DNA is Composed of Complementary Strands Base Pairing
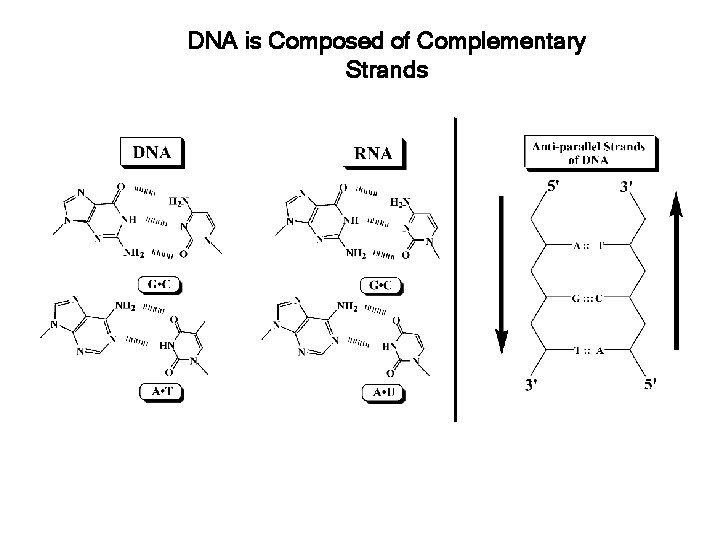
DNA is Composed of Complementary Strands
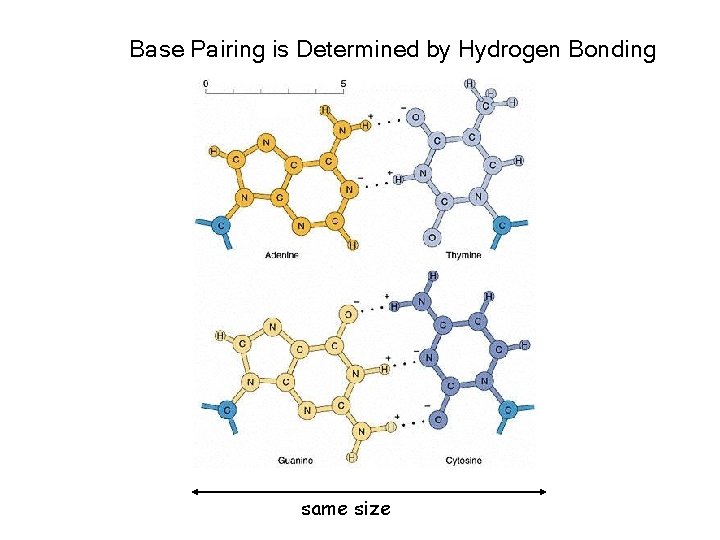
Base Pairing is Determined by Hydrogen Bonding same size

