Phylogeny and Molecular Evolution Character Based Phylogeny 501

Phylogeny and Molecular Evolution Character Based Phylogeny /501

Credit • • • Ron Shamir’s lecture notes Notes by Nir Friedman Dan Geiger, Shlomo Moran, Sagi Snir Durbin et al. Jones and Pevzner’s presentation Bioinformatics Algorithms book by Phillip Compeau and Pavel Pvzner – all book photos shown in this lecture are from there. /502

Roadmap: • • Review where we are at: Character Based Parsimony Two problems: Large versus Small Parsimony Today: Small Parsimony algorithms /503

Type of Tree Reconstruction • Character-based • • Input is a multiple alignment of sequences (one sequence per species). Distance-based • • Input is a matrix of distances between species Distance can be the relative length of the sequence which the two sequences disagree on, or alignment score between them, or (whatever) /624

Parsimony Approach to Evolutionary Tree Reconstruction • Applies Occam’s razor principle to identify the simplest explanation for the data • Assumes observed character differences resulted from the fewest possible mutations • Seeks the tree that yields lowest possible parsimony score - sum of cost of all mutations found in the tree

Two Computational Parsimony Problems: Small versus Large Parsimony /509

Small Parsimony Problem • /5010

versus Large Parsimony • /5011
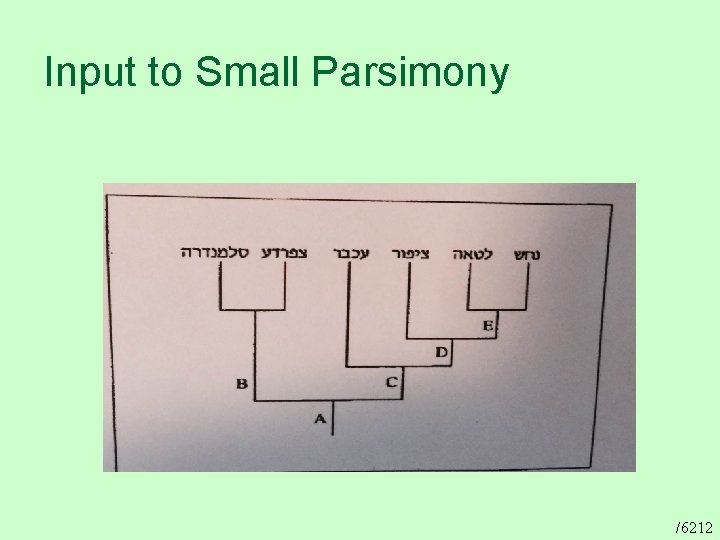
Input to Small Parsimony /6212

Roadmap: • Done with review of where we are at • Next: Small Parsimony Algorithms • Another example to planpractice how to solve Small Parsimony • Fitch Algorithm for Unweighted Small Parsimony • Sankoff Algorithm for Weighted Small Parsimony /5013
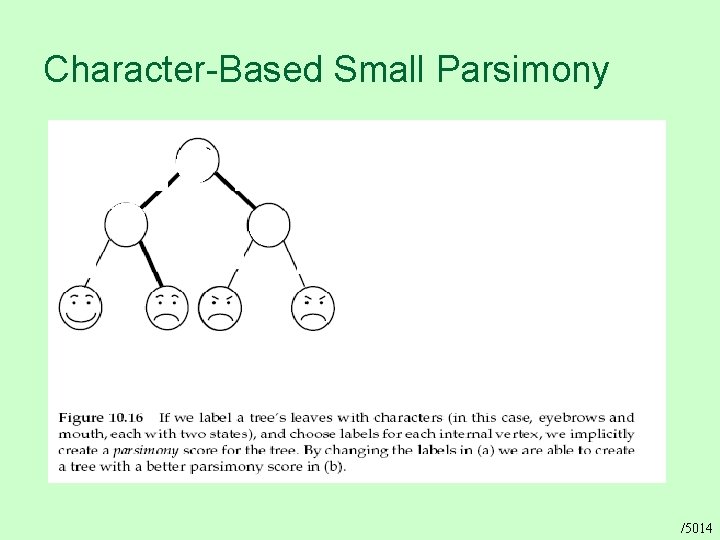
Character-Based Small Parsimony /5014
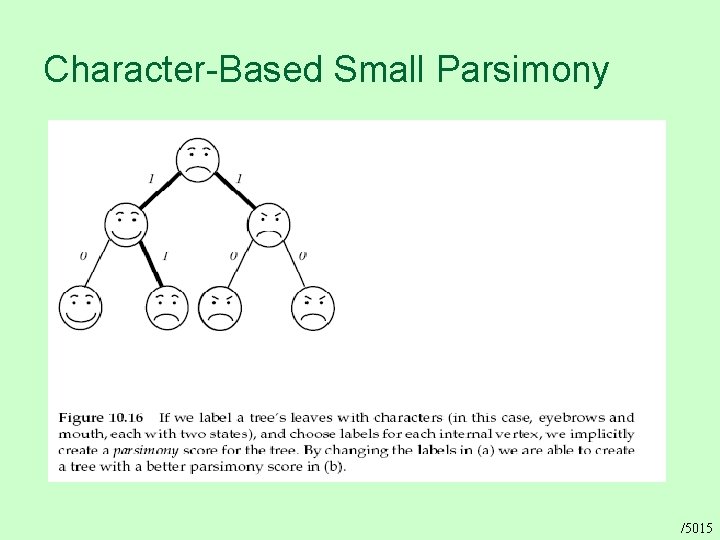
Character-Based Small Parsimony /5015
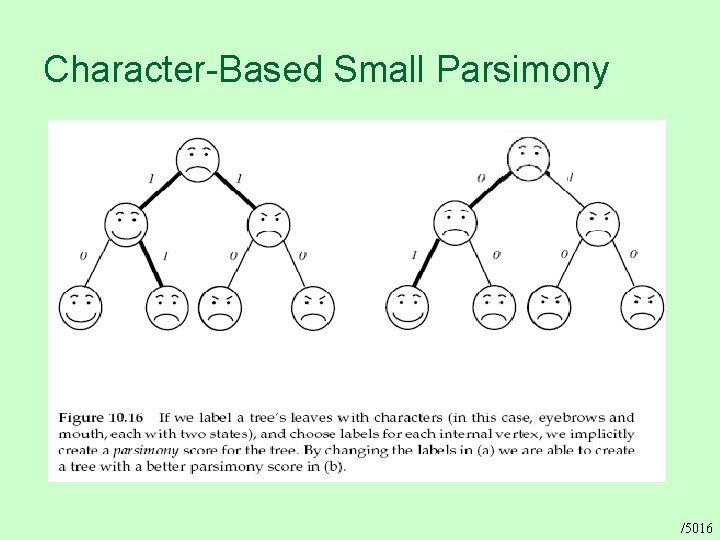
Character-Based Small Parsimony /5016
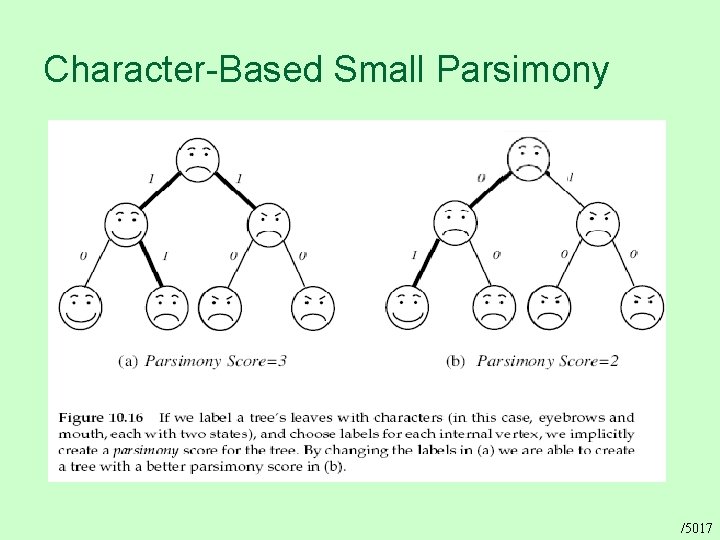
Character-Based Small Parsimony /5017
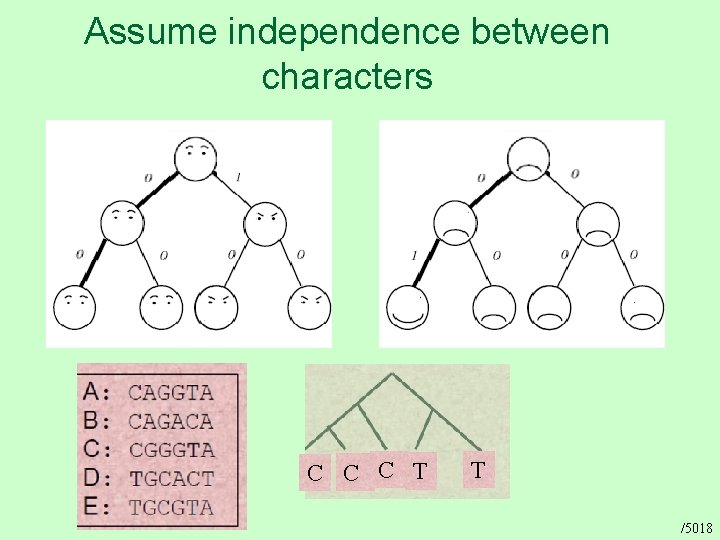
Assume independence between characters C C C T T /5018

