Modelling Proteomes Ram Samudrala University of Washington Overview






- Slides: 16

Modelling Proteomes Ram Samudrala University of Washington

Overview • De novo structure prediction: 3 -6 Å RMSD up to 150 residues A • Comparative modelling: 1 -4 Å RMSD for 50 -20% identity • Predict function based on structure, in conjunction with available experimental data • Predict structures and functions for all tractable proteins • Integrate resulting data with data from single molecule as well as genomic/proteomic studies: Bioverse

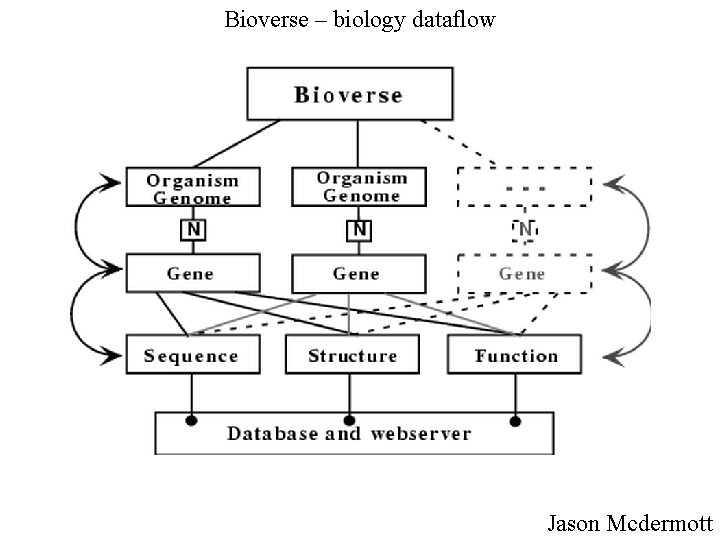
Bioverse – biology dataflow Jason Mcdermott

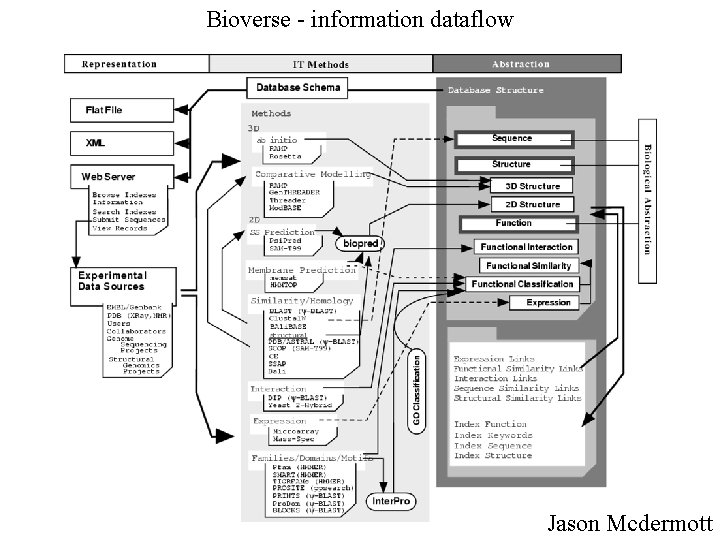
Bioverse - information dataflow Jason Mcdermott

Bioverse – explore relationships among molecules and systems http: //bioverse. compbio. washington. edu Jason Mcdermott

Bioverse – explore relationships among molecules and systems Jason Mcdermott

Bioverse – human protein-protein interaction network Jason Mcdermott/Zach Frazier