Forces stabilizing DNA double helix 1. Hydrogen bonding (2 -3 kcal/mol per base pair) 2. Stacking (hydrophobic) interactions (4 -15 kcal/mol per base pair) 3. Electrostatic forces.
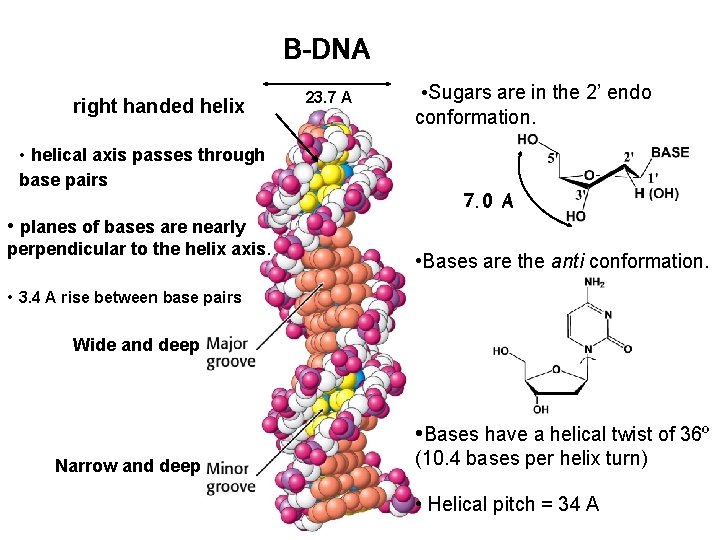
B-DNA right handed helix • helical axis passes through base pairs 23. 7 A • Sugars are in the 2’ endo conformation. 7. 0 A • planes of bases are nearly perpendicular to the helix axis. • Bases are the anti conformation. • 3. 4 A rise between base pairs Wide and deep Narrow and deep • Bases have a helical twist of 36º (10. 4 bases per helix turn) • Helical pitch = 34 A
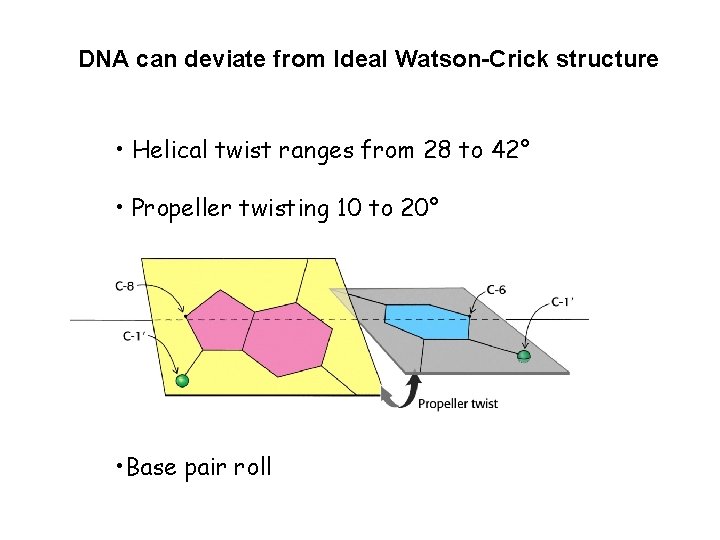
DNA can deviate from Ideal Watson-Crick structure • Helical twist ranges from 28 to 42° • Propeller twisting 10 to 20° • Base pair roll
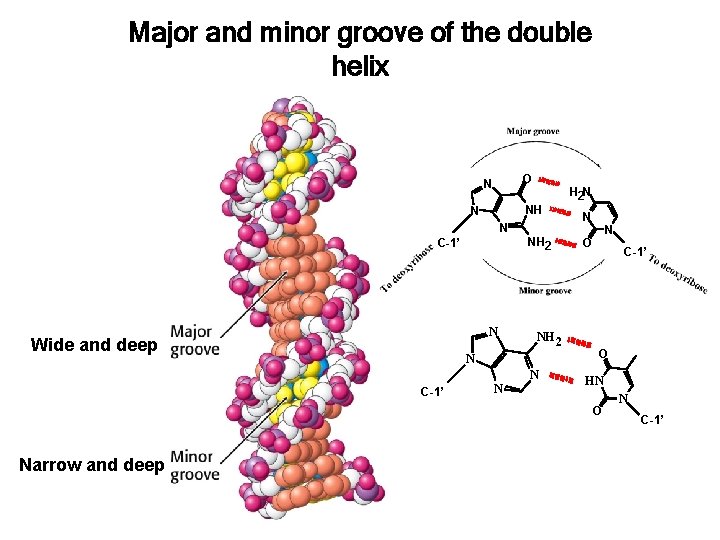
Major and minor groove of the double helix O N H 2 N NH N N NH 2 C-1’ N Wide and deep NH 2 N N C-1’ N N N O C-1’ O HN O Narrow and deep N C-1’
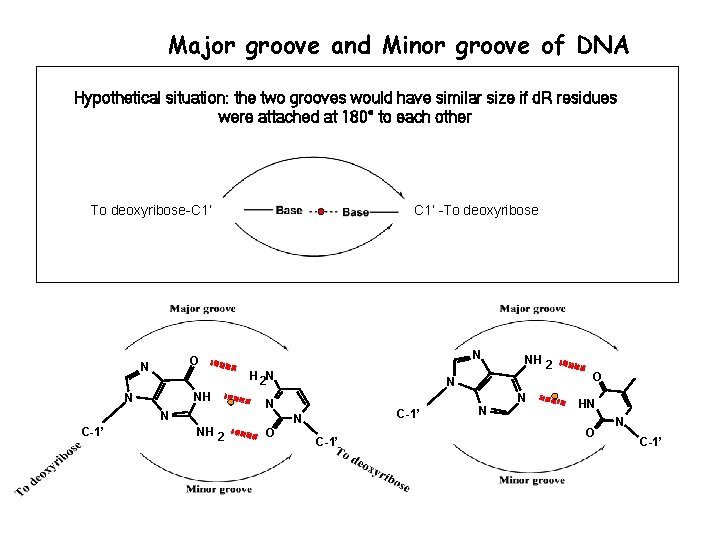
Major groove and Minor groove of DNA Hypothetical situation: the two grooves would have similar size if d. R residues were attached at 180° to each other To deoxyribose-C 1’ N O N H 2 N NH N N C-1’ C 1’ -To deoxyribose N N N C-1’ N NH 2 O NH 2 C-1’ N O HN O N C-1’
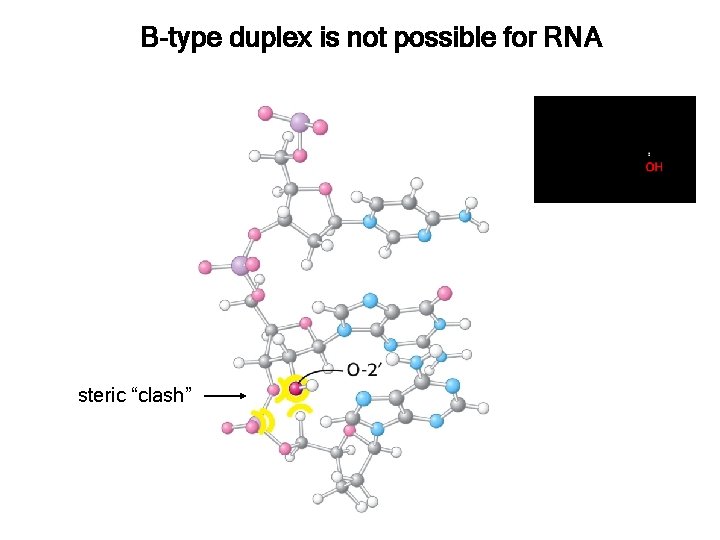
B-type duplex is not possible for RNA steric “clash”
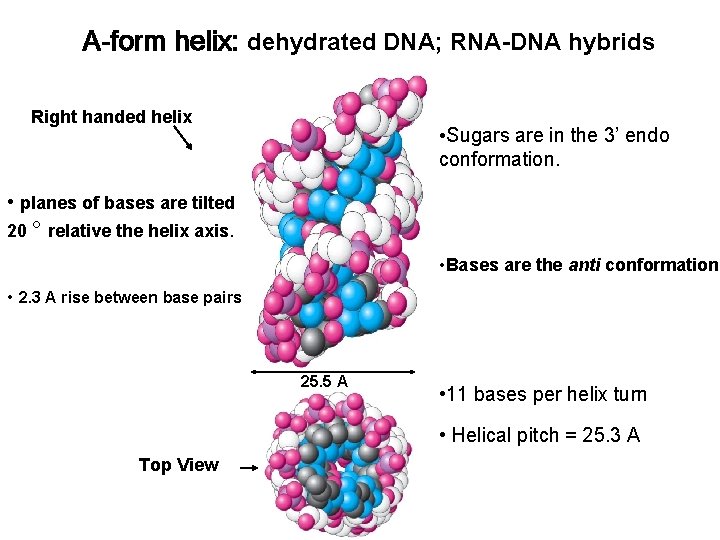
A-form helix: dehydrated DNA; RNA-DNA hybrids Right handed helix • Sugars are in the 3’ endo conformation. • planes of bases are tilted 20 ° relative the helix axis. • Bases are the anti conformation. • 2. 3 A rise between base pairs 25. 5 A • 11 bases per helix turn • Helical pitch = 25. 3 A Top View
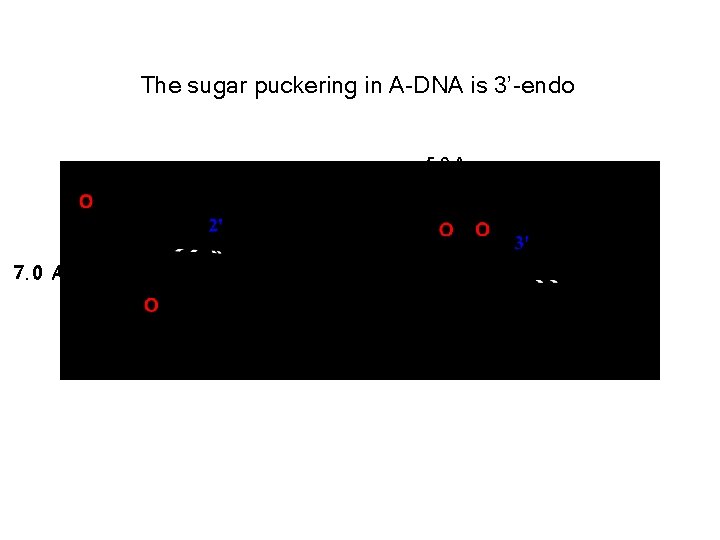
The sugar puckering in A-DNA is 3’-endo 5. 9 A 7. 0 A

