The Whole Genome Sequencing Revolution Martin Wiedmann Gellert
























- Slides: 32

The Whole Genome Sequencing Revolution Martin Wiedmann Gellert Family Professor of Food Safety Department of Food Science Cornell University, Ithaca, NY E-mail: mw 16@cornell. edu Phone: 607 -254 -2838

Outline • Subtyping for disease surveillance: from PFGE to WGS • WGS challenges: when are two isolates the same or different? Can we find identical isolates in different locations? • Looking in the future

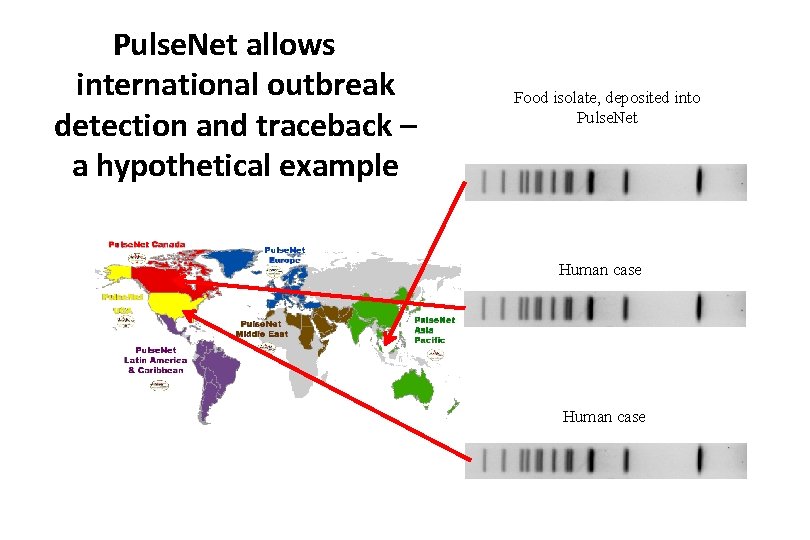
Pulse. Net allows international outbreak detection and traceback – a hypothetical example Food isolate, deposited into Pulse. Net Human case


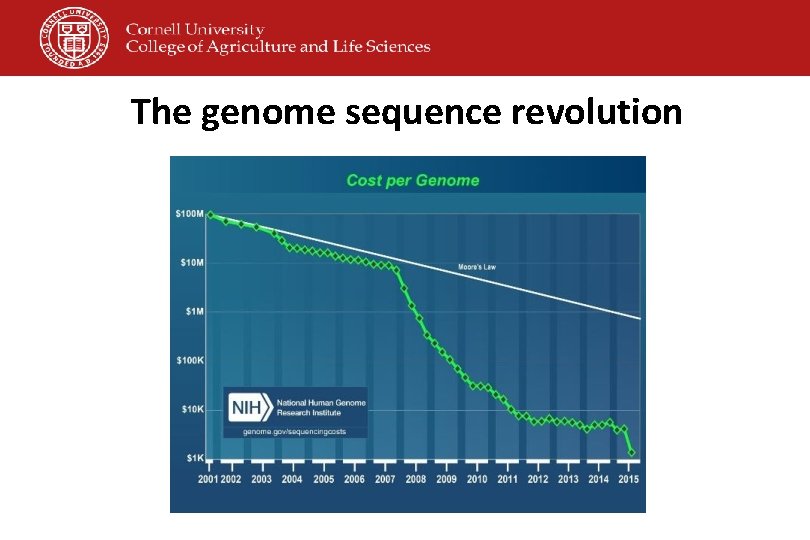
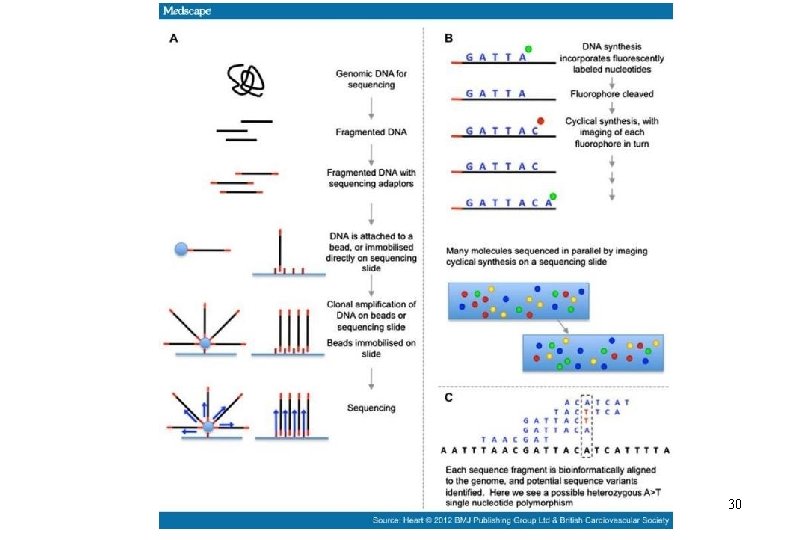
Whole Genome Sequencing • It all started with the human genome project • Sequencing of a bacterial genome is now feasible at costs of <$100/isolate • Costs will continue to drop • Commonly used platforms include • Roche 454 • Illumina Hi. Seq/Mi. Seq • Applied Biosystems SOLi. D Systems • Life Technologies/Thermofisher Ion Torrent; • Pac. Bio RS • Nanopore based systems (e. g. , Oxford Nanopore Min. ION)

