String Matching String matching definition of the problem

String Matching String matching: definition of the problem (text, pattern) • Exact depends on what we have: text or • The patterns ---> Data structures for the matching: patterns • 1 pattern ---> The algorithm depends on |p| patterns • Regular Expressions and | | • Extensions • k patterns ---> The algorithm depends on k, |p| and | | • The text ----> Data structure for the text (suffix tree, . . . ) • Approximate matching: • Dynamic programming • Sequence alignment (pairwise and multiple) • Sequence assembly: hash algorithm • Probabilistic search: Hidden Markov Models

Sequence assembly It is applied to the following topics: DNA sequencing , EST assembly But in the last years new lab technologies , called “next sequencing generation”, have emerge

DNA sequencing There are two techniques: • Hibridization: provide information about l-tuple present in DNA. • Shotgun: DNA sequences are broken into 100 Kb-500 Kb random fragments.

DNA sequencing There are two techniques: • Hibridization: provide information about l-mers present in DNA • Shotgun: DNA sequences are broken into 100 Kb-500 Kb random fragments.

Hybridization Let xxxxxxx be the sequence we want to kno and the hybridization technique gives us the set of 3 -mers that belong t AAC GAT TGC ACG CGG GCC TTG GGA ATT How can the sequence be reconstructed

Hybridization Given the 3 -mers of the sequence: AAC GAT TGC ACG CGG GCC TTG GGA ATT As AAC then and AACG belong to the sequence, belongs to the sequence, because the longest (not proper) suffix of AAC matches the longest (not proper) prefix of AC This relation can be represented with a directed graph AAC ACG
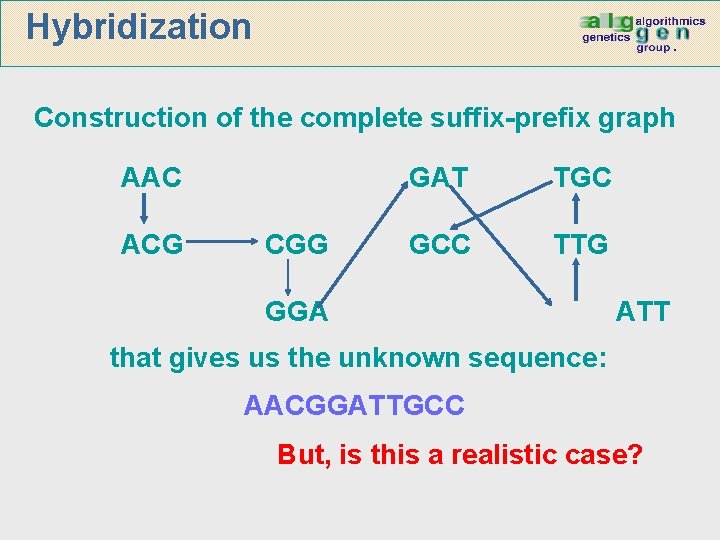
Hybridization Construction of the complete suffix-prefix graph AAC ACG CGG GAT TGC GCC TTG GGA ATT that gives us the unknown sequence: AACGGATTGCC But, is this a realistic case?

Hybridization Let us introduce a more realistic case: AAC CAA GAT TGC ACG CGG GCC TTG GGC GGA CCG ATT and the sequence is given by the Hamiltonian path that is the path that traverses all nodes exactly once and whose cost is NP-Complet! Which is the cost of the hybridization method

Hybridization: cost Cost: 1. Finding the l-mers AAC, CAA, ACG, . . . : There are 4 L l-mers of length L that should be gen 2. Searching for the suffix-prefix matches : If there are m L-mers, then there are O(m 2 L 2 ) com 3. Searching for the Hamiltonian path NP- Complet

Excursió: cost m Linear cost: O(m)10 m 1000 m m Quadratic cost: O(m 210 m ) 1000 m m Exponencial cost: O(2 m 10 m ) 1000 m t = 1000 t = t 1000000 t 1 mseg 10 mseg 1 seg = 1 mseg. = 100 mseg. = 16 min t = 1 ms 210 t = 1 seg 21000 t = 1030 t = 1018 a

