Modelling proteomes Ram Samudrala Assistant Professor Department of




- Slides: 35

Modelling proteomes Ram Samudrala Assistant Professor Department of Microbiology University of Washington How does the genome of an organism specify its behaviour and characteristics?

Proteome – all proteins of a particular system ~60, 000 in human ~60, 000 in rice ~4500 in bacteria like Salmonella and E. coli Several thousand distinct sequence families

Modelling proteomes – understand the structure of individual proteins A few thousand distinct structural folds

Modelling proteomes – understand their individual functions Thousands of possible functions

Modelling proteomes – understand their expression Different expression patterns based on time and location

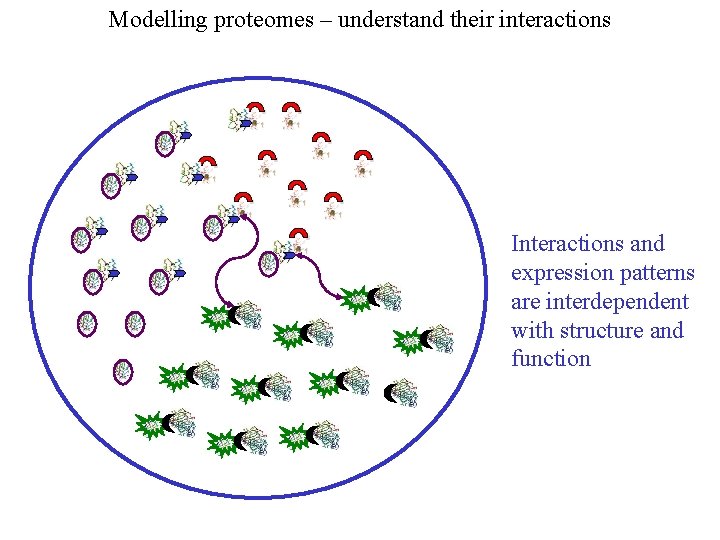
Modelling proteomes – understand their interactions Interactions and expression patterns are interdependent with structure and function

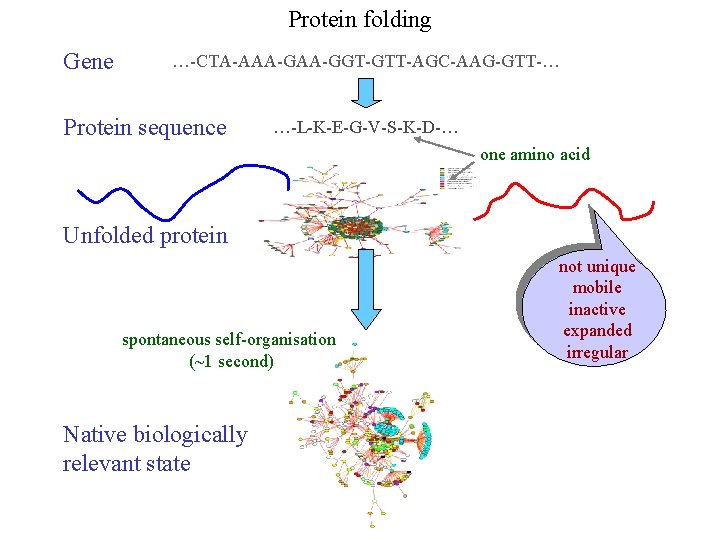
Protein folding Gene …-CTA-AAA-GGT-GTT-AGC-AAG-GTT-… Protein sequence …-L-K-E-G-V-S-K-D-… one amino acid Unfolded protein spontaneous self-organisation (~1 second) Native biologically relevant state not unique mobile inactive expanded irregular

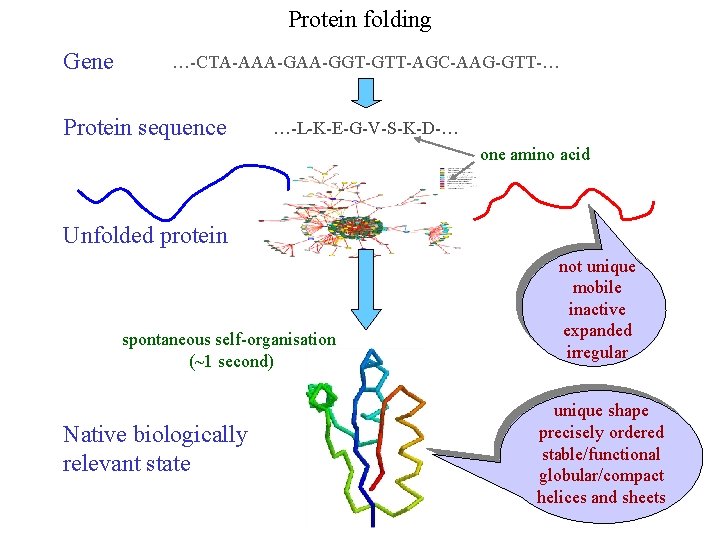
Protein folding Gene …-CTA-AAA-GGT-GTT-AGC-AAG-GTT-… Protein sequence …-L-K-E-G-V-S-K-D-… one amino acid Unfolded protein spontaneous self-organisation (~1 second) Native biologically relevant state not unique mobile inactive expanded irregular unique shape precisely ordered stable/functional globular/compact helices and sheets

