Microarrays with an emphasis on DNA microarrays BE

Microarrays with an emphasis on DNA microarrays BE 4332 Final Project Natalie Derise

Formal Definition A microarray is a hybridization-based technique that allows simultaneous analysis of thousands of samples of biological material on a solid substrate.

Common Types of Microarrays Type of array General function Protein expression and interactions profiling Tissue Compare histologic sections from unique tissues or tumors; tissues from multiple patients on same slide Cellular Reverse transfection and PMHC; study cell responses Antibody Detects antigens and protein expression; many diagnostic applications DNA Measure gene activity and genotyping • • Additional types of assays: Chemical compound microarrays Carbohydrate microarrays Phenotype micro arrays Many more specific types under development

DNA Microarrays • A DNA microarray consists of pre-designed synthetic nucleic acid probes that are immobilized and spatially arrayed on a solid matrix. • DNA microarrays rely on the hybridization between c. DNA that is reverse transcribed from a biological sample to the pre-designed probes on the array.

Common Terms • Array: refers to the glass, plastic, or silicon slide that the DNA probes will be spotted or built on. • Target DNA/RNA*: the nucleic acid (c. DNA or c. RNA) sample that is being identified and/ or measured. • Probe: short sections of oligonucleotides that are attached to the array and hybridize with the target c. DNA or c. RNA • Hybridization: the process of combining two complementary single-stranded DNA or RNA molecules and allowing them to form a single double-stranded molecule through base pairing. *RNA microarrays are very similar to DNA microarrays, but involve RNA probes hybridizing to target c. RNA (also known as anti-sense RNA)

General Process Overview 1. Using PCR or other techniques, synthesize probes specific to sequence(s) of interest and attach them to array; another option is to purchase premade arrays specific to gene(s) of interest 2. Isolate m. RNA from cells of interest 3. Transform this m. RNA into c. DNA by reverse transcriptase using fluorescently labeled nucleotides in order to create labeled c. DNA • for RNA microarrays reverse transcriptase is omitted and the m. RNA is labeled 4. The labeled c. DNA is washed over the microarray and will bind to any matching probes
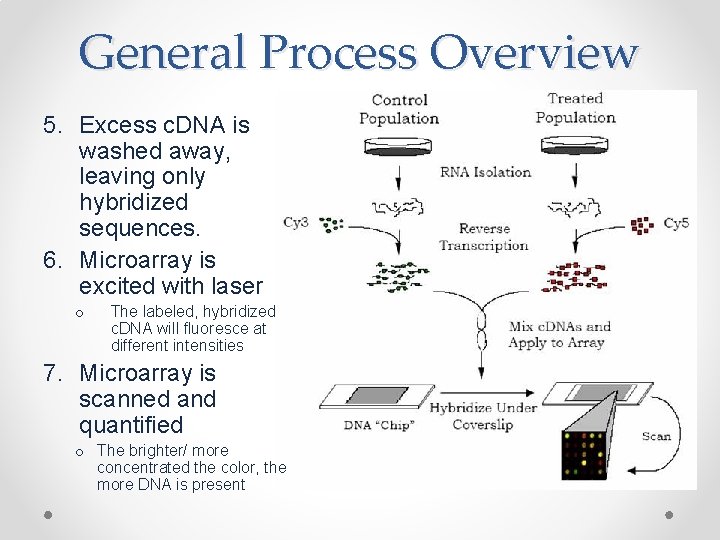
General Process Overview 5. Excess c. DNA is washed away, leaving only hybridized sequences. 6. Microarray is excited with laser o The labeled, hybridized c. DNA will fluoresce at different intensities 7. Microarray is scanned and quantified o The brighter/ more concentrated the color, the more DNA is present
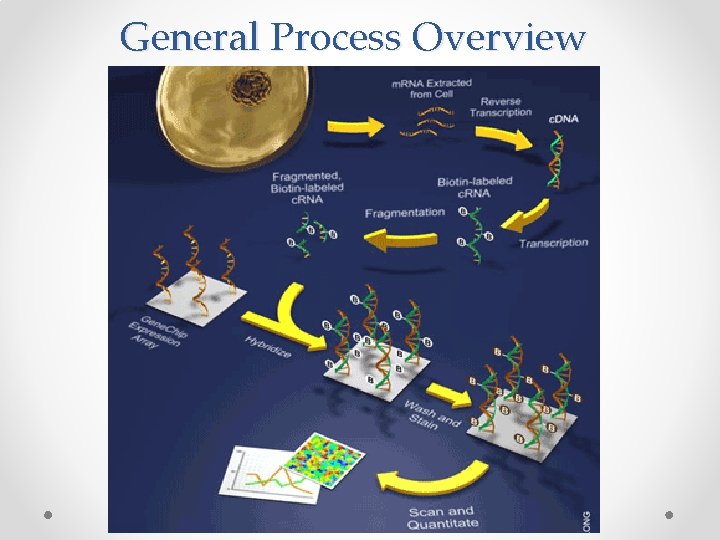
General Process Overview