Hybridization: cost Cost: 1. Finding the l-mers AAC, CAA, ACG, . . . : There are 4 L l-mers of length L that should be gen 2. Searching for the suffix-prefix matches : If there are m L-mers, then there are O(m 2 L 2 ) com 3. Searching for the Hamiltonian path NP- Complet How the NP-completness can be avo
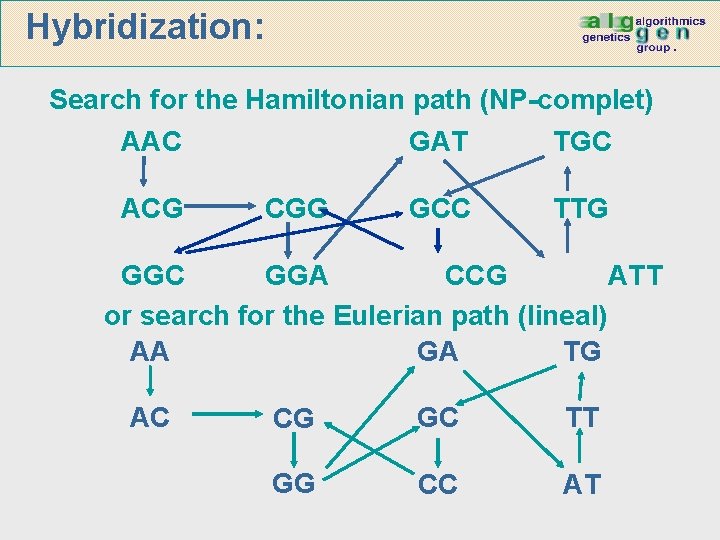
Hybridization: Search for the Hamiltonian path (NP-complet) AAC ACG CGG GAT TGC GCC TTG GGC GGA CCG ATT or search for the Eulerian path (lineal) AA GA TG AC CG GC TT GG CC AT
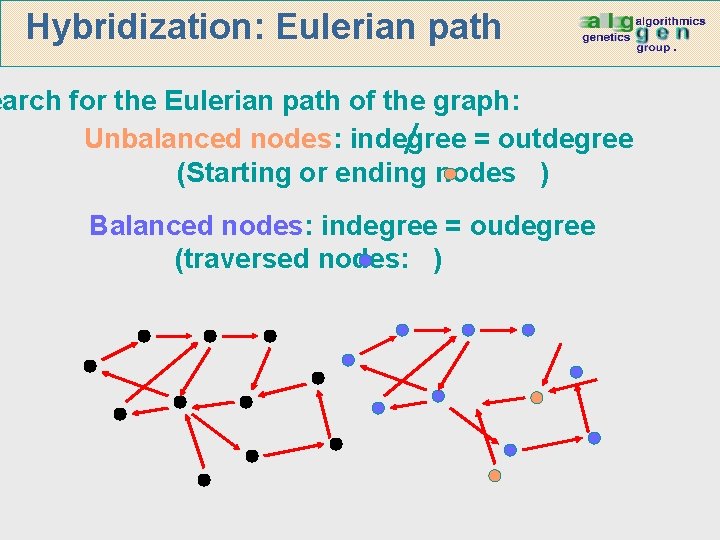
Hybridization: Eulerian path earch for the Eulerian path of the graph: Unbalanced nodes: indegree = outdegree (Starting or ending nodes ) Balanced nodes: indegree = oudegree (traversed nodes: )
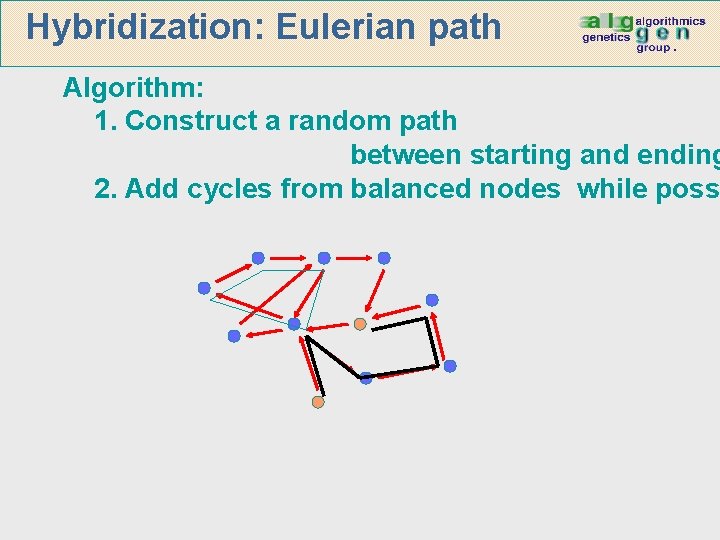
Hybridization: Eulerian path Algorithm: 1. Construct a random path between starting and ending 2. Add cycles from balanced nodes while poss
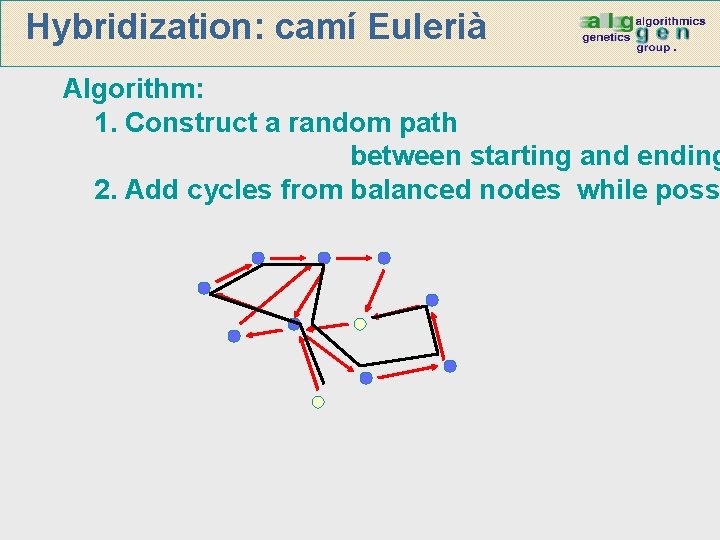
Hybridization: camí Eulerià Algorithm: 1. Construct a random path between starting and ending 2. Add cycles from balanced nodes while poss

