Modelling proteomes Ram Samudrala Department of Microbiology How

Modelling proteomes Ram Samudrala Department of Microbiology How does the genome of an organism specify its behaviour and characteristics?
Proteome – all proteins of a particular system ~60, 000 in human ~60, 000 in rice ~4500 in bacteria like Salmonella and E. coli Several thousand distinct sequence families
Modelling proteomes – understand the structure of individual proteins A few thousand distinct structural folds
Modelling proteomes – understand their individual functions Thousands of possible functions
Modelling proteomes – understand their expression Different expression patterns based on time and location
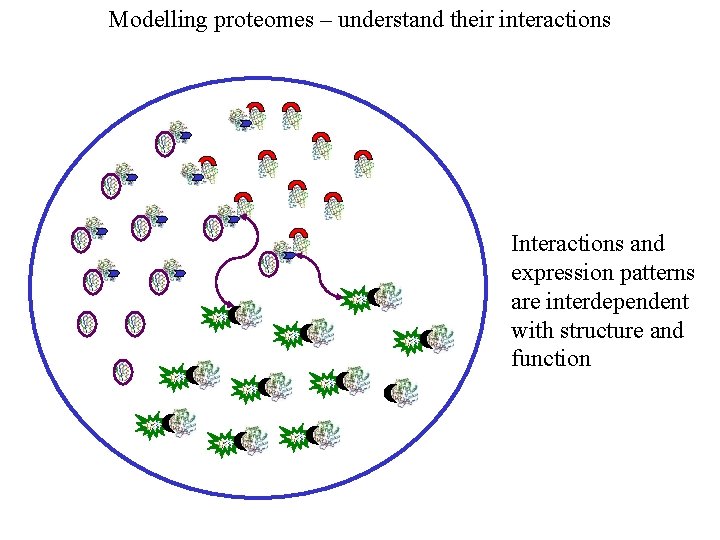
Modelling proteomes – understand their interactions Interactions and expression patterns are interdependent with structure and function
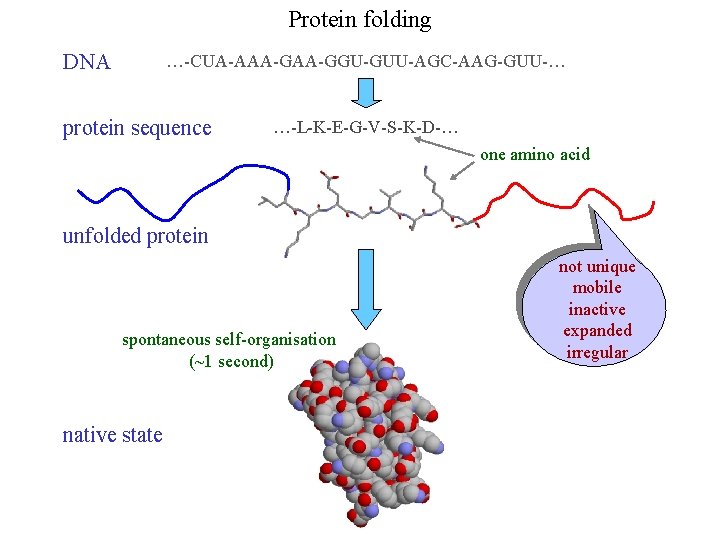
