Exploring Protein Sequences Part 1 Part 2 Patterns























- Slides: 28

Exploring Protein Sequences – Part 1: Part 2: Patterns and Motifs Profiles Hydropathy Plots Transmembrane helices Antigenic Prediction Signal Peptides Repeats Coiled Coils Protein Domains Domain databases Celia van Gelder CMBI Radboud University December 2005 ©CMBI 2005

Patterns and Motifs (1) • In a multiple sequence alignment (MSA) islands of conservation emerge • These conserved regions (motifs, segments, blocks, features) are typically around 10 -20 aa in length • They tend to correspond to the core structural or functional elements of the protein • Their conserved nature allows them to be used to diagnose family membership ©CMBI 2005

Patterns and Motifs (2) • A motif (or pattern or signature) is a regular expression for what residues can be present at any given position. • Motifs can contain - alternative residues - flexible regions C-x(2, 5)-C-x-[GP]-x-P-x(2, 5)-C CXXXCXGXPXXXXXC | | | FGCAKLCAGFPLRRLPCFYG Syntax: A-[BC]-X-D(2, 5)-{EFG}-H Means: A B or C Anything 2 -5 D’s Not E, F or G H

Patterns and Motifs (3) • Motifs can not contain - mismatches exact match or no match at all - gaps C-x(2, 5)-C-x-[GP]-x-P-x(2, 5)-C CXXCXGXPXXXXX-C | ? | | | ? | FGCA-CAGFPLRRLPKCFYG J. Leunissen

PROSITE • PROSITE - A Dictionary of Protein Sites and Patterns • 1328 patterns and 577 profiles/matrices (dec 2005) • For every pattern or profile there is documentation present (e. g. PDOC 00975) - information on taxonomic occurrence - domain architecture, - function, - 3 D structure, - main characteristics of the sequence - some references.

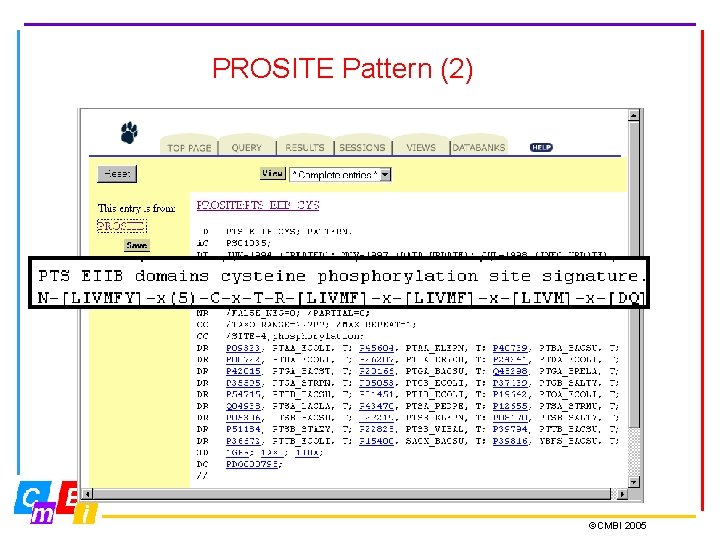
PROSITE Pattern • PROSITE patterns consist of an exact regular expression • Possible patterns occur frequently in proteins; they may not actually be present, such as post-translational modification sites ID ASN_GLYCOSYLATION; PATTERN. DE N-glycosylation site. PA N-{P}-[ST]-{P}. • Notice also in the PROSITE record the number of false positives and false negatives

PROSITE Pattern (2) ©CMBI 2005

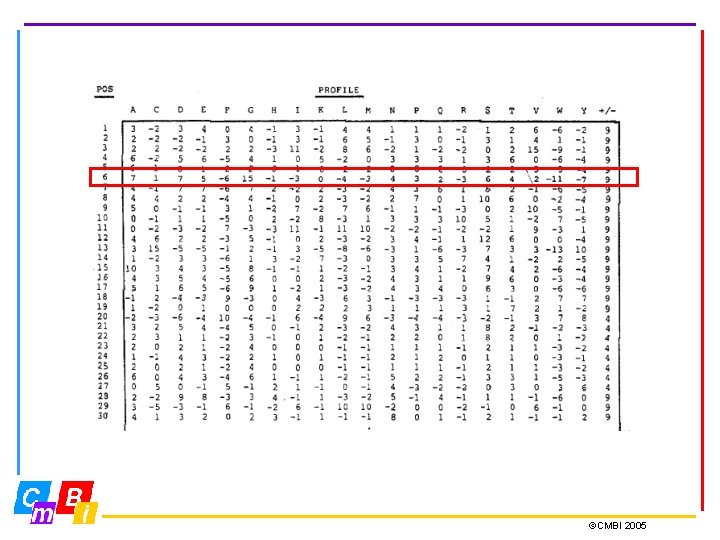
Profiles • If regular expressions fail to define the motif properly we need a profile. • Profiles are specific representations that incorporate the entire information of a multiple sequence alignment. • A profile is a position-specific scoring scheme and holds for each position in the sequence 20 scores for the 20 residue types, and sometimes also two values for gap open and gap elongation. • Profiles provide a sensitive means of detecting distant sequence relationships

