Sequence Alignment Abhishek Niroula Department of Experimental Medical

Sequence Alignment Abhishek Niroula Department of Experimental Medical Science Lund University 2015 -12 -09 1

Sequence alignment • A way of arranging two or more sequences to identify regions of similarity • Shows locations of similarities and differences between the sequences • An 'optimal' alignment exhibits the most similarities and the least differences • The aligned residues correspond to original residue in their common ancestor • Insertions and deletions are represented by gaps in the alignment • Examples Protein sequence alignment MSTGAVLIY--TSILIKECHAMPAGNE-------GGILLFHRTHELIKESHAMANDEGGSNNS * **** Nucleotide sequence alignment attcgttggcaaatcgcccctatccggccttaa att---tggcggatcg-cctctacgggcc---*** 2015 -12 -09 **** ** ****** 2

Sequence alignment: Purpose • Reveal structural, functional and evolutionary relationship between biological sequences • Similar sequences may have similar structure and function • Similar sequences are likely to have common ancestral sequence • Annotation of new sequences • Modelling of protein structures • Design and analysis of gene expression experiments 2015 -12 -09 3

Sequence alignment: Types • Global alignment – Aligns each residue in each sequence by introducing gaps – Example: Needleman-Wunsch algorithm L G P S S K Q T G K G S - S R I W D N L N - I T K S A G K G A I M R L G D A 2015 -12 -09 4

Sequence alignment: Types • Local alignment – Finds regions with the highest density of matches locally – Example: Smith-Waterman algorithm - - - - T G K G - - - - A G K G - - - - 2015 -12 -09 5
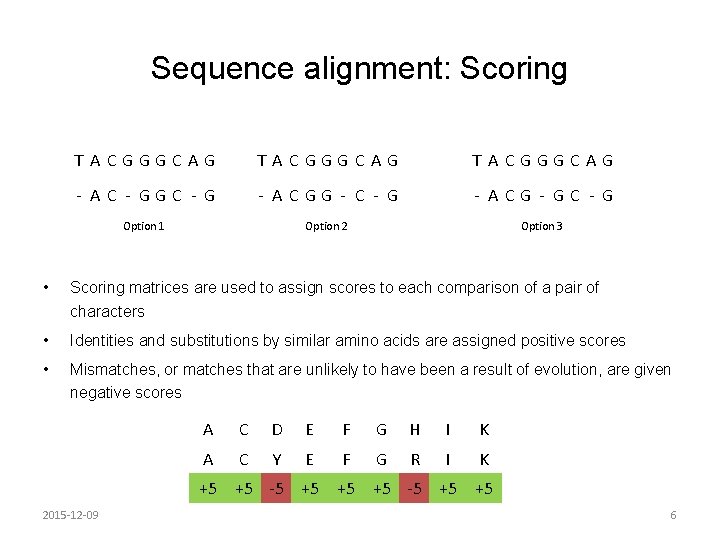
Sequence alignment: Scoring T ACGGGCAG - AC - GGC - G - ACGG - C - G - ACG - GC - G Option 1 Option 2 Option 3 • Scoring matrices are used to assign scores to each comparison of a pair of characters • Identities and substitutions by similar amino acids are assigned positive scores • Mismatches, or matches that are unlikely to have been a result of evolution, are given negative scores A C D E F G H I K A C Y E F G R I K +5 +5 -5 +5 +5 2015 -12 -09 6

Sequence alignment: Scoring • PAM matrices – PAM - Percent Accepted Mutations – PAM gives the probability that a given amino acid will be replaced by any other amino acid – An accepted point mutation in a protein is a replacement of one amino acid by another, accepted by natural selection – Derived from global alignments of closely related sequences – The numbers with the matrix (PAM 40, PAM 100) refer to the evolutionary distance (greater numbers mean greater distances) – 1 -PAM matrix refers to the amount evolution that would change residues/bases (on average) 1% of the – 2 -PAM matrix does NOT refer to change in 2% of residues • Refers 1 -PAM twice • Some variations may change back to original residue 2015 -12 -09 7

PAM-1 2015 -12 -09 8

Sequence alignment: Scoring • BLOSUM matrices – BLOSUM - Blocks Substitution Matrix – Score for each position refers to obtained frequencies of substitutions in blocks of local alignments of protein sequences [Henikoff & Henikoff]. – For example BLOSUM 62 is derived from sequence alignments with no more than 62% identity. 2015 -12 -09 9