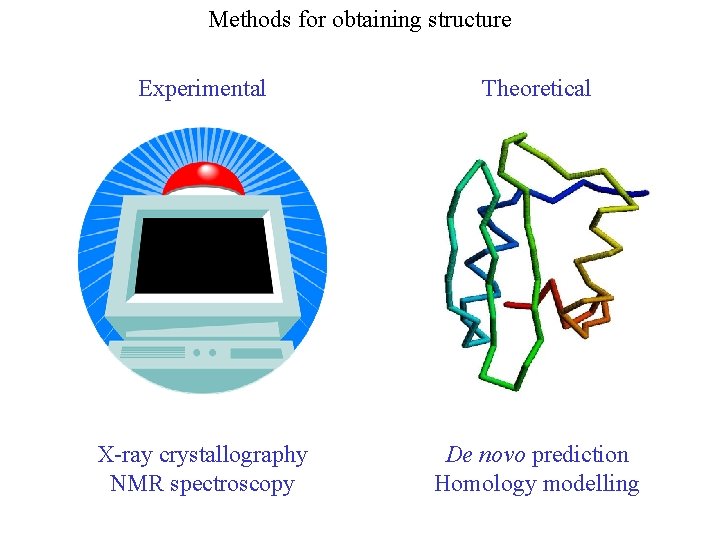
Methods for obtaining structure Experimental Theoretical X-ray crystallography NMR spectroscopy De novo prediction Homology modelling

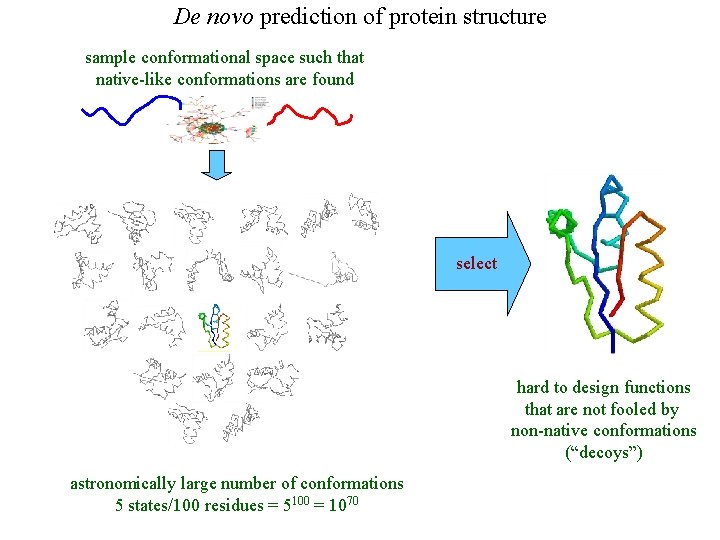
De novo prediction of protein structure sample conformational space such that native-like conformations are found select hard to design functions that are not fooled by non-native conformations (“decoys”) astronomically large number of conformations 5 states/100 residues = 5100 = 1070

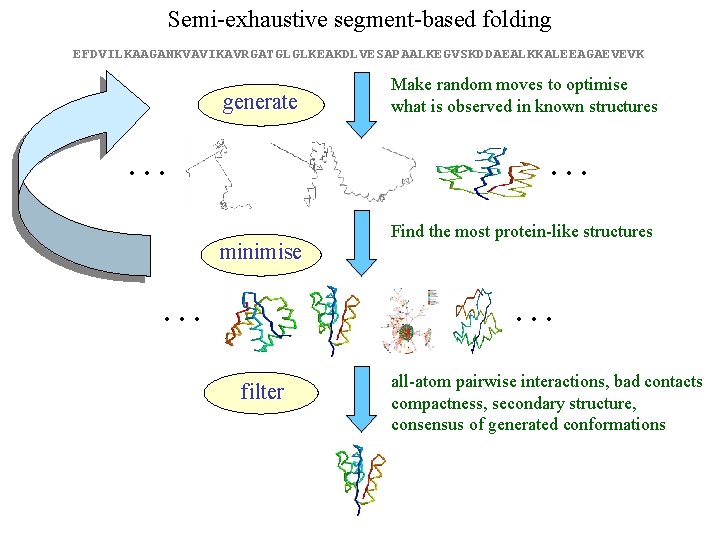
Semi-exhaustive segment-based folding EFDVILKAAGANKVAVIKAVRGATGLGLKEAKDLVESAPAALKEGVSKDDAEALKKALEEAGAEVEVK generate … Make random moves to optimise what is observed in known structures … minimise … Find the most protein-like structures … filter all-atom pairwise interactions, bad contacts compactness, secondary structure, consensus of generated conformations

