String Matching String matching definition of the problem

String Matching String matching: definition of the problem (text, pattern) • Exact matching: depends on what we have: text or patterns • The patterns ---> Data structures for the patterns • 1 pattern ---> The algorithm depends on |p| and | | • k patterns ---> The algorithm depends on k, |p| and | | • Extensions • Regular Expressions • The text ----> Data structure for the text (suffix tree, . . . ) • Approximate matching: • Dynamic programming • Sequence alignment (pairwise and multiple) • Sequence assembly: hash algorithm • Probabilistic search: Hidden Markov Models

2. 2 Pairwise alignment Given two DNA sequences A (a 1 a 2. . . an) and B (b 1 b 2. . . bm) from the alphabet {a, c, t, g} we say that A* and B* from {a, c, t, g, -} are aligned i) A* and B* become A and B if gaps ( – ) are iff removed. ii) |A*|=|B*| iii) For all i, it is not possible that ai = bi = MALIG (an example) How many alignments of two sequences exist? Which is the best alignment?

2. 2 Number of alignments Given two DNA sequences A (a 1 a 2. . . an) and B (b 1 b 2. . . bm) there are: #(a 1 a 2. . . an , b 1 b 2. . . bm) = #(a 1 a 2. . . an-1 , b 1 b 2. . . bm) with (an, -) + #(a 1 a 2. . . an , b 1 b 2. . . bm-1) with (-, bm) b b 2 b 3 + #(a 1 a 2. . . an-1 , b 11 b 2. . . b m-1) with (an, bm) a 1 a 2 a 3 those that end #(a 1, b 1)

2. 2 Number of alignments Given two DNA sequences A (a 1 a 2. . . an) and B (b 1 b 2. . . bm) there are: #(a 1 a 2. . . an , b 1 b 2. . . bm) = #(a 1 a 2. . . an-1 , b 1 b 2. . . bm) with (an, -) + #(a 1 a 2. . . an , b 1 b 2. . . bm-1) with (-, bm) b b 2 b 3 + #(a 1 a 2. . . an-1 , b 11 b 2. . . b m-1) with (an, bm) 1 1 a 2 1 a 3 1 those that end

2. 2 Number of alignments Given two DNA sequences A (a 1 a 2. . . an) and B (b 1 b 2. . . bm) there are: #(a 1 a 2. . . an , b 1 b 2. . . bm) = #(a 1 a 2. . . an-1 , b 1 b 2. . . bm) with (an, -) + #(a 1 a 2. . . an , b 1 b 2. . . bm-1) with (-, bm) b b 2 b 3 + #(a 1 a 2. . . an-1 , b 11 b 2. . . b m-1) with (an, bm) 1 1 a 1 1 3 ? ? a 2 1 a 3 1 those that end

2. 2 Number of alignments Given two DNA sequences A (a 1 a 2. . . an) and B (b 1 b 2. . . bm) there are: #(a 1 a 2. . . an , b 1 b 2. . . bm) = #(a 1 a 2. . . an-1 , b 1 b 2. . . bm) with (an, -) + #(a 1 a 2. . . an , b 1 b 2. . . bm-1) with (-, bm) b b 2 b 3 + #(a 1 a 2. . . an-1 , b 11 b 2. . . b m-1) with (an, bm) 1 1 a 1 1 3 5 7 a 2 1 5 a 3 1 7 ? those that end

2. 2 Number of alignments Given two DNA sequences A (a 1 a 2. . . an) and B (b 1 b 2. . . bm) then: #(a 1 a 2. . . an , b 1 b 2. . . bm) = #(a 1 a 2. . . an-1 , b 1 b 2. . . bm) those that end with ( an , -) + #(a 1 a 2. . . an , b 1 b 2. . . bm-1) those that end with ( - , bm) b b 2 b 3 + #(a 1 a 2. . . an-1 , b 11 b 2. . . b those that end m-1) with ( an , bm)1 1 a 1 1 3 5 7 a 2 1 5 13 25 But, what is the a 3 1 7 25 63 assymptotic

