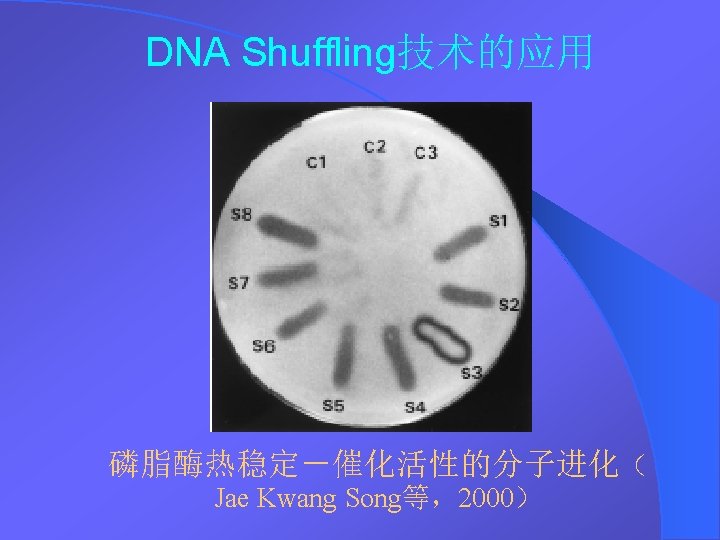
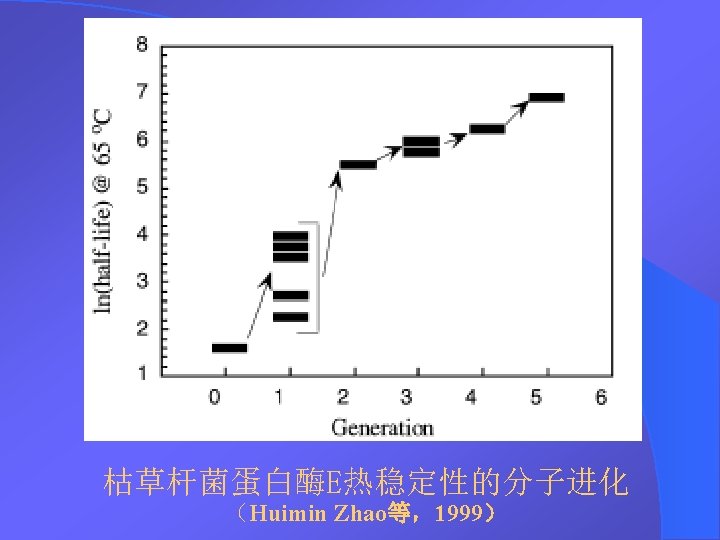
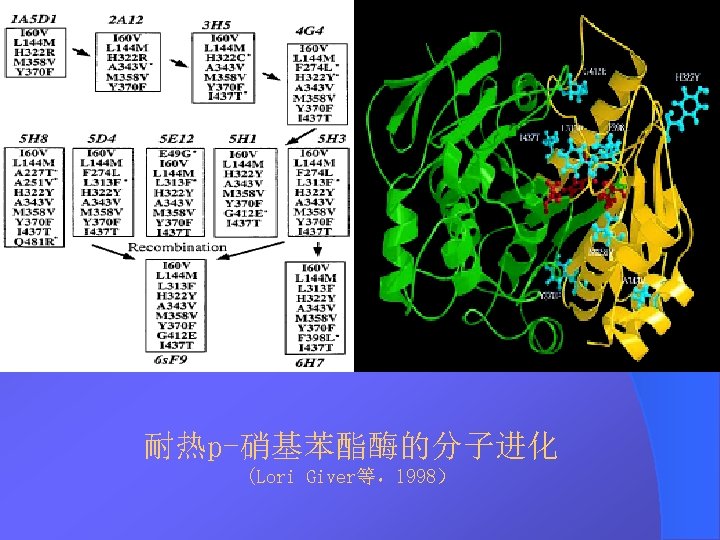
DNA Shuffling DNA DNA Shuffling DNA DNA 1994StemmerDNA

























![OD 540 nm (cell viability) Results-Potency of IFN-as Log 10[IFN-a] (mg/ml) OD 540 nm (cell viability) Results-Potency of IFN-as Log 10[IFN-a] (mg/ml)](https://slidetodoc.com/presentation_image_h2/1b444008c7ee334a8d8fccdc80dade10/image-44.jpg)




