incorporating Well. RED fluorescent dyes into the DNA. The dyes fluoresce and are automatically detected and processed by the system’s software. The system is equipped with an array made up of eight parallel capillaries permitting electrokinetic injections of eight samples at a time. This high throughput setup allows for 96 samples to be processed analyzed by the system's software in 24 hours. On average, sequences of 600 to 850 bases per sample can be generated, depending on primer and template qualities.

Amount of DNA Required
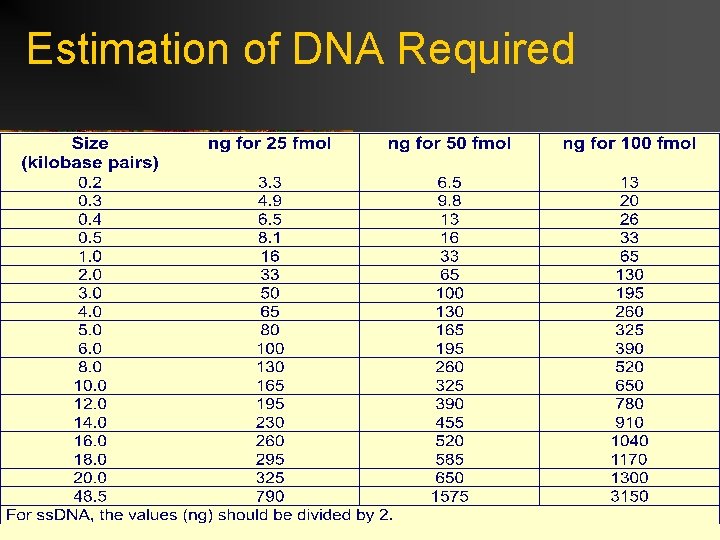
Estimation of DNA Required

Guidelines for DNA Sample Preparation ***REMEMBER: HIGH QUALITY TEMPLATE WILL LEAD TO HIGH QUALITY DATA!!!! IT IS TO YOUR ADVANTAGE TO FOLLOW PROCEDURE!!!! Prepare templates properly: Resuspend in water or Tris-HCl. Do not resuspend in TE or EDTA reagents. Must be purified and free of ionic contaminants Best to use a commercial kit for purification! The Qiagen Qiawell and Qiaprep work well (ds. DNA and ss. DNA) Standard Alkaline Lysis Preps (NO PHENOL!!!) For PCR Products: Qiagen QIAquick PCR purification kit PCR products must be homogeneous Quantitate DNA: Spectrophotometeric Fluorometeric Agarose Gel Custom Primer Considerations: Quality primers necessary Purified by desalting, HPLC or OPC purification preferred Size around 20 bases with 18 bases as a minimum GC content should be less that 50% Avoid primer dimerization Ratio for ds. DNA is greater than 40: 1 primer: template molar ratio (~3. 2 ρmol).