Roadmap: • Done with example to planpractice how to solve Small Parsimony • Next: Fitch Algorithm for Unweighted Small Parsimony /5019
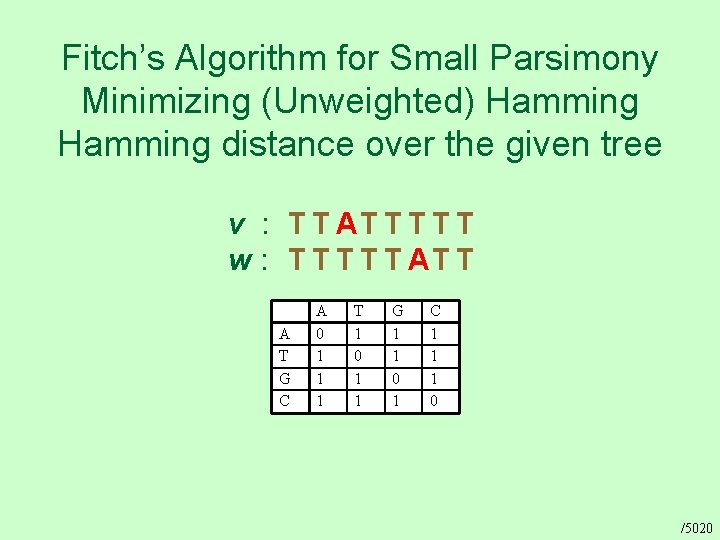
Fitch’s Algorithm for Small Parsimony Minimizing (Unweighted) Hamming distance over the given tree v : T T AT T T w : T T T AT T A T G C A 0 1 1 1 T 1 0 1 1 G 1 1 0 1 C 1 1 1 0 /5020

Fitch Algorithm (Tree is Given) • Work on each position in a string independently. • Start at the leaves. • If two children have common character, parent “inherits” it. • Record union and go up. • After reaching root, go down to fix sets of size > 1. A A C T A /5021

Fitch Algorithm (Tree is Given) • Work on each position in a string independently. • Start at the leaves. • If two children have common character, parent “inherits” it. • Record union and go up. • After reaching root, go down to fix sets of size > 1. A/C A A C T A /5022

Fitch Algorithm (Tree is Given) • Work on each position in a string independently. • Start at the leaves. • If two children have common character, parent “inherits” it. • Record union and go up. • After reaching root, go down to fix sets of size > 1. A A/C C T A /5023

Fitch Algorithm (Tree is Given) • Work on each position in a string independently. • Start at the leaves. • If two children have common character, parent “inherits” it. • Record union and go up. • After reaching root, go down to fix sets of size > 1. A/T A A/C C T A /5024

Fitch Algorithm (Tree is Given) • Work on each position in a string independently. • Start at the leaves. • If two children have common character, parent “inherits” it. • Record union and go up. • After reaching root, go down to fix sets of size > 1. A A/T A A/C C T A /5025

Fitch Algorithm (Tree is Given) • Work on each position in a string independently. • Start at the leaves. • If two children have common character, parent “inherits” it. • Record union and go up. • After reaching root, go down to fix sets of size > 1. A A A/T A/C C T A /5026

Fitch Algorithm (Tree is Given) • Work on each position in a string independently. • Start at the leaves. • If two children have common character, parent “inherits” it. • Record union and go up. • After reaching root, go down to fix sets of size > 1. A A A/T A/C C T A /5027

Fitch Algorithm (Tree is Given) • Work on each position in a string independently. • Start at the leaves. • If two children have common character, parent “inherits” it. • Record union and go up. • After reaching root, go down to fix sets of size > 1. A A A/T A/C C T A /5028

Fitch Algorithm (Tree is Given) • Work on each position in a string independently. • Start at the leaves. • If two children have common character, parent “inherits” it. • Record union and go up. • After reaching root, go down to fix sets of size > 1. A A A/T A/C C T A /5029

Fitch’s Algorithm, More Formally traverse tree from leaves to root determining set of possible states (e. g. nucleotides) for each internal node traverse tree from root to leaves picking ancestral states for internal nodes /5030

Fitch’s Algorithm – Step 1 do a post-order (from leaves to root) traversal of tree Determine possible states Ri of internal node i with children j and k /5031

