Gene network modeling dynamics Luonan Chen Key Laboratory

Gene network modeling (dynamics) Luonan Chen ( 陈洛南 ) Key Laboratory of Systems Biology, Chinese Academy of Sciences;
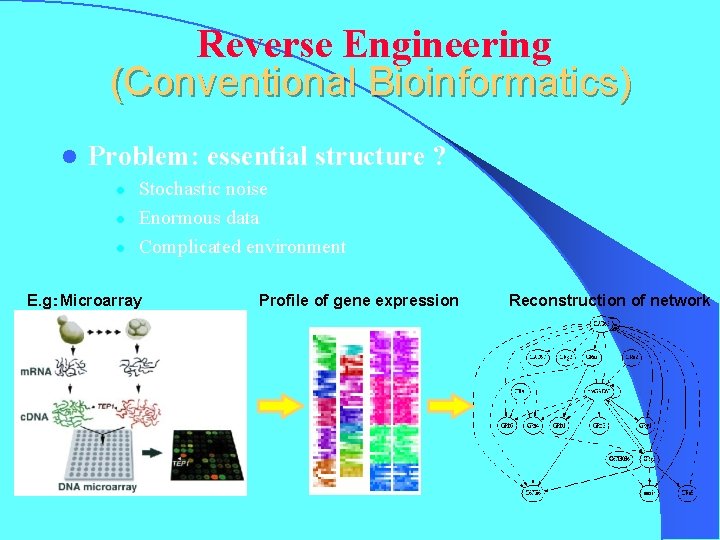
Reverse Engineering (Conventional Bioinformatics) l Problem: essential structure ? l l l Stochastic noise Enormous data Complicated environment E. g:Microarray Profile of gene expression Reconstruction of network
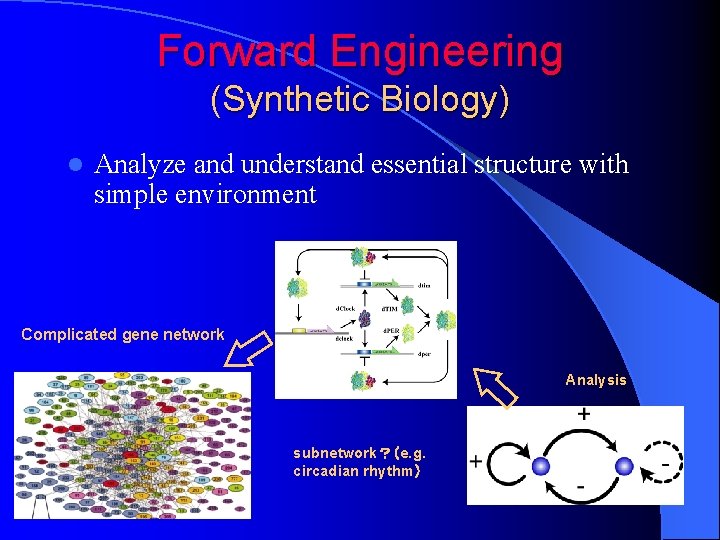
Forward Engineering (Synthetic Biology) l Analyze and understand essential structure with simple environment Complicated gene network Analysis subnetwork?(e. g. circadian rhythm)

Mathematical expression l Master equation (continuous-time discrete-state Markov Chain) l Fokker-Plank equation (continuous approximation) l Langevin equations dx(t)/dt = f(x(t)) + g(x(t)) η(t) l Cumulant eqns (Gaussian assumption) dc(t)/dt = h(c(t)) l Differential eqns (Averaging dynamics) dx(t)/dt = f(x(t)) Algebraic eqns (equilibrium) f(x)=0

Hierarchy of modeling formalisms Graphs Algebraic equations Boolean equations Accuracy Complexity Differential equations Cumulant equations Stochastic differential equations Chemical master equations Reaction-diffusion master equations Abstraction

Modeling Gene Regulatory Network Mathematic Representation (Protein interactions, Metabolic reactions, Gene regulations)
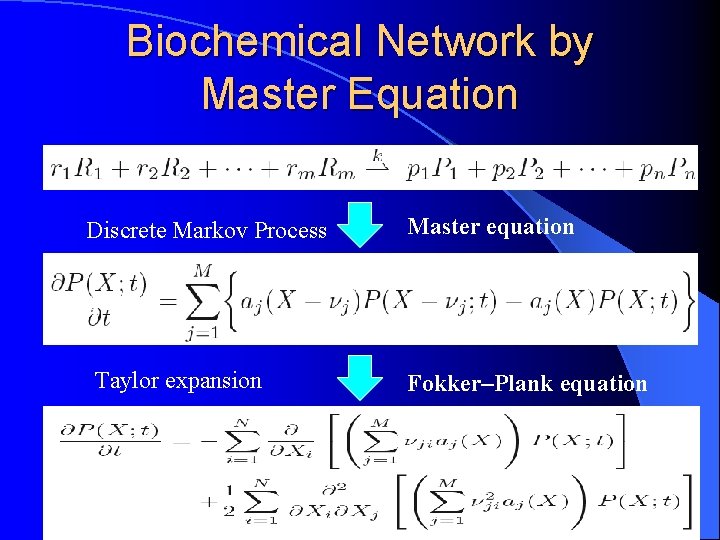
Biochemical Network by Master Equation Discrete Markov Process Taylor expansion Master equation Fokker–Plank equation

Langevin equation Stochastic differential equation Where

Functional Differential Equation L(·) is a linear functional Steady state x : f(x)=0