BLOSUM 62 2015 -12 -09 10

Which scoring matrix to use? For global alignments use PAM matrices. • Lower PAM matrices tend to find short alignments of highly similar regions • Higher PAM matrices will find weaker, longer alignments For local alignments use BLOSUM matrices • BLOSUM matrices with HIGH number, are better for similar sequences • BLOSUM matrices with LOW number, are better for distant sequences 2015 -12 -09 11

Sequence alignment: Methods • Pairwise alignment – Finding best alignment of two sequences – Often used for searching best similar sequences in the sesequence databases • Dot Matrix Analysis • Dynamic Programming (DP) • Short word matching • Multiple Sequence Alignment (MSA) – Alignment of more than two sequences – Often used to find conserved domains, regions or sites among many sequences • Dynamic programming • Progressive methods • Iterative methods • Structural alignments – Alignments based on structure 2015 -12 -09 12
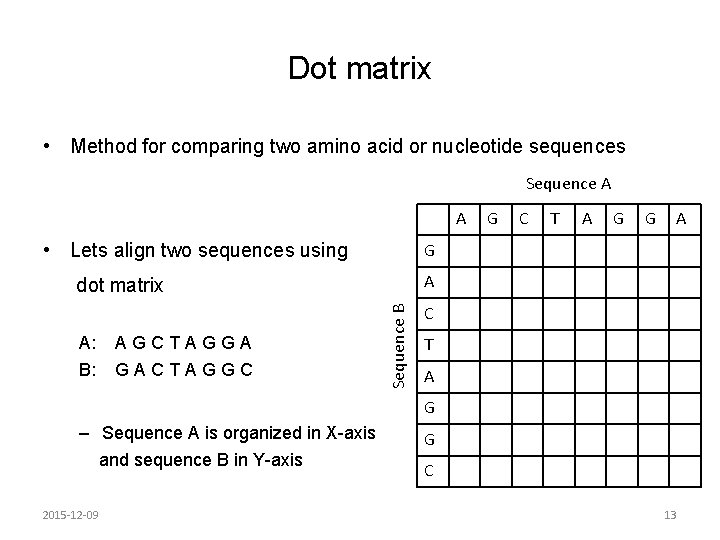
Dot matrix • Method for comparing two amino acid or nucleotide sequences Sequence A A • Lets align two sequences using B: GACTAGGC T A G G A A Sequence B AGCTAGGA C G dot matrix A: G C T A G – Sequence A is organized in X-axis and sequence B in Y-axis 2015 -12 -09 G C 13

Dot matrix – Starting from the first nucleotide in B, move along the first row placing a dot in columns with matching nucleotide – Other isolated dots represent random matches A G – Repeat the procedure for all the nucleotides in B G ● C T A G G ● ● A A Sequence B – Region of similarity is revealed by a diagonal row of dots Sequence A C T A G G C 2015 -12 -09 14

Dot matrix – Starting from the first nucleotide in B, move along the first row placing a dot in columns with matching nucleotide – Other isolated dots represent random matches A G – Repeat the procedure for all the nucleotides in B A Sequence B – Region of similarity is revealed by a diagonal row of dots Sequence A G C T A ● ● ● G G ● ● A ● C T A G G C 2015 -12 -09 15

Dot matrix – Starting from the first nucleotide in B, move along the first row placing a dot in columns with matching nucleotide – Other isolated dots represent random matches A G – Repeat the procedure for all the nucleotides in B A Sequence B – Region of similarity is revealed by a diagonal row of dots Sequence A C ● A G G ● ● A ● ● ● T A T ● C ● ● G ● ● ● C 2015 -12 -09 G ● 16

