Protein Interaction Prediction Network Luonan Chen Key Laboratory

Protein Interaction Prediction -Network. Luonan Chen Key Laboratory of Systems Biology Chinese Academy of Sciences

Overview Basic concept n Probabilistic model n Biological experimental data n LP-based methods n Results of computational experiments n Reconstructing complexes, pathways n Conclusion n
Purpose n Understanding mechanisms of protein interactions and predicting protein functions. n n n Transcription factors Signaling pathways Receptors, etc. Biochemical reactions Gene regulation

Basic concept Predicting interaction of a protein pair based only on sequence data n n Distances Clustering methods

Distance Metrics for Clustering • • • Distances metrics can be classified as • Metric distances • Semi-metric distances Metric distances: 1. dab >= 0 2. dab = dba 3. daa = 0 4. dab <= dac + dcb Semi-metric distances: obey 1) to 3), fail in 4)

Examples of Distance Metrics Minkowski distance If q = 1, d is Manhattan distance (semi-metric distance) If q = 2, d is Euclidean distance (metric distance)
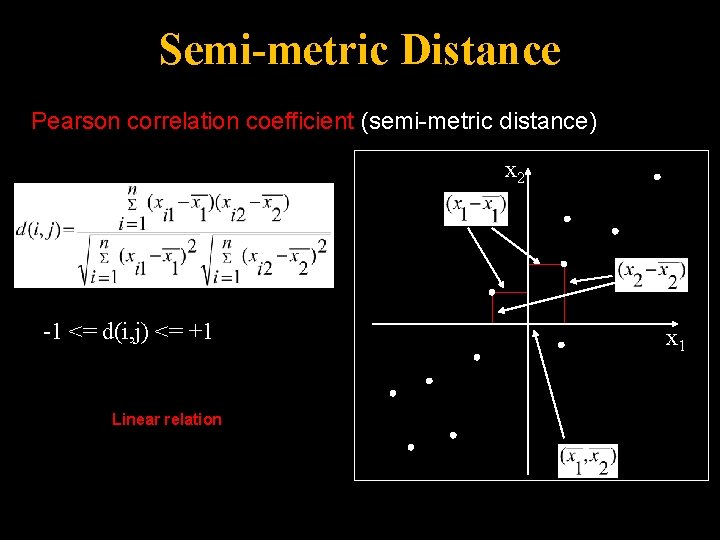
Semi-metric Distance Pearson correlation coefficient (semi-metric distance) x 2 -1 <= d(i, j) <= +1 Linear relation x 1

Other variations of Pearsons correlation coefficient Uncentered Pearson correlation (semi-metric distance) Similar to cos function?
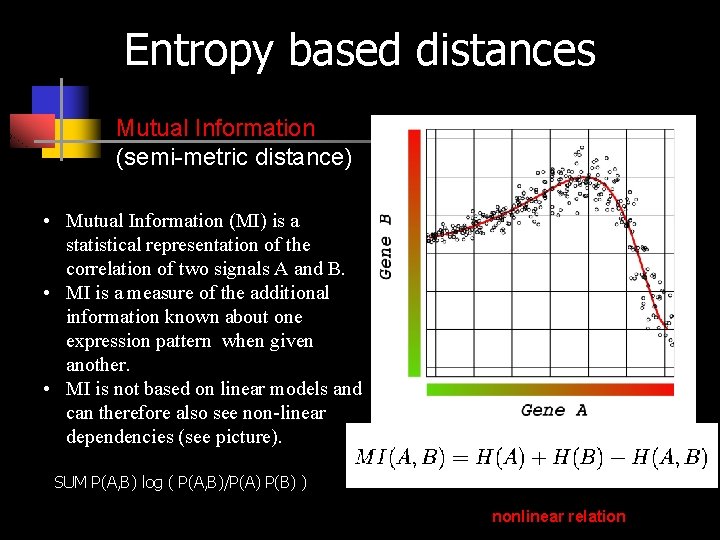
Entropy based distances Mutual Information (semi-metric distance) • Mutual Information (MI) is a statistical representation of the correlation of two signals A and B. • MI is a measure of the additional information known about one expression pattern when given another. • MI is not based on linear models and can therefore also see non-linear dependencies (see picture). SUM P(A, B) log ( P(A, B)/P(A) P(B) ) nonlinear relation

