Modelling Proteomes Ram Samudrala University of Washington Ab
Modelling Proteomes Ram Samudrala University of Washington
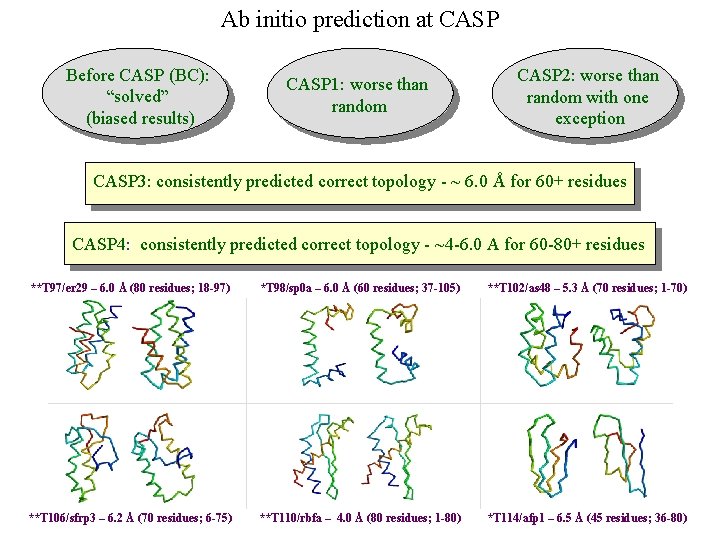
Ab initio prediction at CASP Before CASP (BC): “solved” (biased results) CASP 1: worse than random CASP 2: worse than random with one exception CASP 3: consistently predicted correct topology - ~ 6. 0 Å for 60+ residues CASP 4: consistently predicted correct topology - ~4 -6. 0 A for 60 -80+ residues **T 97/er 29 – 6. 0 Å (80 residues; 18 -97) *T 98/sp 0 a – 6. 0 Å (60 residues; 37 -105) **T 102/as 48 – 5. 3 Å (70 residues; 1 -70) **T 106/sfrp 3 – 6. 2 Å (70 residues; 6 -75) **T 110/rbfa – 4. 0 Å (80 residues; 1 -80) *T 114/afp 1 – 6. 5 Å (45 residues; 36 -80)