Hybridization: cost Cost: 1. Finding the l-mers AAC, CAA, ACG, . . . : There are 4 L l-mers of length L that should be gen 2. Searching for the suffix-prefix matches : If there are m L-mers, then there are O(m 2 L 2 ) com 3. Searching for the Eulerian path Linear cost Now, which is the limiting factor?
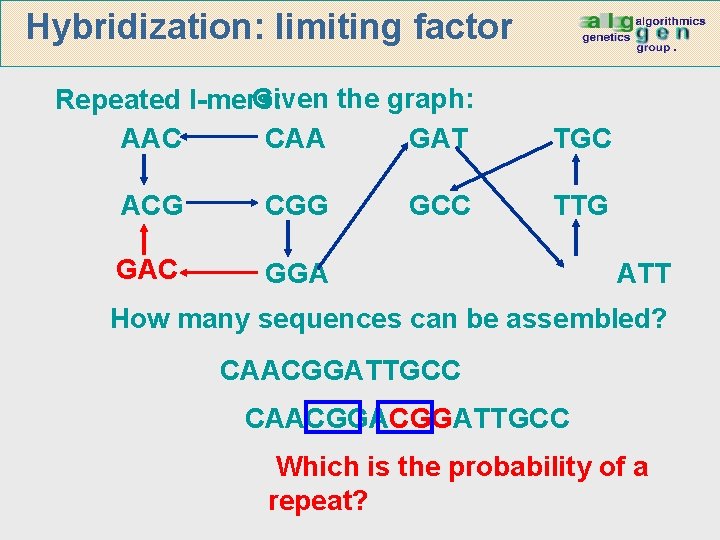
Hybridization: limiting factor Given the graph: Repeated l-mers: AAC CAA GAT ACG CGG GAC GGA GCC TGC TTG ATT How many sequences can be assembled? CAACGGATTGCC Which is the probability of a repeat?

Hybridization: statistical model How the probability of a repeat can be computed? Model: random sequence of length N with identically distributed b Given 2 l-mers, the probability to match is : 4 -L Given 3 l-mers, the expected number of 2 -matches is : (32)4 -L Given m l-mers, the expected number of 2 -matches is: (m 2)4 -L then. L) for L = 8, m =512! If (m 2)4 -L <1 then m<sqr(2· 4 Conclusion: this technique can be applied only to short sequence

DNA sequencing There are two techniques: • Hibridizationació: provide information about l-m present in DNA • Shot gun: DNA sequences are broken into 100 Kb-500 Kb random fragments.

Shotgun With the unknown sequence xxxxxxxxxxxxxxxxxxxxxxx It is possible : • to make some copies • to break it into random and unsorted short segments What can we do?