Fitch’s Algorithm – Step 2 do a pre-order (from root to leaves) traversal of tree select state rj of internal node j with parent i as follows: /5032
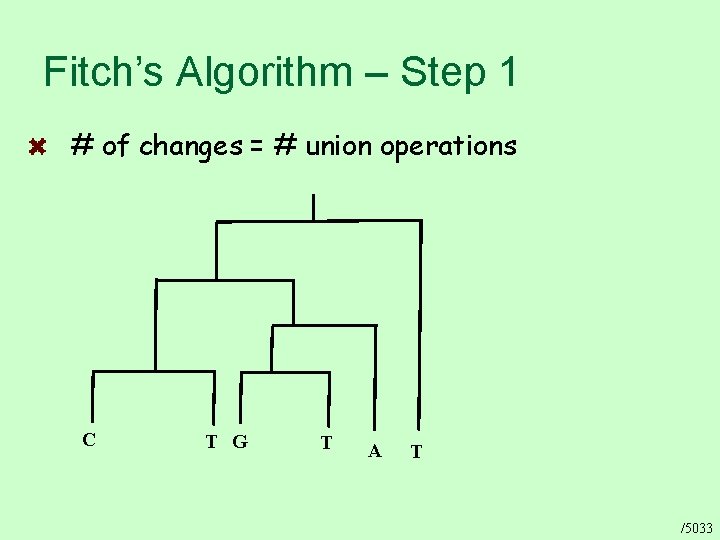
Fitch’s Algorithm – Step 1 # of changes = # union operations C T G T A T /5033
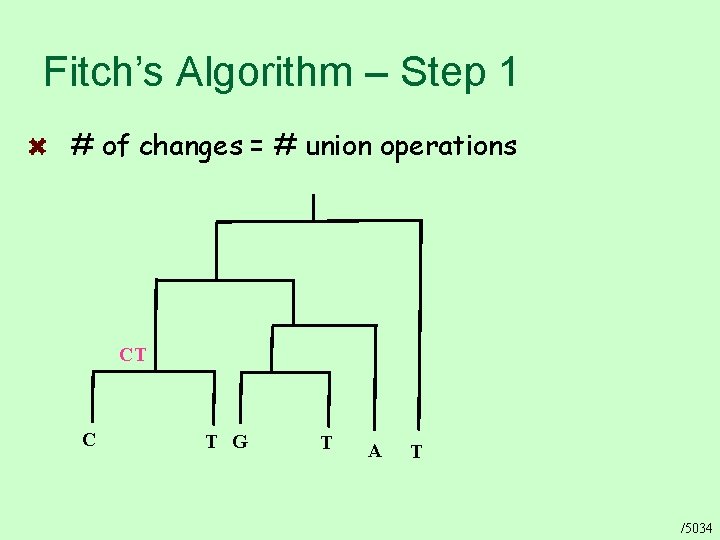
Fitch’s Algorithm – Step 1 # of changes = # union operations CT C T G T A T /5034
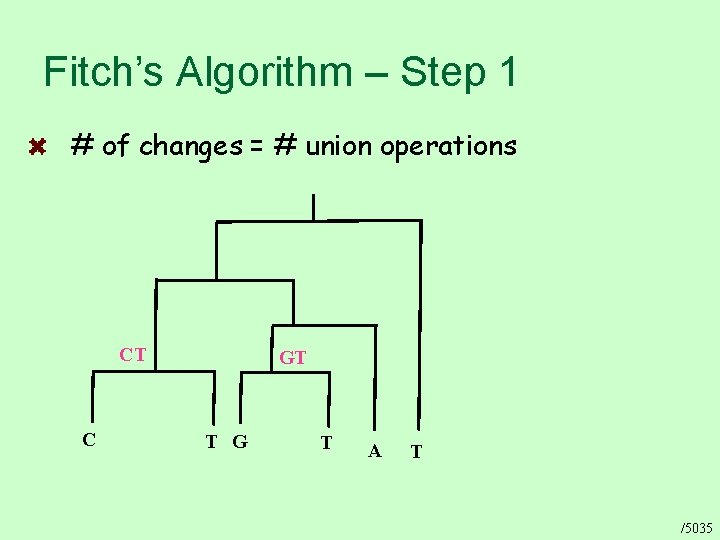
Fitch’s Algorithm – Step 1 # of changes = # union operations CT C GT T G T A T /5035
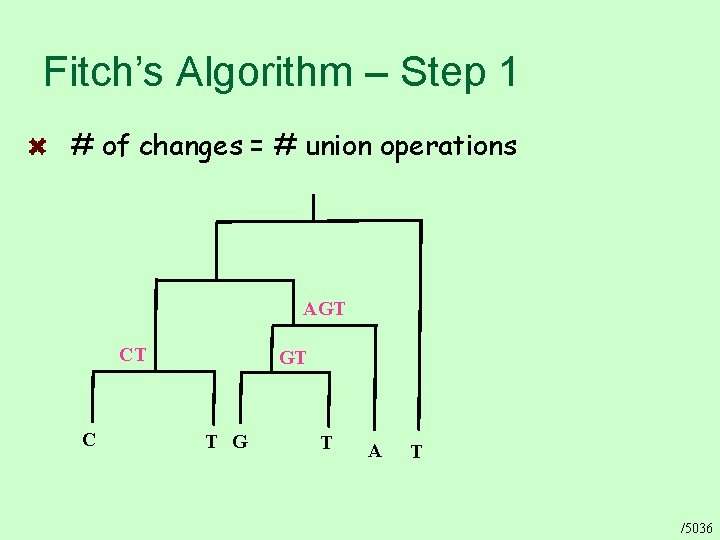
Fitch’s Algorithm – Step 1 # of changes = # union operations AGT CT C GT T G T A T /5036
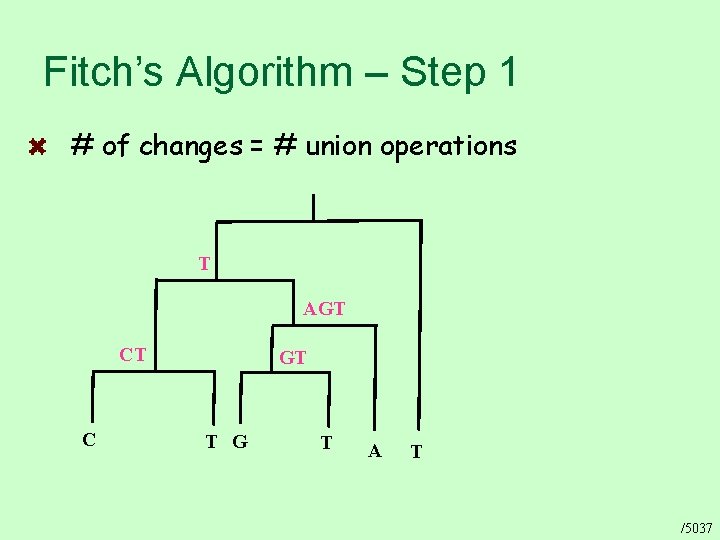
Fitch’s Algorithm – Step 1 # of changes = # union operations T AGT CT C GT T G T A T /5037