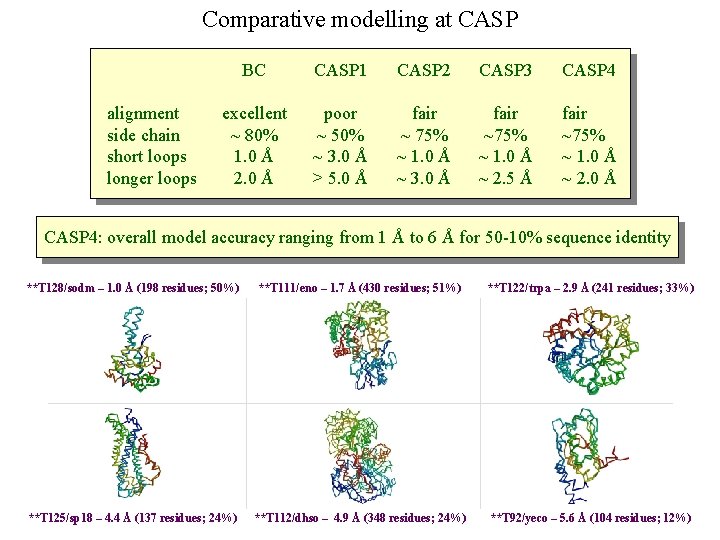
Comparative modelling at CASP alignment side chain short loops longer loops BC CASP 1 CASP 2 CASP 3 CASP 4 excellent ~ 80% 1. 0 Å 2. 0 Å poor ~ 50% ~ 3. 0 Å > 5. 0 Å fair ~ 75% ~ 1. 0 Å ~ 3. 0 Å fair ~75% ~ 1. 0 Å ~ 2. 5 Å fair ~75% ~ 1. 0 Å ~ 2. 0 Å CASP 4: overall model accuracy ranging from 1 Å to 6 Å for 50 -10% sequence identity **T 128/sodm – 1. 0 Å (198 residues; 50%) **T 111/eno – 1. 7 Å (430 residues; 51%) **T 122/trpa – 2. 9 Å (241 residues; 33%) **T 125/sp 18 – 4. 4 Å (137 residues; 24%) **T 112/dhso – 4. 9 Å (348 residues; 24%) **T 92/yeco – 5. 6 Å (104 residues; 12%)