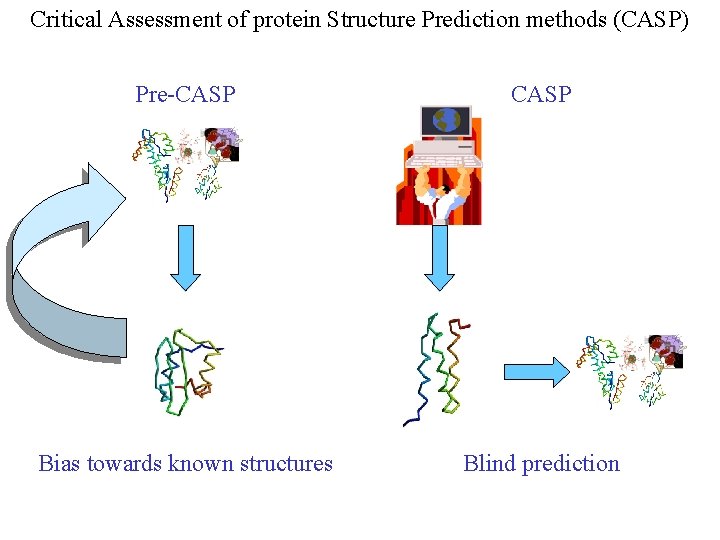
Critical Assessment of protein Structure Prediction methods (CASP) Pre-CASP Bias towards known structures Blind prediction

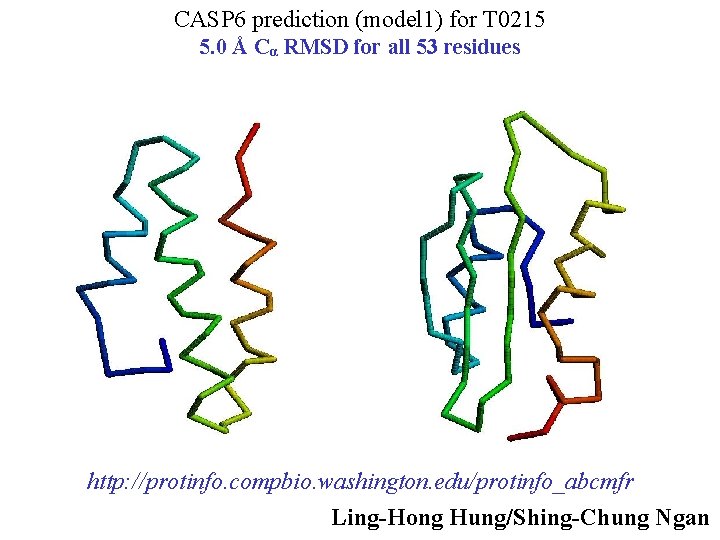
CASP 6 prediction (model 1) for T 0215 5. 0 Å Cα RMSD for all 53 residues http: //protinfo. compbio. washington. edu/protinfo_abcmfr Ling-Hong Hung/Shing-Chung Ngan

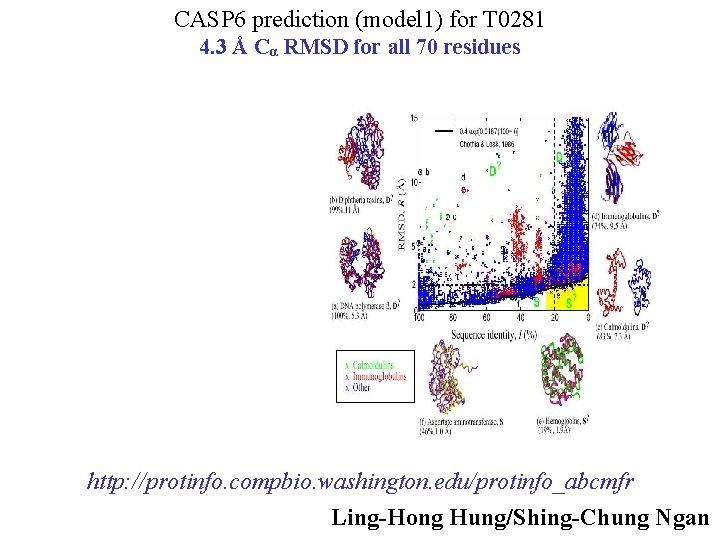
CASP 6 prediction (model 1) for T 0281 4. 3 Å Cα RMSD for all 70 residues http: //protinfo. compbio. washington. edu/protinfo_abcmfr Ling-Hong Hung/Shing-Chung Ngan

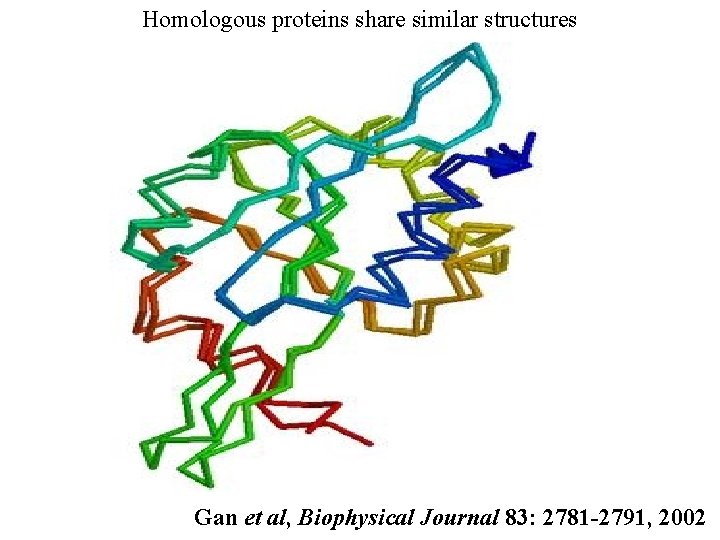
Homologous proteins share similar structures Gan et al, Biophysical Journal 83: 2781 -2791, 2002