©CMBI 2005

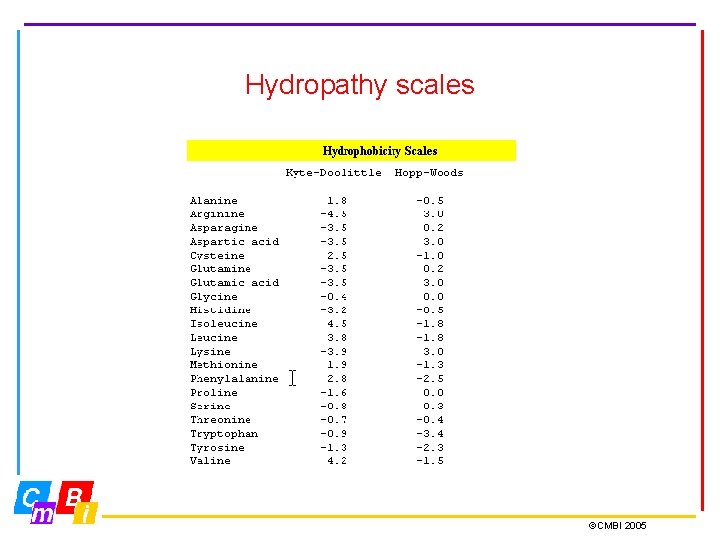
Hydropathy plots are designed to display the distribution of polar and apolar residues along a protein sequence. A positive value indicates local hydrophobicity and a negative value suggests a water-exposed region on the face of a protein. (Kyte-Doolittle scale) Hydropathy plots are generally most useful in predicting transmembrane segments, and N-terminal secretion signal sequences. ©CMBI 2005

Hydropathy scales ©CMBI 2005

Sliding Window Approach Sum amino acid property (e. g. hydrophobicity values) in a given window Plot the value in the middle of the window I L I K E I R 4. 50+3. 80+4. 50 -3. 90 -3. 50+4. 50 -4. 50 = 5. 40 => 5. 4/7=0. 77 Move to the next position in the sequence L I K E I R Q +3. 80+4. 50 -3. 90 -3. 50+4. 50 -4. 50 – 3. 50 = => -2. 6/7=-0. 37 J. Leunissen

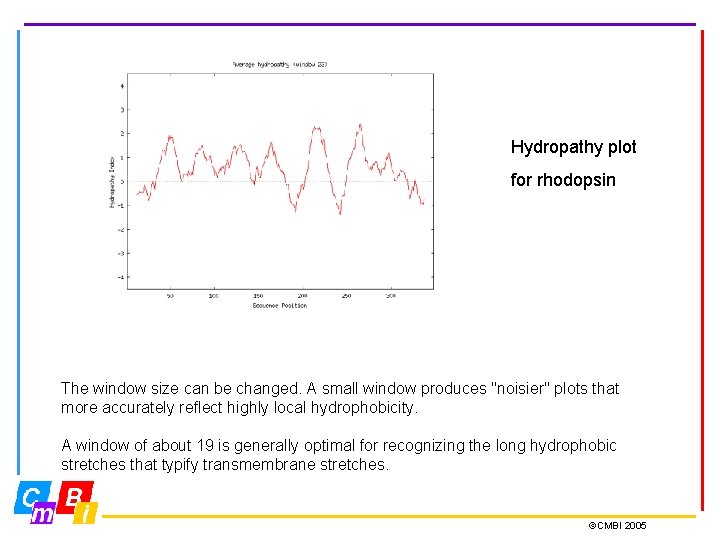
Hydropathy plot for rhodopsin The window size can be changed. A small window produces "noisier" plots that more accurately reflect highly local hydrophobicity. A window of about 19 is generally optimal for recognizing the long hydrophobic stretches that typify transmembrane stretches. ©CMBI 2005

Transmembrane Helices Transmembrane proteins are integral membrane proteins that interact extensively with the membrane lipids. Nearly all known integral membrane proteins span the lipid bilayer Hydropathy analysis can be used to locate possible transmembrane segments The main signal is a stretch of hydrophobic and helix-loving amino acids ©CMBI 2005

Transmembrane Helices (2) In a -helix the rotation is 100 degrees per amino acid The rise per amino acid is 1, 5 Å To span a membrane of 30 Å approx. 30/1, 5 = 20 amino acids are needed ©CMBI 2005

Transmembrane Helix Prediction Servers 1. KDD 2. Tmpred (database Tmbase) 3. DAS 4. Top. Pred II 5. TMHMM 2. 0 6. MEMSAT 2 7. SOSUI 8. HMMTOP 2. 0 ©CMBI 2005

