Modelling Systems Biology Patterns with MATLAB and Sim

Modelling Systems Biology Patterns with MATLAB and Sim. Biology Eckhard Lehmann Application Engineer, The Math. Works © 2007 The Math. Works Gmb. H 12. 10. 2008 CMSB Rostock

The Math. Works at a Glance Headquarters: Natick, Massachusetts USA: California, Michigan, Washington DC, Texas Europe: Asia-Pacific: Korea, Australia, China © 2007 The Math. Works Gmb. H UK, France, Germany, Switzerland, Italy, Spain, The Netherlands, Sweden Earth’s topography on an equidistant cylindrical projection, created with the MATLAB Mapping Toolbox Distributors in 25 countries 2

Overview: MATLAB and Sim. Biology for Systems Biology MATLAB for Systems Biology Modelling – Building and solving Model ODE’s – Using the Optimization Toolbox for Parameter Estimation Parallel Computing Modelling biological systems with Sim. Biology – Building Systems Biology models with Sim. Biology – Model simulation and analysis © 2007 The Math. Works Gmb. H Agenda Wrapup, Questions and Answers 3
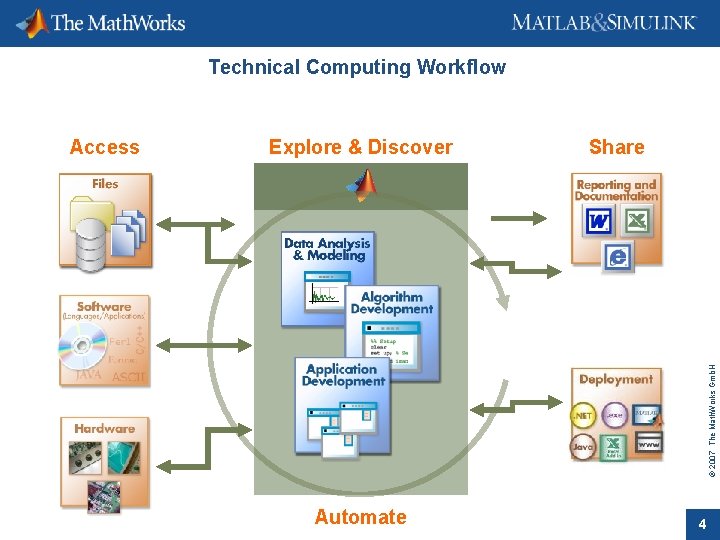
Technical Computing Workflow Explore & Discover Share © 2007 The Math. Works Gmb. H Access Automate 4
Technical computing in Life Sciences Statistics PK/PD Modelling Proteomics HTS/HCS methods Image processing Bioinformatics Systems Biology Fermentation modelling Excel plug-ins © 2007 The Math. Works Gmb. H Creating reports Executables Parallel Computing Database connectivity And more… 5

Key takeaways from this tutorial MATLAB A sophisticated environment for mathemathical modelling and data analysis – Big set of functions and algorithms – Easily extensible and programmable – Sim. Biology An extension for Systems Biology modelling – Graphic modelling capabilities, events, rules and stochastic solvers – Tight integration with MATLAB – © 2007 The Math. Works Gmb. H 6