- Slides: 49



















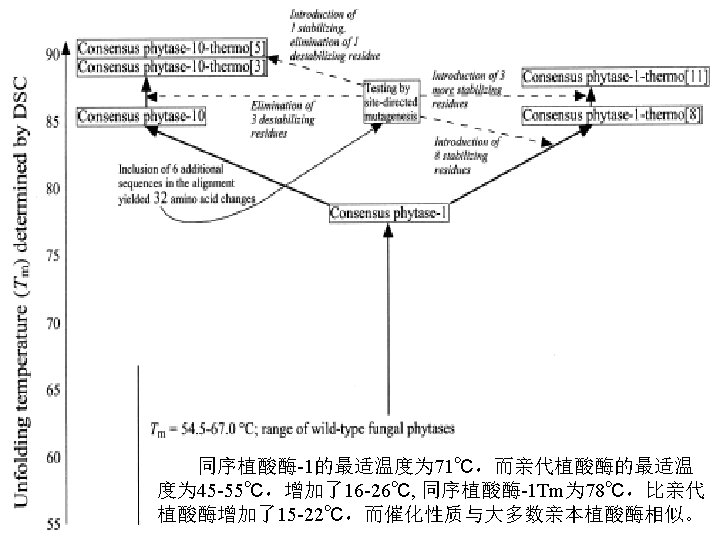
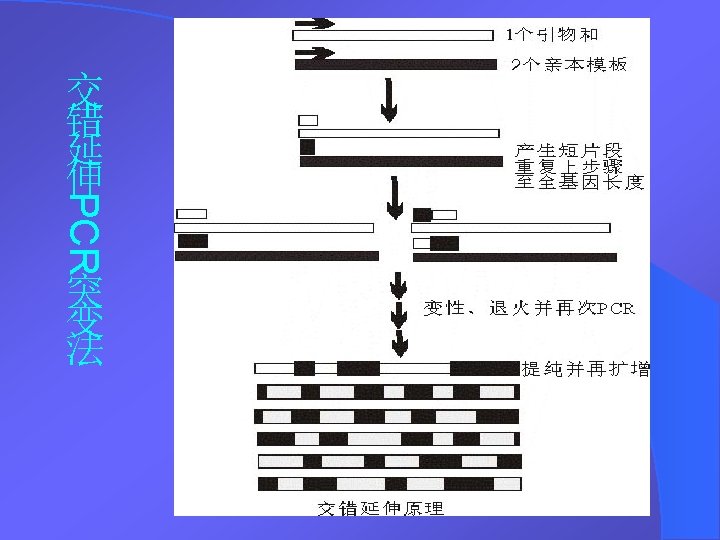
DNA Shuffling技术 DNA改组 DNA Shuffling DNA洗牌 DNA搅乱重排 1994年,Stemmer等,用DNA Shuffling技术体外快 速进化蛋白——有性PCR法 1997年 ,France Aronld研究组将DNA Shuffling技 术做了改进——交错延伸法

DNA Shuffling的内涵 What is DNA shuffling? v crossovers, Darwinian Evolutionv deletions, v insertions, v inversions, Natural selection of v point mutations existing mutations Directed Evolution Targeted selection of created mutations What happens after DNA shuffling Ø Generation of a large library of novel genes (chimeras) Ø Selection for improved/desired bio-functions



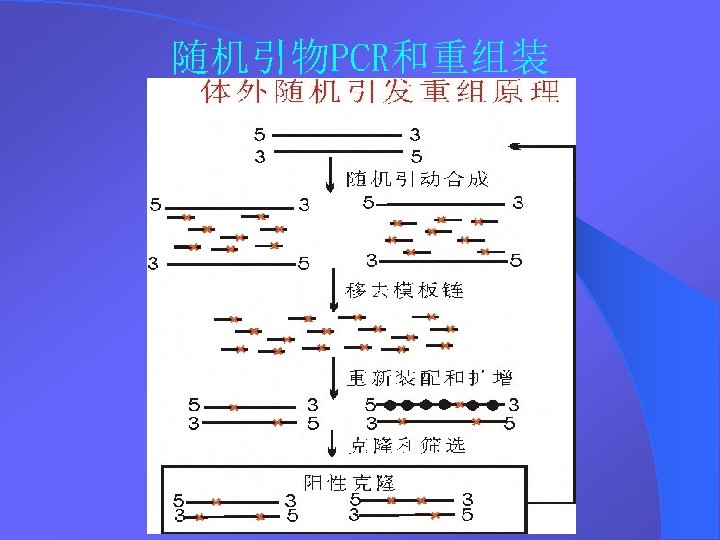
How DNA shuffling works - 1 How DNA shuffling is done in the tube v Random fragmentation of a pool of related genes; v Self-priming polymerase reaction and template switching (causing crossovers); v PCR amplification with primers of reassembled products