Clustering Techniques Several classification criteria of clustering algorithms exist with regard to • Clustering result: hierarchical – not hierarchical • Clustering process: divisive - agglomerative • Clustering criterion: sequential - global
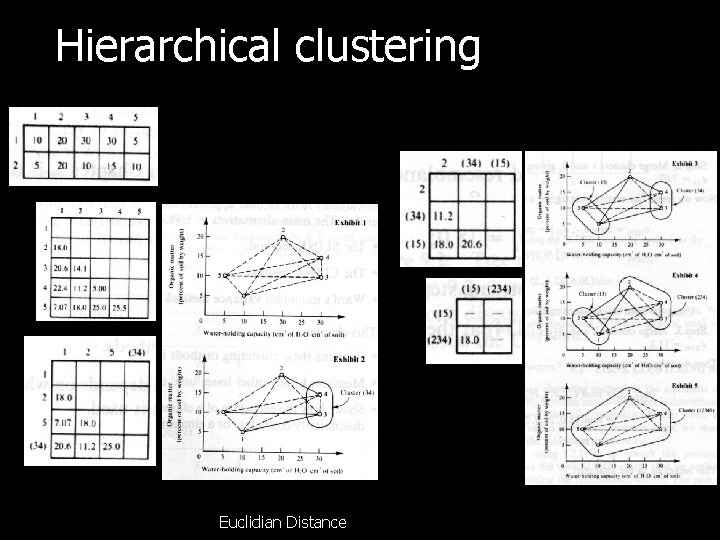
Hierarchical clustering Euclidian Distance
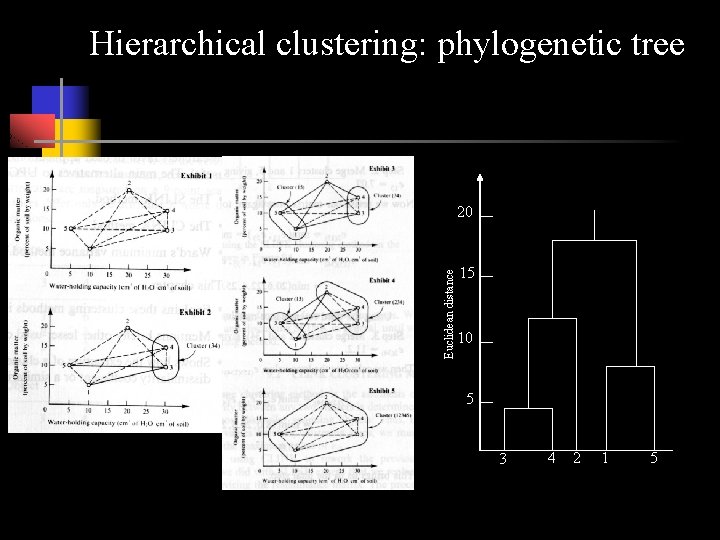
Hierarchical clustering: phylogenetic tree Euclidean distance 20 15 10 5 3 4 2 1 5