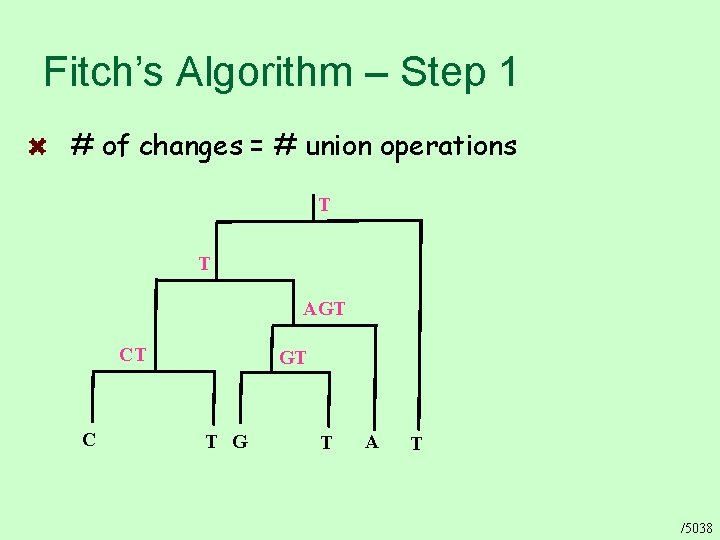
Fitch’s Algorithm – Step 1 # of changes = # union operations T T AGT CT C GT T G T A T /5038
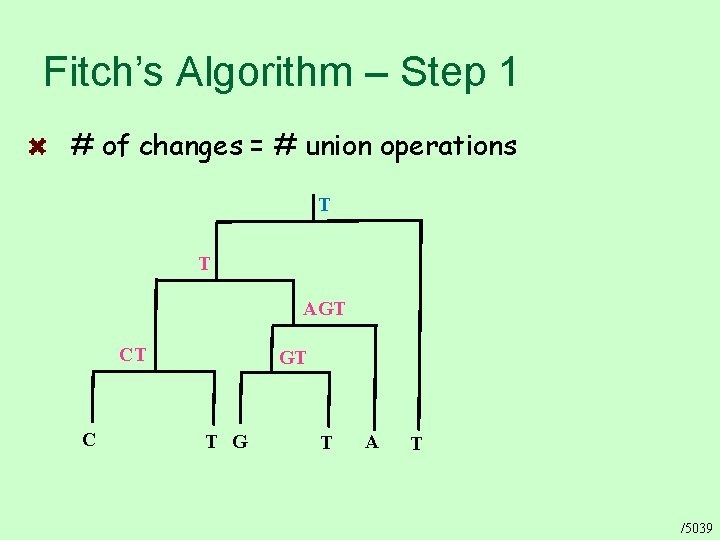
Fitch’s Algorithm – Step 1 # of changes = # union operations T T AGT CT C GT T G T A T /5039
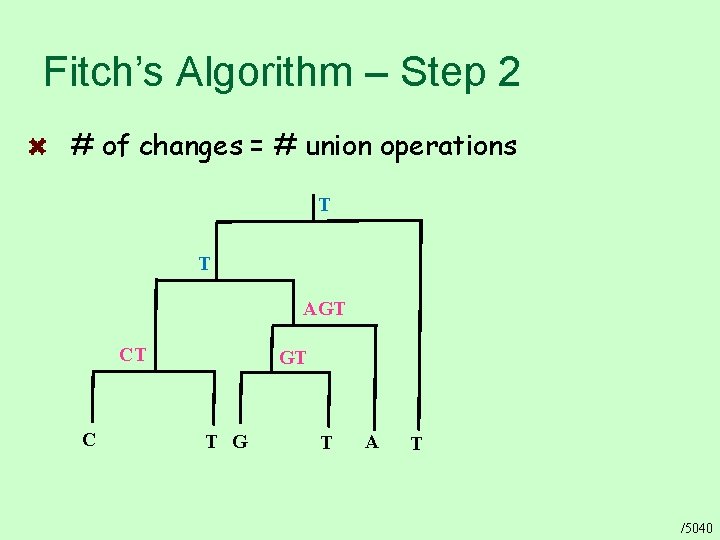
Fitch’s Algorithm – Step 2 # of changes = # union operations T T AGT CT C GT T G T A T /5040
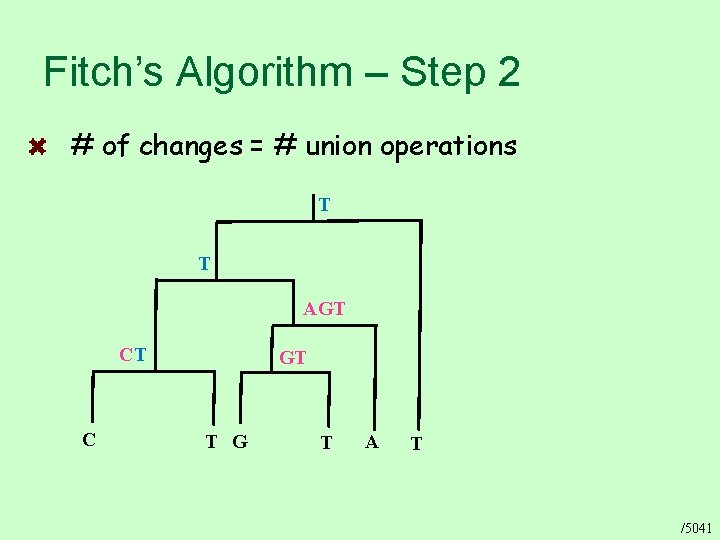
Fitch’s Algorithm – Step 2 # of changes = # union operations T T AGT CT C GT T G T A T /5041
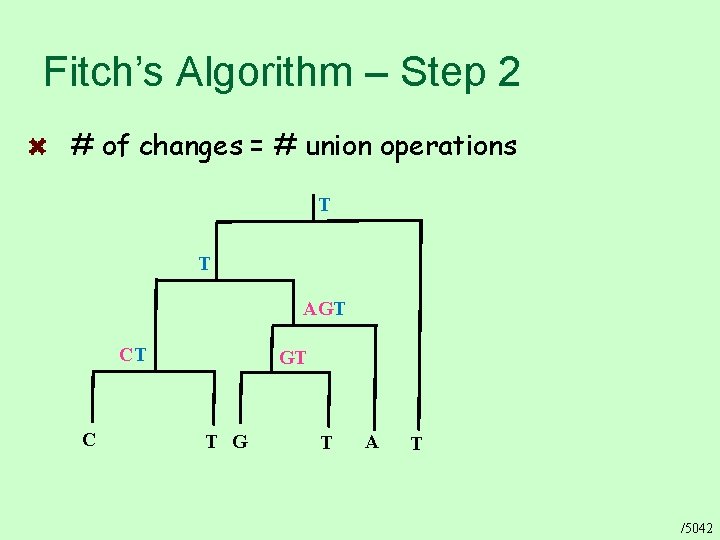
Fitch’s Algorithm – Step 2 # of changes = # union operations T T AGT CT C GT T G T A T /5042
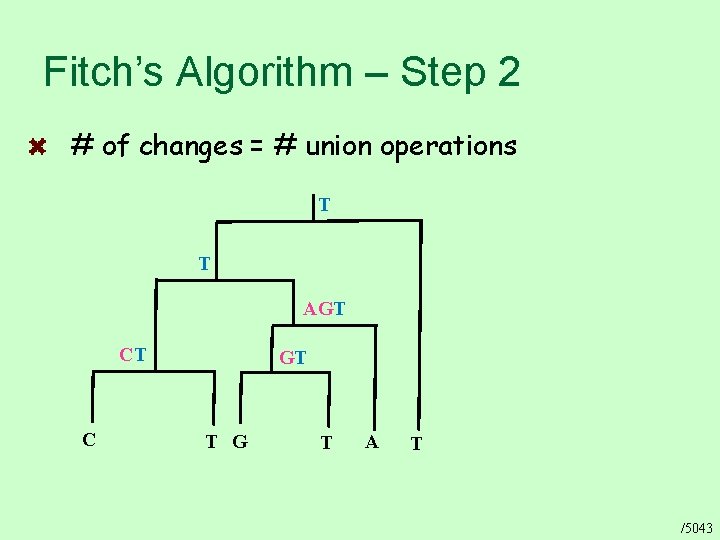
Fitch’s Algorithm – Step 2 # of changes = # union operations T T AGT CT C GT T G T A T /5043