Protein folding DNA …-CUA-AAA-GGU-GUU-AGC-AAG-GUU-… protein sequence …-L-K-E-G-V-S-K-D-… one amino acid unfolded protein spontaneous self-organisation (~1 second) native state not unique mobile inactive expanded irregular
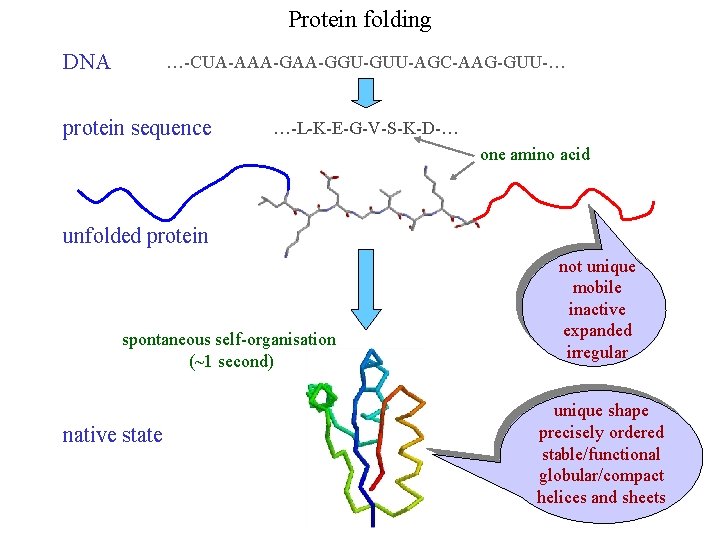
Protein folding DNA …-CUA-AAA-GGU-GUU-AGC-AAG-GUU-… protein sequence …-L-K-E-G-V-S-K-D-… one amino acid unfolded protein spontaneous self-organisation (~1 second) native state not unique mobile inactive expanded irregular unique shape precisely ordered stable/functional globular/compact helices and sheets
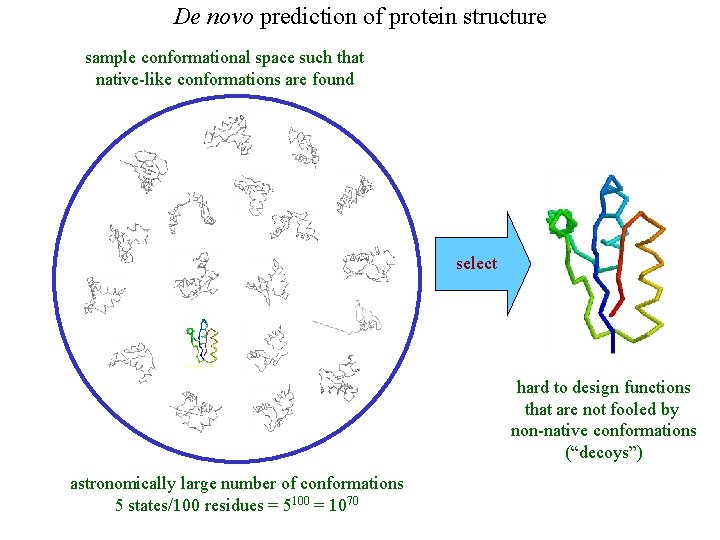
De novo prediction of protein structure sample conformational space such that native-like conformations are found select hard to design functions that are not fooled by non-native conformations (“decoys”) astronomically large number of conformations 5 states/100 residues = 5100 = 1070