Major Reagents/consumables

Sequencing Reaction
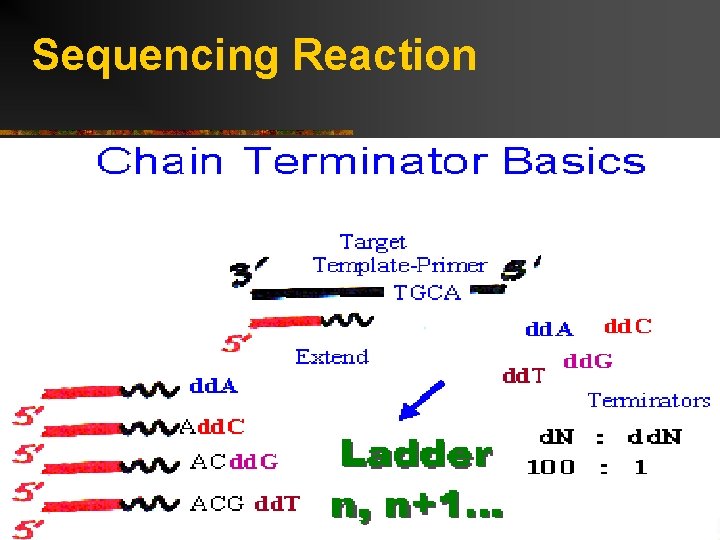
Sequencing Reaction
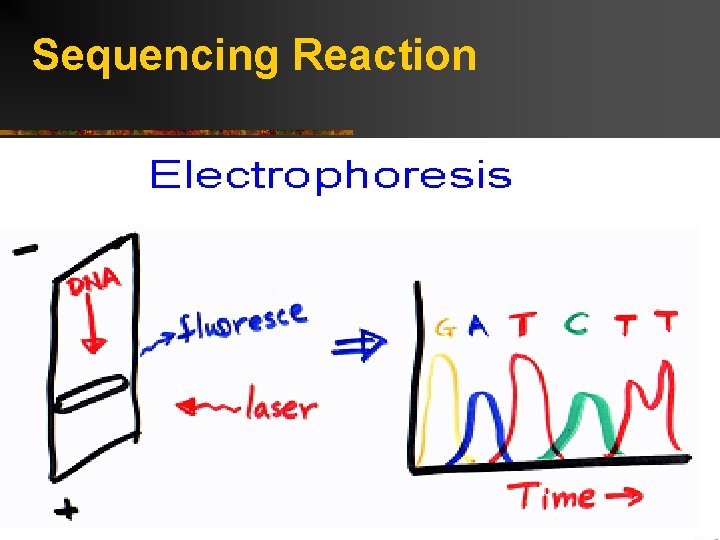
Sequencing Reaction
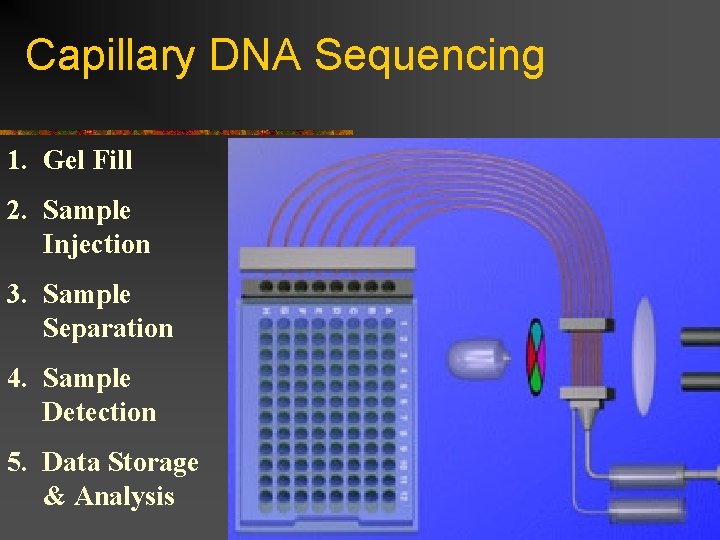
Capillary DNA Sequencing 1. Gel Fill 2. Sample Injection 3. Sample Separation 4. Sample Detection 5. Data Storage & Analysis
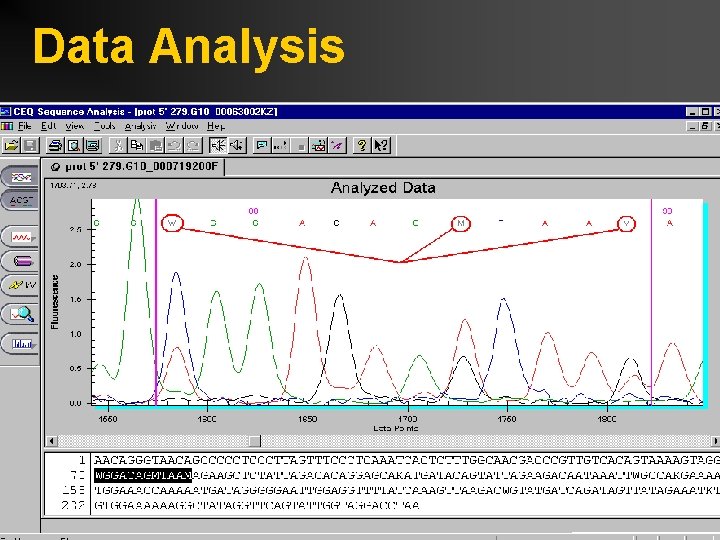
Data Analysis
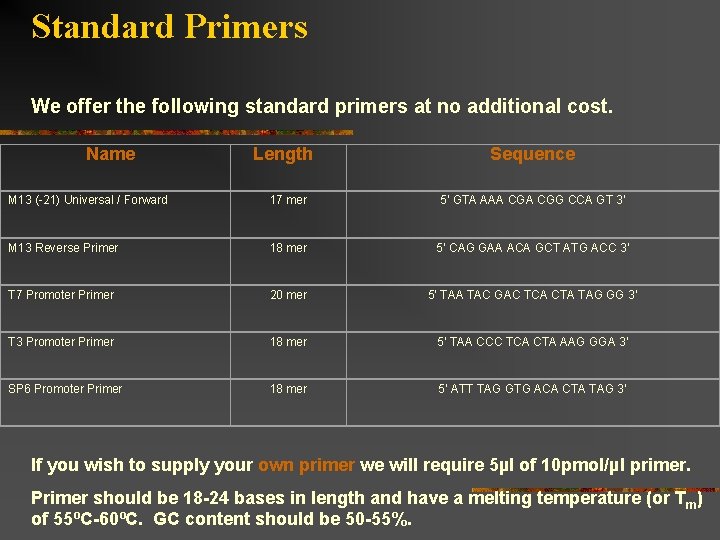
Standard Primers We offer the following standard primers at no additional cost. Name Length Sequence M 13 (-21) Universal / Forward 17 mer 5’ GTA AAA CGG CCA GT 3’ M 13 Reverse Primer 18 mer 5’ CAG GAA ACA GCT ATG ACC 3’ T 7 Promoter Primer 20 mer 5’ TAA TAC GAC TCA CTA TAG GG 3’ T 3 Promoter Primer 18 mer 5’ TAA CCC TCA CTA AAG GGA 3’ SP 6 Promoter Primer 18 mer 5’ ATT TAG GTG ACA CTA TAG 3’ If you wish