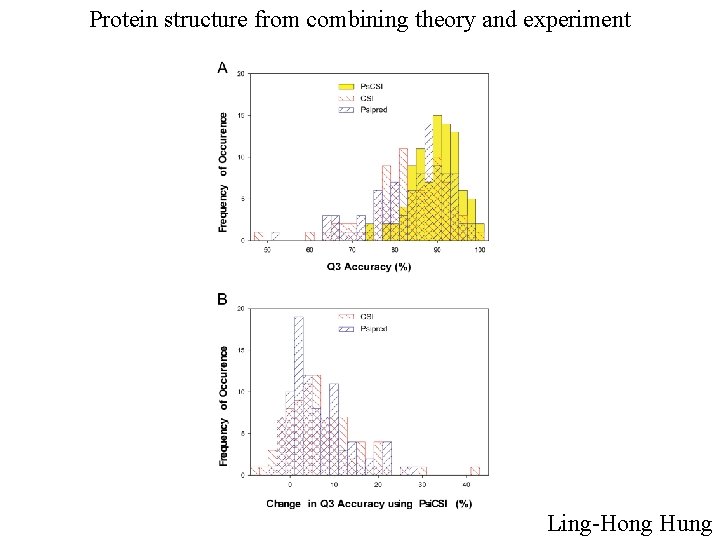
Protein structure from combining theory and experiment Ling-Hong Hung
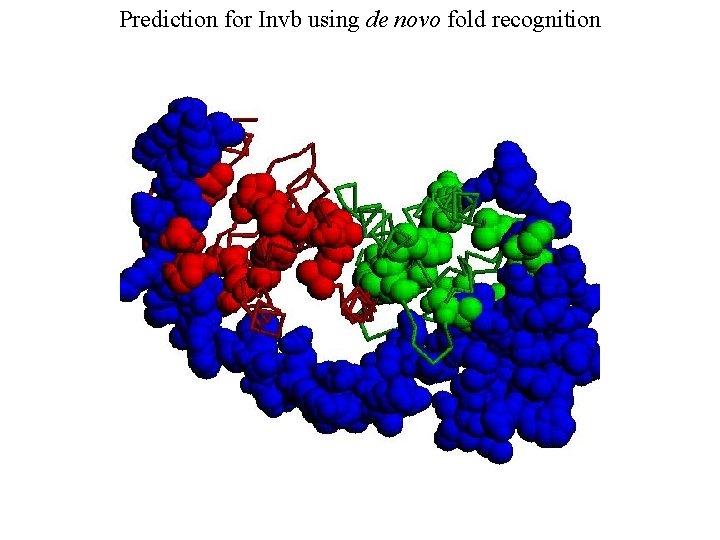
Prediction for Invb using de novo fold recognition
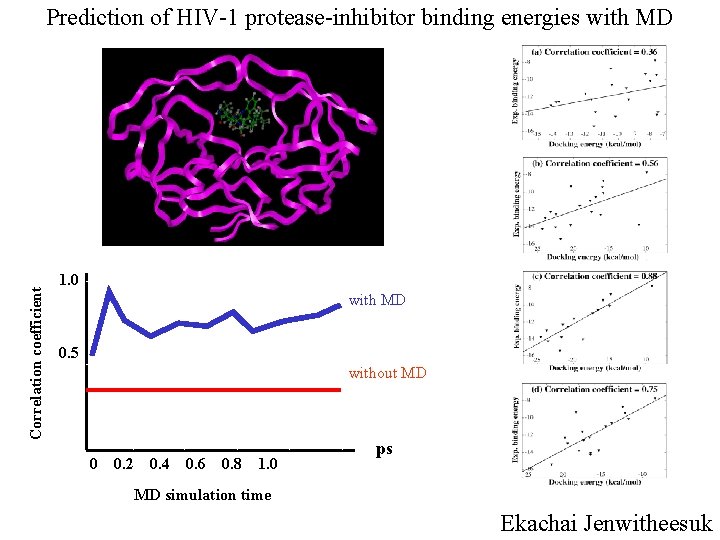
Correlation coefficient Prediction of HIV-1 protease-inhibitor binding energies with MD 1. 0 with MD 0. 5 without MD 0 0. 2 0. 4 0. 6 0. 8 1. 0 ps MD simulation time Ekachai Jenwitheesuk
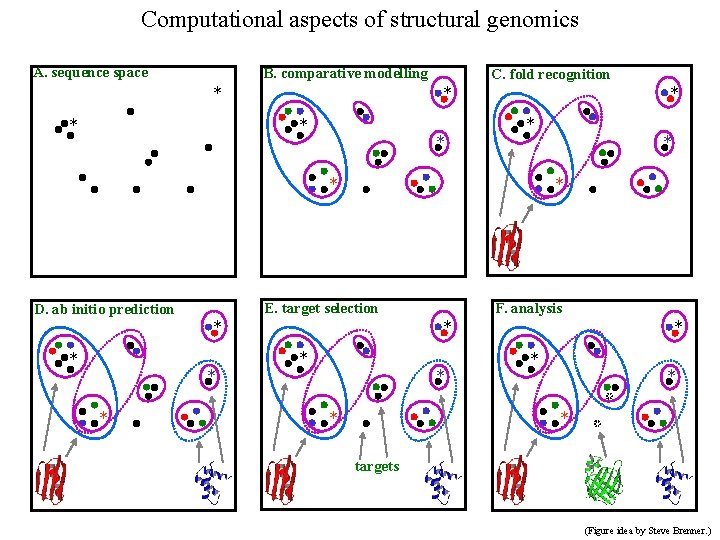
Computational aspects of structural genomics A. sequence space B. comparative modelling * * C. fold recognition * * * * E. target selection D. ab initio prediction * * F. analysis * * * * targets (Figure idea by Steve Brenner. )