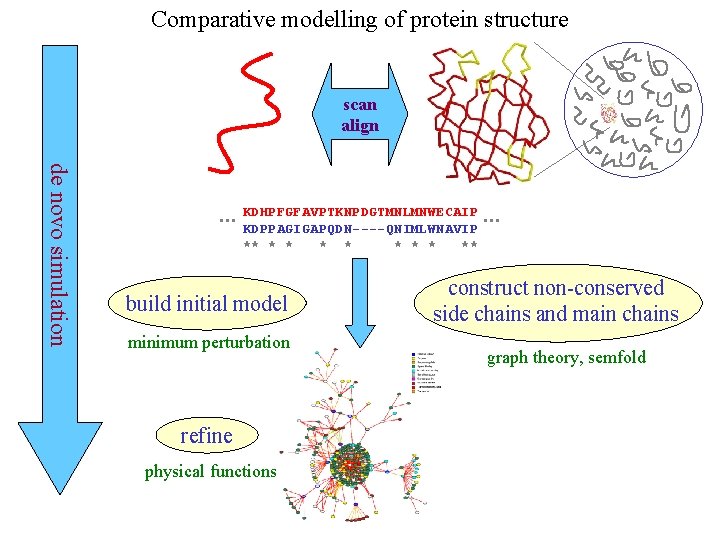
Comparative modelling of protein structure scan align de novo simulation … KDHPFGFAVPTKNPDGTMNLMNWECAIP KDPPAGIGAPQDN----QNIMLWNAVIP ** * * * ** build initial model minimum perturbation refine physical functions … construct non-conserved side chains and main chains graph theory, semfold

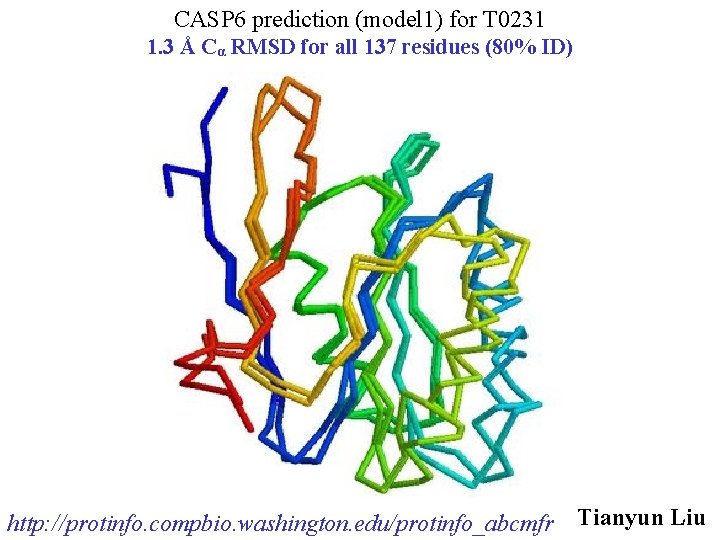
CASP 6 prediction (model 1) for T 0231 1. 3 Å Cα RMSD for all 137 residues (80% ID) http: //protinfo. compbio. washington. edu/protinfo_abcmfr Tianyun Liu

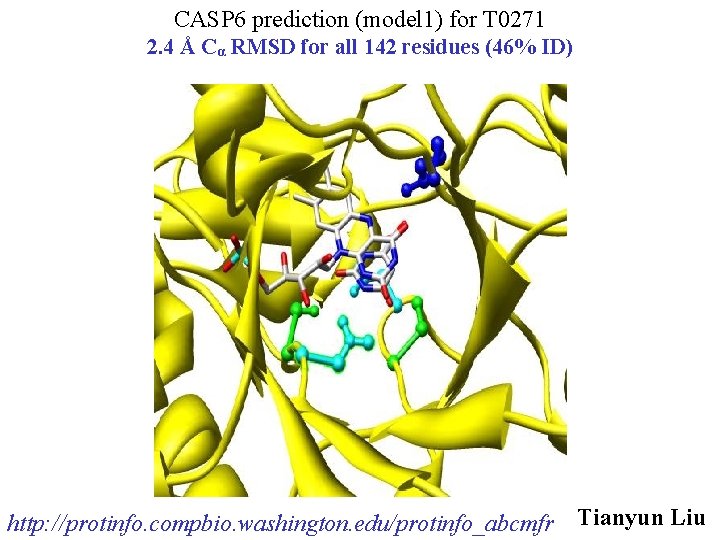
CASP 6 prediction (model 1) for T 0271 2. 4 Å Cα RMSD for all 142 residues (46% ID) http: //protinfo. compbio. washington. edu/protinfo_abcmfr Tianyun Liu