Overview: MATLAB and Sim. Biology for Systems Biology MATLAB for Systems Biology Modelling – Building and solving Model ODE’s – Using the Optimization Toolbox for Parameter Estimation Parallel Computing Modelling biological systems with Sim. Biology – Building Systems Biology models with Sim. Biology – Model simulation and analysis © 2007 The Math. Works Gmb. H Agenda Wrapup, Questions and Answers 7
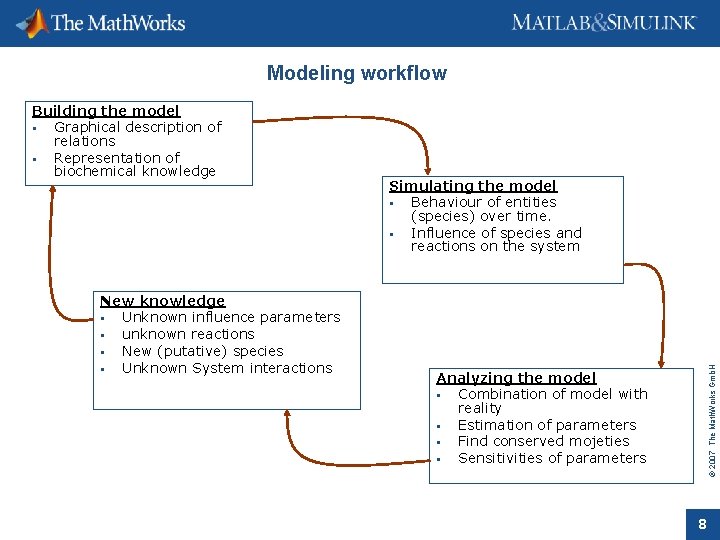
Modeling workflow New knowledge Unknown influence parameters unknown reactions New (putative) species Unknown System interactions Simulating the model Behaviour of entities (species) over time. Influence of species and reactions on the system © 2007 The Math. Works Gmb. H Building the model Graphical description of relations Representation of biochemical knowledge Analyzing the model Combination of model with reality Estimation of parameters Find conserved mojeties Sensitivities of parameters 8
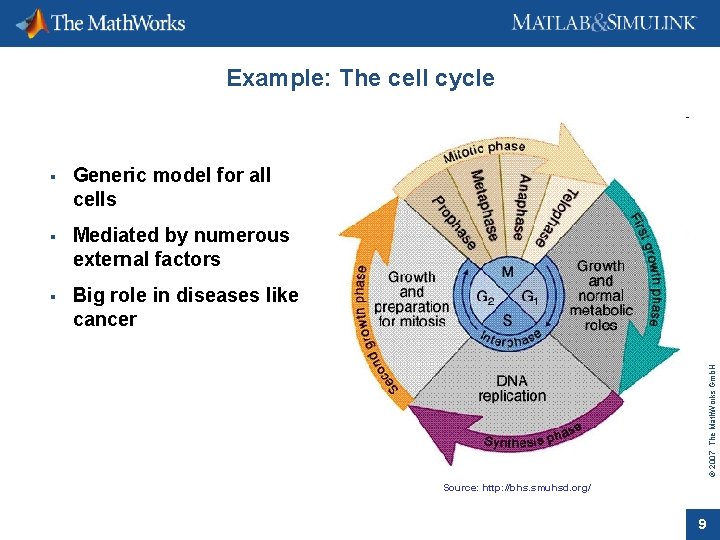
Generic model for all cells Mediated by numerous external factors Big role in diseases like cancer © 2007 The Math. Works Gmb. H Example: The cell cycle Source: http: //bhs. smuhsd. org/ 9
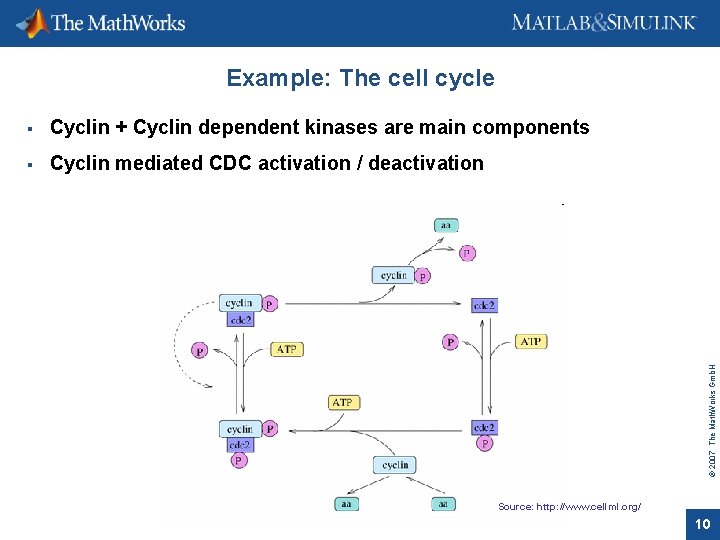
Example: The cell cycle Cyclin + Cyclin dependent kinases are main components Cyclin mediated CDC activation / deactivation © 2007 The Math. Works Gmb. H Source: http: //www. cellml. org/ 10
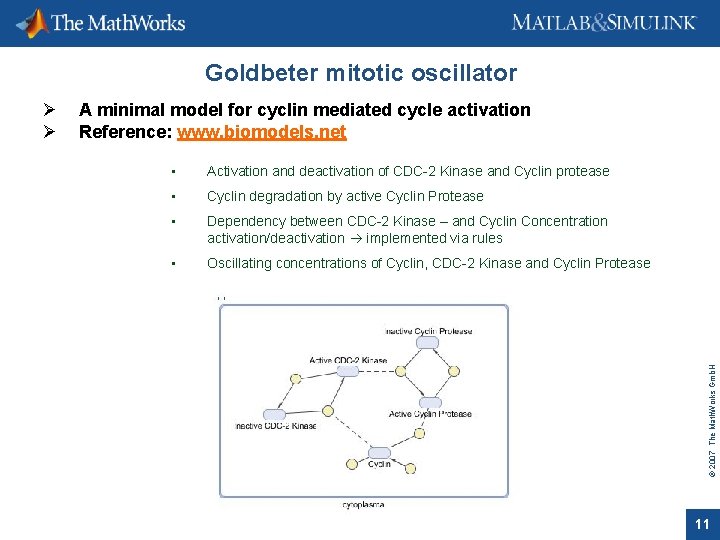
Goldbeter mitotic oscillator A minimal model for cyclin mediated cycle activation Reference: www. biomodels. net • Activation and deactivation of CDC-2 Kinase and Cyclin protease • Cyclin degradation by active Cyclin Protease • Dependency between CDC-2 Kinase – and Cyclin Concentration activation/deactivation implemented via rules • Oscillating concentrations of Cyclin, CDC-2 Kinase and Cyclin Protease © 2007 The Math. Works Gmb. H Ø Ø 11
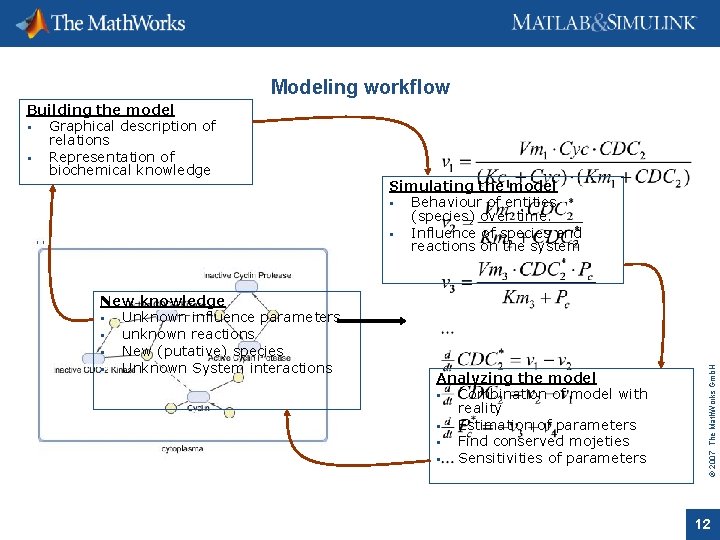
Modeling workflow New knowledge Unknown influence parameters unknown reactions New (putative) species Unknown System interactions Simulating the model Behaviour of entities (species) over time. Influence of species and reactions on the system Analyzing the model Combination of model with reality Estimation of parameters Find conserved mojeties Sensitivities of parameters © 2007 The Math. Works Gmb. H Building the model Graphical description of relations Representation of biochemical knowledge 12
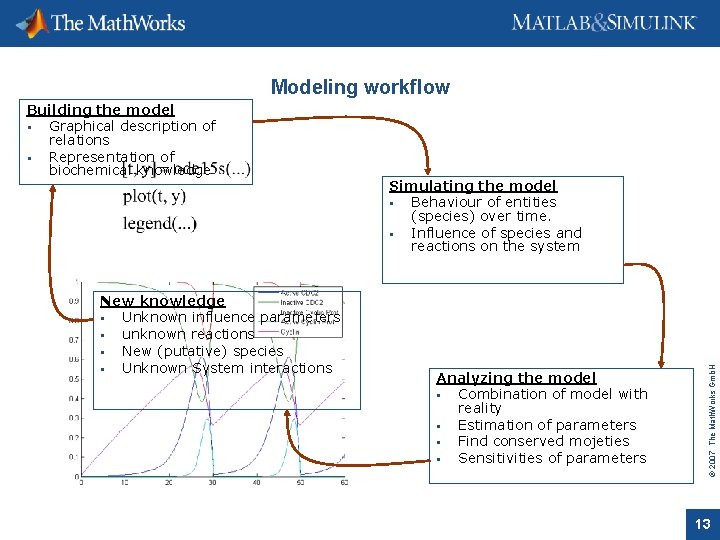
Modeling workflow New knowledge Unknown influence parameters unknown reactions New (putative) species Unknown System interactions Simulating the model Behaviour of entities (species) over time. Influence of species and reactions on the system Analyzing the model Combination of model with reality Estimation of parameters Find conserved mojeties Sensitivities of parameters © 2007 The Math. Works Gmb. H Building the model Graphical description of relations Representation of biochemical knowledge 13