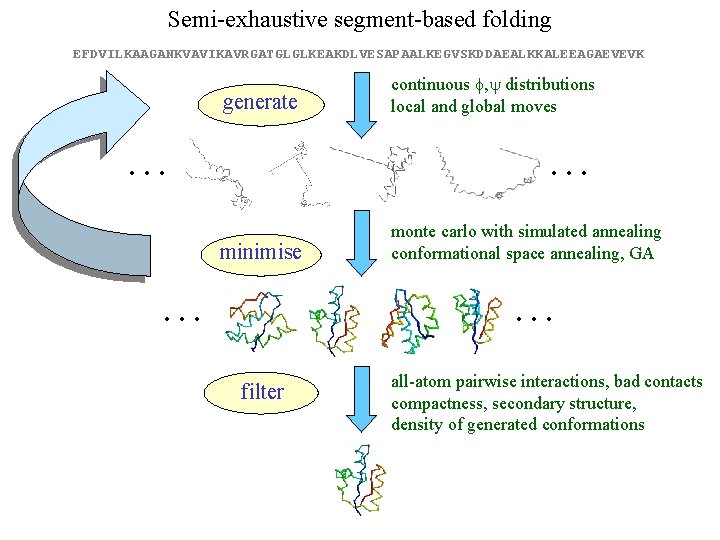
Semi-exhaustive segment-based folding EFDVILKAAGANKVAVIKAVRGATGLGLKEAKDLVESAPAALKEGVSKDDAEALKKALEEAGAEVEVK generate … continuous f, y distributions local and global moves … minimise … monte carlo with simulated annealing conformational space annealing, GA … filter all-atom pairwise interactions, bad contacts compactness, secondary structure, density of generated conformations
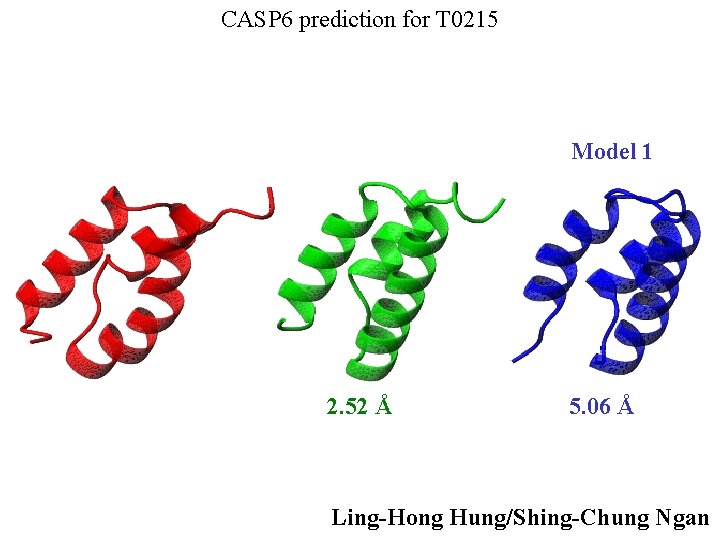
CASP 6 prediction for T 0215 Model 1 2. 52 Å 5. 06 Å Ling-Hong Hung/Shing-Chung Ngan
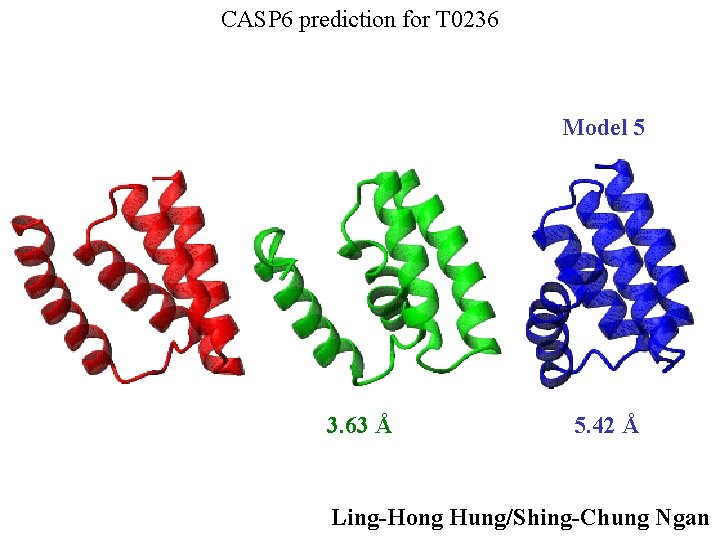
CASP 6 prediction for T 0236 Model 5 3. 63 Å 5. 42 Å Ling-Hong Hung/Shing-Chung Ngan
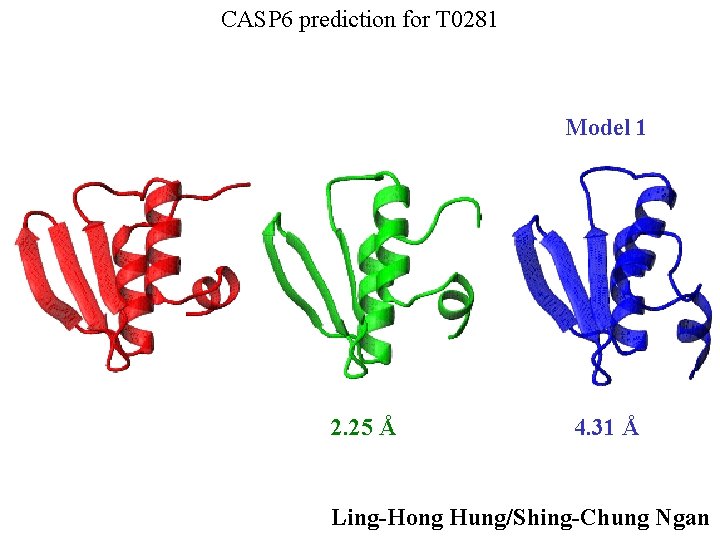
CASP 6 prediction for T 0281 Model 1 2. 25 Å 4. 31 Å Ling-Hong Hung/Shing-Chung Ngan
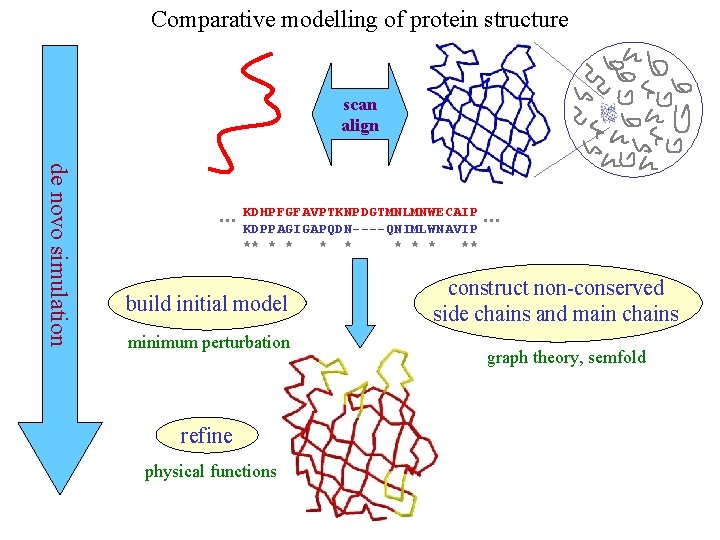