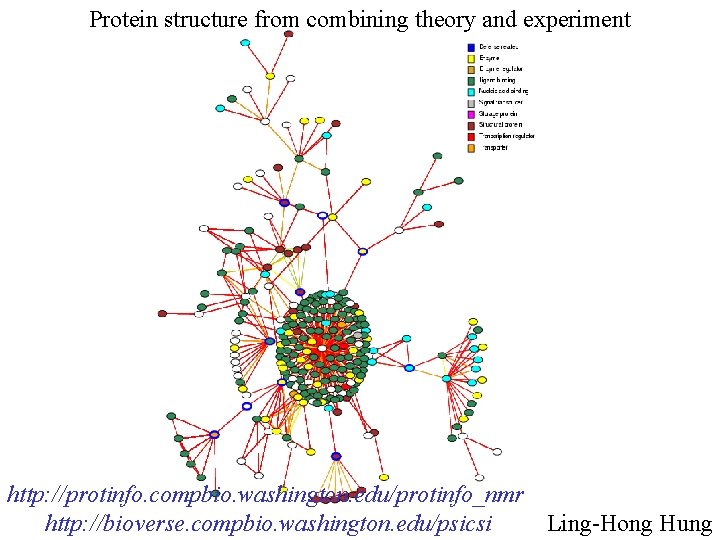
Protein structure from combining theory and experiment http: //protinfo. compbio. washington. edu/protinfo_nmr Ling-Hong Hung http: //bioverse. compbio. washington. edu/psicsi

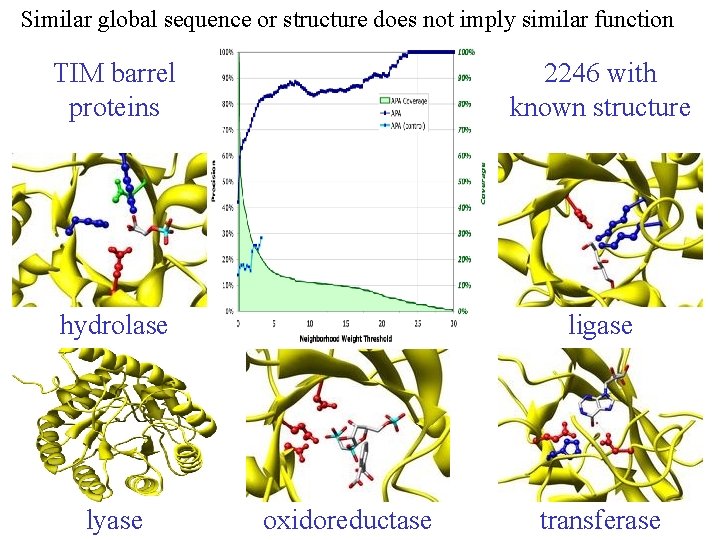
Similar global sequence or structure does not imply similar function TIM barrel proteins 2246 with known structure hydrolase ligase lyase oxidoreductase transferase

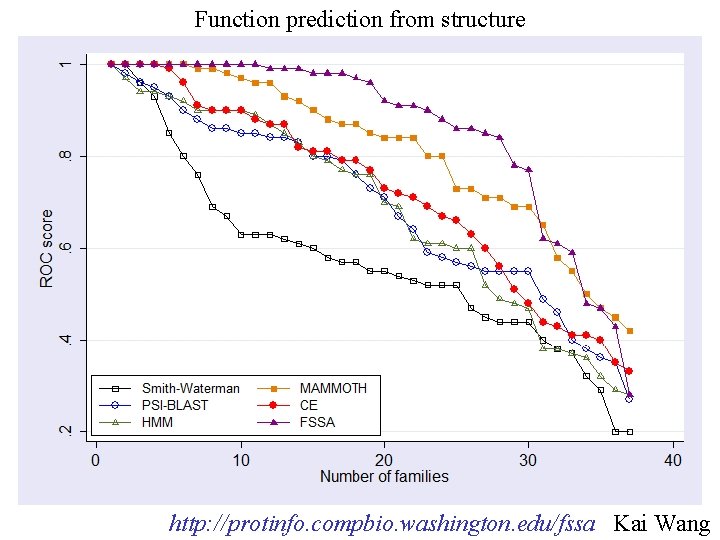
Function prediction from structure http: //protinfo. compbio. washington. edu/fssa Kai Wang

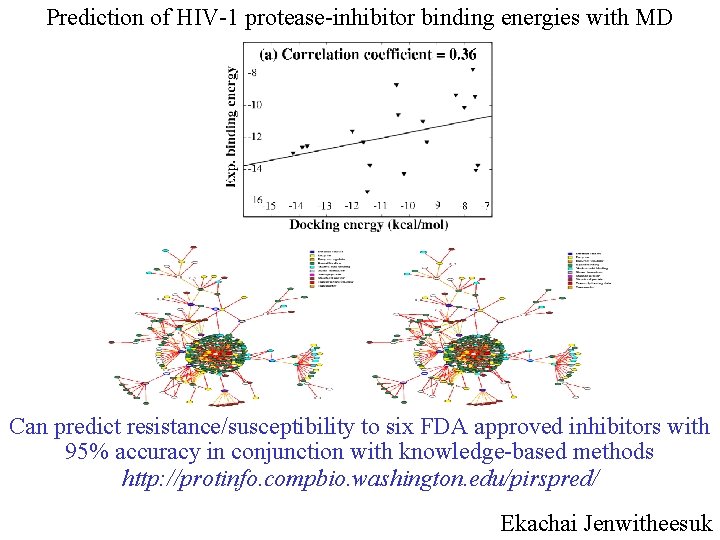
Prediction of HIV-1 protease-inhibitor binding energies with MD Can predict resistance/susceptibility to six FDA approved inhibitors with 95% accuracy in conjunction with knowledge-based methods http: //protinfo. compbio. washington. edu/pirspred/ Ekachai Jenwitheesuk