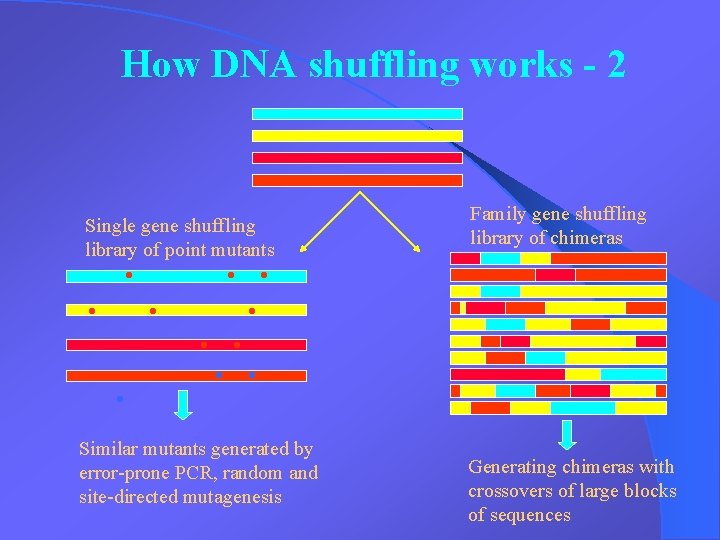
How DNA shuffling works - 2 Single gene shuffling library of point mutants . . . Similar mutants generated by error-prone PCR, random and site-directed mutagenesis Family gene shuffling library of chimeras Generating chimeras with crossovers of large blocks of sequences

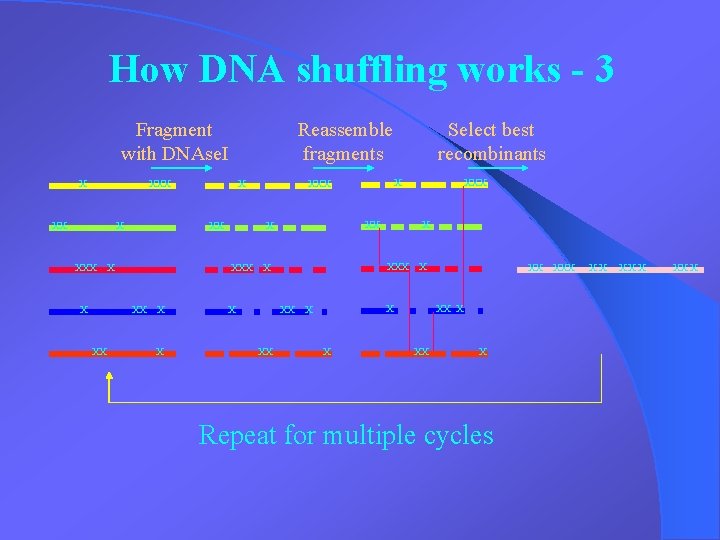
How DNA shuffling works - 3 Fragment with DNAse. I X Reassemble fragments XXX XX X X XXX XX XX X XX XXX X Select best recombinants X XX XX X Repeat for multiple cycles XXX XX X










Evolution of a cytokine using DNA family shuffling Chia-Chun J. Chang ……. Phillip Patten Nature Biotechnology Vol. 17, August, 1999 Aims of the work: Demonstration of usefulness of the DNA shuffling tool for rapidly evolving human a interferon towards improving antiviral property

Interferons (IFNa, IFNb, IFNg) Ø IFNas are produced by monocyte, macrophages and other cells upon stimulation by virus. Ø IFNs are able to protect cells from viral infection by reacting with IFN receptors on the cell which triggers a series of signal transduction events, hence immune response. Ø IFNs suppress an exaggerated growth potential of cancer cells. Ø IFNs can be used as drugs in the treatment of viral infection. Ø Only one natural IFN-a, Hu-IFN-a 2, in clinical studies. Ø Most active engineered IFN-a, alfacon-1, is a consensus of 13 wild type Hu-IFN-a genes that is used in hepatitis C therapy. Reasons for their limited use as drugs: v. Dose-limiting toxicity, v. Receptor cross-reactivity, v. Short serum half-lives

Methods and Materials-1 Ø DNA Shuffling Construction of 1 st round library: 8 cloned Hu-IFN genes were shuffled and assembled insert cloned in the phagemid display vector Construction of 2 nd round 5 libraries: IFN-CH 1. 1 x. IFN-CH 1. 2; IFN-CH 1. 1 x IFN-CH 1. 3; IFN-CH 1. 1 x. IFN-CH 1. 4; IFN-CH 1. 2 x IFN-CH 1. 3; Pair-wise matings IFN-CH 1. 1 x. IFN-CH 1. 2 x IFN-CH 1. 3 x IFN-CH 1. 4 (mating of 4 genes).