Comparative modelling of protein structure scan align de novo simulation … KDHPFGFAVPTKNPDGTMNLMNWECAIP KDPPAGIGAPQDN----QNIMLWNAVIP ** * * * ** build initial model minimum perturbation refine physical functions … construct non-conserved side chains and main chains graph theory, semfold
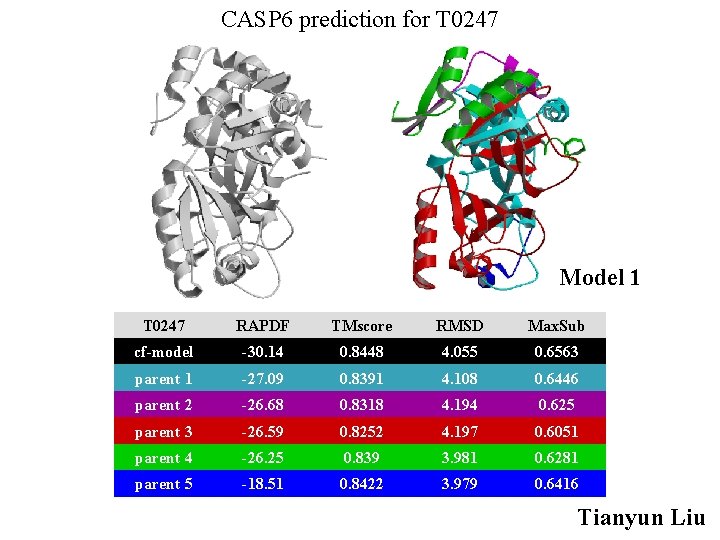
CASP 6 prediction for T 0247 Model 1 T 0247 RAPDF TMscore RMSD Max. Sub cf-model -30. 14 0. 8448 4. 055 0. 6563 parent 1 -27. 09 0. 8391 4. 108 0. 6446 parent 2 -26. 68 0. 8318 4. 194 0. 625 parent 3 -26. 59 0. 8252 4. 197 0. 6051 parent 4 -26. 25 0. 839 3. 981 0. 6281 parent 5 -18. 51 0. 8422 3. 979 0. 6416 Tianyun Liu
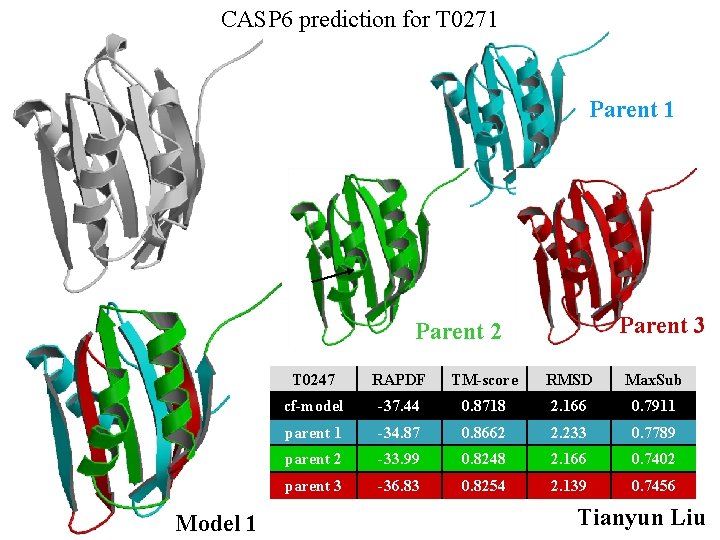
CASP 6 prediction for T 0271 Parent 3 Parent 2 Model 1 T 0247 RAPDF TM-score RMSD Max. Sub cf-model -37. 44 0. 8718 2. 166 0. 7911 parent 1 -34. 87 0. 8662 2. 233 0. 7789 parent 2 -33. 99 0. 8248 2. 166 0. 7402 parent 3 -36. 83 0. 8254 2. 139 0. 7456 Tianyun Liu