2. 2 Assymptotic value As K=n #(a 1 a 2. . . an , b 1 b 2. . . bn) and Σ ( )= (( ) ) n > k=0 k n k 2 n n n! ~ nn e-n (Stirling approximation) then #(a 1 a 2. . . an , b 1 b 2. . . bn) > 22 n

2. 2 Best alignment How can an alignment be scored? catcactactgacgactatcgtagcgcggctatacatctacgccaa- ctac-t-gtgtagatcgccgg c- tgactgc--acgactatcgt- attgcggctacacactacgcacaactactgtatgtcgc-cgg---* * ******* * * ******* * **** *** • Match: favorable • Mismatch: unfavorable • Gap: worst case Then we assign a score for each case, for example 1, -2. How can the best alignment be

2. 2 Best alignment CTACTACTACGT A C T G A The cell contains the score of the best alignment of AC and CTACT.
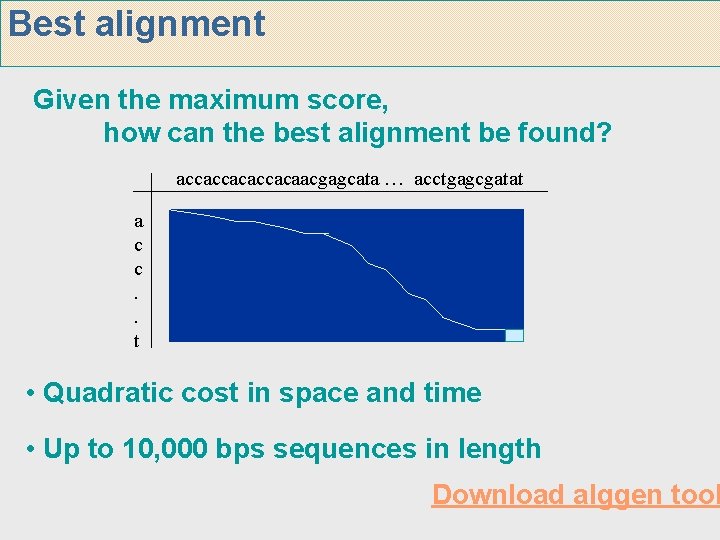
Best alignment Given the maximum score, how can the best alignment be found? accaccacaacgagcata … acctgagcgatat a c c. . t • Quadratic cost in space and time • Up to 10, 000 bps sequences in length Download alggen tool

2. 2 Some slides revisited We have developed theory according to the following principles: 1) Both sequences have a similar length (global). 2) The model of gaps is linear If there are k consecutive gaps the penalty scores k(-2).

2. 2 Semiglobal pairwise alignment Assume that we have sequences with different length S 1 S 2 It is meaningless to introduce gaps until both sequences have similar length …. The most probable alignment should be Initial gaps Final gaps How can these alignments be

2. 2 Semiglobal pairwise alignment Note that Initial gaps CTACTACTACGT A C T Final gaps

2. 2 Semiglobal pairwise alignment Given a cell CTACTACTACGT A C T 0 00 0 0 0 0 The cell contains the score of the best alignment of CTA with the empty sequence.

2. 2 Semiglobal pairwise alignment CTACTACTACGT 0 0 0 0… A C T The contribution of the initial gaps is disregarded, then CTACTACTACGT 0 0 0 0… A 1 C 2 T 3 but, what happens with the final

2. 2 Semiglobal pairwise alignment CTACTACTACGT 0 0 0 0… A 1 C 2 T 3 How does the algorithm search for the best alignment? … by checking the last row for the best score. Practice with the alggen tool.