ØPhagemid display of IFN ØHigh-throughput phagemid preps (Q-BOT colony picker) ØAntiviral assays: Cytopathic effect reduction assay on mouse L 929 cells, challenging virus. EMCV, 96 well plate format, neutral red, cell fixing, colour extracting, spec OD 540 vs IFNa concentration.

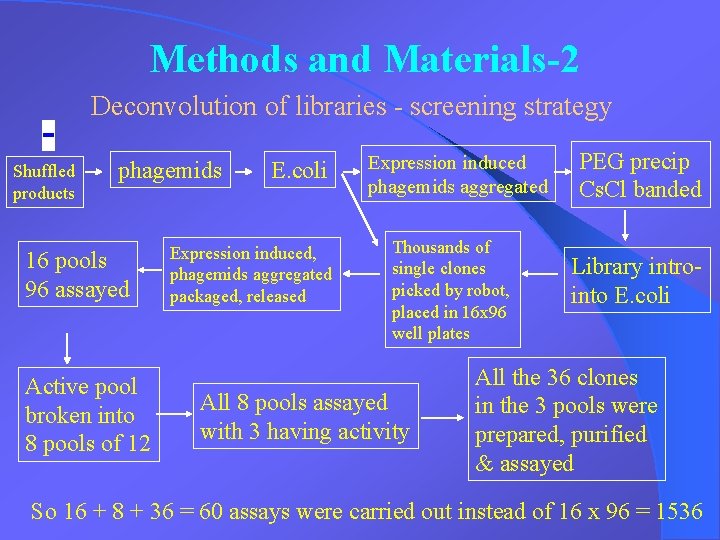
Methods and Materials-2 Deconvolution of libraries - screening strategy Shuffled products phagemids 16 pools 96 assayed Active pool broken into 8 pools of 12 E. coli Expression induced, phagemids aggregated packaged, released Expression induced phagemids aggregated Thousands of single clones picked by robot, placed in 16 x 96 well plates All 8 pools assayed with 3 having activity PEG precip Cs. Cl banded Library introinto E. coli All the 36 clones in the 3 pools were prepared, purified & assayed So 16 + 8 + 36 = 60 assays were carried out instead of 16 x 96 = 1536

Methods and Materials-3 CHO (Chinese Hamster Ovary) expression and purification (His tag) of chimeric IFN-as. Ø High-throughput proliferation assays (inhibition of proliferation) in mouse L 929 cells: Standard [3 H]thymidine incorporation methods--IFN-a samples were mixed with L 929 cells in 96 well plates, incubated and [H 3] thymidine added to each well. Plates were harvested on Harvester-96 and thymidine incorporation was counted on a b counter Ø Human Daudi cell proliferation assays: 4 most active chimeras and Hu-IFN-a 2 a were expressed, purified and assayed for anti-proliferation activity on human Daudi cells

Results-1: structure of shuffled IFNas B: The sequence of one of the cycle 2 chimeras, IFN-CH 2. 2, is aligned with the most potent human and mouse IFN-as. The IFN-a residues that putatively contact the IFN-a receptor are boxed. Residues in Hu-IFN-a 1 that have been shown by site -directed mutagenesis to contribute to activity on mouse cells are shaded.

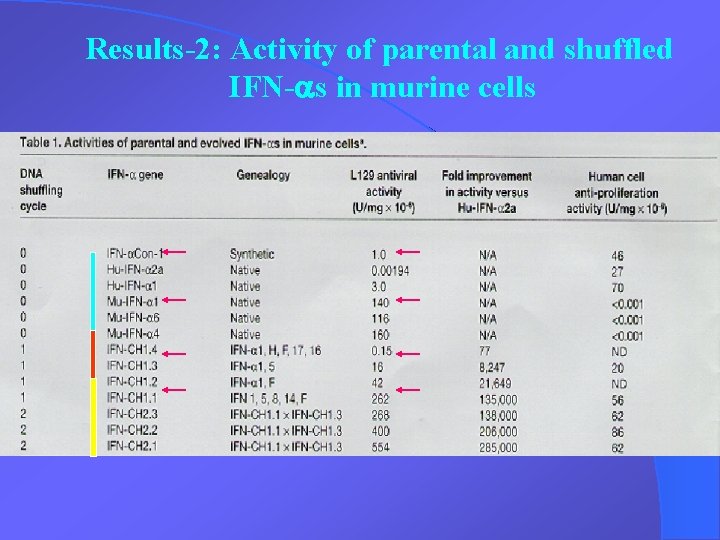
Results-2: Activity of parental and shuffled IFN-as in murine cells