CASP 6 overall summaries Tianyun Liu
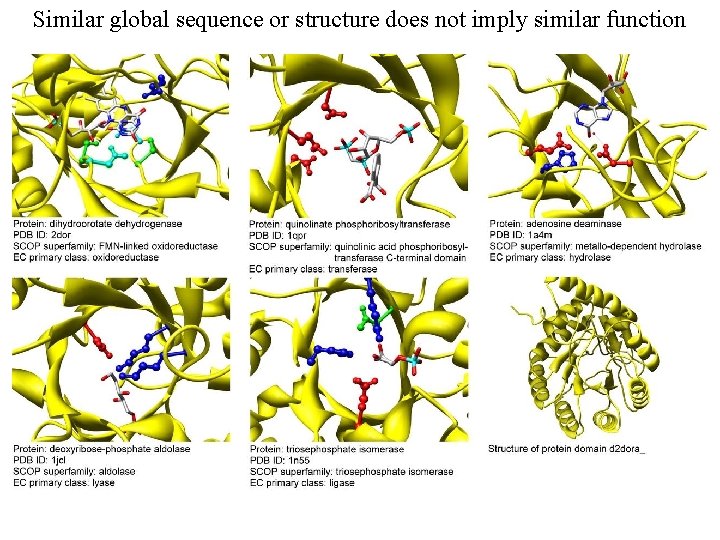
Similar global sequence or structure does not imply similar function
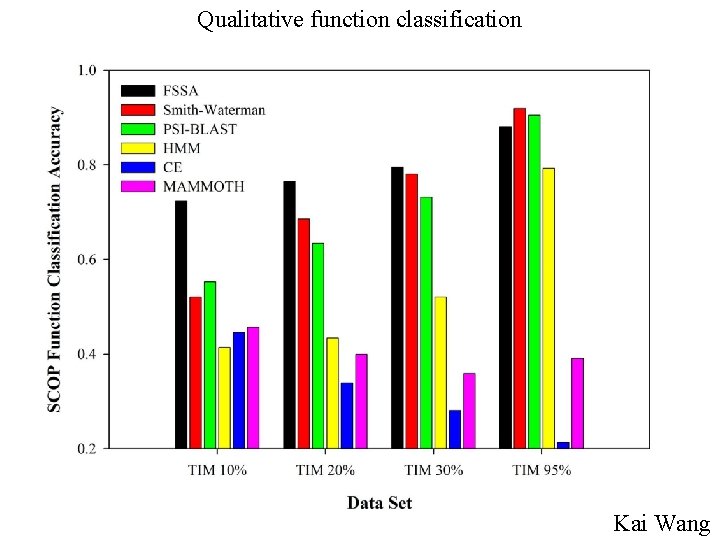
Qualitative function classification Kai Wang
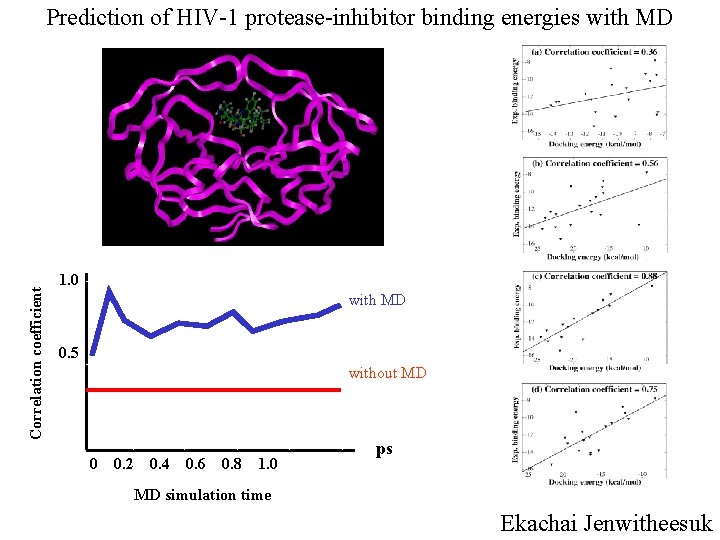
Correlation coefficient Prediction of HIV-1 protease-inhibitor binding energies with MD 1. 0 with MD 0. 5 without MD 0 0. 2 0. 4 0. 6 0. 8 1. 0 ps MD simulation time Ekachai Jenwitheesuk
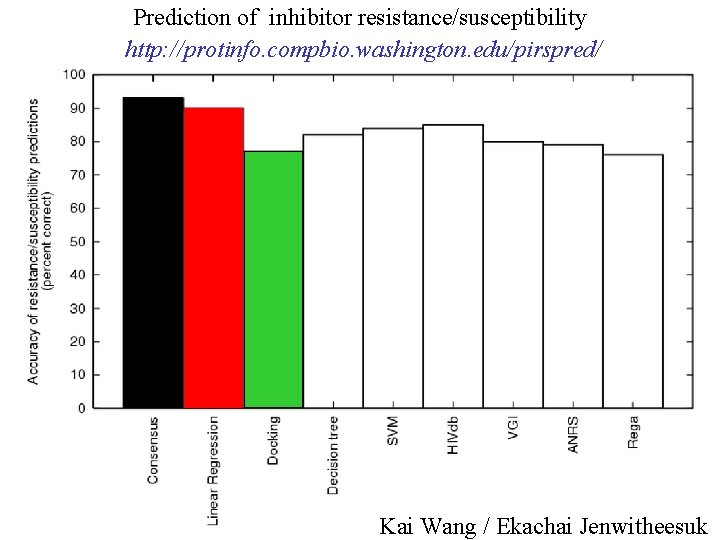
Prediction of inhibitor resistance/susceptibility http: //protinfo. compbio. washington. edu/pirspred/ Kai Wang / Ekachai Jenwitheesuk
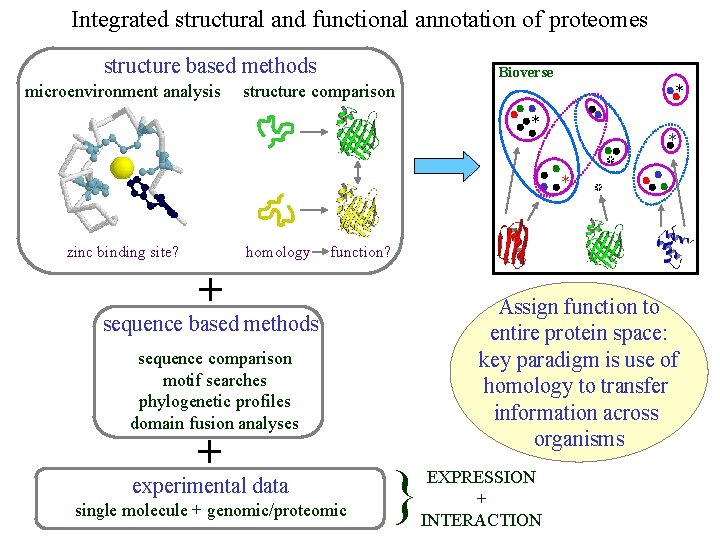
Integrated structural and functional annotation of proteomes structure based methods microenvironment analysis Bioverse structure comparison * * homology zinc binding site? * * function? + Assign function to entire protein space: key