Accurate Whole Human Genome Sequencing using Reversible Terminator

Accurate Whole Human Genome Sequencing using Reversible Terminator Chemistry Wen Hong

Break Down
Introduction to a new sequencing approach Application on whole genome Application on chromosome Method and Mechanism Discussion and Conclusion
Method and Mechanism (further breaking down) Question for you: This sequencing method is A) First generation B) Second generation C) Third generation D) Not a fan of Star Wars Data Analysis Single molecule array Isothermal bridging amplification Sequencing w/ reversible terminators
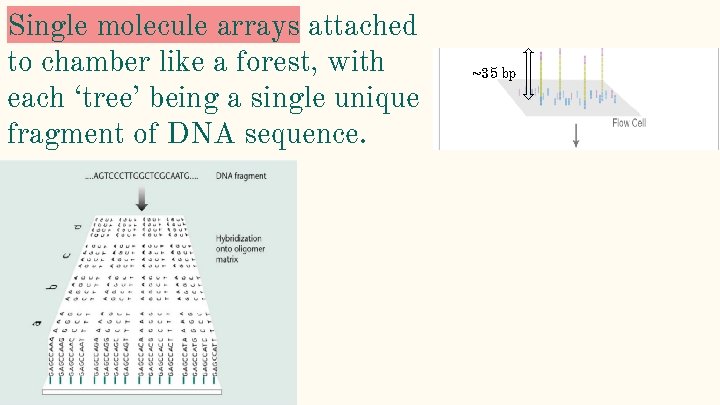
Single molecule arrays attached to chamber like a forest, with each ‘tree’ being a single unique fragment of DNA sequence. ~35 bp
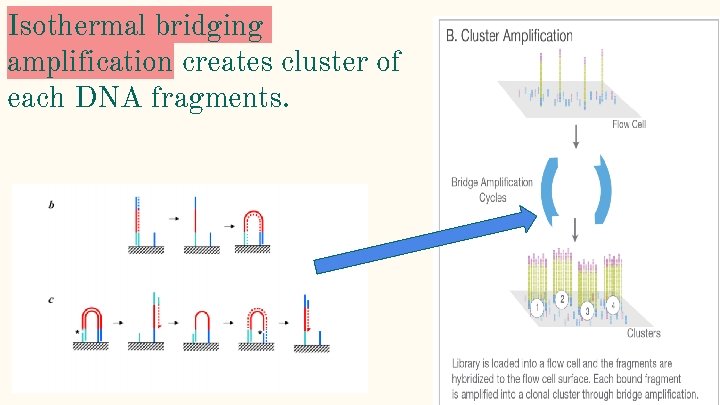
Isothermal bridging amplification creates cluster of each DNA fragments.

$equencing with signature reversible terminators enables massive parallel sequencing for convenience and low cost.

Although this paper did not specify the types of fluorophores, there are two options. 1. Four colors corresponding to each d. NTP → Hiseq 2. Two colors: Red/Green/Red+Green/ Absence → Next. Seq (much cheaper, but more demanding on the camera end)

Data analysis ⇔ application To consult the statistician after an experiment is finished is often merely to ask them to conduct a post mortem examination. They can perhaps say what the experiment died of. -- Ronald Fisher
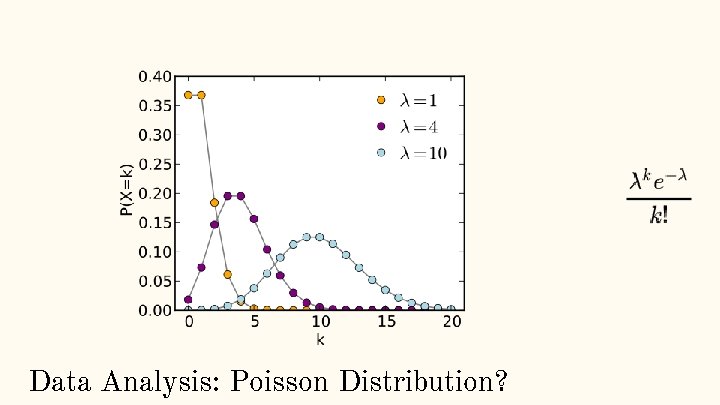
Data Analysis: Poisson Distribution?