The genome sequence revolution

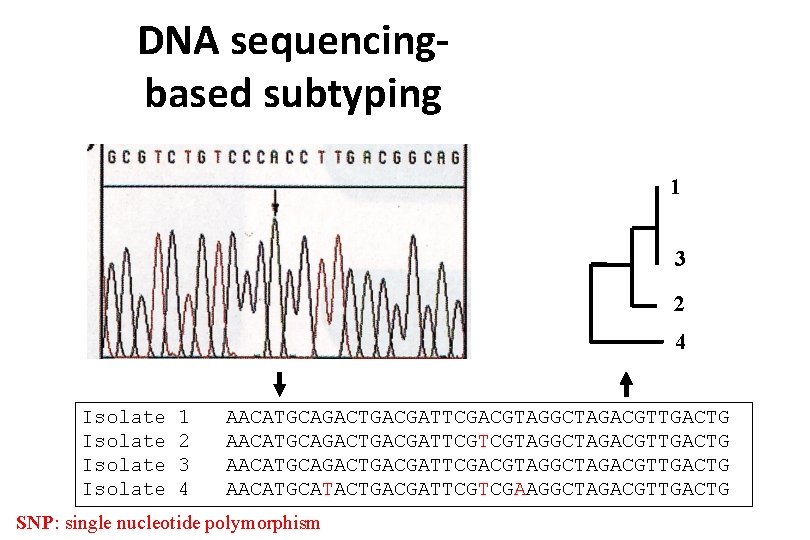
DNA sequencingbased subtyping 1 3 2 4 Isolate 1 2 3 4 AACATGCAGACTGACGATTCGACGTAGGCTAGACGTTGACTG AACATGCAGACTGACGATTCGTCGTAGGCTAGACGTTGACTG AACATGCAGACTGACGATTCGACGTAGGCTAGACGTTGACTG AACATGCATACTGACGATTCGTCGAAGGCTAGACGTTGACTG SNP: single nucleotide polymorphism

Challenges with use of PFGE as a subtyping method in outbreak investigations • Two isolates may show the same PFGE type even though they are genetically distinct • PFGE only interrogates small part of the genome • Two isolates may show “slightly” (? ? - the “ 3 -band rule”) different PFGE patterns despite sharing a very recent common ancestor • Could be due to lateral genes transfer, loss of plasmid, rearrangements, point mutations etc.

Xbal Spe. I Includes isolates form Salmonella outbreak linked to sausages (Rhode Island) and isolates from pistachios L Den Bakker et al. 2011. AEM.