Living Figure – A-DNA http: //bcs. whfreeman. com/biochem 5
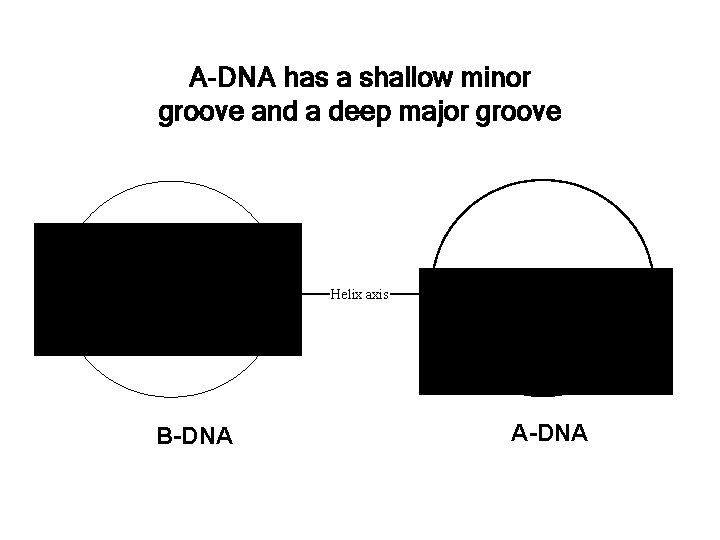
A-DNA has a shallow minor groove and a deep major groove • B-DNA Helix axis • A-DNA
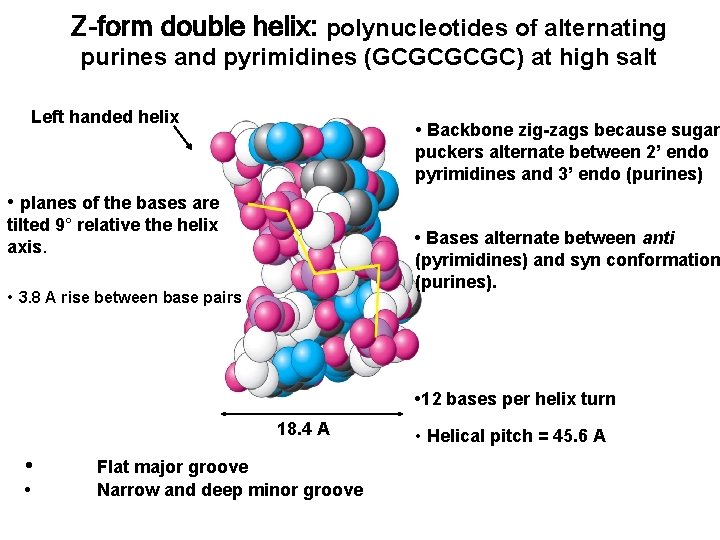
Z-form double helix: polynucleotides of alternating purines and pyrimidines (GCGC) at high salt Left handed helix • Backbone zig-zags because sugar puckers alternate between 2’ endo pyrimidines and 3’ endo (purines) • planes of the bases are tilted 9° relative the helix axis. • Bases alternate between anti (pyrimidines) and syn conformation (purines). • 3. 8 A rise between base pairs • 12 bases per helix turn 18. 4 A • • Flat major groove Narrow and deep minor groove • Helical pitch = 45. 6 A

Sugar and base conformations in Z-DNA alternate: 5’-GCGCGCG 3’-CGCGCGC C: sugar is 2’-endo, base is anti G: sugar is 3’-endo, base is syn

Living Figure – Z-DNA http: //bcs. whfreeman. com/biochem 5
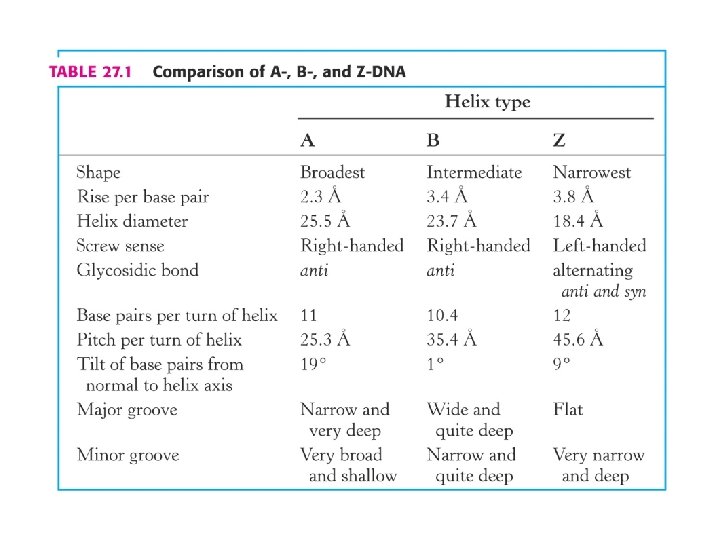
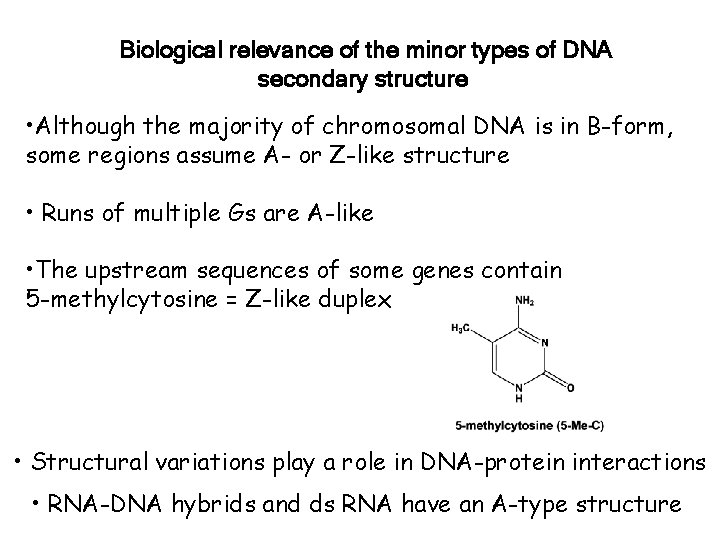
Biological relevance of the minor types of DNA secondary structure • Although the majority of chromosomal DNA is in B-form, some regions assume A- or Z-like structure • Runs of multiple Gs are A-like • The upstream sequences of some genes contain 5 -methylcytosine = Z-like duplex • Structural variations play a role in DNA-protein interactions • RNA-DNA hybrids and ds RNA have an A-type structure
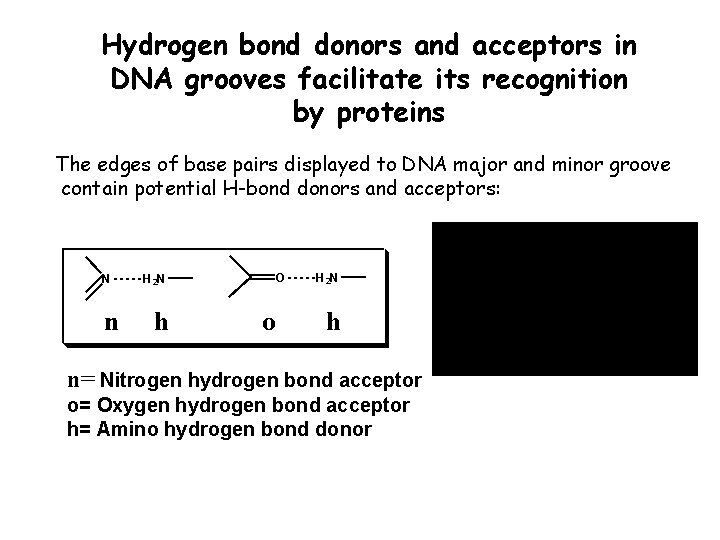
Hydrogen bond donors and acceptors in DNA grooves facilitate its recognition by proteins The edges of base pairs displayed to DNA major and minor groove contain potential H-bond donors and acceptors: N n O H 2 N h o H 2 N h n= Nitrogen hydrogen bond acceptor o= Oxygen hydrogen bond acceptor h= Amino hydrogen bond donor
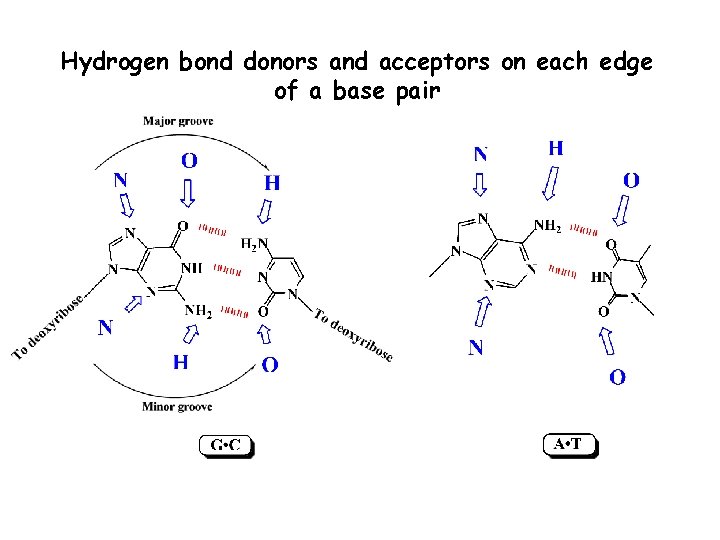
Hydrogen bond donors and acceptors on each edge of a base pair
Structural characteristics of DNA facilitating DNA-Protein Recogtnition 1. Major and major groove of DNA contain sequence 2. dependent patterns of H-bond donors and acceptors. 2. Sequence-dependent duplex structure (A, B, Z, bent 3. DNA). 3. Hydrophobic interactions via intercalation. 4. Ionic interactions with phosphates.
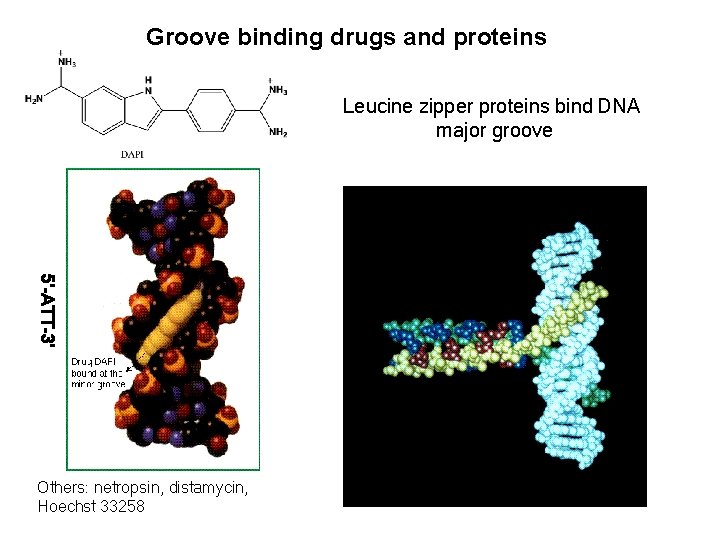
Groove binding drugs and proteins Leucine zipper proteins bind DNA major groove 5’-ATT-3’ Others: netropsin, distamycin, Hoechst 33258
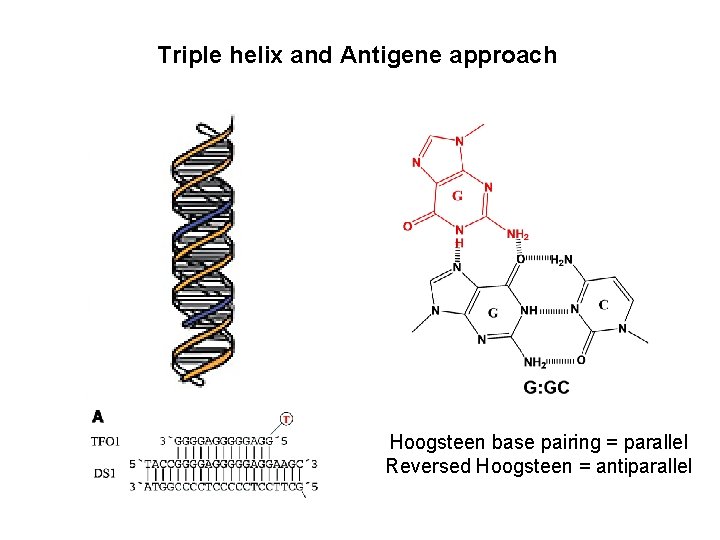
Triple helix and Antigene approach Hoogsteen base pairing = parallel Reversed Hoogsteen = antiparallel