High-level technical computing language Development environment Platform independent Feature areas – – Mathematics Graphics and GUI Design File I/O Call C/C++, Fortran, Java, COM Open and extensible © 2007 The Math. Works Gmb. H The MATLAB Environment 14
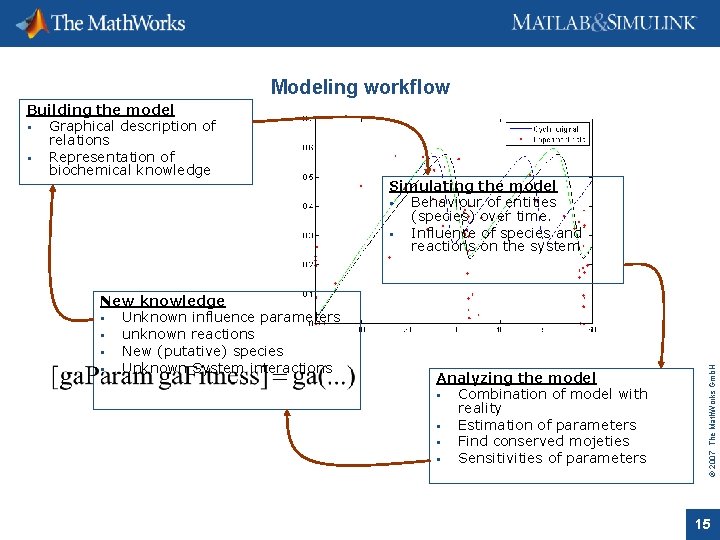
Modeling workflow New knowledge Unknown influence parameters unknown reactions New (putative) species Unknown System interactions Simulating the model Behaviour of entities (species) over time. Influence of species and reactions on the system Analyzing the model Combination of model with reality Estimation of parameters Find conserved mojeties Sensitivities of parameters © 2007 The Math. Works Gmb. H Building the model Graphical description of relations Representation of biochemical knowledge 15