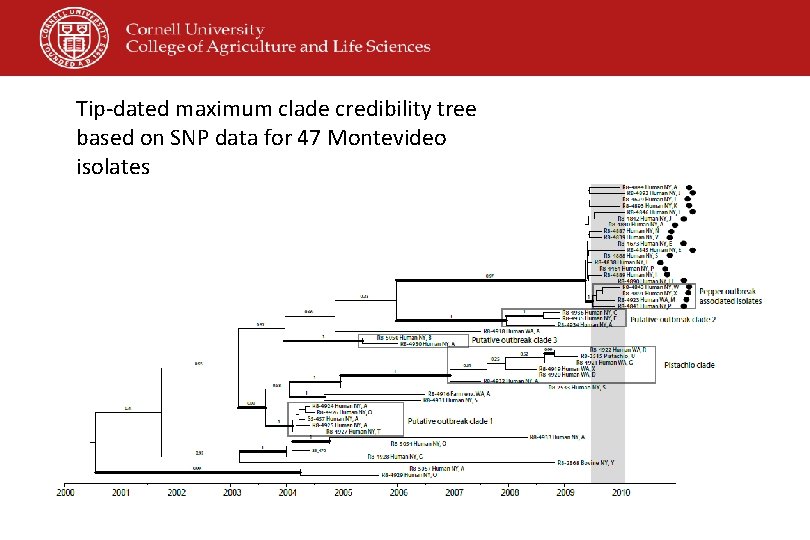
Tip-dated maximum clade credibility tree based on SNP data for 47 Montevideo isolates

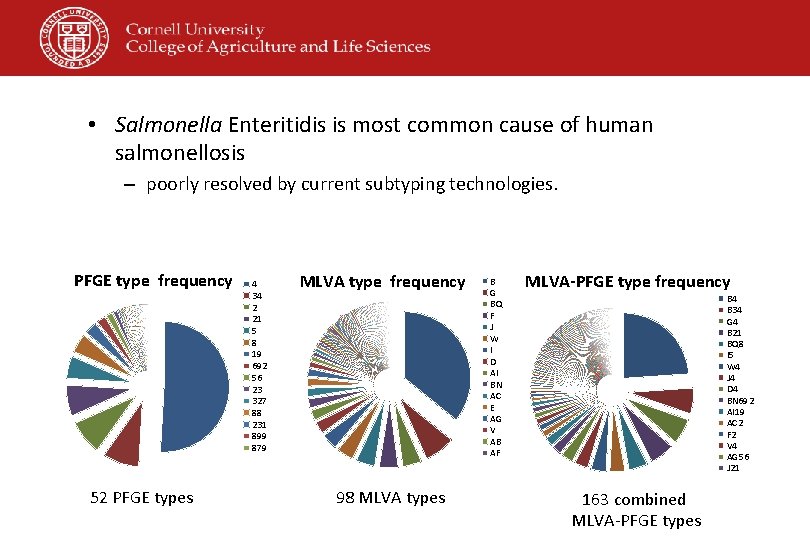
• Salmonella Enteritidis is most common cause of human salmonellosis – poorly resolved by current subtyping technologies. PFGE type frequency 52 PFGE types 4 34 2 21 5 8 19 692 56 23 327 88 231 899 879 MLVA type frequency 98 MLVA types B G BQ F J W I D AI BN AC E AG V AB AF MLVA-PFGE type frequency B 4 B 34 G 4 B 21 BQ 8 I 5 W 4 J 4 D 4 BN 692 AI 19 AC 2 F 2 V 4 AG 56 J 21 163 combined MLVA-PFGE types


Full genome sequencing identified the following differences between these isolates: (i) 28 single nucleotide polymorphisms (SNPs) and (ii) three indels, including a 33 kbp prophage that accounted for the observed difference in Asc. I PFGE patterns. Both isolates were found to harbor a 50 kbp putative mobile genomic island encoding translocation and efflux functions that has not been observed in other Listeria genomes. Gilmour et al. BMC Genomics 2010, 11: 120


In addition, whole genome sequencing showed that 5 Listeria isolates collected in 2010 from the same facility were also closely related genetically to isolates from ill people.