Shotgun: algorisme Assume xxxxx|xxxxxxx|xxxxxx|xxx xxxx|xxxxxx|xxxxxxx The algorithm is: 1 st. Compare all pairs searching for suffix-prefix approximate matches. 2 nd. Construct the graph suffix-prefix 3 th. Find the path

Shotgun Given the three copies xxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxx The shotgun brokes it into the following segments accgt, aggt, acgatac, accttta, tttaac, gataca, accgtacc, ggt, acaggt, taacgat, accg, tacctt

Shotgun The pairwise comparison that searchs for suffixprefix approximate matching can be done with: • Dynamic programming ( quadratic cost) the pairs suspected to be assembled • two • Find steps: (Linear cost with the hash algorithm) • Assembly them with dynamic programming.
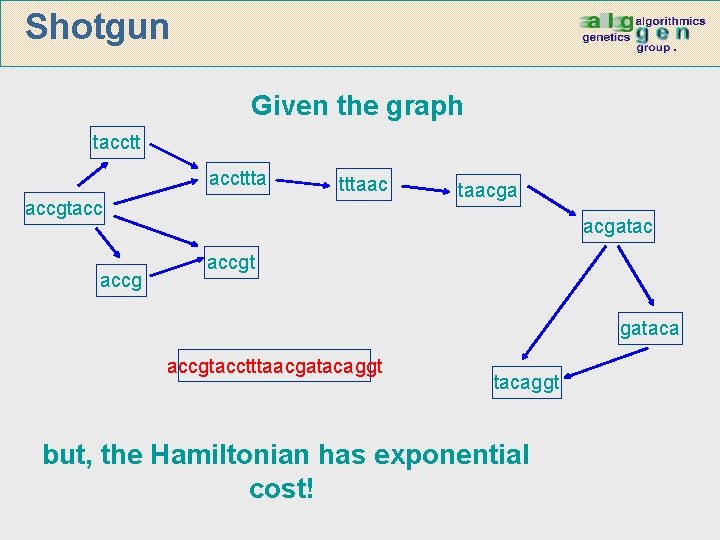
Shotgun Given the graph taccttta tttaac accgtacc accg taacgatac accgt gataca accgtacctttaacgatacaggt but, the Hamiltonian has exponential cost!
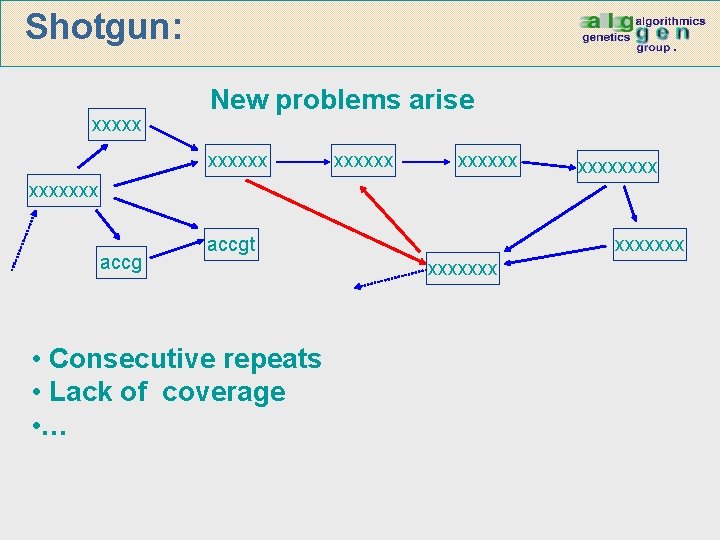
Shotgun: xxxxx New problems arise xxxxxxxx xxxxxxx accgt • Consecutive repeats • Lack of coverage • … xxxxxxx

Shotgun: properties of the coverage Given the coverage: Some questions arisess: • What is the percentage of coverage? • How many contigs we have to expect? • What is the mean length of contigs?

Shotgun: percentage of coverage Given the model L d Degree of coverage N d / L N We assume that segments are randomly distributed. The probability that a base was covered by k segments is given by the binomial dsitribution (N Prob{X=k}=N k (d/L)k (1 -d/L)n-k

Shotgun: percentage of coverage What is the limit of the binomial distribution n i p 0 having np= Distribució de Poisson P( ) k Prob{X=k}= e- k! the probability that at least one segment covers a base Prob{X>0}= 1 -Prob{X=0}= 1 - e=- 1 - e(N d / L) Then, with N d / L = 4. 6 we obtain a 99% of coverag and with N d / L = 6. 9 weobtain a 99. 9% of cove
- Slides: 29