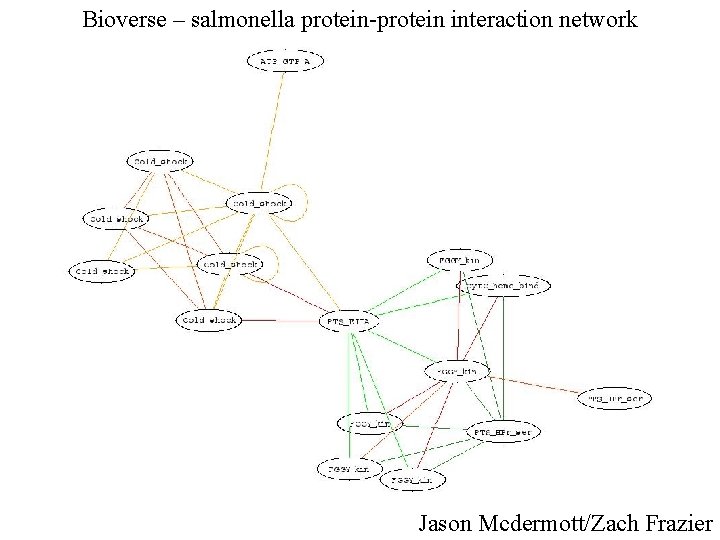
Bioverse – salmonella protein-protein interaction network Jason Mcdermott/Zach Frazier

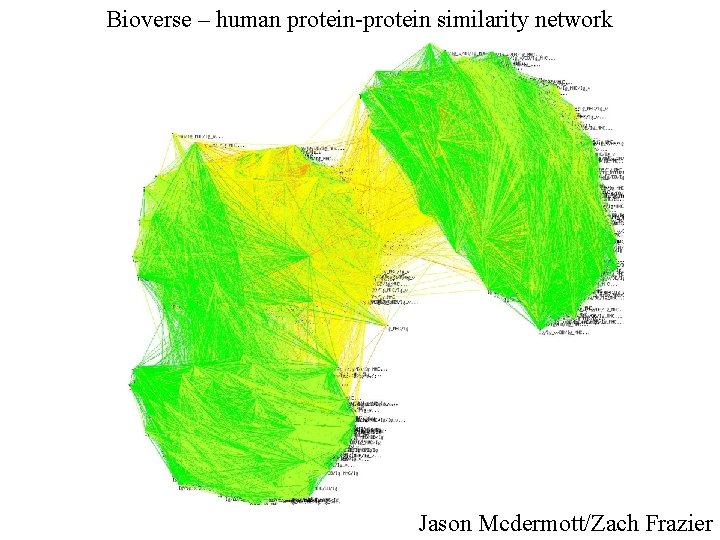
Bioverse – human protein-protein similarity network Jason Mcdermott/Zach Frazier

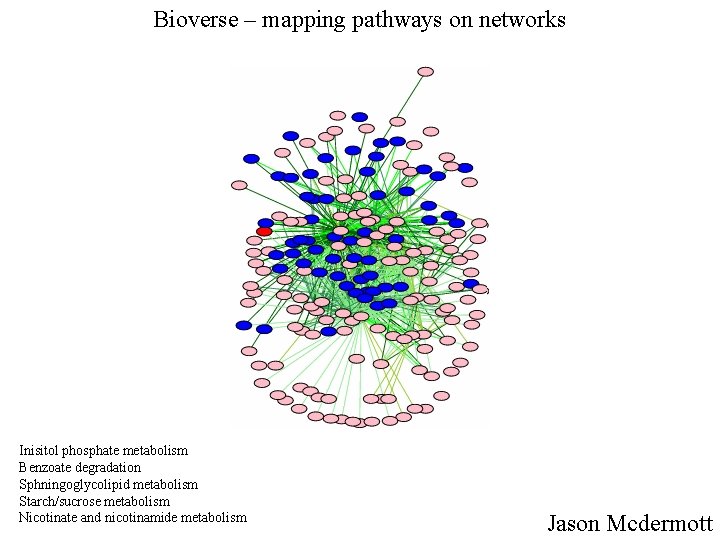
Bioverse – mapping pathways on networks Inisitol phosphate metabolism Benzoate degradation Sphningoglycolipid metabolism Starch/sucrose metabolism Nicotinate and nicotinamide metabolism Jason Mcdermott