Biophysical properties of DNA • Facile denaturation (melting) and re-association of the duplex are important for DNA’s biological functions. In the laboratory, melting can be induced by heating. • Single strands T° duplex • Hybridization techniques are based on the affinity of complementary DNA strands for each other. • Duplex stability is affected by DNA length, % GC base pairs, ionic strength, the presence of organic solvents, p. H • Negative charge – can be separated by gel electrophoresis

Separation of DNA fragments by gel electrophoresis Polyacrylamide gel: • DNA strands are negatively charged – migrate towards the anode • Migration time ~ ln (number of base pairs)

DNA Topology DNA has to be coiled to fit inside the cell Organism Number of base pairs Contour length, m E. Coli bacteria 4, 600, 000 1, 360 SV 40 virus 5, 100 1. 7 Human chromosomes 48, 000240, 000 1. 6 – 8. 2 cm DNA polymers must be folded to fit into the cell or nucleus (tertiary structure).

DNA Topology:

DNA Topology: linking number
Consider a 260 bp B-duplex:
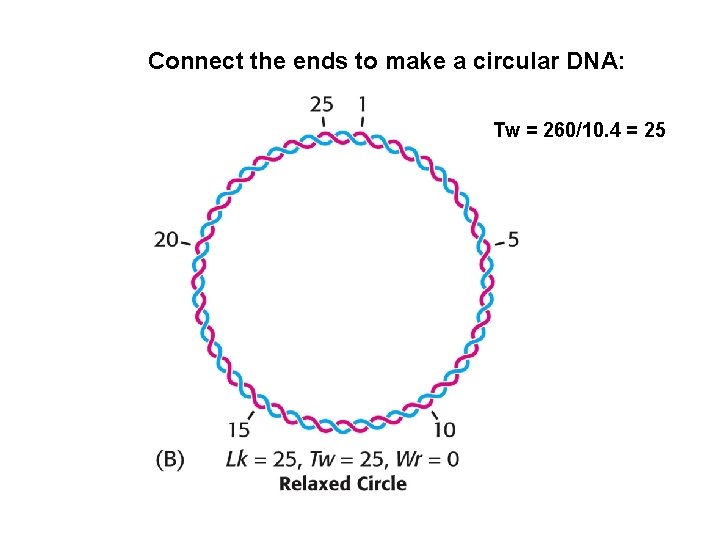
Connect the ends to make a circular DNA: Tw = 260/10. 4 = 25