Modeling Biochemical Reaction (Mass Action Law ) k 1 l A+B C (kinetics) k-1 d[C]/dt=k 1[A][B]-k-1[C] d[A]/dt=k-1[C]-k 1[A][B] Protein Interaction Network k 2 k 1 l E+S ES E+P (Enzyme reaction) k-1 fast Approximation d[ES]/dt=k 1[E][S]-(k-1+k 2)[ES]≈0 Metabolic d[P]/dt=k 2[ES]=k 2[Et][S]/(km+[S]) Network km=(k-1+k 2)/k 1: Michaelis-Mention term [Et]=[E]+[ES]; S: substrate, E: enzyme; P: product
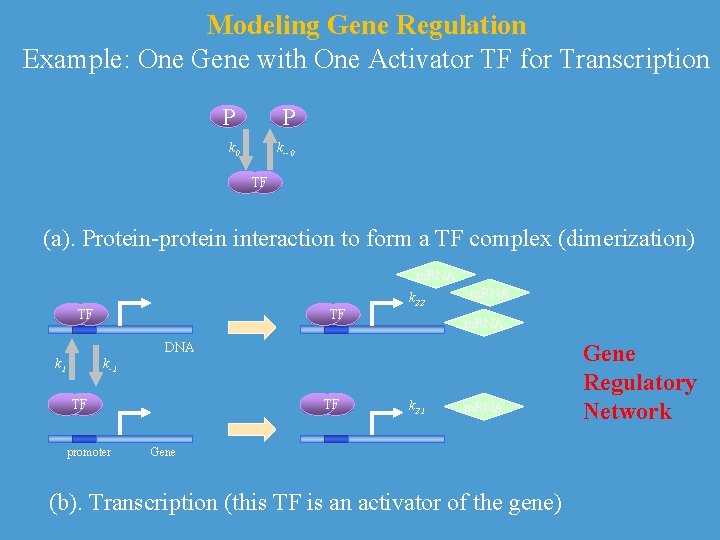
Modeling Gene Regulation Example: One Gene with One Activator TF for Transcription P P k 0 k--0 TF (a). Protein-protein interaction to form a TF complex (dimerization) m. RNA TF TF k 22 m. RNA DNA k 1 k-1 TF promoter TF k 21 m. RNA Gene (b). Transcription (this TF is an activator of the gene) Gene Regulatory Network
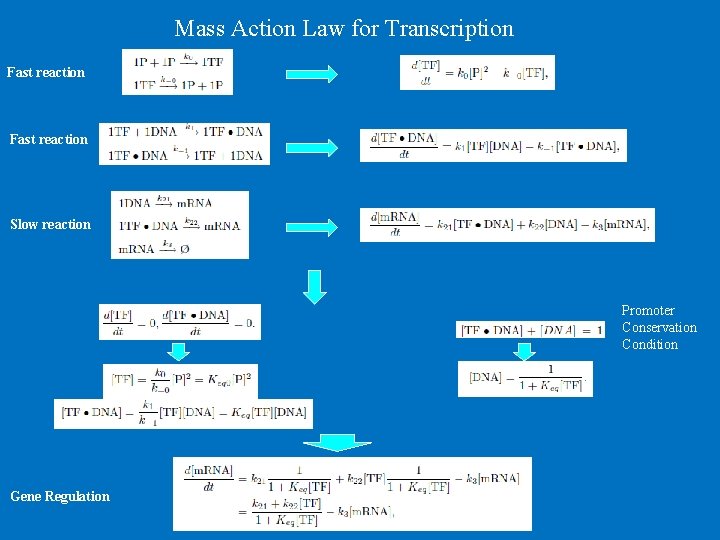
Mass Action Law for Transcription Fast reaction Slow reaction Promoter Conservation Condition Gene Regulation
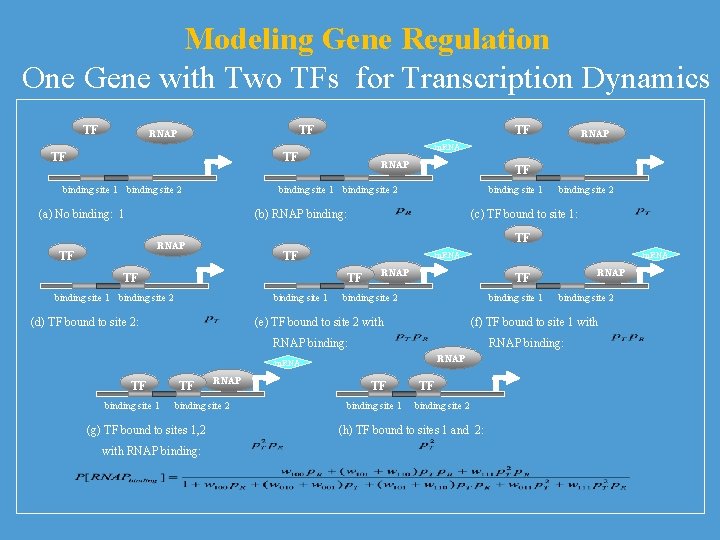
Modeling Gene Regulation One Gene with Two TFs for Transcription Dynamics TF TF RNAP TF TF TF binding site 1 binding site 2 RNAP TF binding site 1 binding site 2 (a) No binding: 1 binding site 1 (b) RNAP binding: (c) TF bound to site 1: TF TF m. RNA TF binding site 1 binding site 2 binding site 1 (d) TF bound to site 2: binding site 1 TF RNAP binding site 2 (g) TF bound to sites 1, 2 with RNAP binding: m. RNAP TF binding site 2 binding site 1 binding site 2 (e) TF bound to site 2 with (f) TF bound to site 1 with RNAP binding: RNAP m. RNA TF binding site 2 TF RNAP m. RNA TF binding site 1 TF binding site 2 (h) TF bound to sites 1 and 2:

Based on conservation condition Gene Regulation
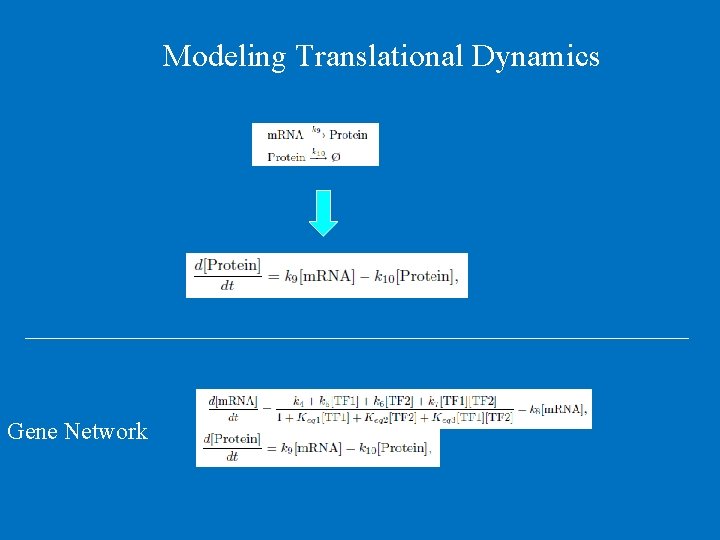
Modeling Translational Dynamics Gene Network