Dot matrix – Starting from the first nucleotide in B, move along the first row placing a dot in columns with matching nucleotide – Other isolated dots represent random matches A G – Repeat the procedure for all the nucleotides in B A Sequence B – Region of similarity is revealed by a diagonal row of dots Sequence A C ● A G G ● ● A ● ● ● T A T ● C ● ● G ● ● ● C 2015 -12 -09 G ● 17
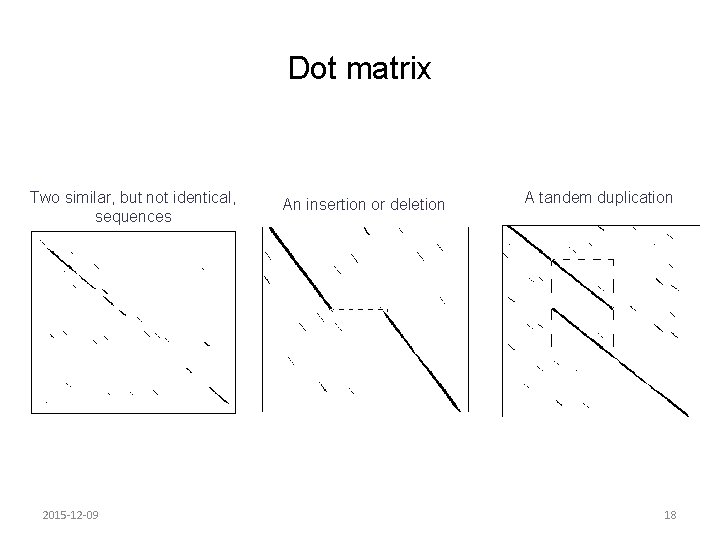
Dot matrix Two similar, but not identical, sequences 2015 -12 -09 An insertion or deletion A tandem duplication 18
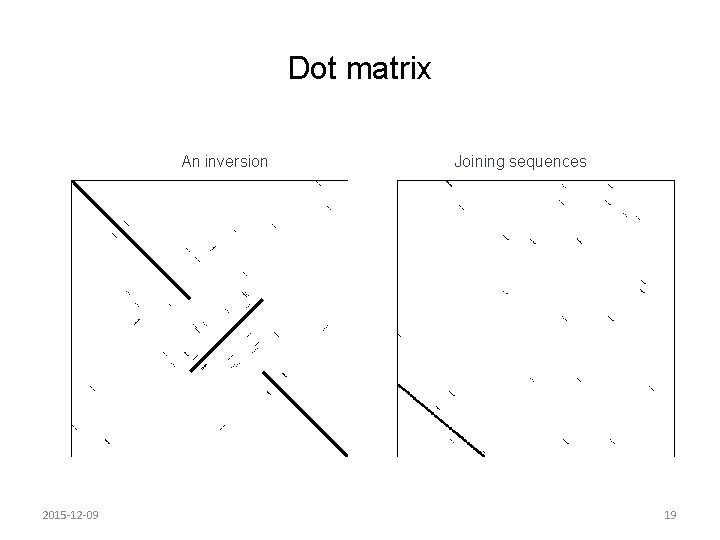
Dot matrix An inversion 2015 -12 -09 Joining sequences 19

Limitations of dot matrix • Sequences with low-complexity regions give false diagonals – Sequence regions with little diversity • Noisy and space inefficient • Limited to 2 sequences 2015 -12 -09 20
Dotplot exercise • Use the following three tools to generate dot plots for the given two sequences • YASS: : genomic similarity search tool – http: //bioinfo. lifl. fr/yass. php • Lalign/Palign – http: //fasta. bioch. virginia. edu/fasta_www 2/fasta_www. cgi? rm=lalign • multi-z. Picture – http: //zpicture. dcode. org/ 2015 -12 -09 21

Dynamic programming • Breaks down the alignment problem into smaller problems • Example – Needleman-Wunsch algorithm: global alignment – Smith-Waterman algorithm: local alignment • Three steps – Initialization – Scoring – Traceback 2015 -12 -09 22

Gap penalties • Insertion of gaps in the alignment • Gaps should be penalized • Gap opening should be penalized higher than gap extension (or at least equal) • In BLOSUM 62 – Gap opening score = -11 – Gap extension score = -1 AAAGAGAAA AAA - - AAAA Gap initiation 2015 -12 -09 Gap extention 23
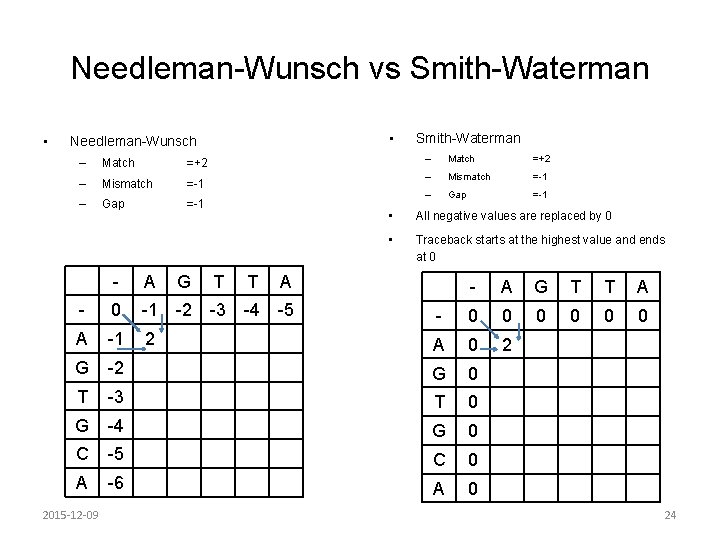
Needleman-Wunsch vs Smith-Waterman • • Needleman-Wunsch – Match =+2 – Mismatch =-1 – Gap =-1 - A G T T A - 0 -1 -2 -3 -4 -5 A -1 2 G Smith-Waterman – Match =+2 – Mismatch =-1 – Gap =-1 • All negative values are replaced by 0 • Traceback starts at the highest value and ends at 0 - A G T T A - 0 0 0 A 0 2 -2 G 0 T -3 T 0 G -4 G 0 C -5 C 0 A -6 A 0 2015 -12 -09 24