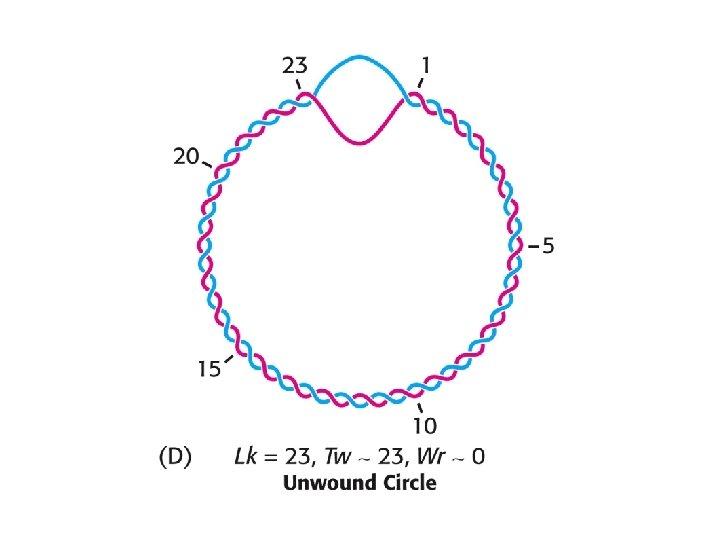
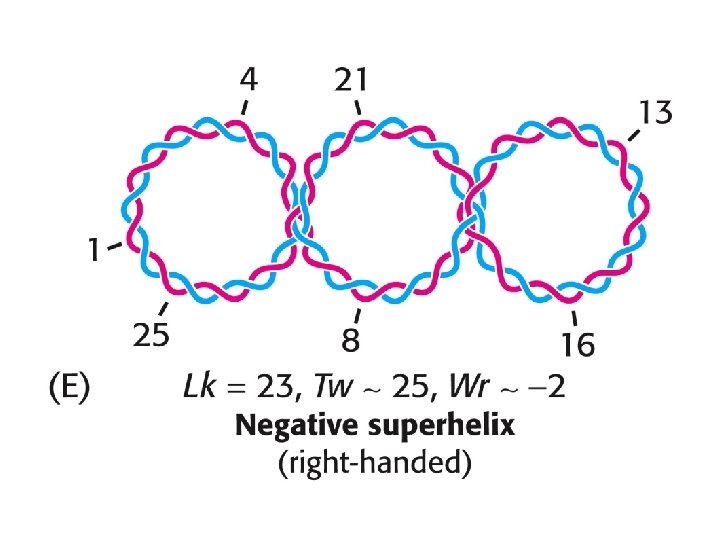
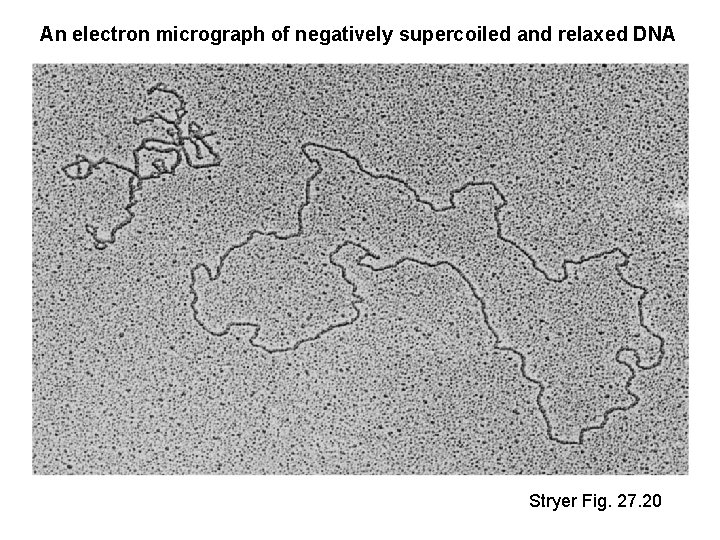
An electron micrograph of negatively supercoiled and relaxed DNA Stryer Fig. 27. 20
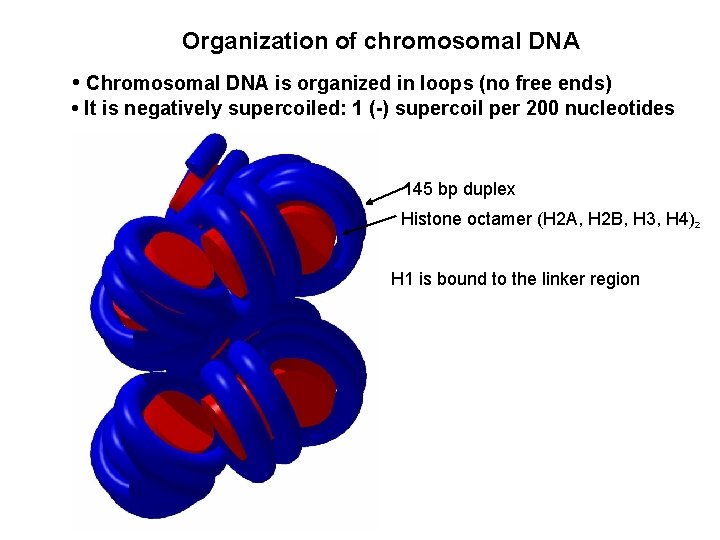
Organization of chromosomal DNA • Chromosomal DNA is organized in loops (no free ends) • It is negatively supercoiled: 1 (-) supercoil per 200 nucleotides 145 bp duplex Histone octamer (H 2 A, H 2 B, H 3, H 4)2 H 1 is bound to the linker region

Enzymes that control DNA supercoiling: DNA Topoisomerases Change the linking number (Lk) of DNA duplex by concerted breakage and re-joining DNA strands Topoisomerase enzymes Topoisomerases II Relax DNA supercoiling by increments of 1 (cleave one strand) Change DNA supercoiling by the increments of 2 (break both strands) Usually introduce negative supercoiling
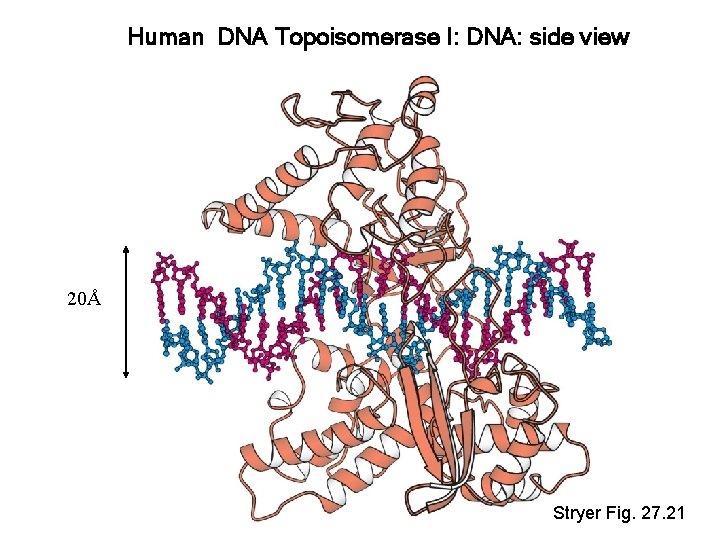
Human DNA Topoisomerase I: DNA: side view 20Å Stryer Fig. 27. 21
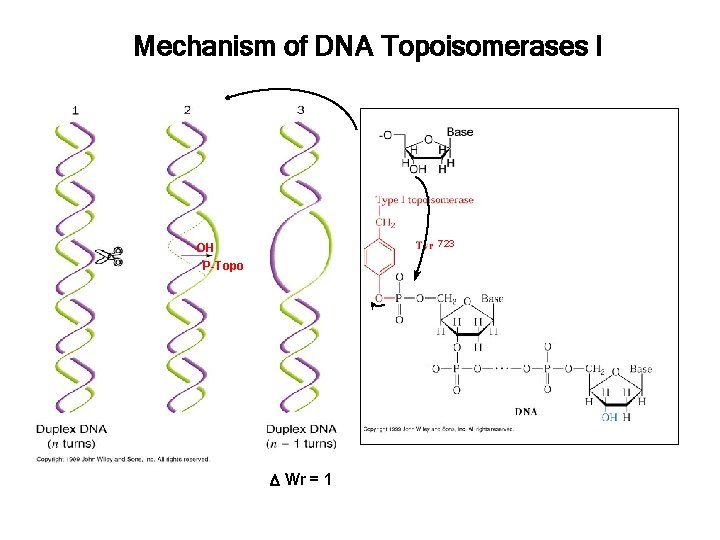
Mechanism of DNA Topoisomerases I 723 OH P-Topo Wr = 1
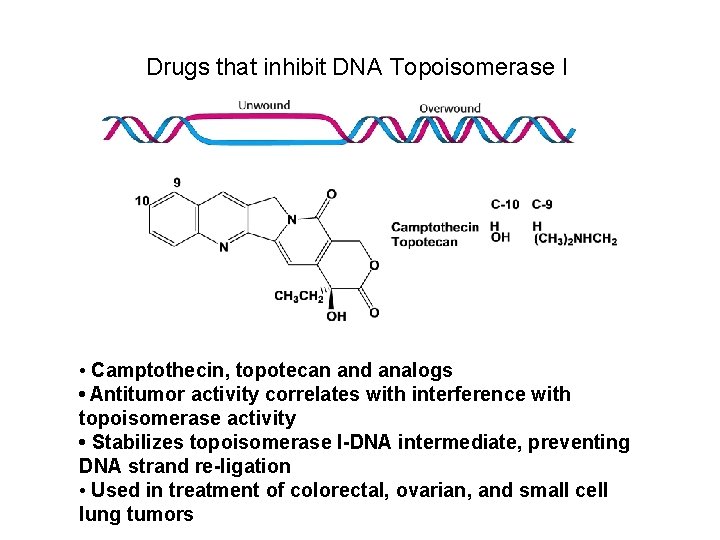
Drugs that inhibit DNA Topoisomerase I • Camptothecin, topotecan and analogs • Antitumor activity correlates with interference with topoisomerase activity • Stabilizes topoisomerase I-DNA intermediate, preventing DNA strand re-ligation • Used in treatment of colorectal, ovarian, and small cell lung tumors