paradigm is use of homology to transfer information across organisms sequence based methods sequence comparison motif searches phylogenetic profiles domain fusion analyses + experimental data single molecule + genomic/proteomic } EXPRESSION + INTERACTION

Bioverse – explore relationships among molecules and systems http: //bioverse. compbio. washington. edu Jason Mc. Dermott/Michal Guerquin/Zach Frazier

Bioverse – explore relationships among molecules and systems http: //bioverse. compbio. washington. edu Jason Mc. Dermott/Michal Guerquin/Zach Frazier
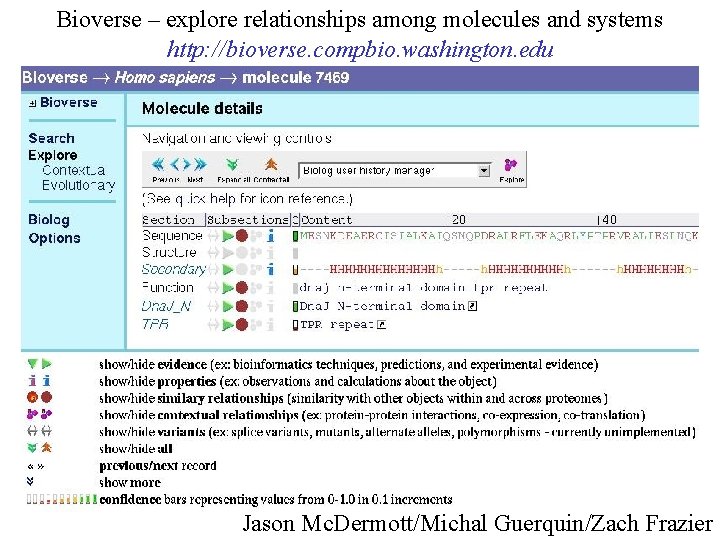
Bioverse – explore relationships among molecules and systems http: //bioverse. compbio. washington. edu Jason Mc. Dermott/Michal Guerquin/Zach Frazier

Bioverse – explore relationships among molecules and systems http: //bioverse. compbio. washington. edu Jason Mc. Dermott/Michal Guerquin/Zach Frazier