Modeling a general gene regulatory process with mi TFs and sij binding sites for j-th TF Based on Equilibrium of Thermodynamics, we have a general gene regulatory network Closed form Where

Modeling Gene Regulatory Networks (GRN) network: represented by directed graphs: Nodes: states of system, (gene or gene product) edges: interactions or feedback between two states, activation or inhibition (oriented) Label : type of interaction (- repression, + activation) Circuit or Loop: A sequence of nodes that are linked into a cycle (Positive circuit: a circuit with an even number of inhibition edges. Negative circuit: a circuit with an odd number of inhibition edges. )
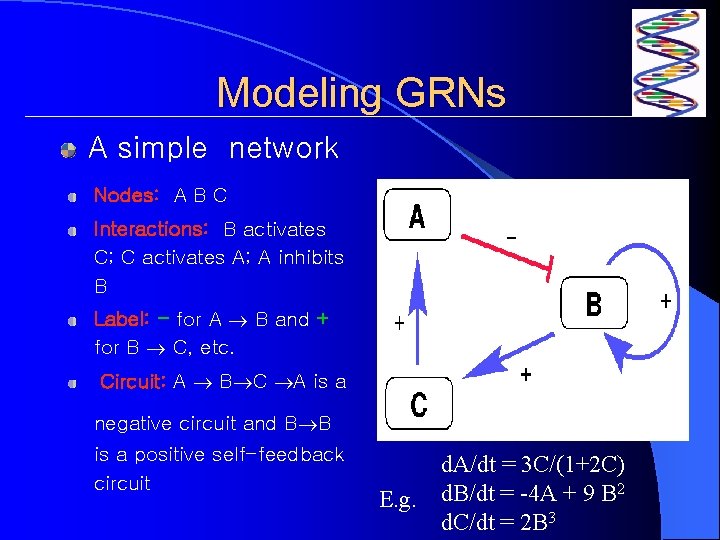
Modeling GRNs A simple network Nodes: A B C Interactions: B activates C; C activates A; A inhibits B Label: - for A B and + for B C, etc. Circuit: A B C A is a negative circuit and B B is a positive self-feedback circuit d. A/dt = 3 C/(1+2 C) 2 E. g. d. B/dt = -4 A + 9 B d. C/dt = 2 B 3

Modeling GRNs GRN as a dynamical system What is a dynamical system? a set of possible states, together with a rule that determines t he present state in terms of past states When a dynamical system is deterministic or stochastic? If the present state can be determined uniquely from the past states (no randomness is allowed), it is deterministic, otherwise, it is stochastic

Modeling GRNs What mathematical formalism? Logical / binary approach (Boolean or logical networks, discrete) On/off states Looks at how each influences others Chemical kinetics /rate equation (FDEs, continuous) Treat state concentrations as continuous variables governed by differential equations, use laws of mass action for interactions Deterministic (no randomness) Stochastic kinetics (ME or F-PE) Treat state concentrations as individual entities, dynamics is governed by individual motions Master or Fokker-Planck equations Adds randomness with noisy terms

Modeling GRNs FDEs : major formalism for modeling GRNs Represent the concentration of state of system by R, R is governed by the rates of formation and degradation: Rate of change of R = rate of formation of R – rate of degradation of R When written as a formular d. R/dt= F(R)-D(R)

Terminology Implication for biology Trajectory – series of state transition Attractor - an attractor is a set of states that repeats itself. Steady state or equilibrium: production exactly balance degradation, i. e. F(R)=D(R) A steady state is linearly stable if the system returns to it after a small perturbation, if it is unstable, it will not be observed in reality. The stability of equilibrium can be determined by the eigenvalues of its Jacobian matrix at equilibrium, in particular, the equilibrium is stable if all eigenvalues of Jacobian matrix have nagative real parts.
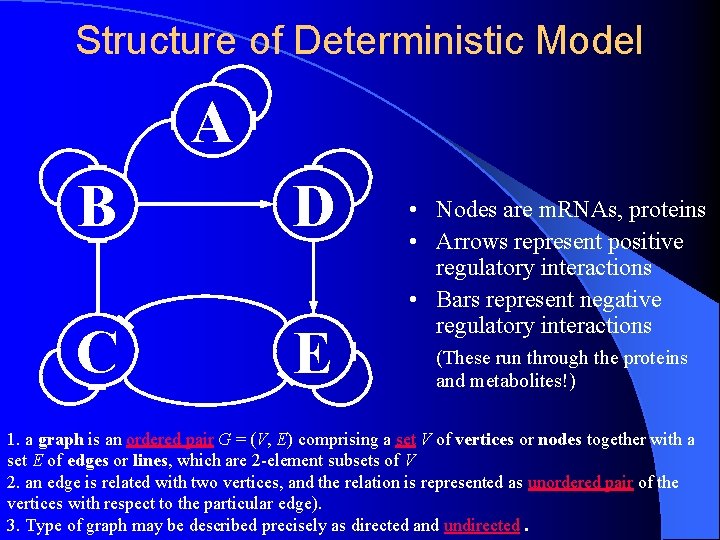
Structure of Deterministic Model A B D C E • Nodes are m. RNAs, proteins • Arrows represent positive regulatory interactions • Bars represent negative regulatory interactions (These run through the proteins and metabolites!) 1. a graph is an ordered pair G = (V, E) comprising a set V of vertices or nodes together with a set E of edges or lines, which are 2 -element subsets of V 2. an edge is related with two vertices, and the relation is represented as unordered pair of the vertices with respect to the particular edge). 3. Type of graph may be described precisely as directed and undirected.
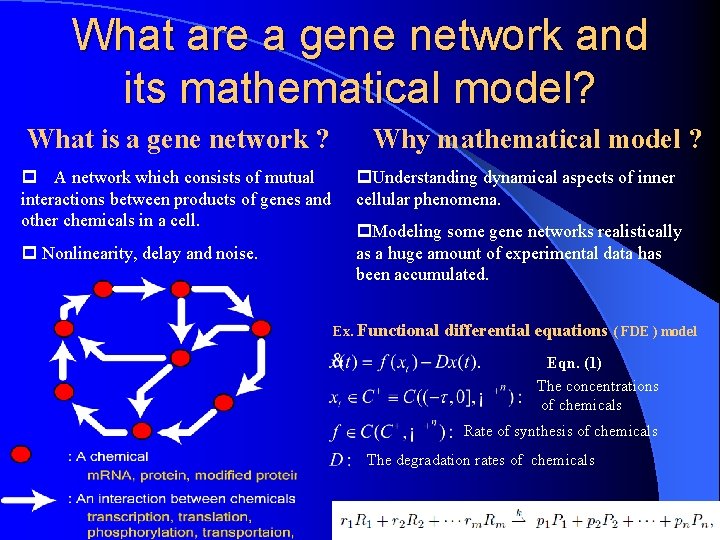
What are a gene network and its mathematical model? What is a gene network ? Why mathematical model ? p A network which consists of mutual interactions between products of genes and other chemicals in a cell. p Nonlinearity, delay and noise. p. Understanding dynamical aspects of inner cellular phenomena. p. Modeling some gene networks realistically as a huge amount of experimental data has been accumulated. Ex. Functional differential equations ( FDE ) model Eqn. (1) The concentrations of chemicals Rate of synthesis of chemicals The degradation rates of chemicals
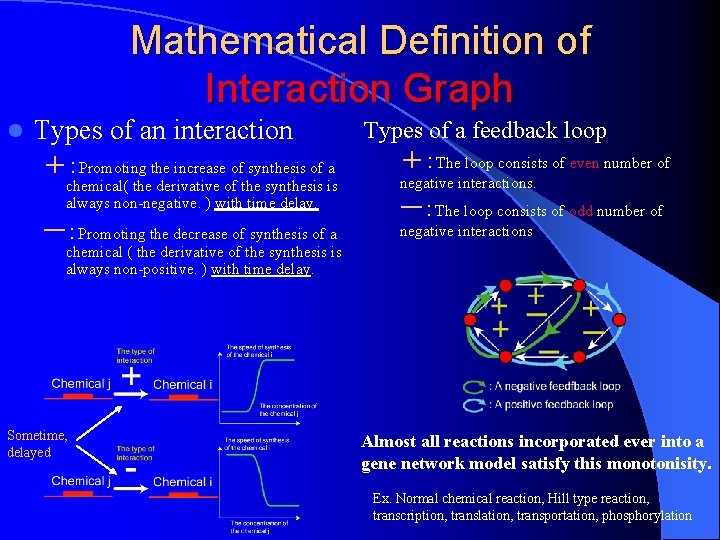
Mathematical Definition of Interaction Graph l Types of an interaction +:Promoting the increase of synthesis of a Types of a feedback loop +:The loop consists of even number of chemical( the derivative of the synthesis is always non-negative. ) with time delay. negative interactions. ー:Promoting the decrease of synthesis of a negative interactions ー:The loop consists of odd number of chemical ( the derivative of the synthesis is always non-positive. ) with time delay. Sometime, delayed Almost all reactions incorporated ever into a gene network model satisfy this monotonisity. Ex. Normal chemical reaction, Hill type reaction, transcription, translation, transportation, phosphorylation