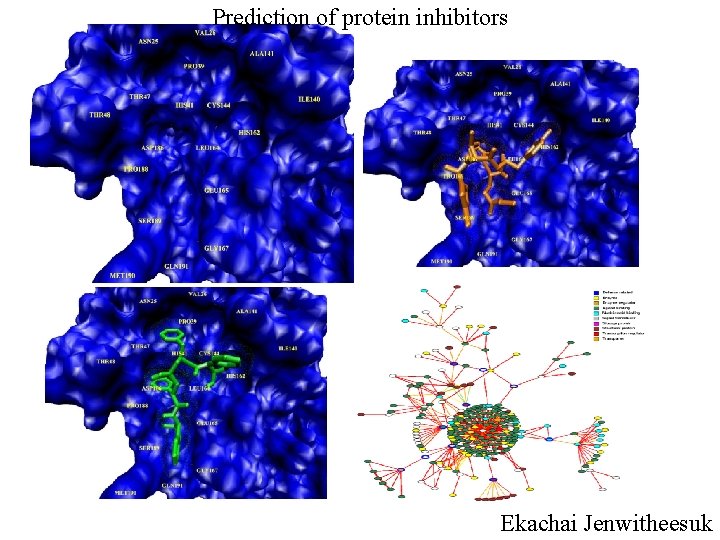
Prediction of protein inhibitors Ekachai Jenwitheesuk

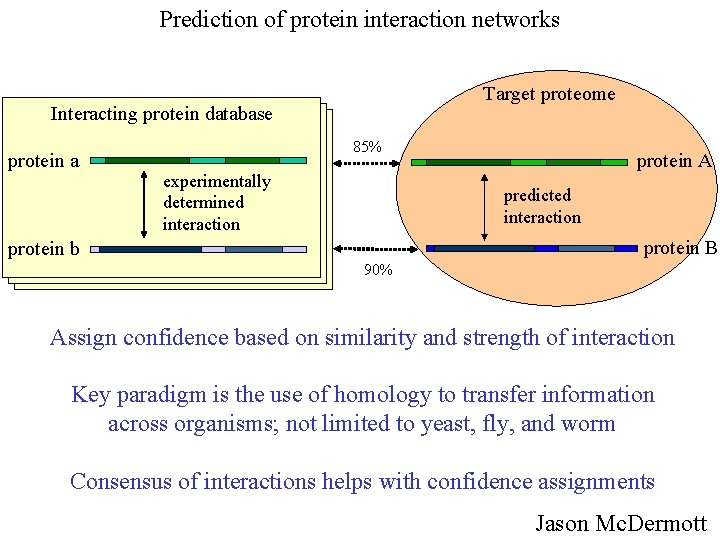
Prediction of protein interaction networks Target proteome Interacting protein database protein a 85% experimentally determined interaction protein A predicted interaction protein B protein b 90% Assign confidence based on similarity and strength of interaction Key paradigm is the use of homology to transfer information across organisms; not limited to yeast, fly, and worm Consensus of interactions helps with confidence assignments Jason Mc. Dermott

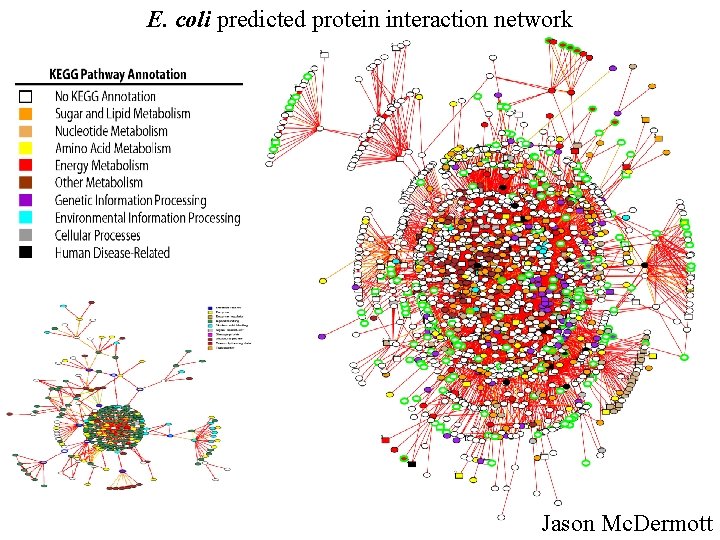
E. coli predicted protein interaction network Jason Mc. Dermott