Needleman-Wunsch vs Smith-Waterman Sequence alignment teacher (http: //melolab. org/websoftware/web/? sid=3) 2015 -12 -09 25

Dynamic programming: example • http: //www. avatar. se/molbioinfo 2001/dynprog/dynamic. html • Scoring – Match = +2 – Mismatch = -2 – Gap = -1 2015 -12 -09 26

Dynamic programming exercise • Generate a scoring matrix for nucleotides (A, C, G, and T) • Align two sequences using dynamic programming • Align two sequences using following tools – EMBOSS Needle • http: //www. ebi. ac. uk/Tools/psa/emboss_needle/ – EMBOSS Water • http: //www. ebi. ac. uk/Tools/psa/emboss_water/ 2015 -12 -09 27

Multiple sequence alignment • A multiple sequence alignment (MSA) is an alignment of three or more sequences • Why MSA? – – – 2015 -12 -09 To identify patterns of conservation across more than 2 sequences To characterize protein families and generate profiles of protein families To infer relationships within and among gene families To predict secondary and tertiary structures of new sequences To perform phylogenetic studies 28
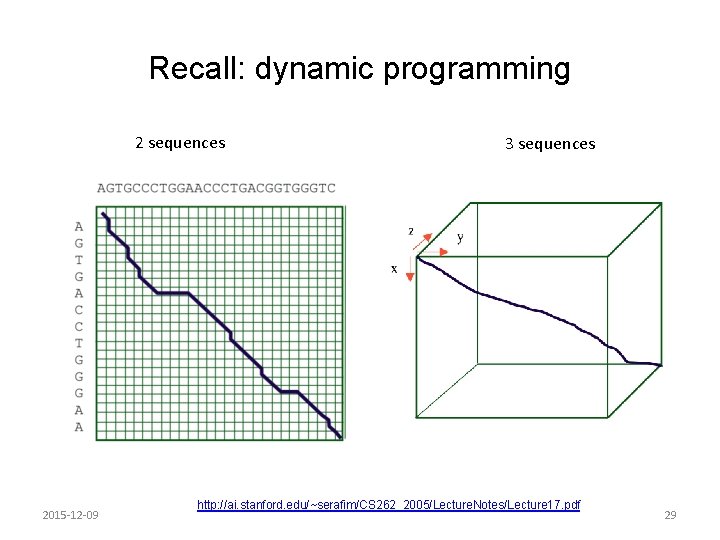
Recall: dynamic programming 2 sequences 2015 -12 -09 3 sequences http: //ai. stanford. edu/~serafim/CS 262_2005/Lecture. Notes/Lecture 17. pdf 29

MSA methods • Dynamic programming – Align each pair of sequences – Sum scores for each pair at each position • Progressive sequence alignment – Hierarchical or tree based method – E. g. Clustal. W, T-Coffee • Iterative sequence alignment – Improved progressive alignment – Realigns the sequences repeatedly – E. g. MUSCLE 2015 -12 -09 30