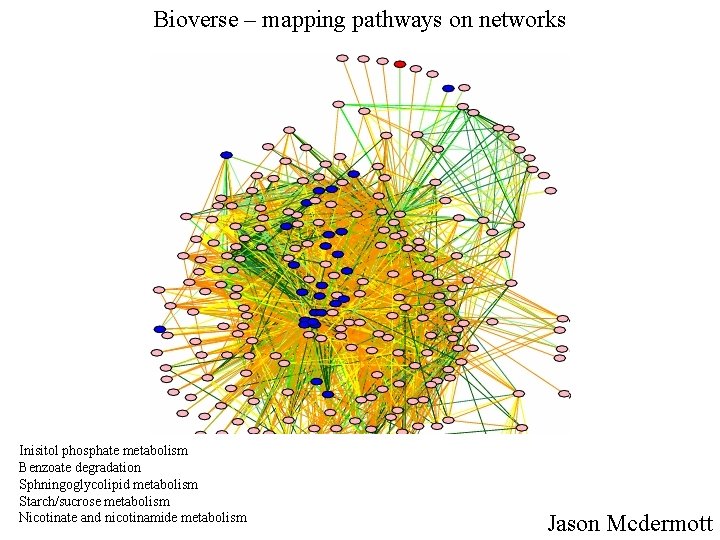
Bioverse – mapping pathways on networks Inisitol phosphate metabolism Benzoate degradation Sphningoglycolipid metabolism Starch/sucrose metabolism Nicotinate and nicotinamide metabolism Jason Mcdermott

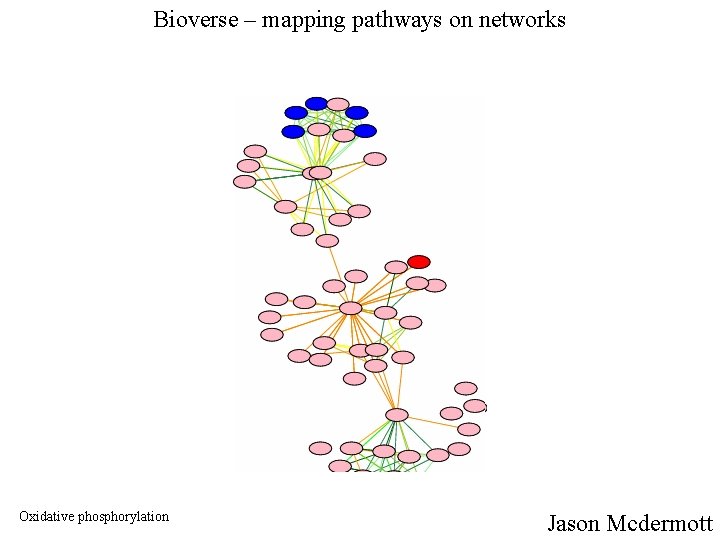
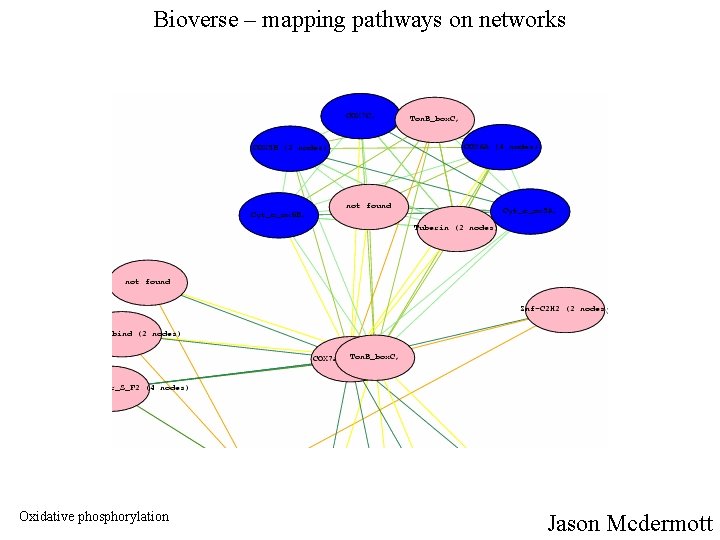
Bioverse – mapping pathways on networks Oxidative phosphorylation Jason Mcdermott

Bioverse – mapping pathways on networks Oxidative phosphorylation Jason Mcdermott

Bioverse – viewer Galen Wilkerson

Bioverse – issues • Algorithms – graph theory algorithms (connected components, cliques, articulation points, condensation) • Integration – how to normalise and weight different combinations of data (neural networks, SOMs) • Database – flexible searching of large amounts of data (OODBMS)

Take home message Prediction of protein structure and function can be used to model whole genomes to understand organismal function and evolution Acknowledgements Group members