Enzymes that control DNA supercoiling: DNA Topoisomerases Change the linking number (Lk) of DNA duplex by concerted breakage and re-joining DNA strands Topoisomerase enzymes Topoisomerases II Relax DNA supercoiling by increments of 1 (cleave one strand) Change DNA supercoiling by the increments of 2 (break both strands) Usually introduce negative supercoiling
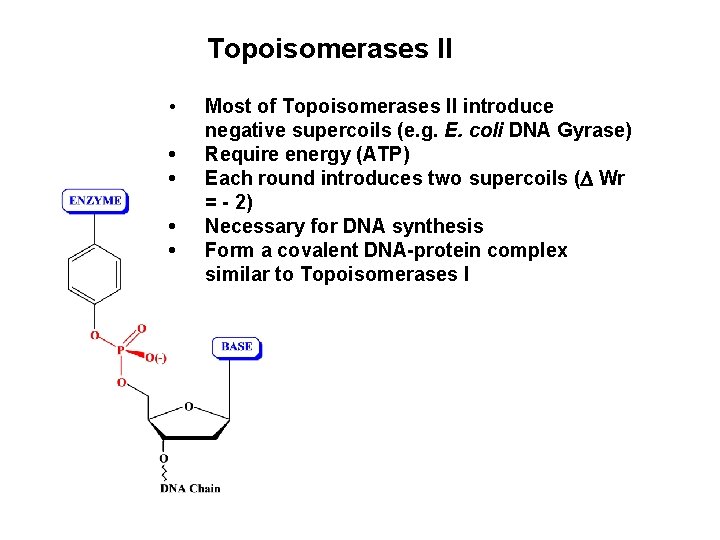
Topoisomerases II • • • Most of Topoisomerases II introduce negative supercoils (e. g. E. coli DNA Gyrase) Require energy (ATP) Each round introduces two supercoils ( Wr = - 2) Necessary for DNA synthesis Form a covalent DNA-protein complex similar to Topoisomerases I
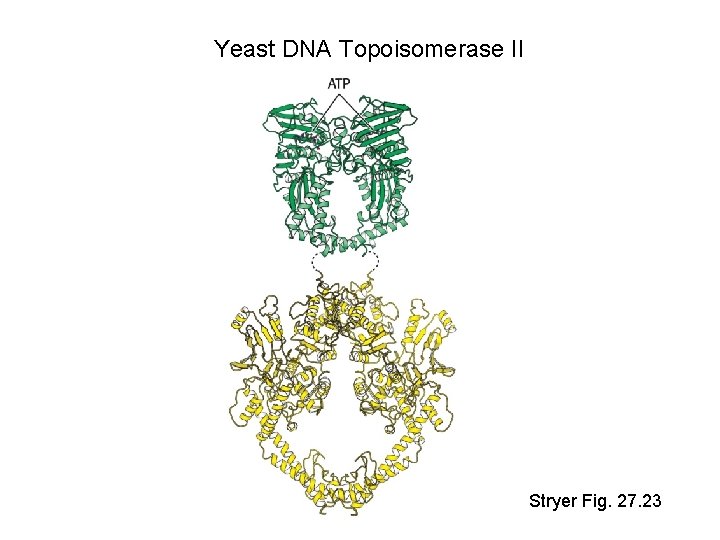
Yeast DNA Topoisomerase II Stryer Fig. 27. 23
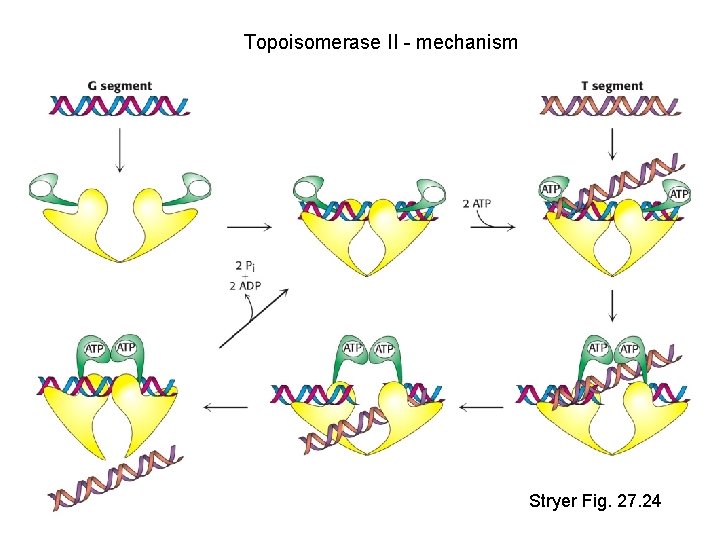
Topoisomerase II - mechanism Stryer Fig. 27. 24
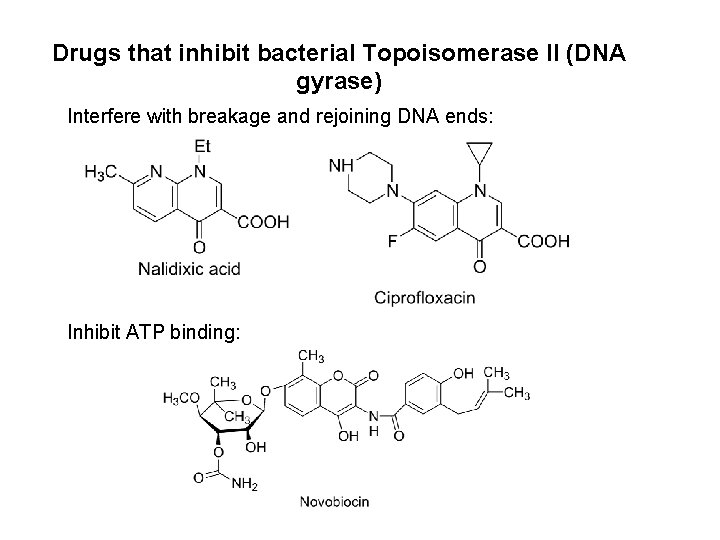
Drugs that inhibit bacterial Topoisomerase II (DNA gyrase) Interfere with breakage and rejoining DNA ends: Inhibit ATP binding:
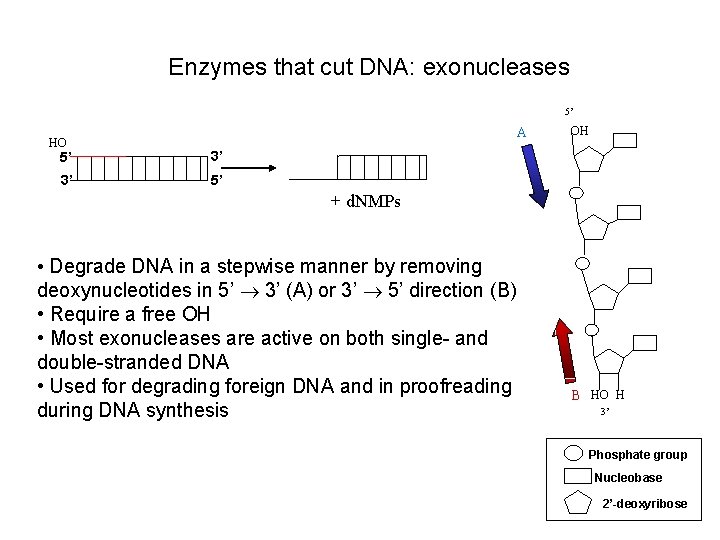
Enzymes that cut DNA: exonucleases 5’ HO A 5’ 3’ 3’ 5’ OH + d. NMPs • Degrade DNA in a stepwise manner by removing deoxynucleotides in 5’ 3’ (A) or 3’ 5’ direction (B) • Require a free OH • Most exonucleases are active on both single- and double-stranded DNA • Used for degrading foreign DNA and in proofreading during DNA synthesis B HO H 3’ Phosphate group Nucleobase 2’-deoxyribose
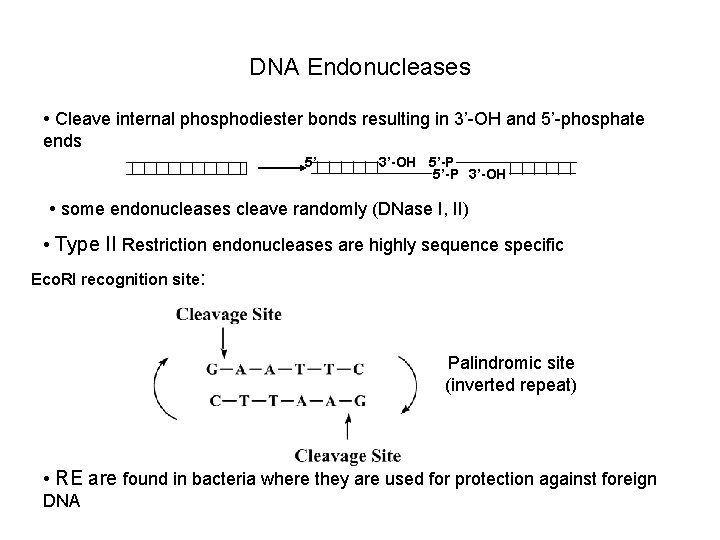
DNA Endonucleases • Cleave internal phosphodiester bonds resulting in 3’-OH and 5’-phosphate ends 5’ 3’-OH 5’-P 3’-OH • some endonucleases cleave randomly (DNase I, II) • Type II Restriction endonucleases are highly sequence specific Eco. RI recognition site: Palindromic site (inverted repeat) • RE are found in bacteria where they are used for protection against foreign DNA
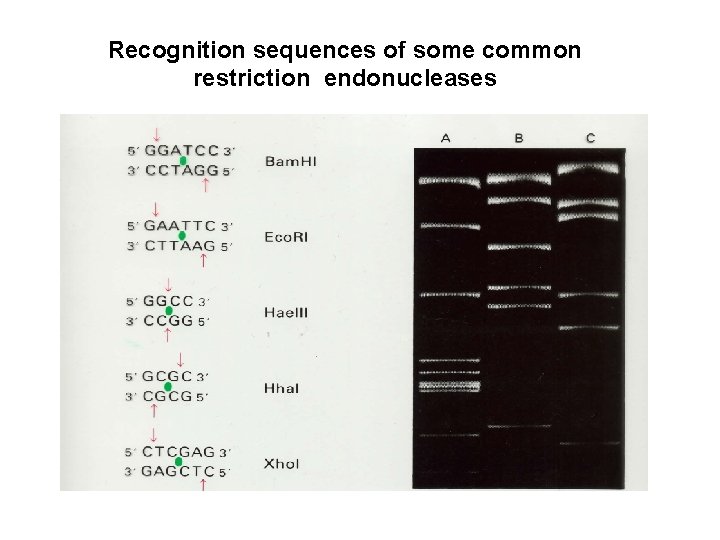
Recognition sequences of some common restriction endonucleases
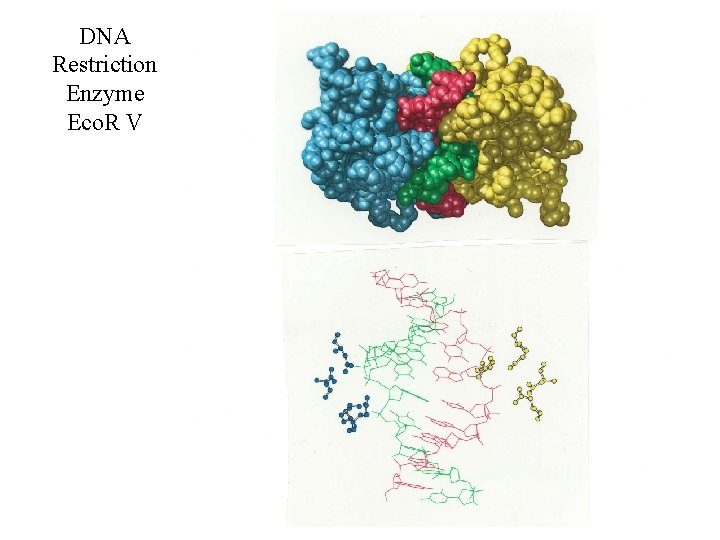
DNA Restriction Enzyme Eco. R V