Terminology Implication for biology periodic orbit, stable periodic orbit is in the form of limit cycle, which is the mathematical form of periodic behaviors in biological systems. Circadian oscillation play a key role in the adaptation to periodically varying environment Bifurcation - qualitative change of cell state, for example, from steady state to periodic behavior or from periodic behavior to steady state Bifurcation analysis is an effective way to produce periodic oscillation, althouth it is locally, but when some conditions are satisfied, the periodic osillation will be global

Terminology Implication for biology Time delays – simplified representations of time delays caused by transcription, translation, translocation and diffusion process Structural stability – nearby system has qualitatively the same dynamics, allow the cell to maintain functional state within the tissue even under perturbation
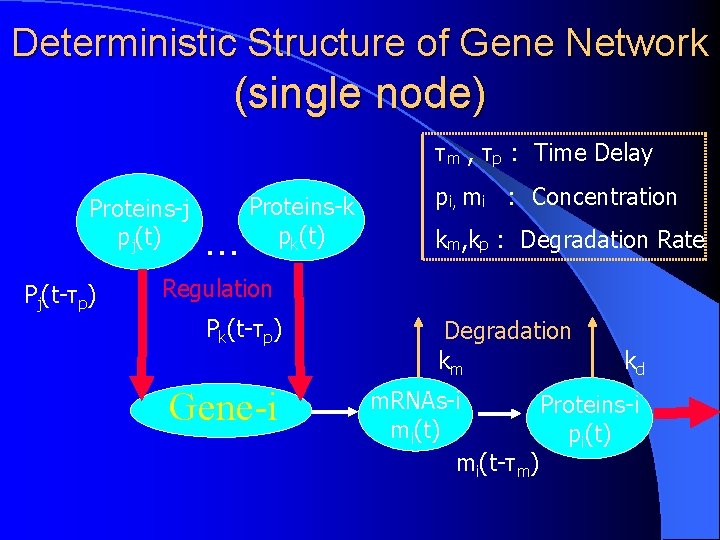
Deterministic Structure of Gene Network (single node) τm , τp : Time Delay Proteins-j pj(t) Pj(t-τp) Proteins-k pk(t) ・・・ Regulation Pk(t-τp) Gene-i pi, mi : Concentration km, kp : Degradation Rate Degradation km kd m. RNAs-i Proteins-i mi(t) pi(t) mi(t-τm)
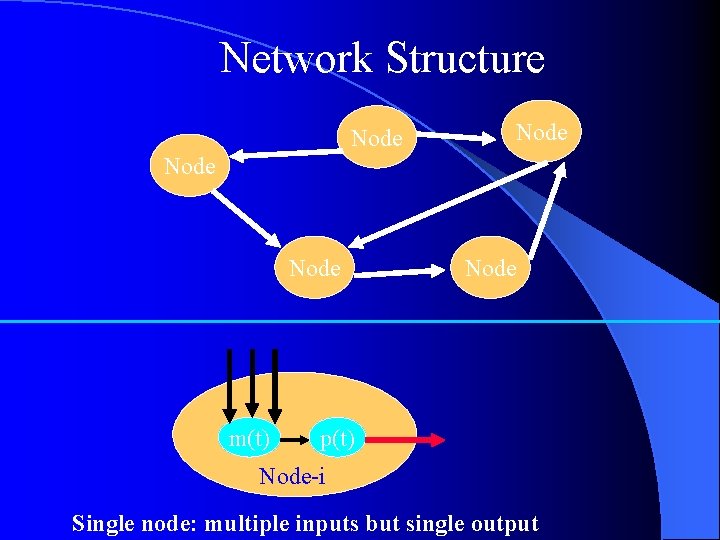
Network Structure Node m(t) Node p(t) Node-i Single node: multiple inputs but single output