2. 2 Affine-gap model score Given the following alignments that have the same score … agtaccccgtag agt- cc- -gta- agtaccccgtag agt- c-c -gta- agtaccccgtag agt- c- -cgta- agtaccccgtag agt- -cc -gta- agtaccccgtag agt- -c -cgta- agtaccccgtag agt- -- ccgta- Which is the most reliable case from a biological point of view?
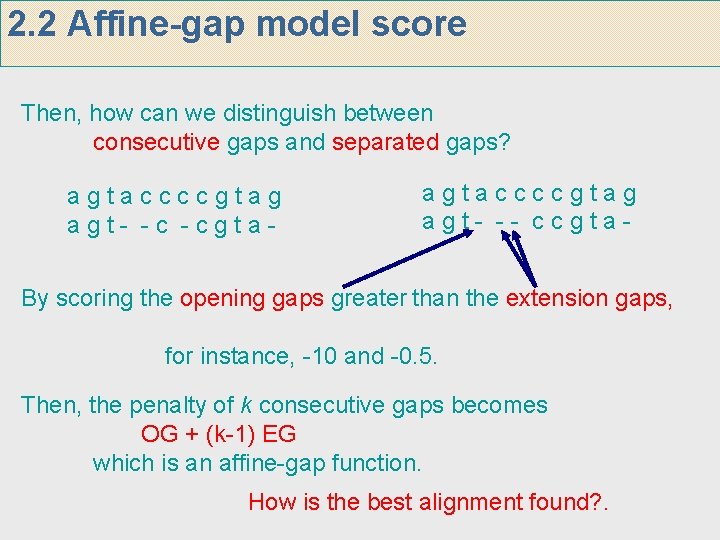
2. 2 Affine-gap model score Then, how can we distinguish between consecutive gaps and separated gaps? agtaccccgtag agt- -c -cgta- agtaccccgtag agt- -- ccgta- By scoring the opening gaps greater than the extension gaps, for instance, -10 and -0. 5. Then, the penalty of k consecutive gaps becomes OG + (k-1) EG which is an affine-gap function. How is the best alignment found? .
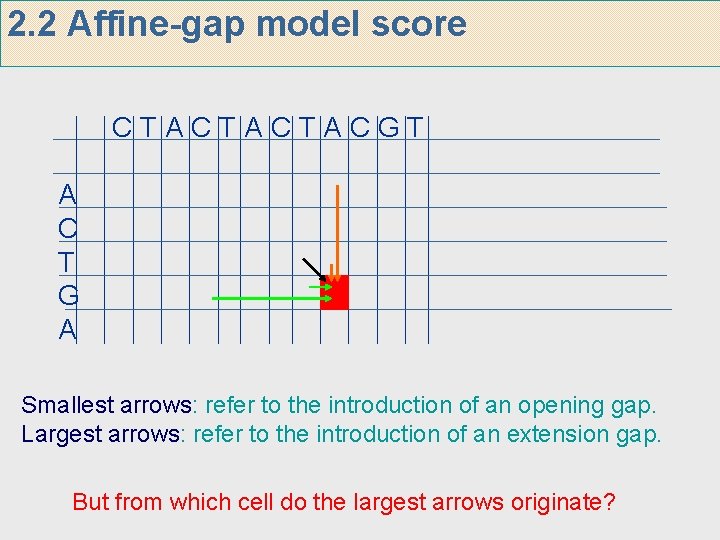
2. 2 Affine-gap model score CTACTACTACGT A C T G A Smallest arrows: refer to the introduction of an opening gap. Largest arrows: refer to the introduction of an extension gap. But from which cell do the largest arrows originate?

2. 2 Local alignment Given two sequences, we can consider the alignments of all their substrings… …how can the best of them be found? Two questions arise: - how can the alignments be compared? - how can the best one be selected?

2. 2 Local alignment accaccacaacgagcata … acctgagcgatat Given a path a c c. . t Imagine the graph of the scores: can the best subalignments be detected? … It suffices to compare the value of each cell with zero!
- Slides: 22