Fitch’s Algorithm (cont’d) Another example: a c t a /5044
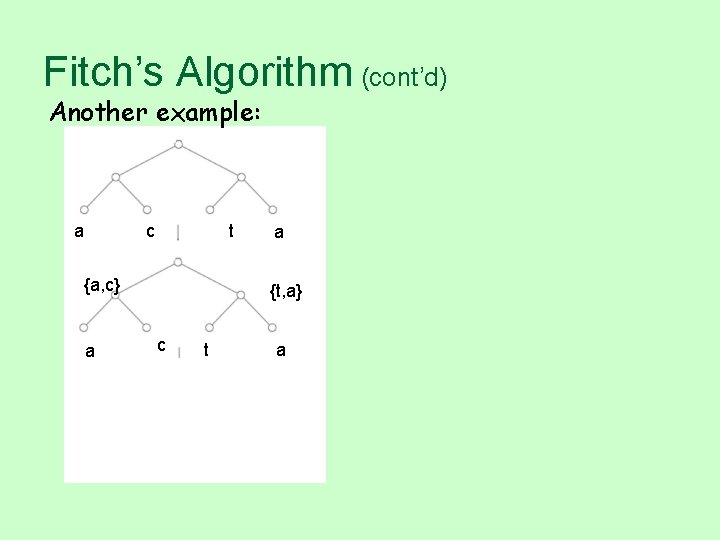
Fitch’s Algorithm (cont’d) Another example: a c t {a, c} a a {t, a} c t a
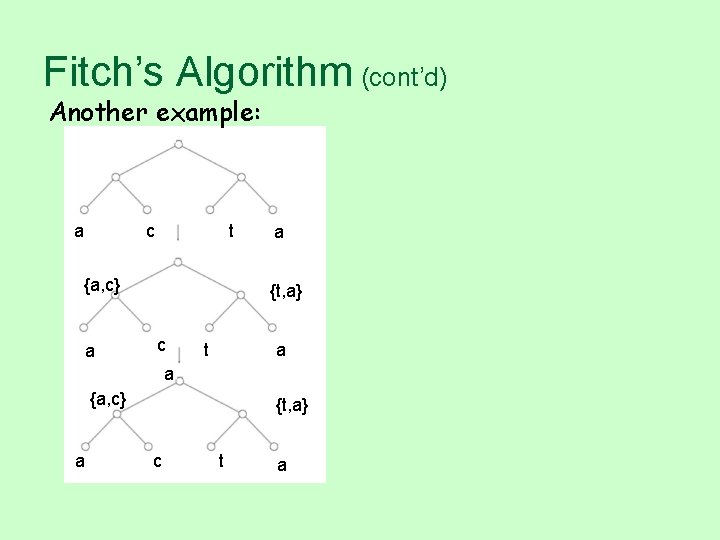
Fitch’s Algorithm (cont’d) Another example: a c t {a, c} a a {t, a} c t a a {a, c} a {t, a} c t a
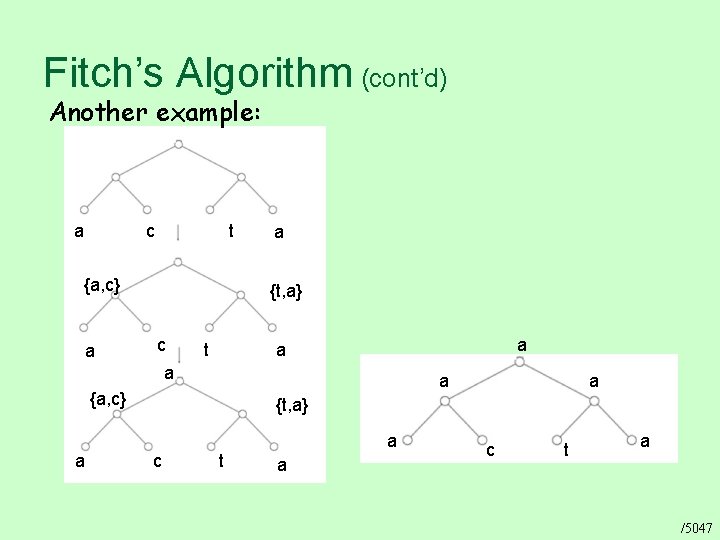
Fitch’s Algorithm (cont’d) Another example: a c t {a, c} a a {t, a} c t a a c t a /5047