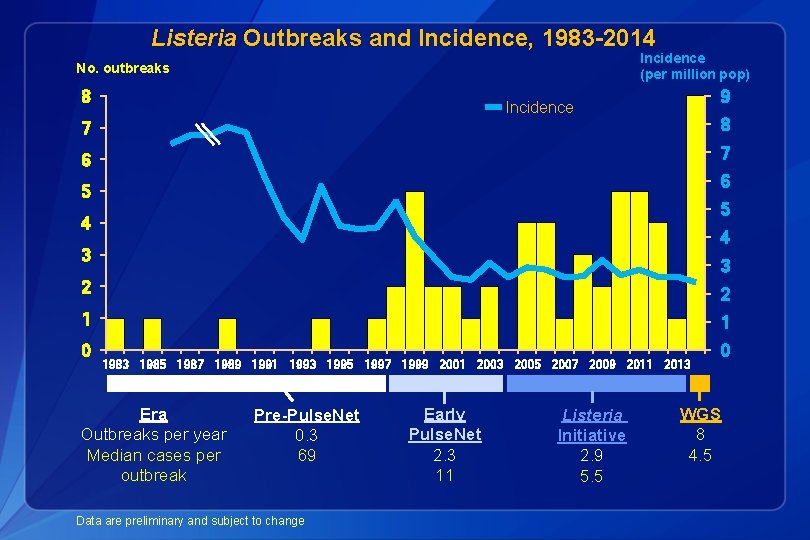
Listeria Outbreaks and Incidence, 1983 -2014 Incidence (per million pop) No. outbreaks 8 9 Incidence 7 8 6 7 6 5 5 4 4 3 3 2 2 1 1 0 1983 1985 1987 1989 1991 1993 1995 1997 1999 2001 2003 2005 2007 2009 2011 2013 Era Outbreaks per year Median cases per outbreak Pre-Pulse. Net 0. 3 69 Data are preliminary and subject to change Early Pulse. Net 2. 3 11 Listeria Initiative 2. 9 5. 5 0 WGS 8 4. 5

March 2015: Listeriosis cases linked to Blue Bell ice cream

Outline • Subtyping for disease surveillance: from PFGE to WGS • WGS challenges: when are two isolates the same or different? Can we find identical isolates in different locations? • Looking in the future

The challenge • Identical bacteria (100% match over the whole genome) can be found in different places that can be potential sources of foodborne disease outbreaks

The theoretical background • Bacteria divide asexually: Bacterial populations can be seen as large populations of “identical twins” • Mutation rate during replication is low: extremes of the suggested mutation rates range from 2. 25 × 10 -11 to 4. 50 × 10 -10 per bp per generation – With a genome size of around 5 Million bp per bacterial genome (5 × 106) between approx. 450 and 9, 000 generations are needed for a single SNP difference – Eyre et al. estimated evolutionary rate of 0. 74 SNVs per successfully sequenced genome per year for C. difficile (N. Engl. J. Med. 2013) • “Whole-genome sequencing … identified 13% of cases that were genetically related (≤ 2 SNVs) but without any evidence of plausible previous contact through a hospital, residential area, or family doctor. ” – Unknown bacterial generation time in different environments complicates interpretation

2000 US outbreak - Environmental persistence of L. monocytogenes • 1988: one human listeriosis case linked to hot dogs produced by plant X • 2000: 29 human listeriosis cases linked to sliced turkey meats from plant X

Real world observations


Real world observations In one case, isolates with < 3 SNP differences were found in retail delis in there different states

Conclusions • Even with WGS, epidemiological data are still essential • Number of SNP differences/allele differences that is meaningful differs by organism, strain, outbreak/cluster, and growth environment – Number of bacterial generations per calendar year can differ hugely (think dry environment versus active infection in an animal population) • Best way to determine “meaningful” SNP differences is through combination of phylogenetic and epidemiological data

Looking in the future • WGS will get cheaper and will be used more – STEC next, probably Salmonella Enteritidis after that – Detection of more clusters and outbreaks • WGS database will grow rapidly with inclusion of environmental isolates – More outbreak will be linked to source by using WGS matches between food or environmental isolates and human isolates as stating point • More broad application of WGS by private labs, maybe customers and consumers?


Conclusions • WGS is a game changer and will significantly improve detection of outbreaks, adulteration, etc. – False alarms will occur though • Pathogen detection in environments, by regulatory agencies, will lead to inclusion of WGS data in CDC/FDA/USDA databases (Genome. Trakr) – Environmental pathogen monitoring by industry will become even more important


30

Analysis of genome wide SNPs (wg. SNPs) • Identifies all high confidence SNPs over whole genome (approx. 3 to 5 million nucleotides)

Whole genome multilocus sequence typing (MLST) • Allows for simpler analysis and clear naming of subtypes • Performs comparison on a gene by gene level Isolate A Isolate B Isolate C Gene 1 1 Gene 2 8 8 12 Gene 3 5 5 2 Gene 1, 005 4 4 4 wg. MLST type A A B Etc.