FDE Formalism: time-invariant General Form of Gene Network Degradation terms m. RNAs: Proteins : Degradation rates: d: diag matrix Synthesis terms

Time-varying General Gene Network (with cell cycle) - Cell volume factor: v(t) = V(t) /V 0 - Host cell volume: V(t) - V 0 is the host cell volume at the beginning of its growth phase, thereby 1 < v(t) < 2. - m, p: numbers of m. RNA and Protein m -> m/2; p -> p/2; v -> v/2 at division instant

Gene Network in an Eukaryotic Cell (Impulsive Differential Equations)
![Gene Network in a Prokaryotic Cell (Ordinary Differential Equations) [m]: relative concentration Design and Gene Network in a Prokaryotic Cell (Ordinary Differential Equations) [m]: relative concentration Design and](http://slidetodoc.com/presentation_image/5f0b94de937bfd0199ccff068cf01dd8/image-33.jpg)
Gene Network in a Prokaryotic Cell (Ordinary Differential Equations) [m]: relative concentration Design and analyze biomolecular networks based on such formalism
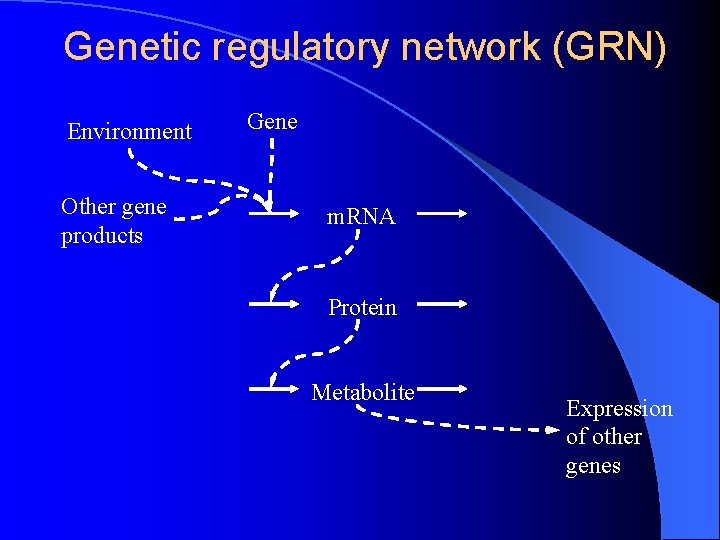
Genetic regulatory network (GRN) Environment Other gene products Gene m. RNA Protein Metabolite Expression of other genes

Design and Analysis of Gene Switches DNA Double Helix
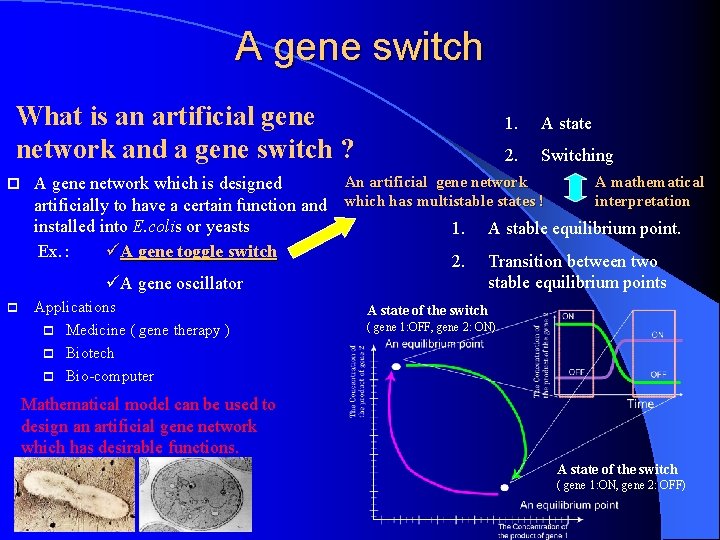
A gene switch What is an artificial gene network and a gene switch ? p A state 2. Switching An artificial gene network A mathematical A gene network which is designed interpretation artificially to have a certain function and which has multistable states ! installed into E. colis or yeasts 1. A stable equilibrium point. Ex. : üA gene toggle switch 2. Transition between two stable equilibrium points üA gene oscillator p 1. Applications p Medicine ( gene therapy ) p Biotech p Bio-computer A state of the switch ( gene 1: OFF, gene 2: ON) Mathematical model can be used to design an artificial gene network which has desirable functions. A state of the switch ( gene 1: ON, gene 2: OFF)