Optimization Toolbox Graphical and command line tools and solvers for: – – – Linear and nonlinear programming Quadratic programming Nonlinear least-squares, data fitting, and nonlinear equations Multiobjective optimization Binary integer programming © 2007 The Math. Works Gmb. H 16

Overview: MATLAB and Sim. Biology for Systems Biology MATLAB for Systems Biology Modelling – Building and solving Model ODE’s – Using the Optimization Toolbox for Parameter Estimation Parallel Computing Modelling biological systems with Sim. Biology – Building Systems Biology models with Sim. Biology – Model simulation and analysis Wrapup, Questions and Answers © 2007 The Math. Works Gmb. H Agenda 17
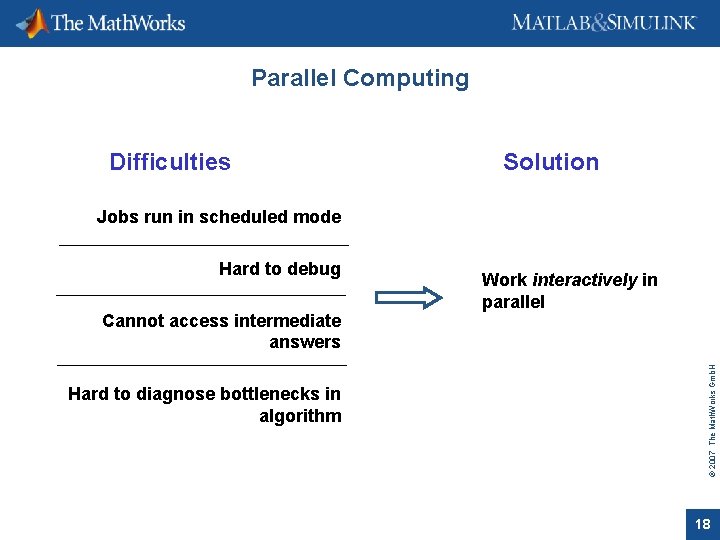
Parallel Computing Difficulties Solution Jobs run in scheduled mode Cannot access intermediate answers Hard to diagnose bottlenecks in algorithm Work interactively in parallel © 2007 The Math. Works Gmb. H Hard to debug 18
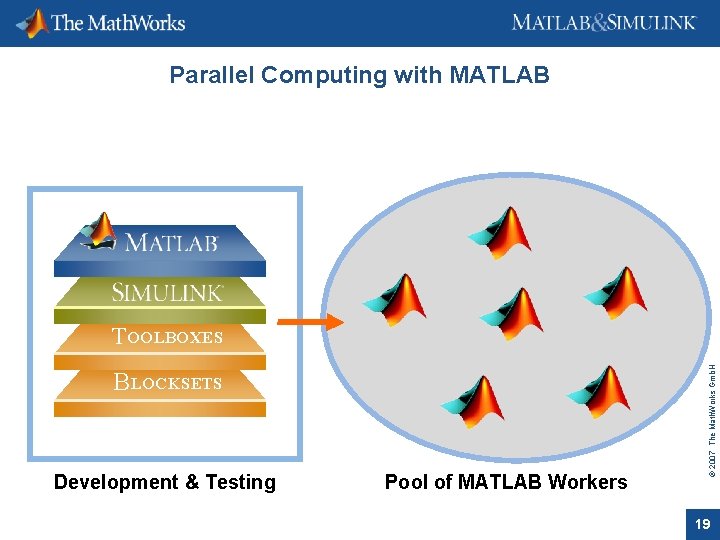
Parallel Computing with MATLAB BLOCKSETS Development & Testing Pool of MATLAB Workers © 2007 The Math. Works Gmb. H TOOLBOXES 19