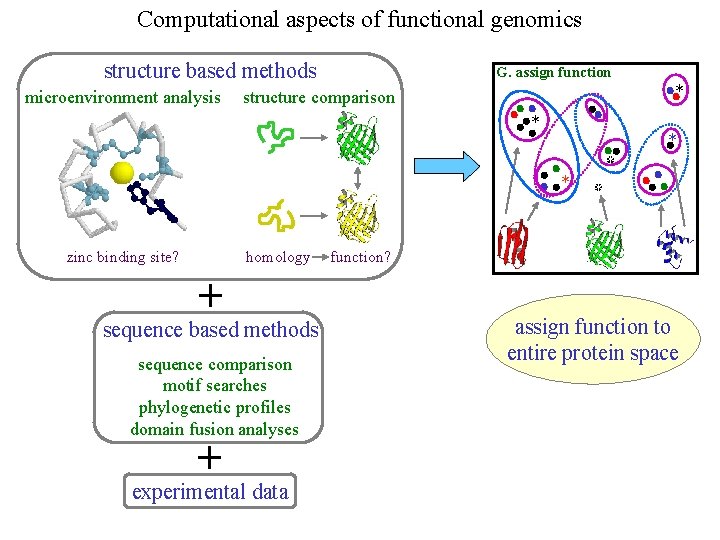
Computational aspects of functional genomics structure based methods microenvironment analysis G. assign function * structure comparison * * * zinc binding site? homology + sequence based methods sequence comparison motif searches phylogenetic profiles domain fusion analyses + experimental data * * function? assign function to entire protein space

Bioverse – explore relationships among molecules and systems http: //bioverse. compbio. washington. edu Jason Mcdermott

Bioverse – explore relationships among molecules and systems Jason Mcdermott
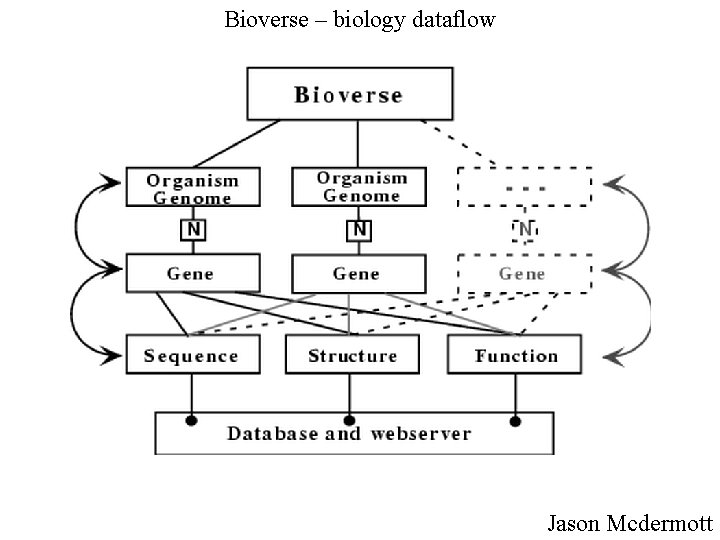
Bioverse – biology dataflow Jason Mcdermott