Applications of Restriction Endonucleases in Molecular Biology 1. DNA fingerprinting (restriction fragment length polymorphism). 2. 2. Molecular cloning (isolation and amplification of genes).
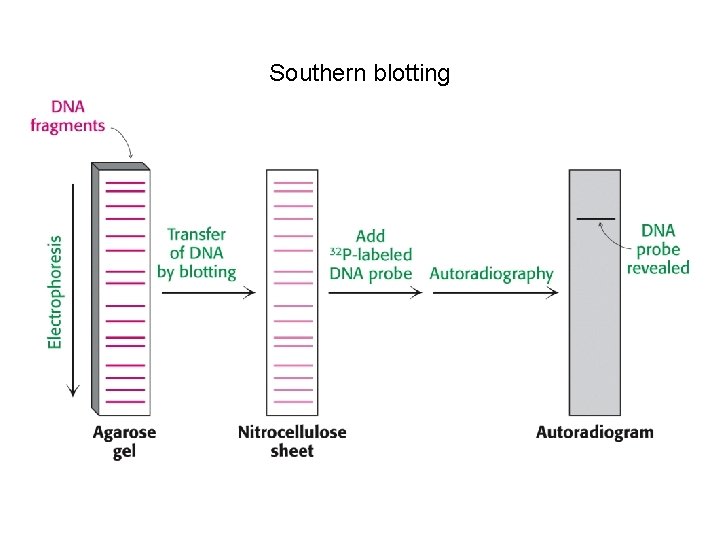
Southern blotting
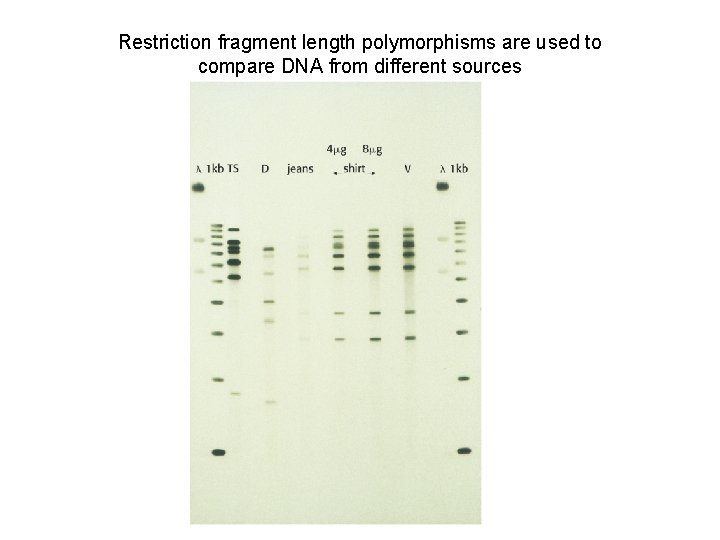
Restriction fragment length polymorphisms are used to compare DNA from different sources
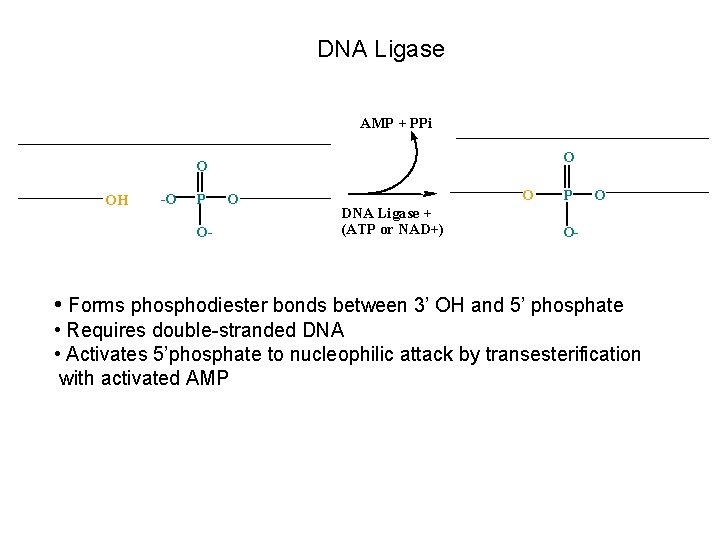
DNA Ligase AMP + PPi O O OH -O P O- O O DNA Ligase + (ATP or NAD+) P O O- • Forms phosphodiester bonds between 3’ OH and 5’ phosphate • Requires double-stranded DNA • Activates 5’phosphate to nucleophilic attack by transesterification with activated AMP
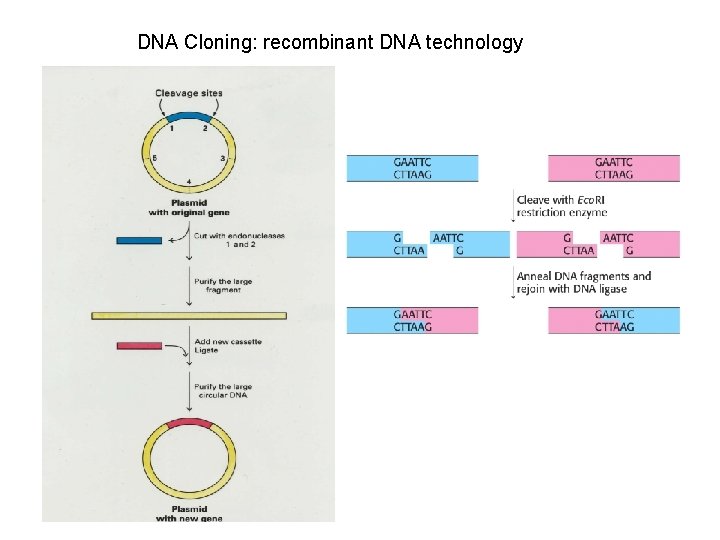
DNA Cloning: recombinant DNA technology