Tools for MSA 2015 -12 -09 31

Clustal. W • Progressive sequence alignment • Basic steps – Calculate pairwise distances based on pairwise alignments between the sequences – Build a guide tree, which is an inferred phylogeny for the sequences – Align the sequences 2015 -12 -09 32
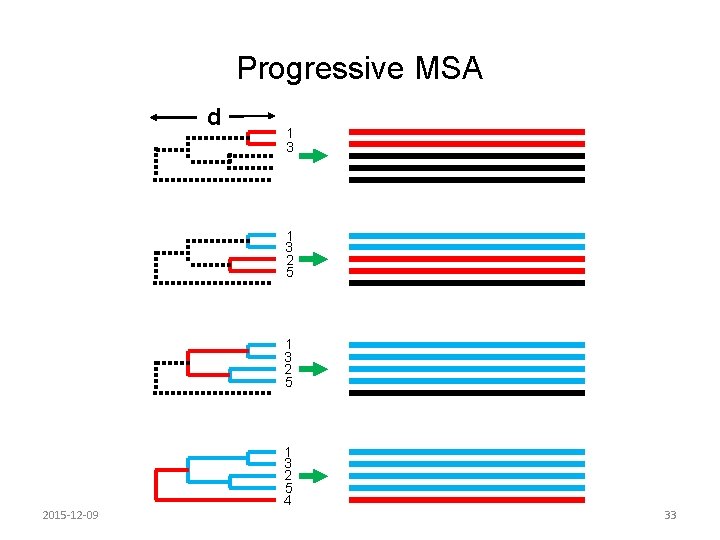
Progressive MSA d 1 3 2 5 2015 -12 -09 1 3 2 5 4 33
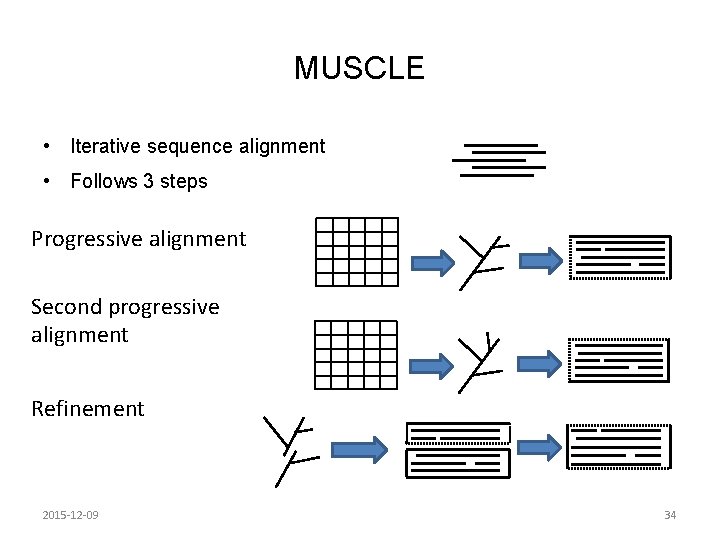
MUSCLE • Iterative sequence alignment • Follows 3 steps Progressive alignment Second progressive alignment Refinement 2015 -12 -09 34

Phylogenetic tree • A phylogenetic tree shows evolutionary relationships between the sequences • Types: – Rooted • Nodes represent most recent common ancestor • Edge lengths represents time estimates – Unrooted • No ancestry and time estimates • Algorithms to generate phylogenetic tree – Neighbor-joining – Unweighted Pair Group Method with Arithmetic Mean (UPGMA) – Maximum parsimony 2015 -12 -09 35
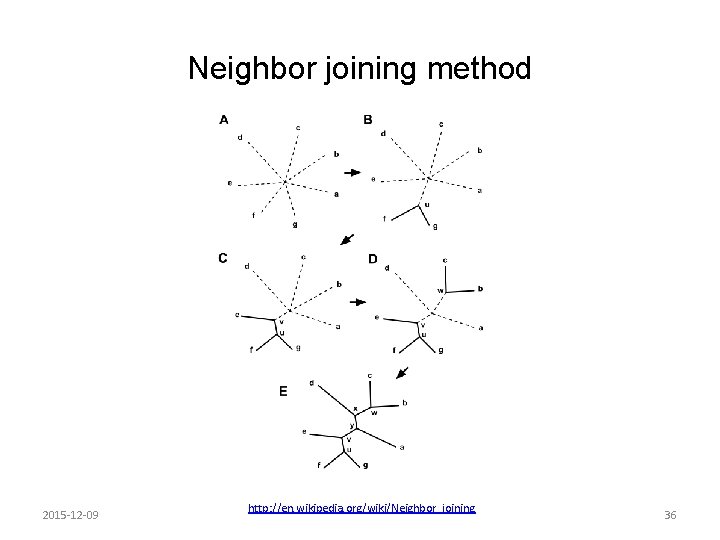
Neighbor joining method 2015 -12 -09 http: //en. wikipedia. org/wiki/Neighbor_joining 36