Fitch’s Algorithm (cont’d) Time Complexity? /5048

Fitch’s Algorithm (cont’d) Time Complexity? Note that in the biomolecular sequences we mentioned so far the alphabet size is constant, but there are some bioinformatic applications where it is not. For example, when considering an alphabet of gene orthology families in bacterial genomes the alphabet size is typically O(n). /5049

Fitch’s Algorithm (cont’d) Correctness? Each set-union operation corresponds to exactly one required mutation /5050

/5051

Roadmap: • Done with Fitch Algorithm for Unweighted Small Parsimony • Next: Sankoff Algorithm for Weighted Small Parsimony /5052

Weighted Small Parsimony score: • Each change is weighted by a score c(a, b). • The weighted parsimony score reduces to the parsimony score when c(a, a)=0 and c(a, b)=1 for all b ≠ a. A T G C A 0 3 4 9 T 3 0 2 4 G 4 2 0 4 C 9 4 4 0 /5053
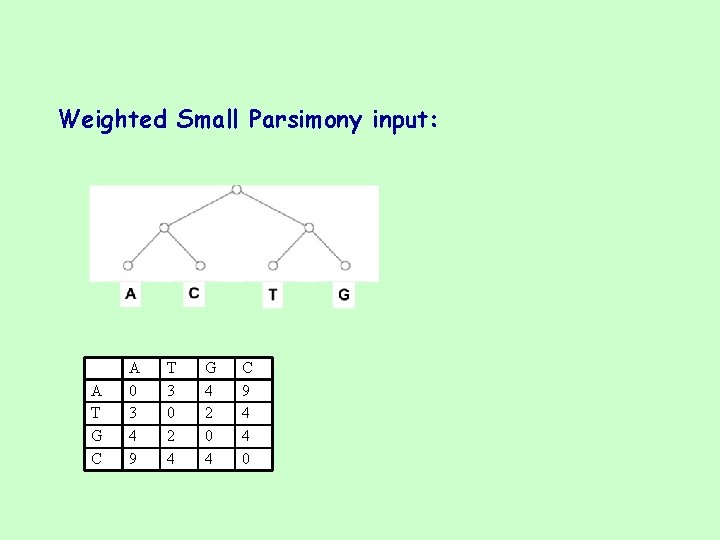
Weighted Small Parsimony input: A T G C A 0 3 4 9 T 3 0 2 4 G 4 2 0 4 C 9 4 4 0
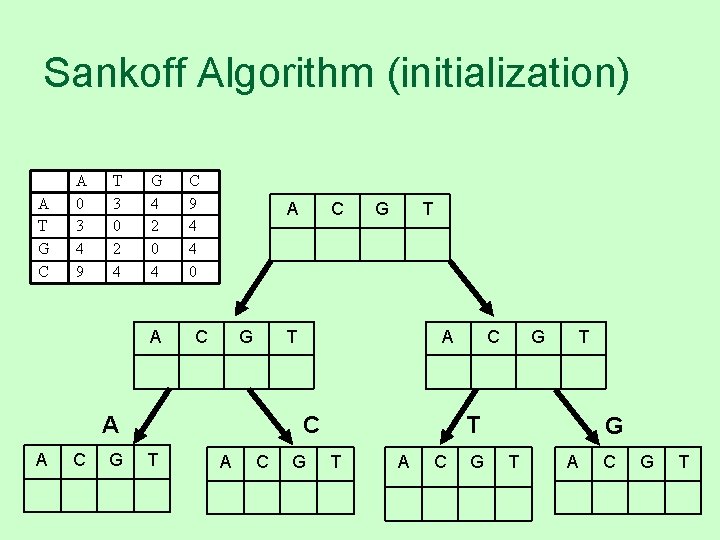
Sankoff Algorithm (initialization) A T G C A 0 3 4 9 T 3 0 2 4 G 4 2 0 4 A C 9 4 4 0 A C G G T T A A C T A C G T T T A C G G T A C G T
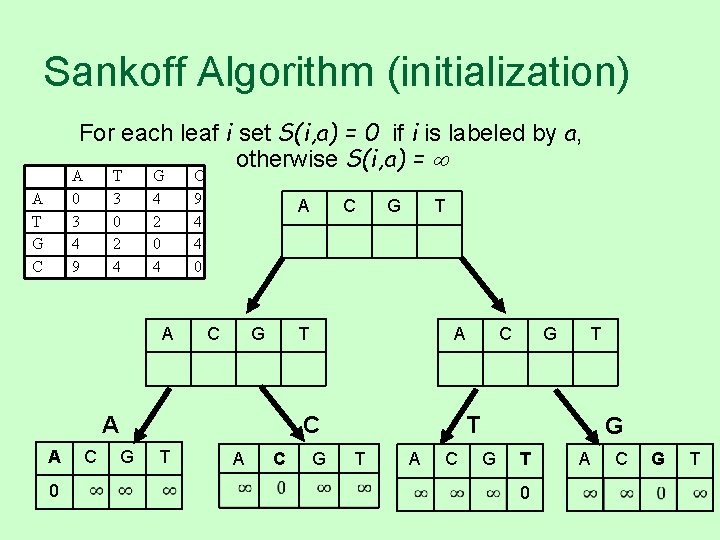
Sankoff Algorithm (initialization) For each leaf i set S(i, a) = 0 if i is labeled by a, otherwise S(i, a) = A 0 3 4 9 A T G C T 3 0 2 4 G 4 2 0 4 A C 9 4 4 0 A C G 0 C G T T A A C G T T T A C G G T 0 A C G T