Antigenic Prediction General Remarks Antibodies are a powerful tool for life science research They find multiple application in a variety of areas including biotechnology, medicine and diagnosis. Antibodies can recognize either linear or 3 D epitopes There are rules to predict what peptide fragments from a protein are likely to be antigenic ©CMBI 2005

Antigenic Prediction 1. Antigenic peptides should be located in solvent accessible regions and contain both hydrophobic and hydrophilic residues • • Determine solvent accessibility in case 3 D coordinates are available. If you have only a sequence, predict the accessibilities. 2. The peptide should also adopt a conformation that mimics its shape when contained within the protein. • • Preferably select peptides lying in long loops connecting secondary structure motifs. Neither the peptide stand-alone, nor the peptide in the full protein should be helical. ©CMBI 2005

Rules of thumb in antigenic prediction • N- and C- terminal peptides sometimes work better than peptides elsewhere in the protein. • Avoid peptides with internal sequence repeats or near repeats. • Avoid sequences that look funny (i. e. avoid low complexity sequences). • Try to avoid prolines and cysteines. • Last, but not least, use antigenicity prediction programs. ©CMBI 2005

Signal Peptides Proteins have intrinsic signals that govern their transport and localization in the cell (nucleus, ER, mitochondria, chloroplasts) Specific amino acid sequences determine whether a protein will pass through a membrane into a particular organelle, become integrated into the membrane, or be exported out of the cell. ©CMBI 2005

Signal Peptides (2) The common structure of signal peptides from various proteins is described as: • a positively charged (N-terminal) n-region • followed by a hydrophobic h-region (which can adopt an -helical conformation in an hydrophobic environment) • and a neutral but polar c-region (cleavage region; the signal sequence is cleaved off here after delivering the protein at the right site). The (-3, -1) rule states that the residues at positions – 3 and – 1 (relative to the cleavage site) must be small and neutral for cleavage to occur correctly. ©CMBI 2005

Signal Peptides (3) Marlinda. Hupkes 2004

Prediction of Signal Peptides Prokaryotes and Eukaryotes: Signal. P 3. 0 SPScan Sig. Cleave PSORT Eukaryotes: SIGFIND Target. P Specific localization signals: Predict. NLS - Nuclear Localization Signals Chloro. P – Chloroplast transit peptides Net. Nes – Nuclear Export Signals ©CMBI 2005

Repeats in proteins • Although they are usually found in non-coding genomic regions, repeating sequences are also found within genes. • Ranging from repeats of a single amino acid, through three residue short tandem repeats (e. g. in collagen), to the repetition of homologous domains of 100 or more residues. • Duplicated sequence segments occur in 14 % of all proteins, but eukaryotic proteins are three times more likely to have internal repeats than prokaryotic proteins ©CMBI 2005

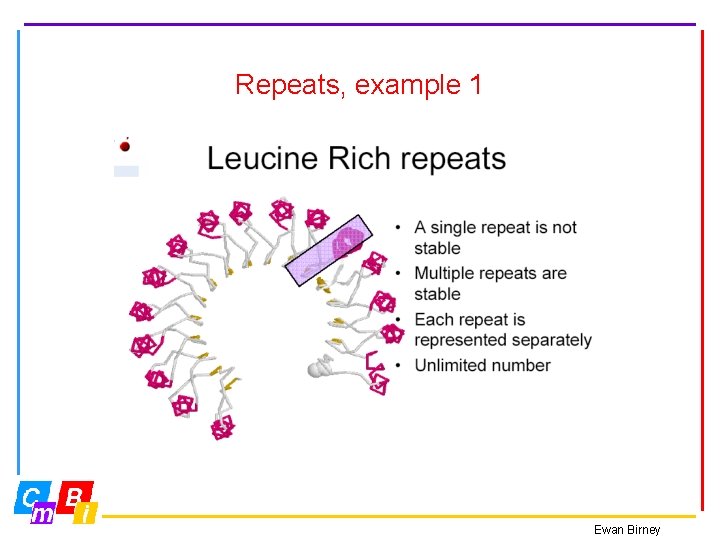
Repeats, example 1 Ewan Birney

Repeats, example 2 ©CMBI 2005

Prediction of Repeats 1. Repsim (a database of simple repeats) 2. Rep (Searches a protein sequence for repeats) 3. RADAR (Rapid Automatic Detection and Alignment of Repeats in protein sequences. ) 4. REPRO (De novo repeat detection in protein sequences) 5. Other? ©CMBI 2005

Coiled-Coils The coiled-coil is a ubiquitous protein motif that is often used to control oligomerisation. It is found in many types of proteins, including transcription factors, viral fusion peptides, and certain t. RNA synthetases. Most coiled-coil sequences contain heptad repeats - seven residue patterns denoted abcdefg in which the a and d residues (core positions) are generally hydrophobic. A number of programs are available to predict coiled-coil regions in a protein: COILS, PAIRCOILS, MULTICOILS. ©CMBI 2005