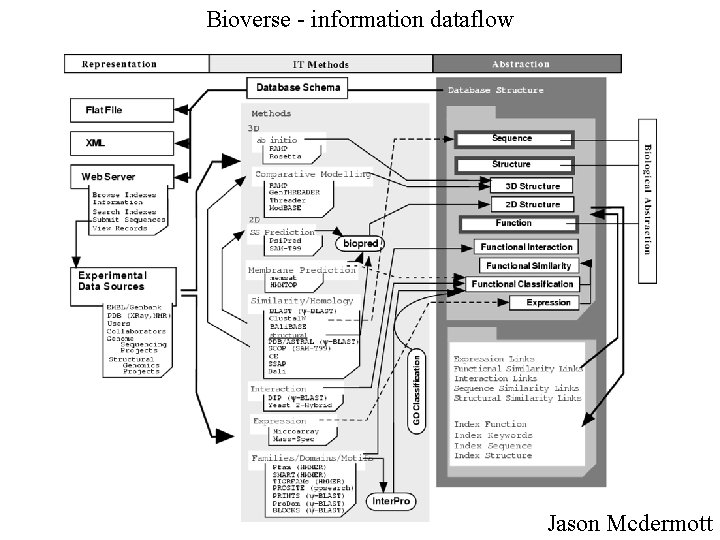
Bioverse - information dataflow Jason Mcdermott

Bioverse – human protein-protein interaction network Jason Mcdermott/Zach Frazier
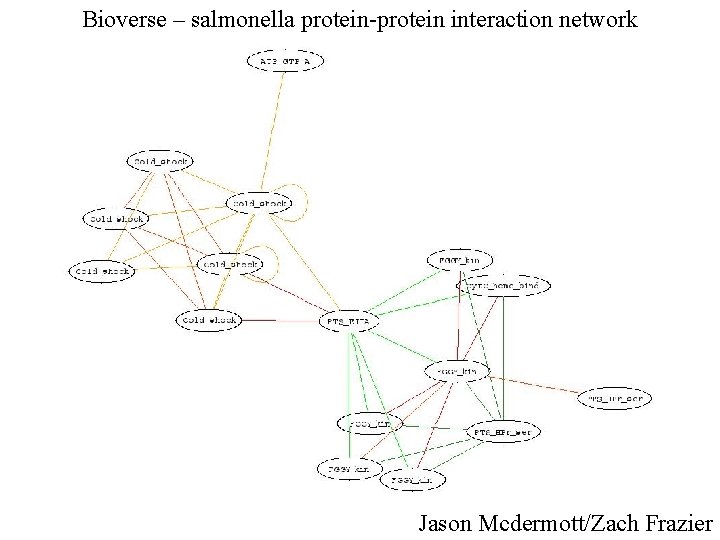
Bioverse – salmonella protein-protein interaction network Jason Mcdermott/Zach Frazier
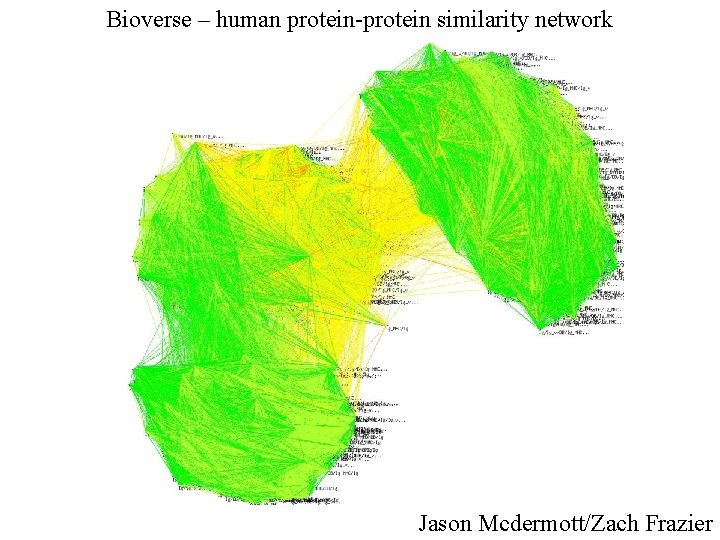
Bioverse – human protein-protein similarity network Jason Mcdermott/Zach Frazier
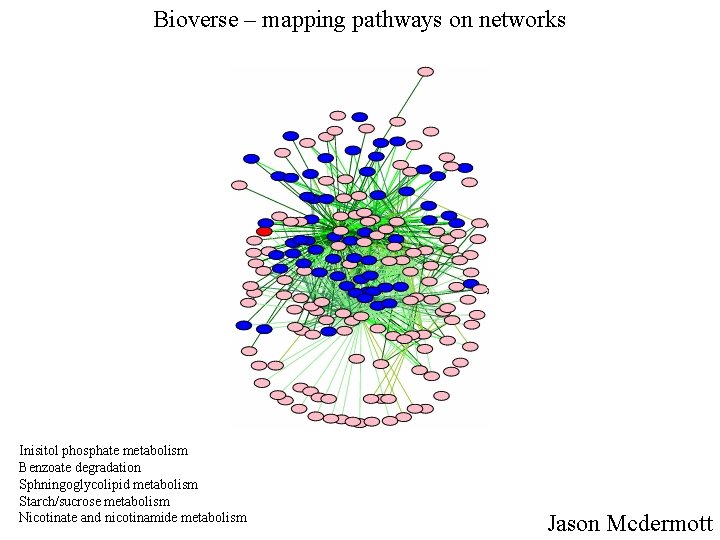
Bioverse – mapping pathways on networks Inisitol phosphate metabolism Benzoate degradation Sphningoglycolipid metabolism Starch/sucrose metabolism Nicotinate and nicotinamide metabolism Jason Mcdermott
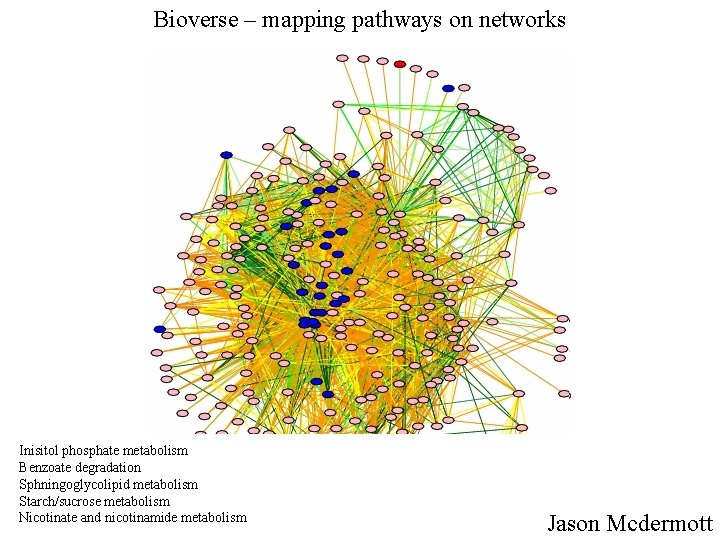