Toggle Switch : two states
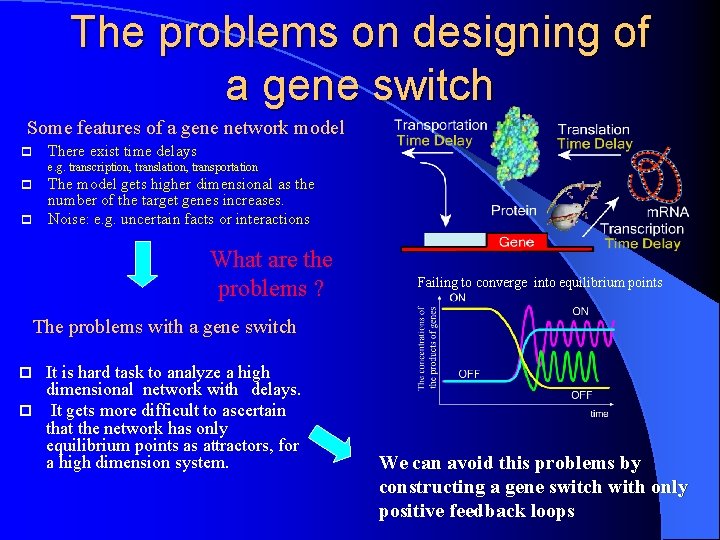
The problems on designing of a gene switch Some features of a gene network model p There exist time delays e. g. transcription, translation, transportation The model gets higher dimensional as the number of the target genes increases. p Noise: e. g. uncertain facts or interactions p What are the problems ? Failing to converge into equilibrium points The problems with a gene switch It is hard task to analyze a high dimensional network with delays. p It gets more difficult to ascertain that the network has only equilibrium points as attractors, for a high dimension system. p We can avoid this problems by constructing a gene switch with only positive feedback loops
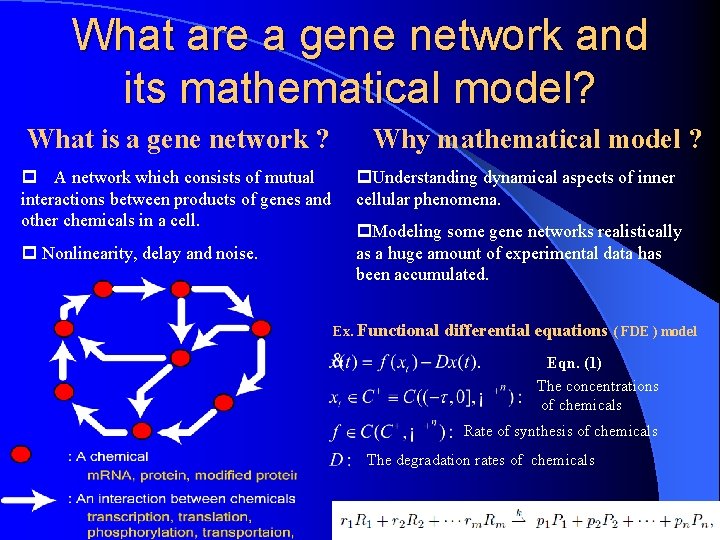
What are a gene network and its mathematical model? What is a gene network ? Why mathematical model ? p A network which consists of mutual interactions between products of genes and other chemicals in a cell. p Nonlinearity, delay and noise. p. Understanding dynamical aspects of inner cellular phenomena. p. Modeling some gene networks realistically as a huge amount of experimental data has been accumulated. Ex. Functional differential equations ( FDE ) model Eqn. (1) The concentrations of chemicals Rate of synthesis of chemicals The degradation rates of chemicals
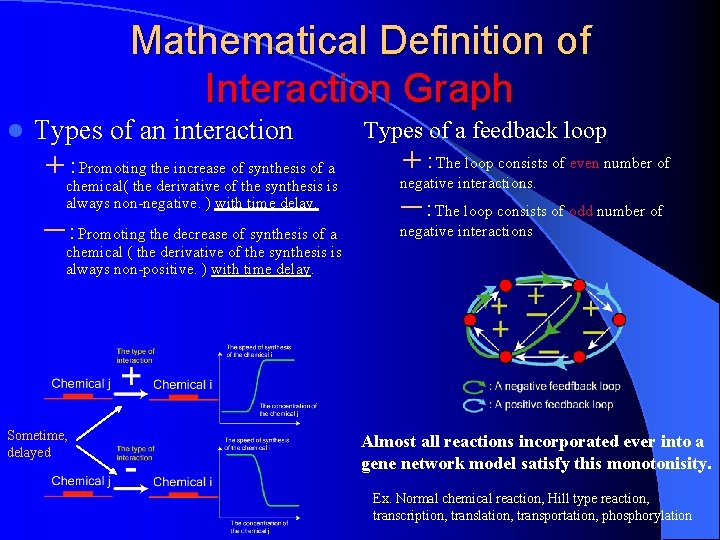
Mathematical Definition of Interaction Graph l Types of an interaction +:Promoting the increase of synthesis of a Types of a feedback loop +:The loop consists of even number of chemical( the derivative of the synthesis is always non-negative. ) with time delay. negative interactions. ー:Promoting the decrease of synthesis of a negative interactions ー:The loop consists of odd number of chemical ( the derivative of the synthesis is always non-positive. ) with time delay. Sometime, delayed Almost all reactions incorporated ever into a gene network model satisfy this monotonisity. Ex. Normal chemical reaction, Hill type reaction, transcription, translation, transportation, phosphorylation

Types of Interactions l Genetic network: dx(t)/dt = f(x(t)) – D x(t) Positive interaction; No interaction; Negative interaction hold for all x in X

Types of Feedback Loops An interaction graph is a directed graph whose nodes represent individual components and whose edges represent interaction between the two nodes. Positive loop 1 Negative loop -1