Pros: ● Easy to use → R language, python, MATLAB, etc all have pre-installed packages to check for poisson distribution ● No over-fitting → only one parameter in the function, and it is also the mean value. ● Prevalent → in situations where a lot of independent events occur randomly at a fixed amount of time, poisson distribution is your best friend. Data Analysis: Why Poisson Distribution?
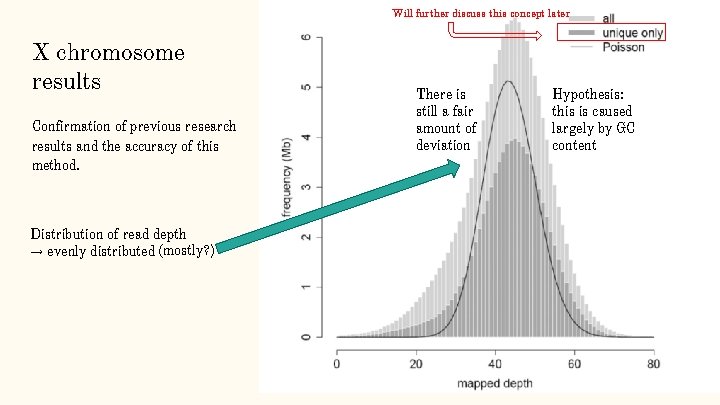
Will further discuss this concept later X chromosome results Confirmation of previous research results and the accuracy of this method. Distribution of read depth → evenly distributed (mostly? ) There is still a fair amount of deviation Hypothesis: this is caused largely by GC content
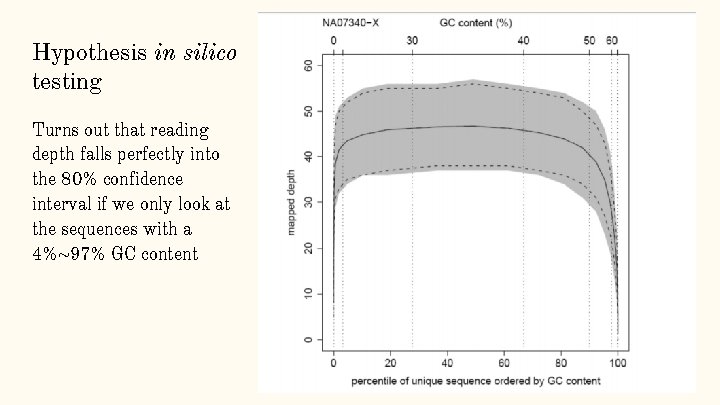
Hypothesis in silico testing Turns out that reading depth falls perfectly into the 80% confidence interval if we only look at the sequences with a 4%~97% GC content
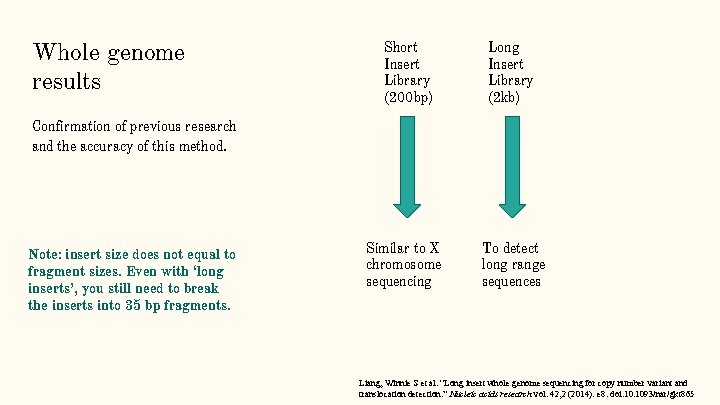
Whole genome results Short Insert Library (200 bp) Long Insert Library (2 kb) Confirmation of previous research and the accuracy of this method. Note: insert size does not equal to fragment sizes. Even with ‘long inserts’, you still need to break the inserts into 35 bp fragments. Similar to X chromosome sequencing To detect long range sequences Liang, Winnie S et al. “Long insert whole genome sequencing for copy number variant and translocation detection. ” Nucleic acids research vol. 42, 2 (2014): e 8. doi: 10. 1093/nar/gkt 865
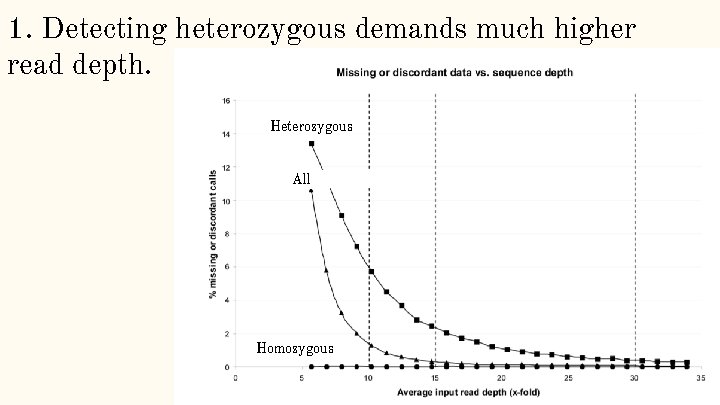
1. Detecting heterozygous demands much higher read depth. Heterozygous Homozygous All Homozygous

2. The choice of alignment algorithm is particularly tricky for this method: MAQ and ELAND have different strategies to deal with non-unique alignments (those that have more than one possible positions). MAQ assign the sequence randomly and give a zero quality score; ELAND rejects the sequence completely.

Take Home Messages 1. Single molecule array + isothermal bridge amplification + reversible terminator sequencing. 2. Reversible terminators → parallel sequencing → cost effective. 3. Two sequence alignment algorithms complement each other, but still misses some shots. 4. Poisson distribution is useful for biological data regression (depends on the system you’re modeling). 5. Don’t forget about figure legends.
- Slides: 17