Bioverse – mapping pathways on networks Inisitol phosphate metabolism Benzoate degradation Sphningoglycolipid metabolism Starch/sucrose metabolism Nicotinate and nicotinamide metabolism Jason Mcdermott
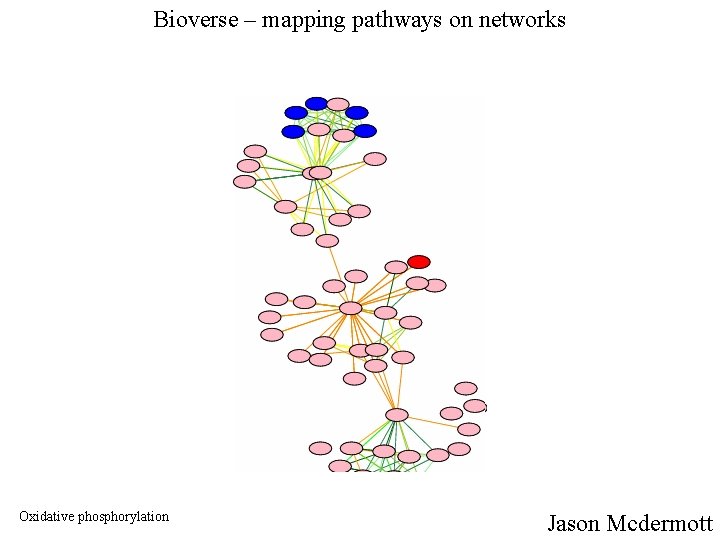
Bioverse – mapping pathways on networks Oxidative phosphorylation Jason Mcdermott
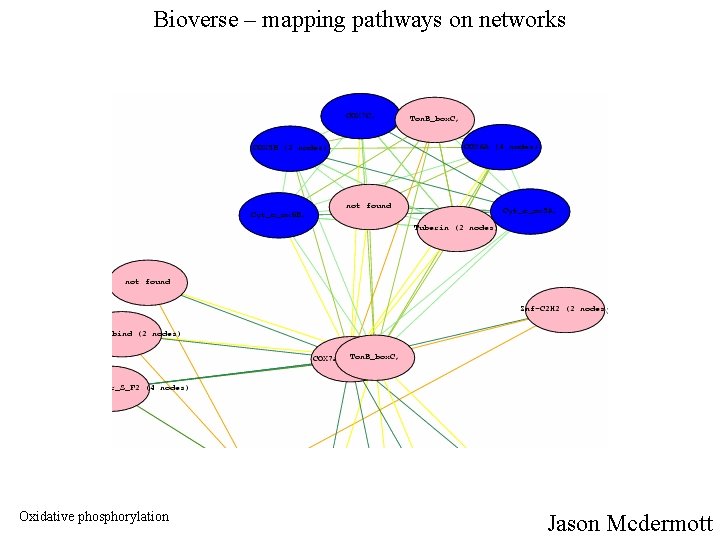
Bioverse – mapping pathways on networks Oxidative phosphorylation Jason Mcdermott

Bioverse – viewer Galen Wilkerson
Take home message Prediction of protein structure and function can be used to model whole genomes to understand organismal function and evolution Acknowledgements Group members
- Slides: 21