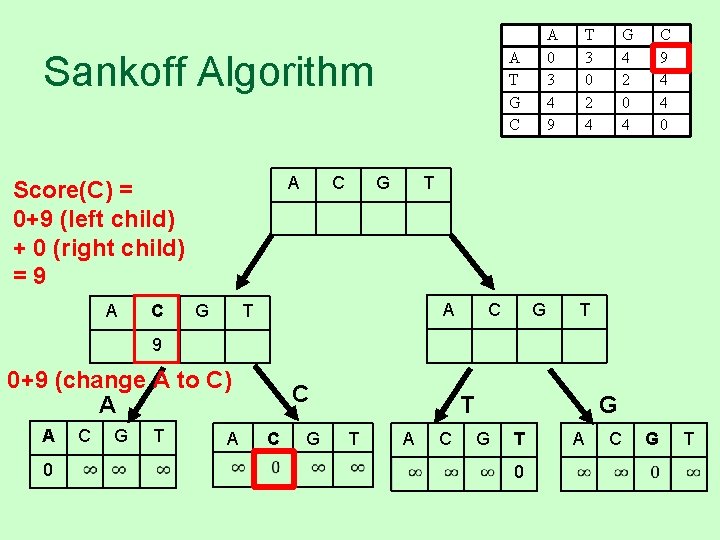
A T G C Sankoff Algorithm A Score(C) = 0+9 (left child) + 0 (right child) =9 A C G C A 0 3 4 9 G T 3 0 2 4 G 4 2 0 4 C 9 4 4 0 T A T C G T 9 0+9 (change A to C) A A 0 C G T A C C G T T A C G G T 0 A C G T
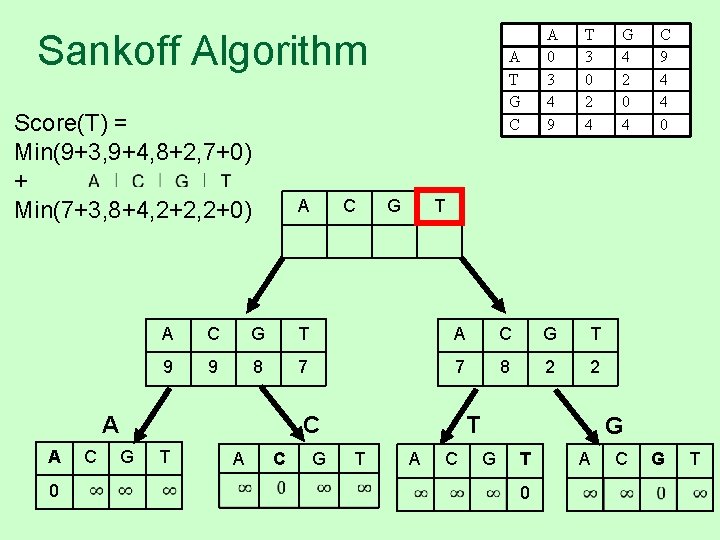
Sankoff Algorithm Score(T) = Min(9+3, 9+4, 8+2, 7+0) + Min(7+3, 8+4, 2+2, 2+0) A 0 C G T A C G T 9 9 8 7 7 8 2 2 T A C G T T G 4 2 0 4 C 9 4 4 0 T C G T 3 0 2 4 A A A C A T G C A 0 3 4 9 A C G G T 0 A C G T
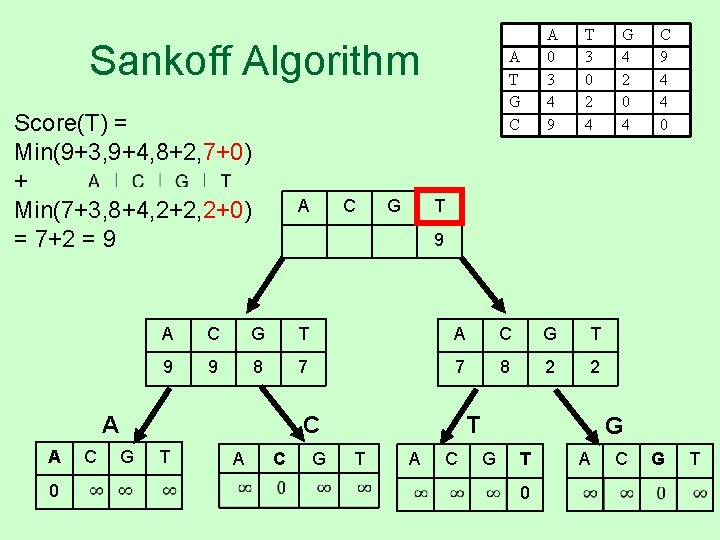
Sankoff Algorithm Score(T) = Min(9+3, 9+4, 8+2, 7+0) + Min(7+3, 8+4, 2+2, 2+0) = 7+2 = 9 A 0 C G A C G T 9 9 8 7 7 8 2 2 T A G 4 2 0 4 C 9 4 4 0 T C G T 3 0 2 4 9 A A C A T G C A 0 3 4 9 C G T T A C G G T 0 A C G T
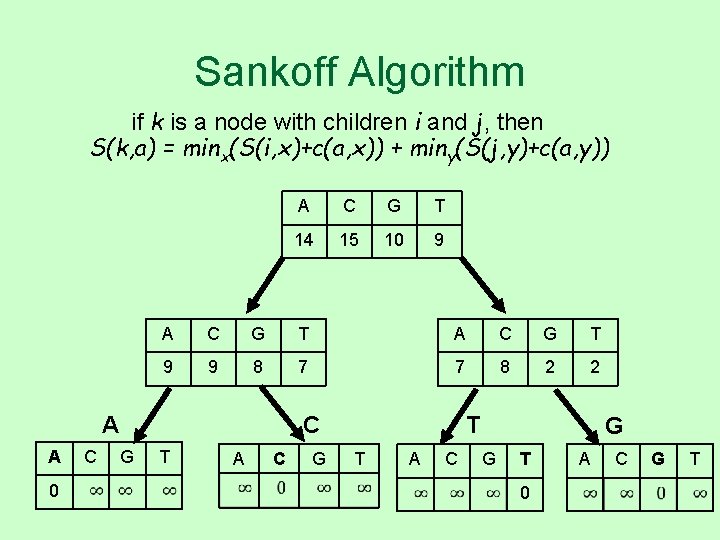
Sankoff Algorithm if k is a node with children i and j, then S(k, a) = minx(S(i, x)+c(a, x)) + miny(S(j, y)+c(a, y)) 0 C C G T 14 15 10 9 A C G T 9 9 8 7 7 8 2 2 A A A C G T T A C G G T 0 A C G T
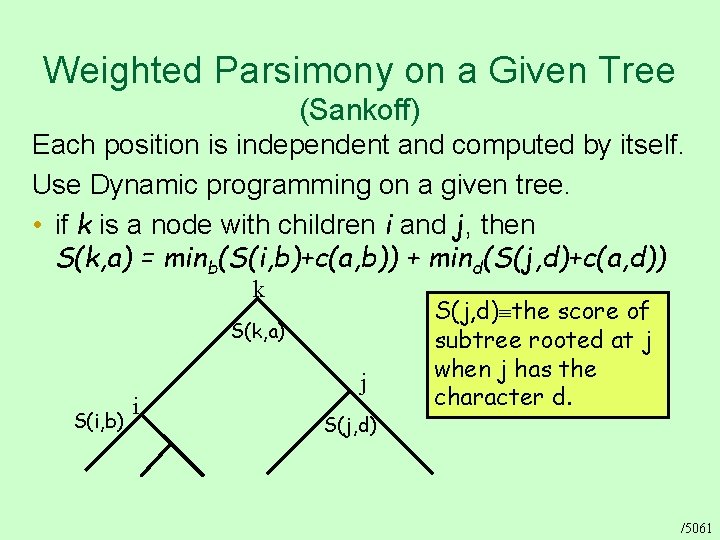
Weighted Parsimony on a Given Tree (Sankoff) Each position is independent and computed by itself. Use Dynamic programming on a given tree. • if k is a node with children i and j, then S(k, a) = minb(S(i, b)+c(a, b)) + mind(S(j, d)+c(a, d)) k S(k, a) S(i, b) i j S(j, d) the score of subtree rooted at j when j has the character d. /5061