MSA exercise • Align the protein sequences SET 1 and SET 2 using MSA tools and compare the alignments • Clustalw 2 – http: //www. ebi. ac. uk/Tools/msa/clustalw 2/ • MUSCLE – http: //www. ebi. ac. uk/Tools/msa/muscle/ 2015 -12 -09 37

What to align: DNA or protein sequence? If ORF exists, then always align at protein level • Many mis-matches in DNA sequences are synonymous • DNA sequences contain non-coding regions, which should be avaided in homology searching • Matches are more reliable in protein sequence – Probability to occur randomly at any position in a sequence • • • Amino acids: 1/20 = 0. 05 Nucleotides: 1/4 = 0. 25 Searcing at protein level: In case of frameshifts, the alignment score for protein sequence may be very low even though the DNA sequence are similar 2015 -12 -09 ACT TTT CAT GGG . . . Thr Phe His Gly . . . ACT TTT TCA TGG G. . Thr Phe Ser Trp 38

Searching bioinformatics databases using: keywords and, sequences 2015 -12 -09 39

Search strategy • Keyword search – Find information related to specific keywords – Each bioinformatics database has its own search tool – Some search tools have a wide spectrum which access multiple databases and gather results together – Gquery, EBI search • Sequence search – Use a sequence of interest to find more information about the sequence – BLAST, FASTA 2015 -12 -09 40

Keyword search • Find information related to specific keywords • Gquery – A central search tool to find information in NCBI databases – Searches in large number of NCBI databases and shows them in one page – http: //www. ncbi. nlm. nih. gov/gquery • EBI search – Search tool to find infroamtion from databases developed, managed and hosted by EMBL-EBI – http: //www. ebi. ac. uk/services 2015 -12 -09 41
Gquery 2015 -12 -09 42
EBI search 2015 -12 -09 43
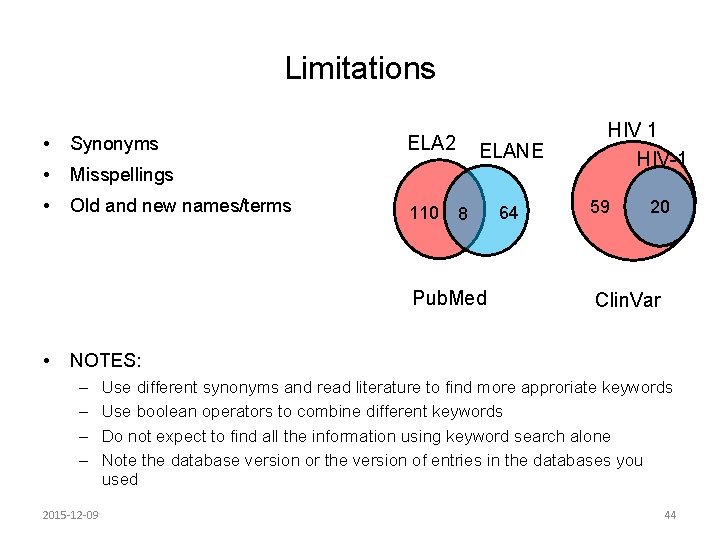
Limitations • Synonyms • Misspellings • Old and new names/terms ELA 2 110 ELANE 8 Pub. Med • 64 HIV 1 HIV-1 59 20 Clin. Var NOTES: – – 2015 -12 -09 Use different synonyms and read literature to find more approriate keywords Use boolean operators to combine different keywords Do not expect to find all the information using keyword search alone Note the database version or the version of entries in the databases you used 44

Gene nomenclature • HUGO Gene Nomenclature Committee (HGNC) – Assigns standardized nomenclature to human genes – Each symbol is unique and each gene is given only one name • Species specific nomenclature committees – Mouse Genome Informatics Database • http: //www. informatics. jax. org/mgihome/nomen/ – Rat Genome Database • http: //rgd. mcw. edu/nomen. shtml 2015 -12 -09 45

HGNC symbol report • Approved symbol • Approved name • Synonyms – Terms used in literature to indicate the gene – HGNC, Ensembl, Entrez Gene, OMIM • Previous symbols and names – Previous HGNC approved symbol • NOTE: HGNC does not approve protein names. Usually genes and proteins have the same name and gene names are written in italics. 2015 -12 -09 46
HGNC search 2015 -12 -09 47

Keyword search • Exercise 2015 -12 -09 48
- Slides: 48