Human Genetic Polymorphisms • Human genome size: 3. 2 x 109 base pairs • 30, 000 genes • 2 -4 % of total sequence codes for proteins • Human genetic variation: 1 sigle nucleotide polymorphism (SNP) per 1, 300 bp
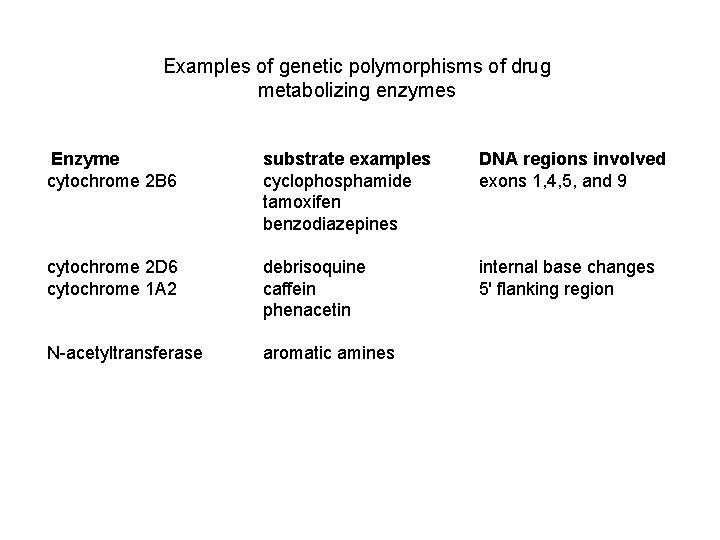
Examples of genetic polymorphisms of drug metabolizing enzymes Enzyme cytochrome 2 B 6 substrate examples cyclophosphamide tamoxifen benzodiazepines DNA regions involved exons 1, 4, 5, and 9 cytochrome 2 D 6 cytochrome 1 A 2 debrisoquine caffein phenacetin internal base changes 5' flanking region N-acetyltransferase aromatic amines

DNA Structure: Take Home Message 1. Genetic information is stored in DNA. 2. DNA is a double stranded biopolymer containing repeating units of nitrogen base, deoxyribose sugar, and phosphate. 3. DNA can be arranged in 3 types of duplexes which contain major and minor grooves. 4. DNA can adopt several topological forms. 5. There are enzymes that will cut DNA, ligate DNA, and change the topology of DNA. 6. Human genome contains about 3. 2 billion base pairs. Interindividual differences are observed at about 1 per 1, 000 nucleotides.
- Slides: 55