Monotone Dynamical Systems Positive Network (The Proposed Model) l A network with only positive feedback loops
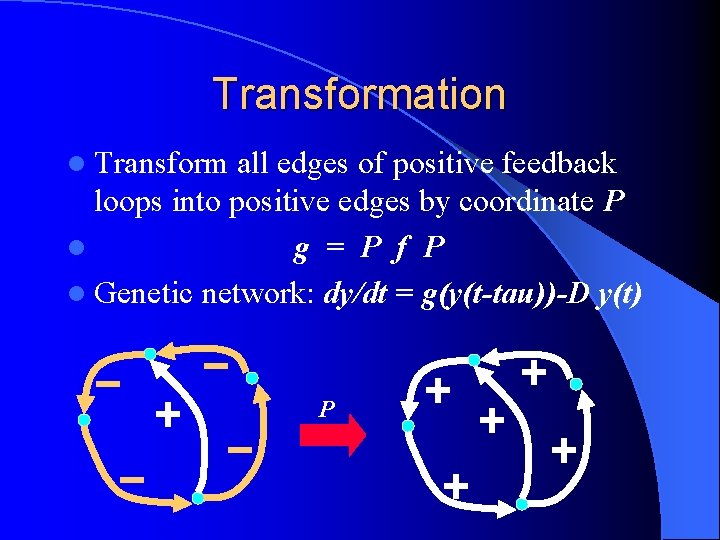
Transformation l Transform all edges of positive feedback loops into positive edges by coordinate P l g = P f P l Genetic network: dy/dt = g(y(t-tau))-D y(t) P
![The special features for positive networks: Part 1 [1] The almost all trajectories of The special features for positive networks: Part 1 [1] The almost all trajectories of](http://slidetodoc.com/presentation_image/5f0b94de937bfd0199ccff068cf01dd8/image-45.jpg)
The special features for positive networks: Part 1 [1] The almost all trajectories of the network fall into equilibria under several mild conditions It is automatically ascertained that the network has no attractor other than stable equilibrium points. p This result is valid for a high dimensional network with delays. p The conditions p All feedback loops in a network are positive ( Weaker condition is sufficient ). p The Jacobian of Eqn (1) is always irreducible. p f maps bounded subsets of C+ to bounded subsets of R+n. p For each initial conditions φin C+ , the solution of Eqn. (1) is always defined for all t >= 0. p The orbit of Eqn. (1) is bounded in C+ p For each compact subet A of C+ There exists a bounded set B of C+ such that the omega limit set of Enq. (1) is included in B for every initial conditions in A.
![The special features for positive networks: Part 2 [2] The equilibria of FDE correspond The special features for positive networks: Part 2 [2] The equilibria of FDE correspond](http://slidetodoc.com/presentation_image/5f0b94de937bfd0199ccff068cf01dd8/image-46.jpg)
The special features for positive networks: Part 2 [2] The equilibria of FDE correspond to those of ODE one by one, and the stability is identical with that of FDE. l The gene switch can be analyzed and designed with ODE ( The asymptotic behavior of a switch is preserved ). The transient behavior of a switch is not necessarily preserved.
![The special features for positive networks: Part 3 [3] It is possible to reduce The special features for positive networks: Part 3 [3] It is possible to reduce](http://slidetodoc.com/presentation_image/5f0b94de937bfd0199ccff068cf01dd8/image-47.jpg)
The special features for positive networks: Part 3 [3] It is possible to reduce the dimensions of the gene switch model keeping the equilibria and their stability. Reduction Simpler minimal Reduction l A switch can be reduced if a component has no auto regulatory direct feedback loop. Mathematically, this reduction is accomplished by changing a ODE to an algebraic equation.
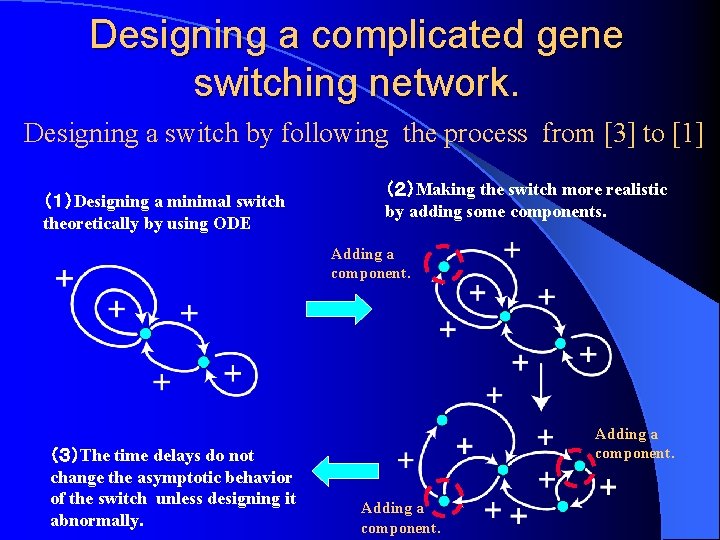
Designing a complicated gene switching network. Designing a switch by following the process from [3] to [1] (1)Designing a minimal switch theoretically by using ODE (2)Making the switch more realistic by adding some components. Adding a component. (3)The time delays do not change the asymptotic behavior of the switch unless designing it abnormally. Adding a component.
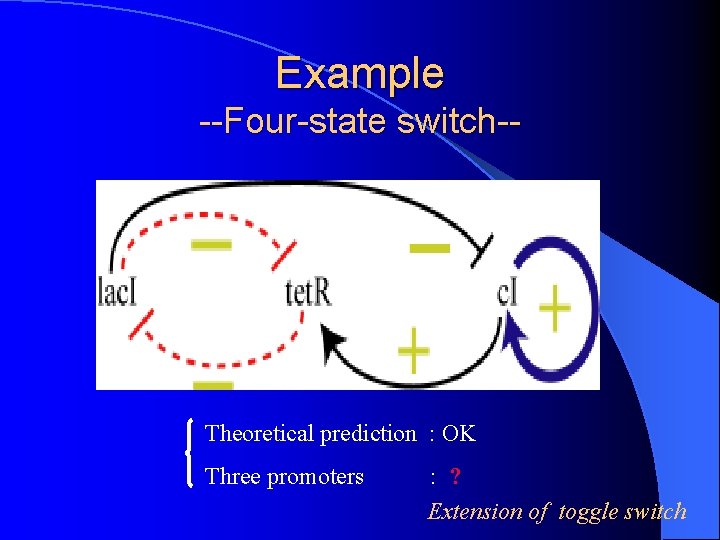
Example --Four-state switch-- Theoretical prediction : OK Three promoters : ? Extension of toggle switch

Under experiment Implementation Tetramer Dimer Operons : avoid constructing logical gates PRM : binding site OR 3 is mutated RBS : 1 and 2 for different RBS Toggle switch : PLtet and Ptrc-2 with lac. I and tet. R
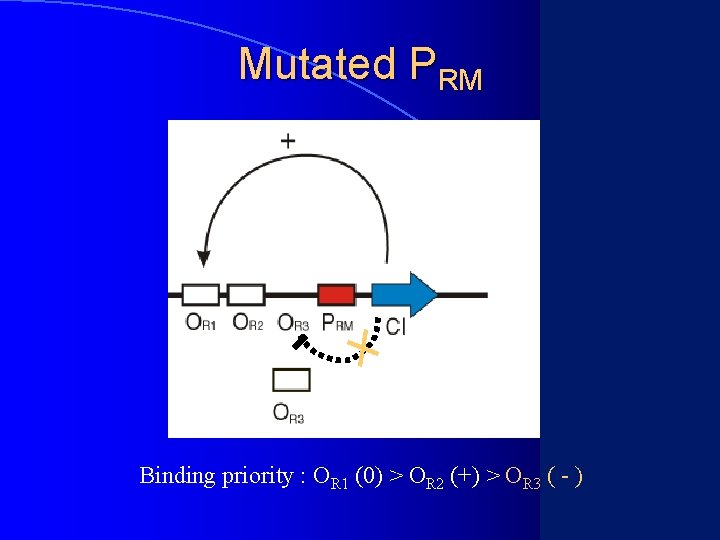
Mutated PRM Binding priority : OR 1 (0) > OR 2 (+) > OR 3 ( - )

Necessary Conditions of 3 or 4 States Switch l (1) Toggle switch has two stable equilibria l (2) c. I has low expression level No problem for condition (1)

Problem: Activation of c. I is too strong ! Inefficiency of poly-cistronic transcription (transcriptional efficiency of the second gene at downstream of promoter can be as low as 1/100 as that of the first gene) c. I is located at the second place in operons