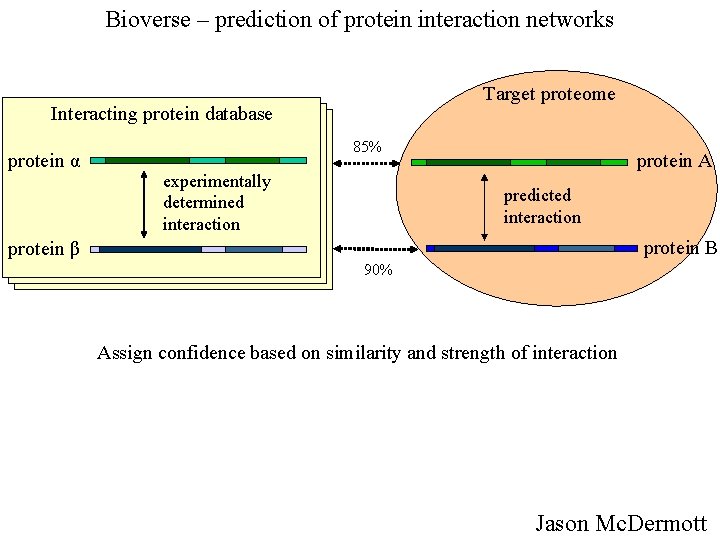
Bioverse – prediction of protein interaction networks Target proteome Interacting protein database protein α 85% experimentally determined interaction protein A predicted interaction protein B protein β 90% Assign confidence based on similarity and strength of interaction Jason Mc. Dermott
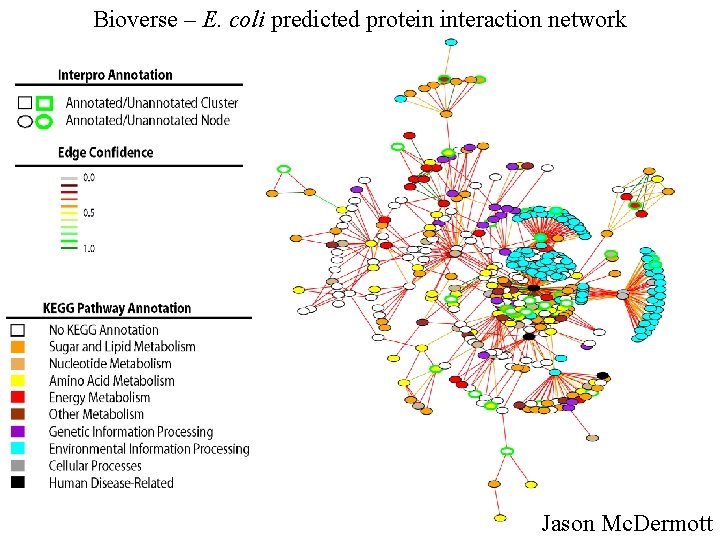
Bioverse – E. coli predicted protein interaction network Jason Mc. Dermott
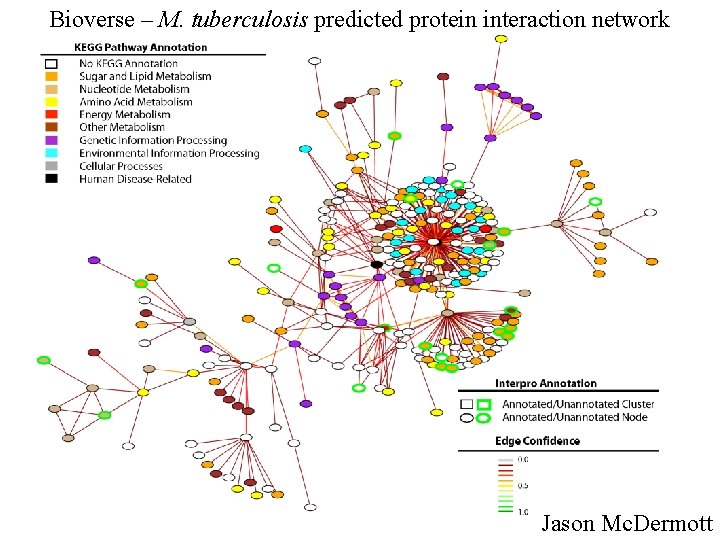
Bioverse – M. tuberculosis predicted protein interaction network Jason Mc. Dermott
Bioverse – C. elegans predicted protein interaction network Jason Mc. Dermott

Bioverse – H. sapiens predicted protein interaction network Jason Mc. Dermott
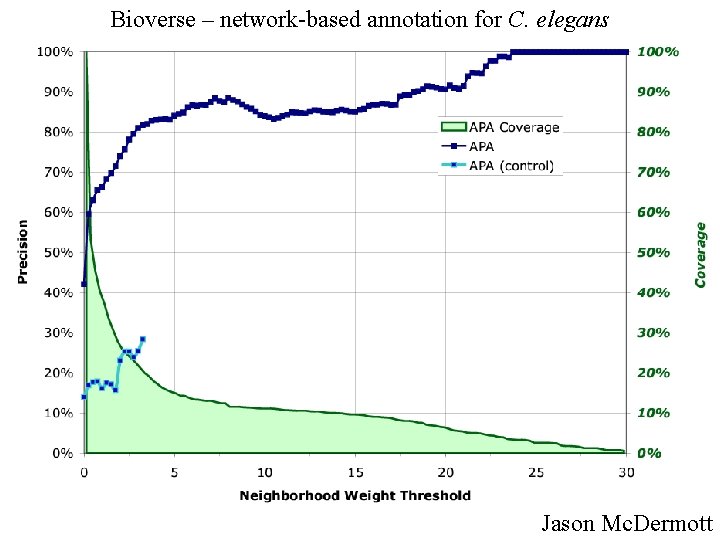
Bioverse – network-based annotation for C. elegans Jason Mc. Dermott
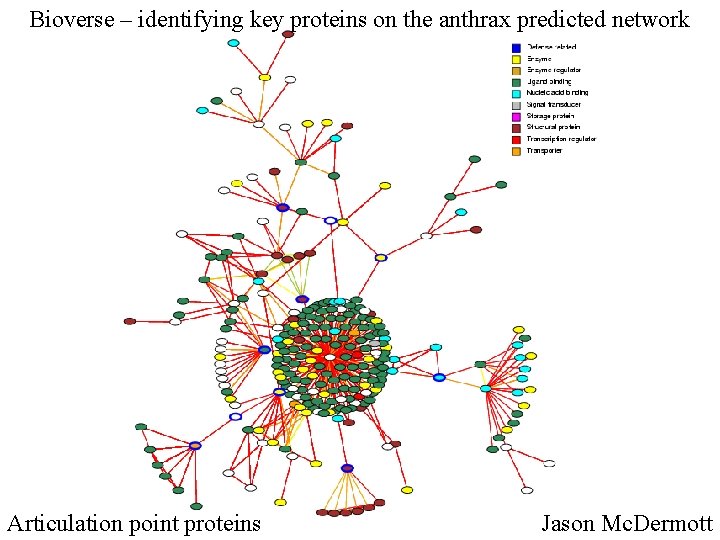