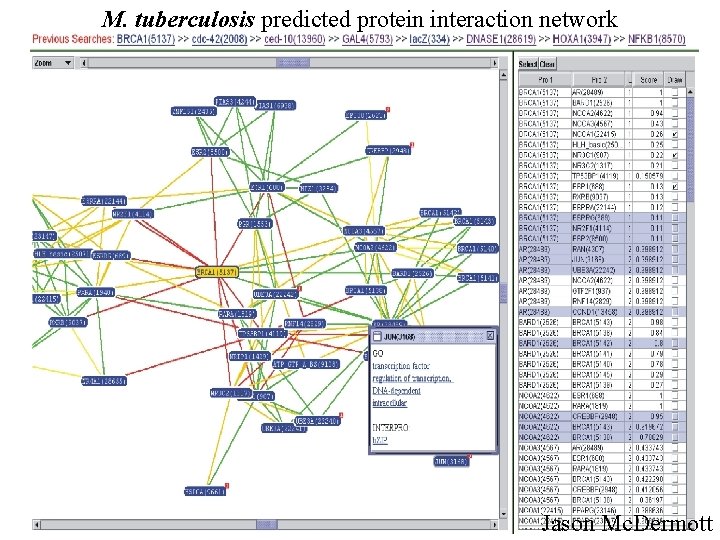
M. tuberculosis predicted protein interaction network Jason Mc. Dermott

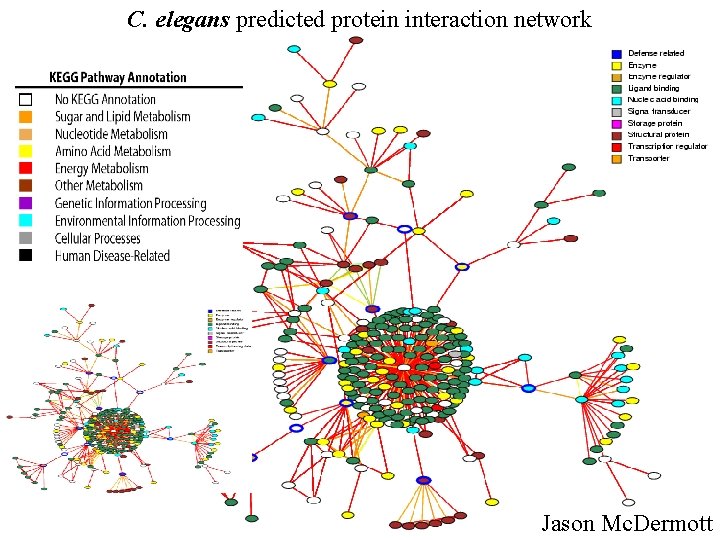
C. elegans predicted protein interaction network Jason Mc. Dermott

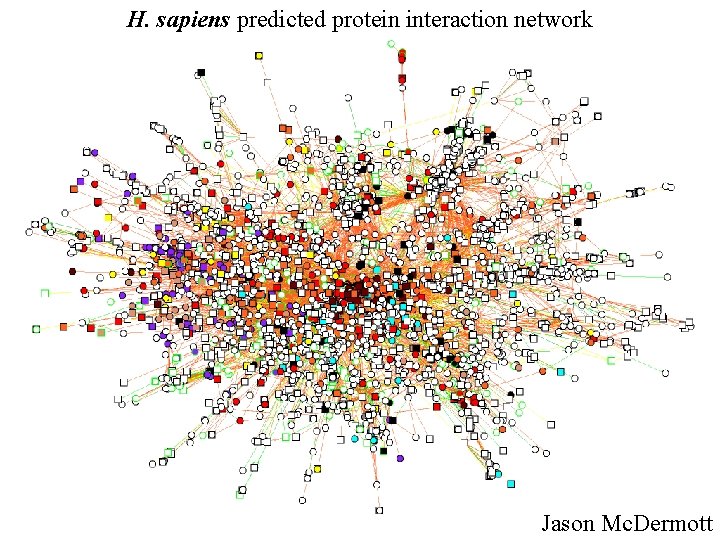
H. sapiens predicted protein interaction network Jason Mc. Dermott

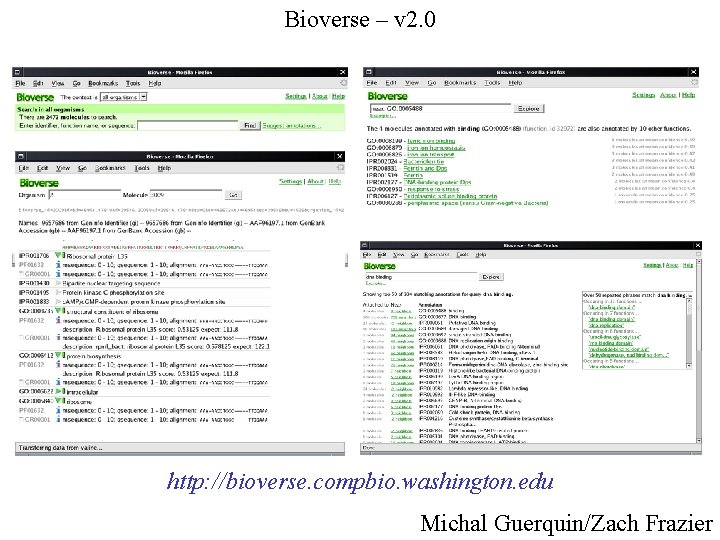
Bioverse – v 2. 0 http: //bioverse. compbio. washington. edu Michal Guerquin/Zach Frazier