to supply your own primer we will require 5µl of 10 pmol/µl primer. Primer should be 18 -24 bases in length and have a melting temperature (or T m) of 55ºC-60ºC. GC content should be 50 -55%.
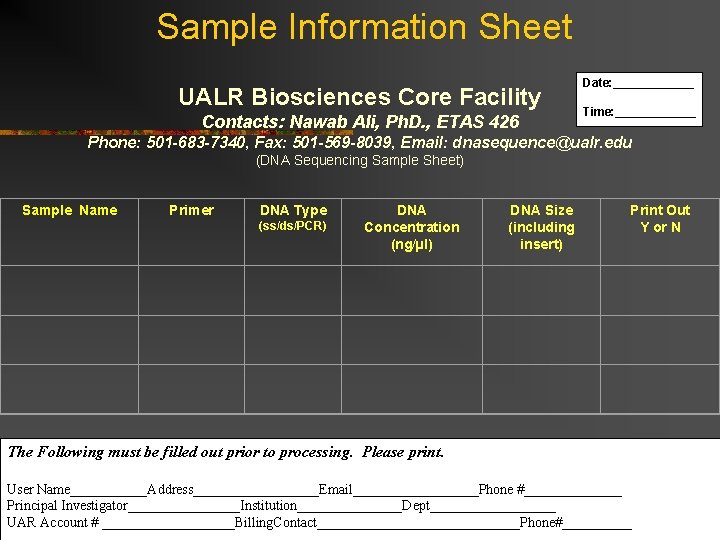
Sample Information Sheet UALR Biosciences Core Facility Contacts: Nawab Ali, Ph. D. , ETAS 426 Date: ______ Time: ______ Phone: 501 -683 -7340, Fax: 501 -569 -8039, Email: dnasequence@ualr. edu (DNA Sequencing Sample Sheet) Sample Name Primer DNA Type (ss/ds/PCR) DNA Concentration (ng/µl) DNA Size (including insert) Print Out Y or N The Following must be filled out prior to processing. Please print. User Name______Address_________Email_________Phone #_______ Principal Investigator________Institution________Dept_________ UAR Account # __________Billing. Contact_______________Phone#_____

- Slides: 15