Easier Parallel Programming Example: Transposing a Distributed Matrix Using MATLAB and MPI P>> D = distributed(A) P>> E = D’ © 2007 The Math. Works Gmb. H Using Fortran and MPI Using Distributed Arrays 20

Parallel for-Loops Mix task-parallel and serial code in the same function Run loops on a pool of MATLAB resources Iterations must be order-independent M-Lint analysis helps in converting existing for-loops into to parfor-loops © 2007 The Math. Works Gmb. H parfor i = 1 : n % do something with i end 21
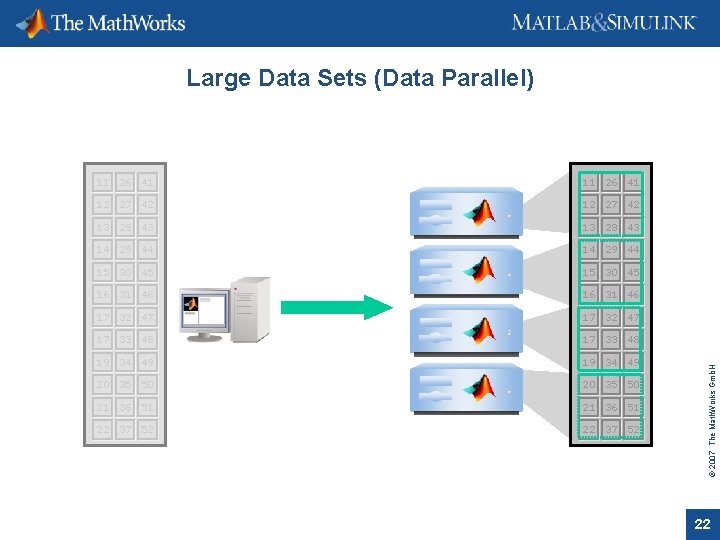
11 26 41 12 27 42 13 28 43 14 29 44 15 30 45 16 31 46 17 32 47 17 33 48 19 34 49 20 35 50 21 36 51 22 37 52 © 2007 The Math. Works Gmb. H Large Data Sets (Data Parallel) 22

Demo: Monte Carlo Simulation of Coin Tossing 10 Simulations of Flipping 20 Coins at a Time 7 12 15 7 9 9 Number of Heads Out of 20 7 8 12 © 2007 The Math. Works Gmb. H 11 23