Network-based annotation for C. elegans Jason Mc. Dermott

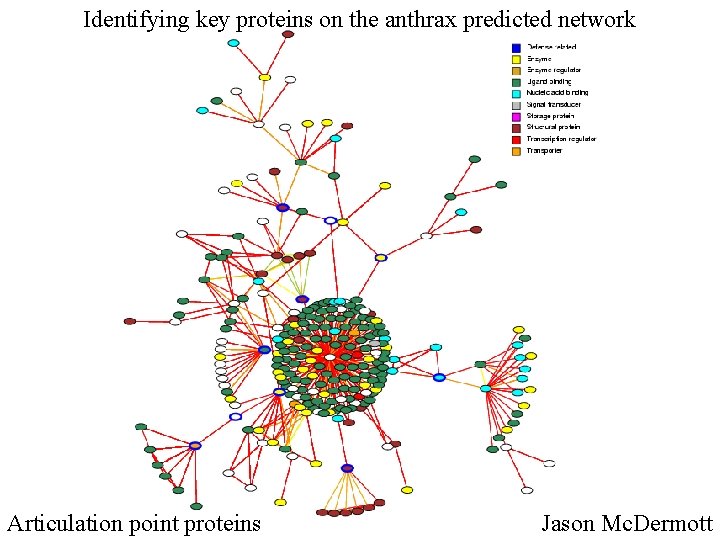
Identifying key proteins on the anthrax predicted network Articulation point proteins Jason Mc. Dermott

Identification of virulence factors Jason Mc. Dermott

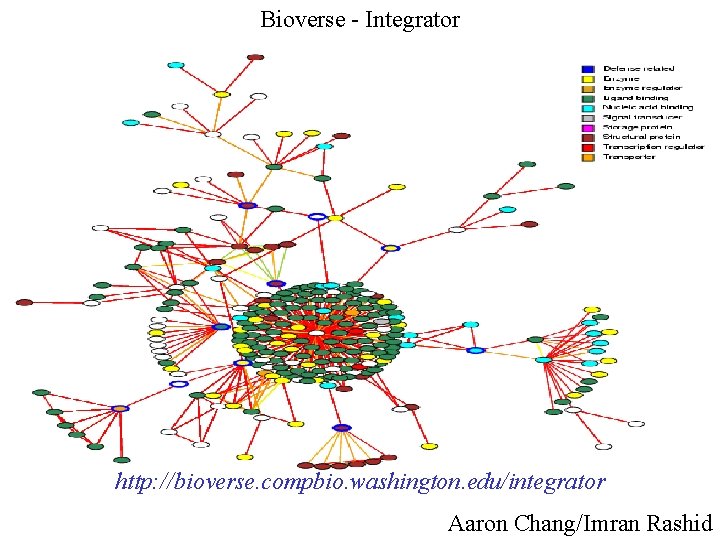
Bioverse - Integrator http: //bioverse. compbio. washington. edu/integrator Aaron Chang/Imran Rashid

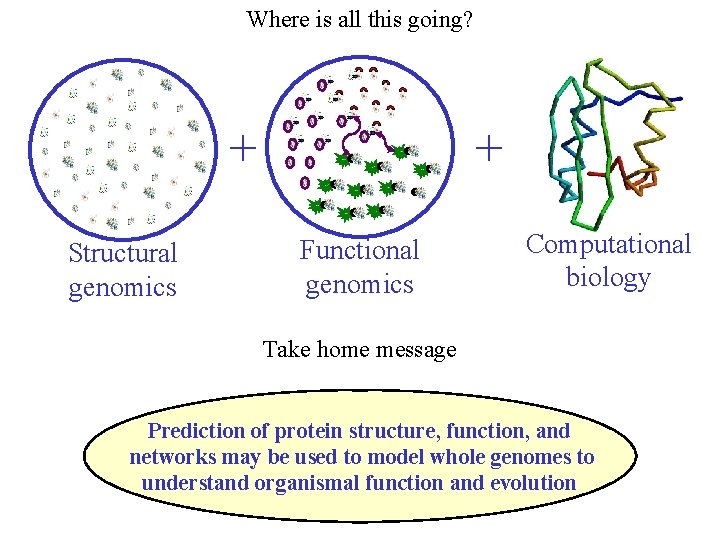
Where is all this going? + Structural genomics + Functional genomics Computational biology Take home message Prediction of protein structure, function, and networks may be used to model whole genomes to understand organismal function and evolution

Acknowledgements Current group members: Past group members: • Andrew Nichols • Aaron Chang • Baishali Chanda • Marissa La. Madrid • Chuck Mader • Mike Inouye • David Nickle • Sarunya Suebtragoon • Duangdao Wichadakul • Duncan Milburn • Ersin Emre Oren Funding agencies: • Ekachai Jenwitheesuk • National Institutes of Health • Gong Cheng • National Science Foundation • Jason Mc. Dermott • Searle Scholars Program • Jeremy Horst • Puget Sound Partners in Global Health • Kai Wang • UW Advanced Technology Initiative • Ling-Hong Hung • Michal Guerquin • Shing-Chung Ngan • Somsak Phattarasukol http: //protinfo. compbio. washington. edu • Stewart Moughon http: //bioverse. compbio. washington. edu • Tianyun Liu • Weerayuth Kittichotirat • Zach Frazier • Kristina Montgomery, Program Manager