Attaching the Probe to the Array • DNA fragments are chemically tethered to glass, plastic, nylon, or silicon biochips (also known as DNA chips) • This is done in different ways according to which type of microarray is being used. The two common types are: o Spotted arrays o In situ synthesized arrays

Spotted Arrays • The nucleotide sequences of interest are generated by PCR and “spotted” onto the array surface by a robot. • Sequences are ~1 kb in length • Relatively low costs and easy to synthesize for smaller gene sets http: //www. youtube. com/watch? v=3 ZXq_a. Df. SB 8

In situ Synthesized Arrays • Also known as photolithographic arrays • Sequences of interest are synthesized directly on the array surface http: //www. youtube. com/watch? v=Mu. N 54 ecf. HPw (view up to 30 second mark) • Sequences are ~30 -70 bp long more specific binding and more probes per slide • Much more expensive than spotted o The individual masks must be designed and manufactured. This is the most tedious step, and is what makes the process pricey. o To combat this cost, Digital Light Processing (DLP) has been developed; this process uses a set of movable micromirrors to apply light to certain areas on the array. This computerized process bypasses the need for the masks.
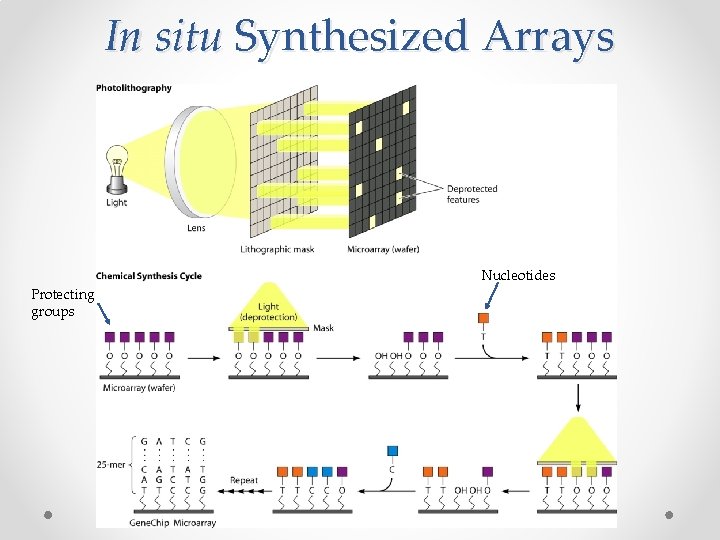
In situ Synthesized Arrays Nucleotides Protecting groups

Types of Signal Detection Two color • Control and experimental samples labeled with two different fluorophores • Hybridized on same array • Allows for direct comparisons • Cheaper (also commonly used with spotted) • Can compare ratio of two genes to determine if their expression levels are related • Sensitive to error, more complex design One color • Each sample labeled with same fluorophore • Expression is compared between multiple arrays • Usually used with in situ; also requires twice as many arrays $$ • Less prone to error • Simpler design
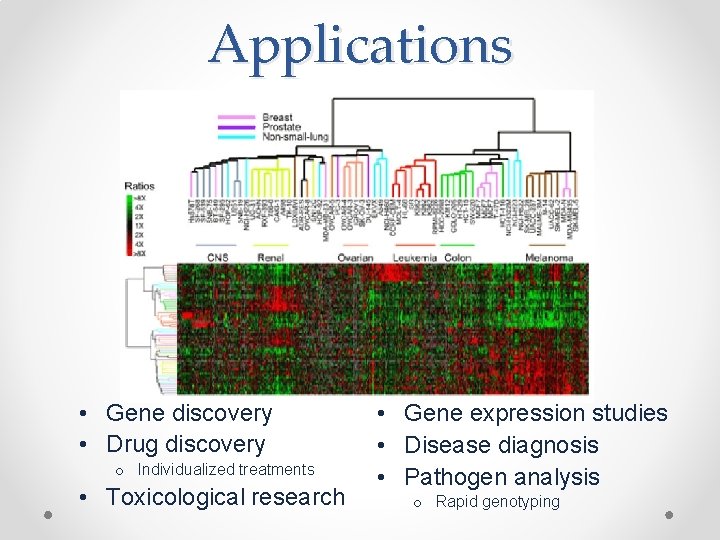
Applications • Gene discovery • Drug discovery o Individualized treatments • Toxicological research • Gene expression studies • Disease diagnosis • Pathogen analysis o Rapid genotyping

Microarray Pros and Cons Pros • Many pre-made probes available in market • Prior knowledge to gene sequence is not required • Large c. DNA works well with hybridization • Spotting technology is readily available • Process is fast and flexible • Can be used for all organisms • Study multiple genes at once Cons • Contamination potential extremely high • Cross-hybridization with similar, repetitive gene families can create false positives • Not extremely quantitative • In situ can become very expensive • Many possible errors during scanning due to background noise

In Summary: Complete process overview: http: //www. youtube. com/watch? v=3 j. X_08 zd. YCE Microarray technology is a very powerful tool used for many applications and if often paired with PCR. This field is continuing to grow, leading to cheaper prices as well as increasingly specialized applications.
- Slides: 16