Demo: Monte Carlo Simulation of Coin Tossing 10 Simulations of Flipping 20 Coins at a Time 11 11 7 12 15 9 9 13 7 8 12 Change in Number of Heads -- ↓ ↑ ↑ ↓ → ↑ ↓ ↑ ↑ © 2007 The Math. Works Gmb. H Number of Heads Out of 20 25
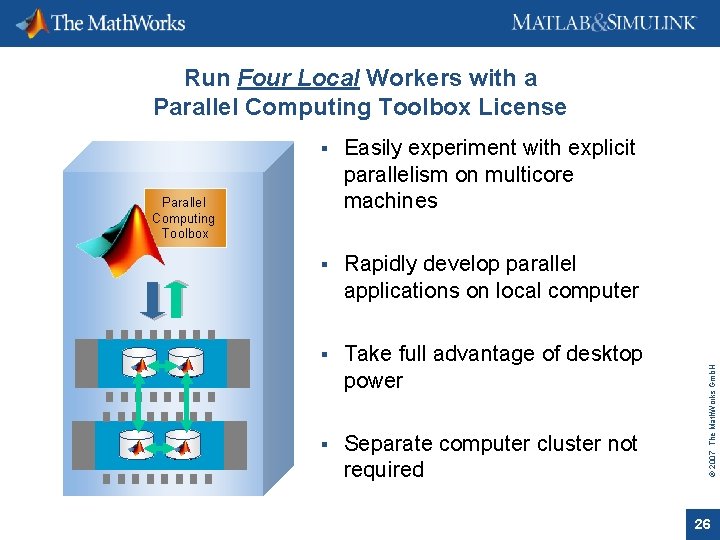
Easily experiment with explicit parallelism on multicore machines Rapidly develop parallel applications on local computer Take full advantage of desktop power Separate computer cluster not required Parallel Computing Toolbox © 2007 The Math. Works Gmb. H Run Four Local Workers with a Parallel Computing Toolbox License 26
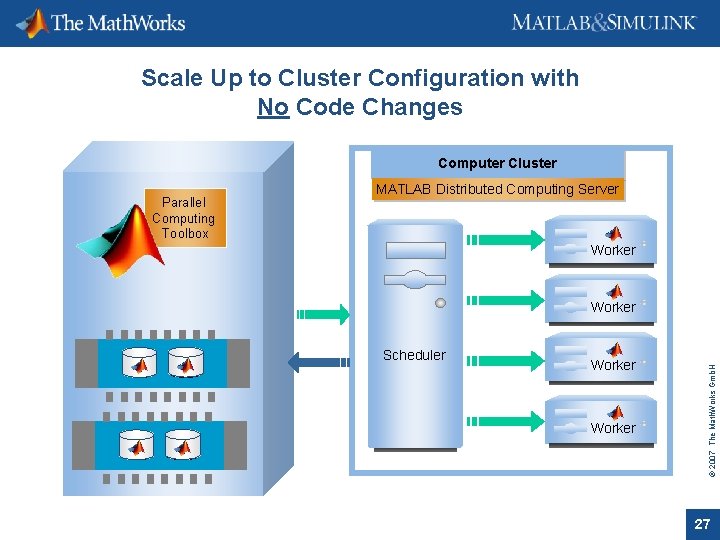
Scale Up to Cluster Configuration with No Code Changes Computer Cluster MATLAB Distributed Computing Server CPU Worker Scheduler CPU Worker © 2007 The Math. Works Gmb. H Parallel Computing Toolbox 27
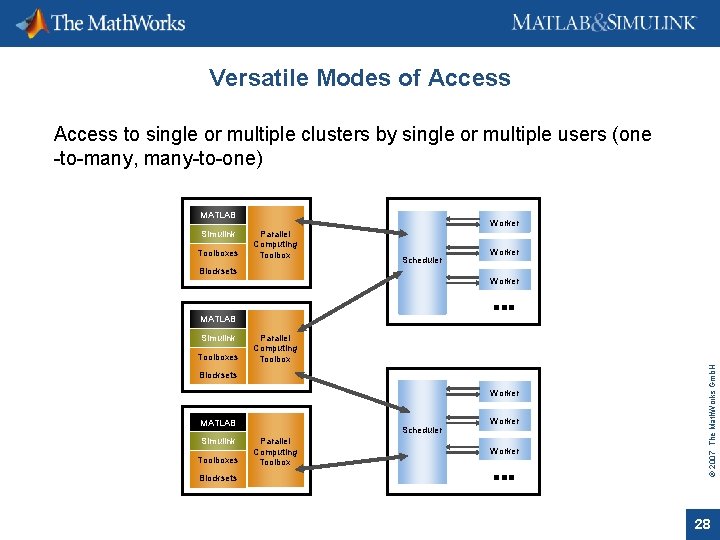
Versatile Modes of Access to single or multiple clusters by single or multiple users (one -to-many, many-to-one) MATLAB Simulink Toolboxes Worker Parallel Computing Toolbox Scheduler Blocksets Worker MATLAB Toolboxes Parallel Computing Toolbox Blocksets Worker MATLAB Simulink Toolboxes Blocksets Scheduler Parallel Computing Toolbox Worker © 2007 The Math. Works Gmb. H Simulink 28

Overview: MATLAB and Sim. Biology for Systems Biology MATLAB for Systems Biology Modelling – Building and solving Model ODE’s – Using the Optimization Toolbox for Parameter Estimation Parallel Computing Modelling biological systems with Sim. Biology – Building Systems Biology models with Sim. Biology – Model simulation and analysis Wrapup, Questions and Answers © 2007 The Math. Works Gmb. H Agenda 29