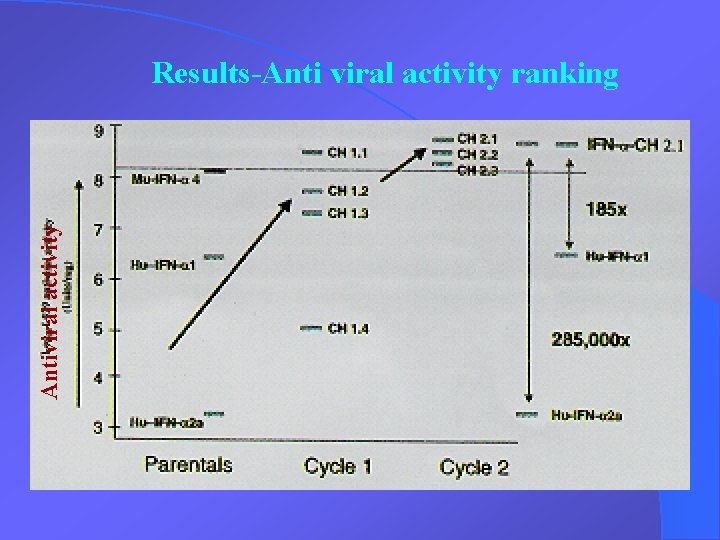
Antiviral activity Results-Anti viral activity ranking
![OD 540 nm cell viability ResultsPotency of IFNas Log 10IFNa mgml OD 540 nm (cell viability) Results-Potency of IFN-as Log 10[IFN-a] (mg/ml)](https://slidetodoc.com/presentation_image_h2/1b444008c7ee334a8d8fccdc80dade10/image-44.jpg)
OD 540 nm (cell viability) Results-Potency of IFN-as Log 10[IFN-a] (mg/ml)

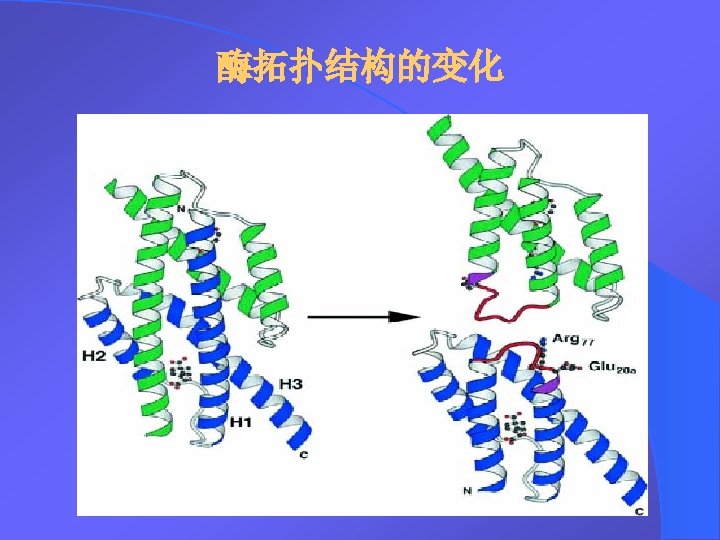
Results-continued Structural modelling of IFNCH 2. 2 based on the NRM structure of Hu-IFN-a 2 a. The protein backbone is coloured to indicate the native Hu-IFN-a segments (5 Hu-IFN-as from which it is derived). The side chains of putative murine IFN-a receptor contacting residues Lys 121 and Arg 125 are shown

Conclusions 1. Unlike site-directed mutagenesis and other protein engineering methods which produce mainly point mutations, DNA family shuffling can generate recombinants with blocks of sequences from more than two parental genes, hence enhancing genetic diversity. 2. DNA family shuffling of Human-IFN-a genes generates recombinants with potent IFN-a activity on a distantly related species (mouse). 3. Evolution of commercial genes and proteins may be practical, even when very complex, timeconsuming, or expensive assays are required.