Parameter Design FDE for operon model Reduced to a two-dimensional ODE
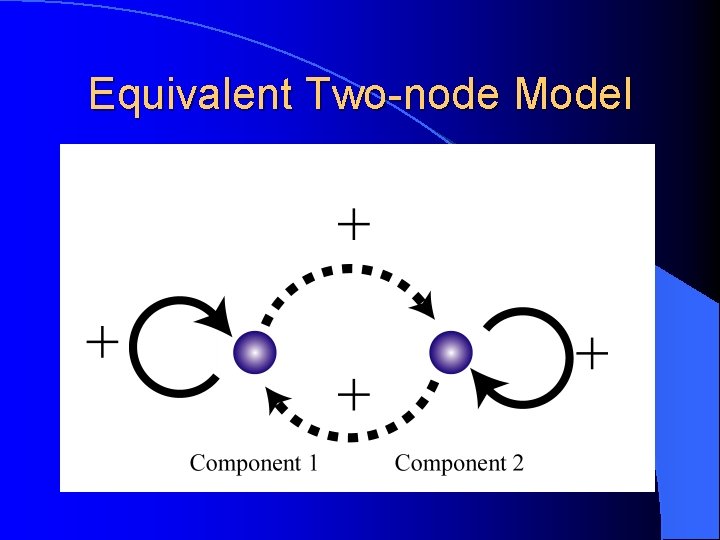
Equivalent Two-node Model
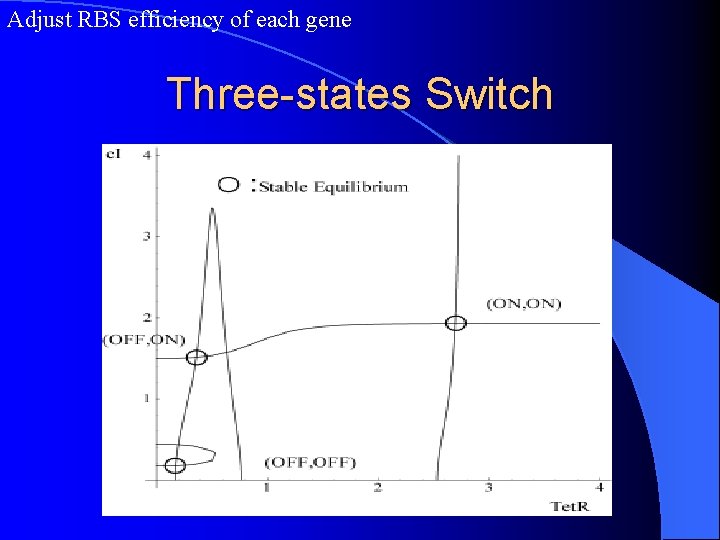
Adjust RBS efficiency of each gene Three-states Switch
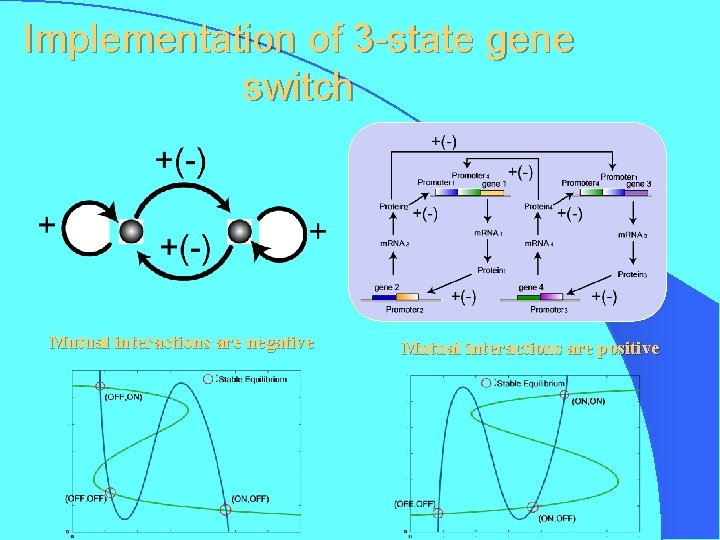
Implementation of 3 -state gene switch Mutual interactions are negative Mutual interactions are positive
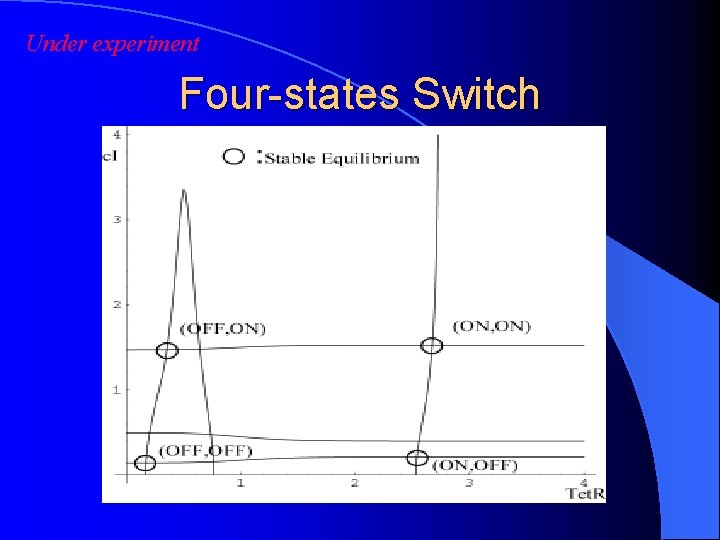
Under experiment Four-states Switch

Conclusion l Design genetic switch network with multiple genes and uncertain delays ---- complicated logical gates ---l Develop a procedure of equivalent reduction l Biological plausible example and simulation l General settings

Acknowledgment l Chinese Academy of Sciences: Prof. X-S. Zhang, Drs. L-Y. Wu, Y. Wang, Z-P. Li, etc. l University of Tokyo: Prof. K. Aihara, Dr. XM. Zhao, Dr. T. Kobayashi, etc. l Shanghai University: Dr. R-Q. Wang, Prof. Z. Liu, Prof. T. Wen, Prof. Zhang, etc. l Osaka Sangyo University: Dr. R-S. Wang

Books Network ! l L. Chen, R. Wang, X. Zhang, Biomolecular Network: Methods and Applications in Systems Biology, John Wiley & Sons, 2009 Dynamics ! l L. Chen, R. Wang, C. Li, K. Aihara, Modelling Biomolecular Networks in Cells: Structures and Dynamics. Springer-Verlag, 2010
- Slides: 61