Systems biology with MATLAB: Sim. Biology™ Extend the MATLAB® platform for systems biologists Gain insight into dynamic behavior of pathways and biological systems Communicate and exchange dynamic models © 2007 The Math. Works Gmb. H A platform and environment for modelling, simulating and analyzing pathways and other biological systems 30
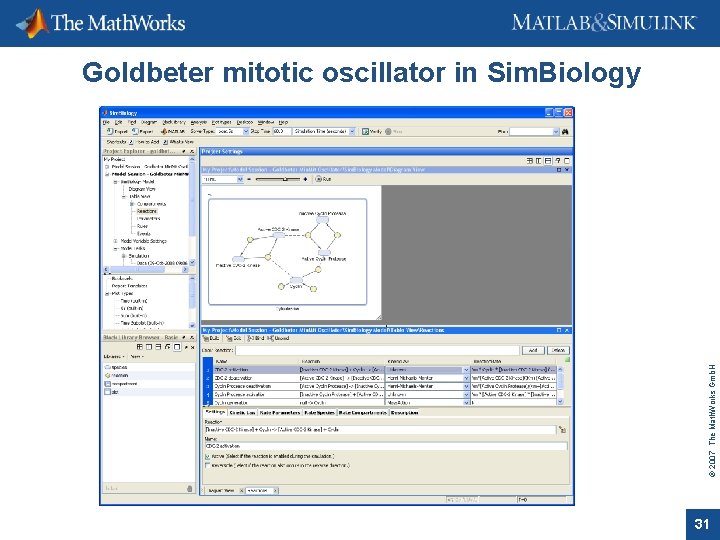
© 2007 The Math. Works Gmb. H Goldbeter mitotic oscillator in Sim. Biology 31
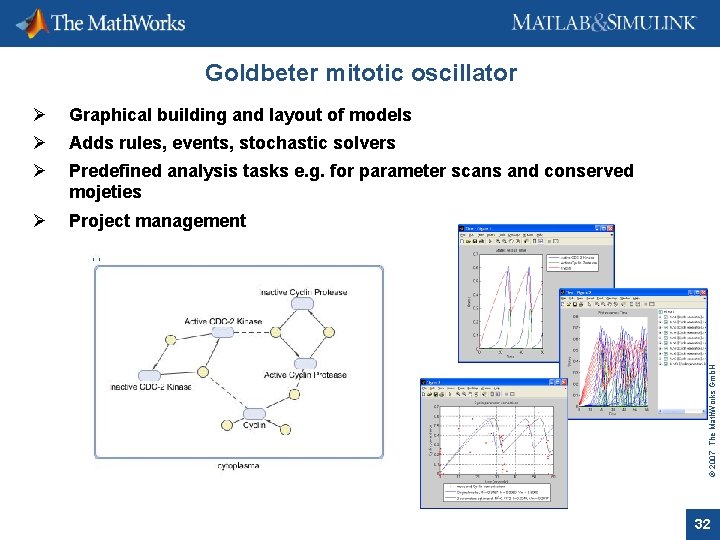
Ø Graphical building and layout of models Ø Adds rules, events, stochastic solvers Ø Predefined analysis tasks e. g. for parameter scans and conserved mojeties Ø Project management © 2007 The Math. Works Gmb. H Goldbeter mitotic oscillator 32

What else can you do? Ø SBML v 2. 1 import and export Ø Use predefined reaction rates (Michaelis Menten, Ping. Pong etc) and define your own kinetics equations Ø Define events and rules with callbacks Ø Automatically layout model components Ø Programatically create models Ø Speed up model analysis with distributed computing © 2007 The Math. Works Gmb. H Sim. Biology is a powerful environment for systems biologists 33