Evaluating Parsimony Scores Dynamic programming on a given tree Initialization: • For each leaf i set S(i, a) = 0 if i is labeled by a, otherwise S(i, a) = Iteration: • if k is a node with children i and j, then S(k, a) = minx(S(i, x)+c(a, x)) + miny(S(j, y)+c(a, y)) Termination: • The cost of the tree is minx. S(r, x) where r is the root Traceback: • If we keep in each node for each character “a” the two characters x, y that bring about the minimum, then we can trace the best assignment to all internal nodes. /5062
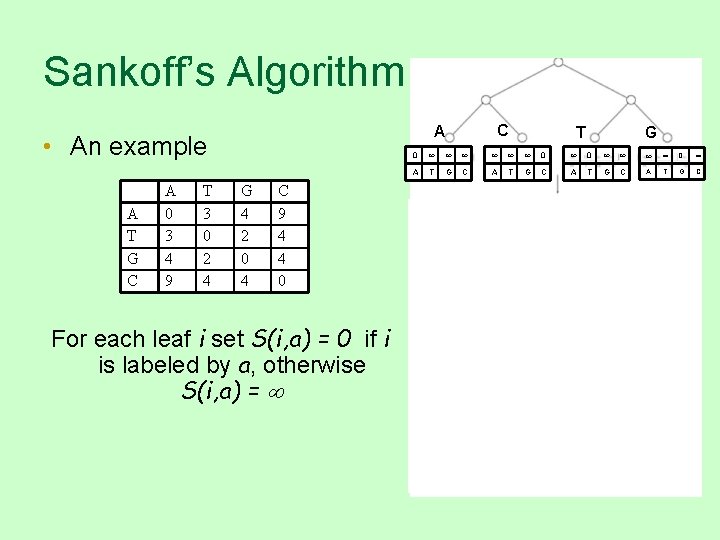
Sankoff’s Algorithm • An example A T G C A 0 3 4 9 T 3 0 2 4 C A G 4 2 0 4 C 9 4 4 0 For each leaf i set S(i, a) = 0 if i is labeled by a, otherwise S(i, a) = T G 0 0 0 0 A T G C
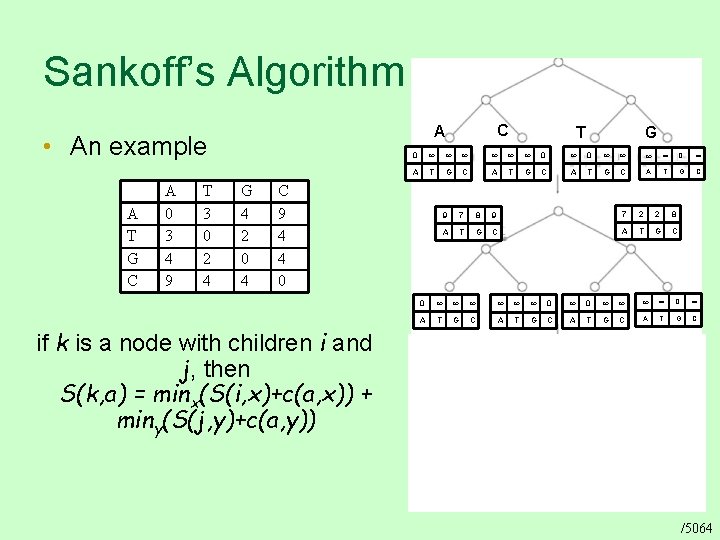
Sankoff’s Algorithm • An example A T G C A 0 3 4 9 T 3 0 2 4 C A G 4 2 0 4 G 0 0 0 0 A T G C C 9 4 4 0 if k is a node with children i and j, then S(k, a) = minx(S(i, x)+c(a, x)) + miny(S(j, y)+c(a, y)) T 9 7 8 9 7 2 2 8 A T G C 0 0 0 0 A T G C 14 9 10 15 A T G C 9 7 8 9 7 2 2 8 A T G C 0 0 0 0 A T G C /5064
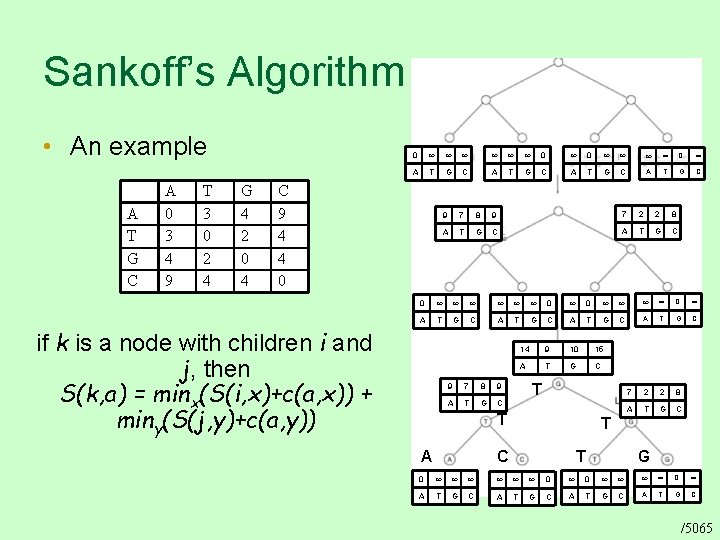
Sankoff’s Algorithm • An example A T G C A 0 3 4 9 T 3 0 2 4 G 4 2 0 4 0 0 0 0 A T G C C 9 4 4 0 if k is a node with children i and j, then S(k, a) = minx(S(i, x)+c(a, x)) + miny(S(j, y)+c(a, y)) 9 7 8 9 7 2 2 8 A T G C 0 0 0 0 A T G C 14 9 10 15 A T G C 7 2 2 8 A T G C 9 7 8 9 A T G C T T A T C T G 0 0 0 0 A T G C /5065
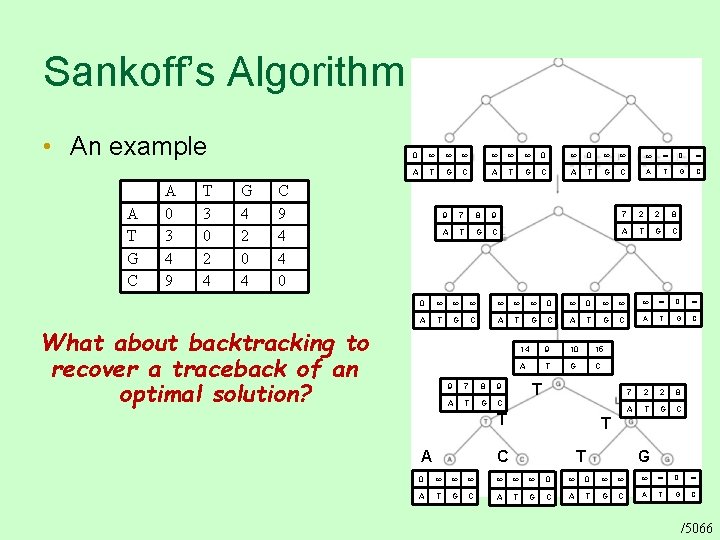
Sankoff’s Algorithm • An example A T G C A 0 3 4 9 T 3 0 2 4 G 4 2 0 4 0 0 0 0 A T G C C 9 4 4 0 What about backtracking to recover a traceback of an optimal solution? 9 7 8 9 7 2 2 8 A T G C 0 0 0 0 A T G C 14 9 10 15 A T G C 7 2 2 8 A T G C 9 7 8 9 A T G C T T A T C T G 0 0 0 0 A T G C /5066

Complexity of Evaluating (Small) Weighted Parsimony If there are n nodes, m characters, and s possible values for each character, then what is the complexity of Sankoff’s algorithm for Small Parsimony? /5067

Complexity of Evaluating (Small) Weighted Parsimony If there are n nodes, m characters, and s possible values for each character, then complexity is O(nms 2). Of course, in Large Parsimony we still need to search over possible trees and find the best one. One usually resorts to heuristic search techniques. /5068

/5069

Food for thought: Sankoff algorithm versus Fitch: • The weighted parsimony score reduces to the parsimony score when c(a, a)=0 and c(a, b)=1 for all b ≠ a. • But the time complexity of the two algorithms differs by a factor of s… • What happens if you run the Sankoff algorithm using a table where c(a, a)=0 and c(a, b)=1 for all b ≠ a … is the output equivalent to that of the Fitch algorithm? Is there any additional information provided by the Sankoff algorithm? /5070
- Slides: 67