Bioverse – identifying key proteins on the anthrax predicted network Articulation point proteins Jason Mc. Dermott
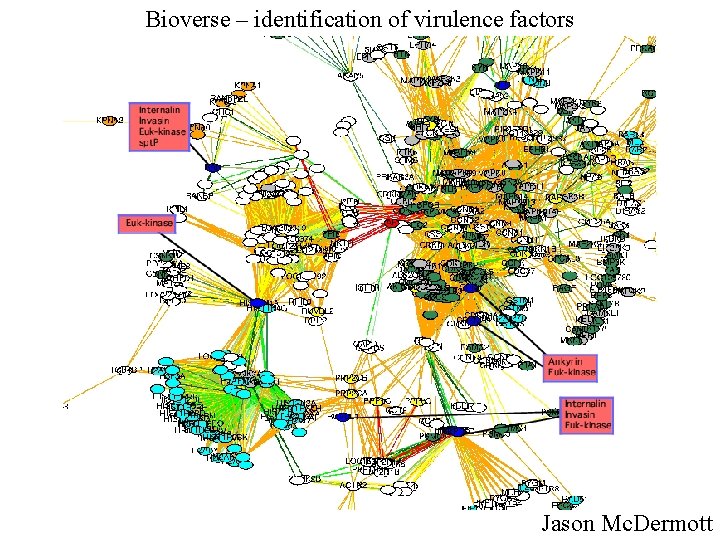
Bioverse – identification of virulence factors Jason Mc. Dermott
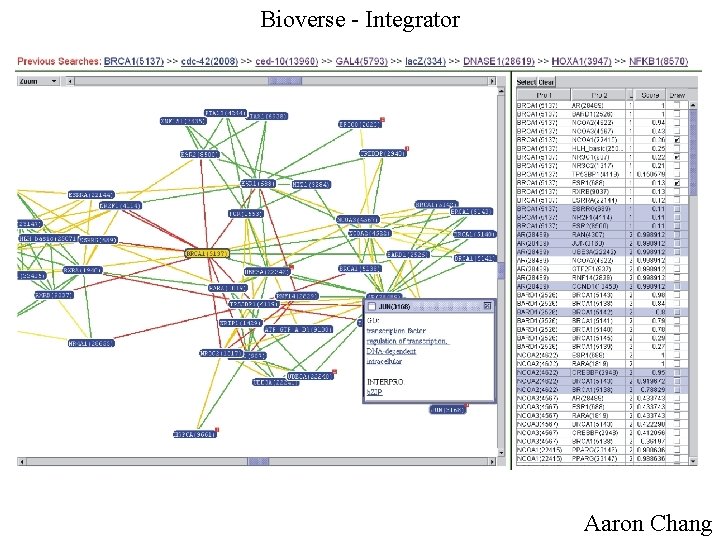
Bioverse - Integrator Aaron Chang

Take home message Prediction of protein structure, function, and networks may be used to model whole genomes to understand organismal function and evolution

Acknowledgements Aaron Chang Chuck Mader David Nickle Ekachai Jenwitheesuk Gong Cheng Jason Mc. Dermott Kai Wang Ling-Hong Hung Mike Inouye Michal Guerquin Stewart Moughon Shing-Chung Ngan Tianyun Liu Zach Frazier National Institutes of Health National Science Foundation Searle Scholars Program (Kinship Foundation) UW Advanced Technology Initiative in Infectious Diseases http: //bioverse. compbio. washington. edu http: //protinfo. compbio. washington. edu
- Slides: 37