Overview: MATLAB and Sim. Biology for Systems Biology MATLAB for Systems Biology Modelling – Building and solving Model ODE’s – Using the Optimization Toolbox for Parameter Estimation Parallel Computing Modelling biological systems with Sim. Biology – Building Systems Biology models with Sim. Biology – Model simulation and analysis Wrapup, Questions and Answers © 2007 The Math. Works Gmb. H Agenda 34
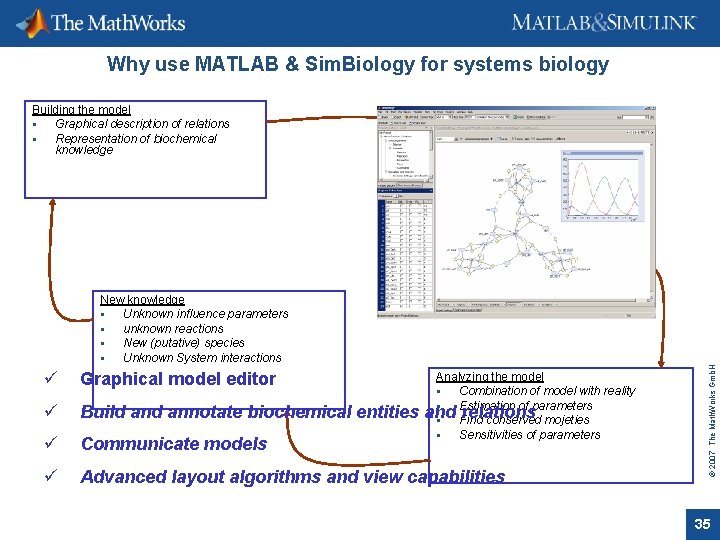
Why use MATLAB & Sim. Biology for systems biology Building the model Graphical description of relations Representation of biochemical knowledge Simulating the model Behaviour of entities (species) over time. Influence of species and reactions on the system ü Graphical model editor ü Build annotate biochemical entities ü Communicate models ü Advanced layout algorithms and view capabilities Analyzing the model Combination of model with reality Estimation of parameters and relations Find conserved mojeties Sensitivities of parameters © 2007 The Math. Works Gmb. H New knowledge Unknown influence parameters unknown reactions New (putative) species Unknown System interactions 35
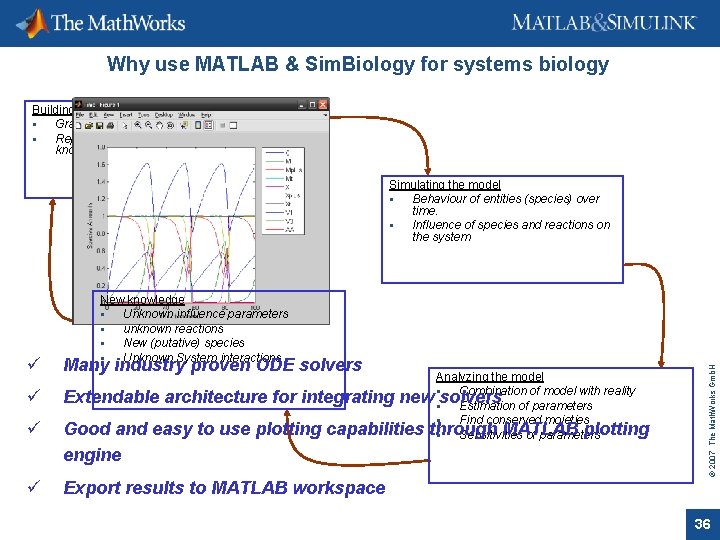
Why use MATLAB & Sim. Biology for systems biology Building the model Graphical description of relations Representation of biochemical knowledge Simulating the model Behaviour of entities (species) over time. Influence of species and reactions on the system ü ü Many industry proven ODE solvers Analyzing the model Combination of model with reality Extendable architecture for integrating new solvers Estimation of parameters Find conserved mojeties Good and easy to use plotting capabilities through MATLAB plotting Sensitivities of parameters engine ü © 2007 The Math. Works Gmb. H ü New knowledge Unknown influence parameters unknown reactions New (putative) species Unknown System interactions Export results to MATLAB workspace 36

Why use MATLAB & Sim. Biology for systems biology ü the Advanced optimization techniques Building model Graphical description of relations üRepresentation of biochemical Standard analysis techniques built knowledge out of the box into the Sim. Biology Desktop ü Work with simulation results in MATLAB ü Easy to extend and customize Simulating the model Behaviour of entities (species) over analysis capabilities time. Influence of species and reactions on the system Analyzing the model Combination of model with reality Estimation of parameters Find conserved mojeties Sensitivities of parameters © 2007 The Math. Works Gmb. H New knowledge Unknown influence parameters unknown reactions New (putative) species Unknown System interactions 37

The Math. Works Web Resources Training Calendar Solution Database Documentation Books about MATLAB File Exchange Newsletters Newsgroup www. mathworks. de Events © 2007 The Math. Works Gmb. H Information & Downloads 38

39 © 2007 The Math. Works Gmb. H
- Slides: 38