Other Clustering Techniques • Single-linkage clustering, nearest-neighbour • Complete-linkage, furthest-neighbour • Average-linkage, unweighted pair-group method average (UPGMA) • Weighted-pair group average, UPGMA weighted by cluster sizes • Within-groups clustering • Ward‘s method
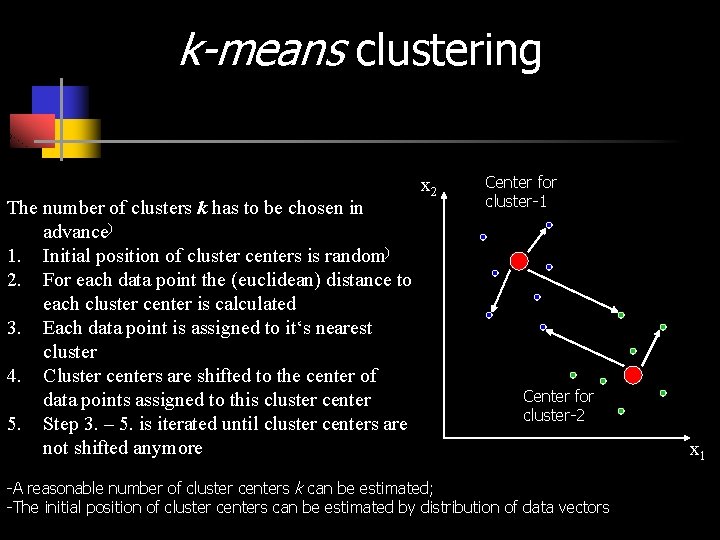
k-means clustering The number of clusters k has to be chosen in advance) 1. Initial position of cluster centers is random) 2. For each data point the (euclidean) distance to each cluster center is calculated 3. Each data point is assigned to it‘s nearest cluster 4. Cluster centers are shifted to the center of data points assigned to this cluster center 5. Step 3. – 5. is iterated until cluster centers are not shifted anymore x 2 Center for cluster-1 Center for cluster-2 -A reasonable number of cluster centers k can be estimated; -The initial position of cluster centers can be estimated by distribution of data vectors x 1
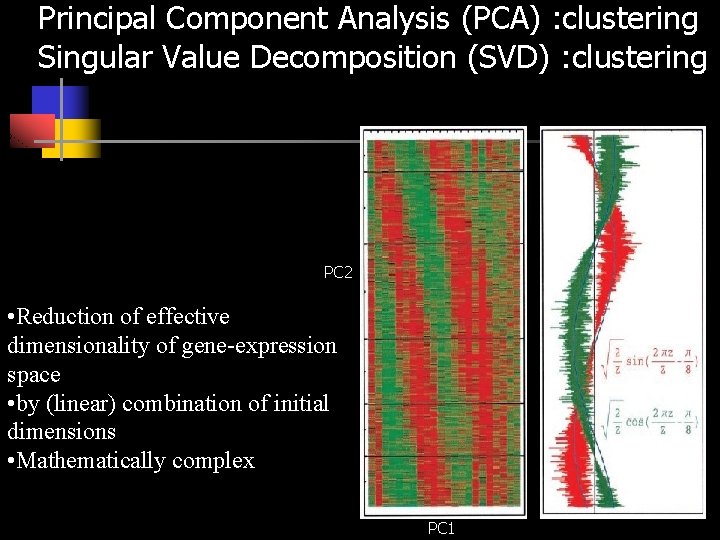
Principal Component Analysis (PCA) : clustering Singular Value Decomposition (SVD) : clustering PC 2 • Reduction of effective dimensionality of gene-expression space • by (linear) combination of initial dimensions • Mathematically complex PC 1

Comparison of Clustering Techniques • No clustering technique is better than another • Different clustering techniques have shown to lead to reasonable results depending on the measured data: • Organism • Tissue • Experimental condition, e. g. mutants or time course experiments • Distance metric used

Various Heuristic Methods for PPI n Heuristic Various methods were developed for inferring PPI and protein functions n Gene fusion/Rosetta stone (Enright et al. and Marcotte et al. 1999), Phylogenetic profile, protein structure n Theoretical n Number of possible genes to be applied is limited (coverage). Molecular dynamics n n Long CPU time, small molecules Difficult to predict precisely

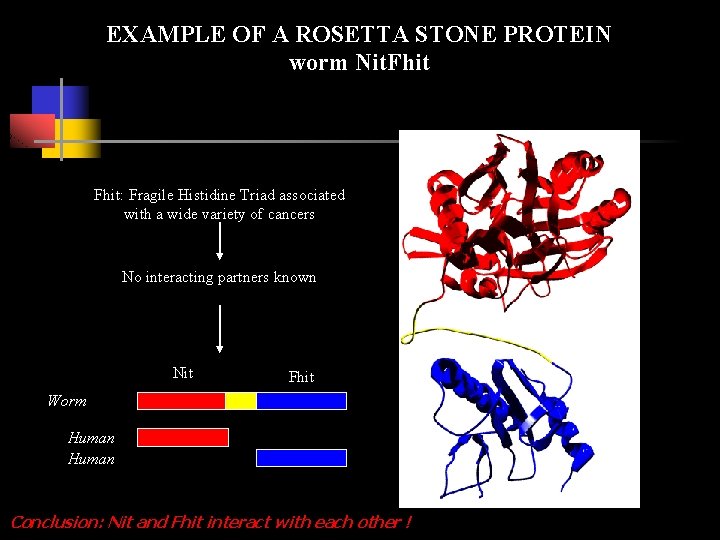
EXAMPLE OF A ROSETTA STONE PROTEIN worm Nit. Fhit: Fragile Histidine Triad associated with a wide variety of cancers No interacting partners known Nit Fhit Worm Human Conclusion: Nit and Fhit interact with each other !
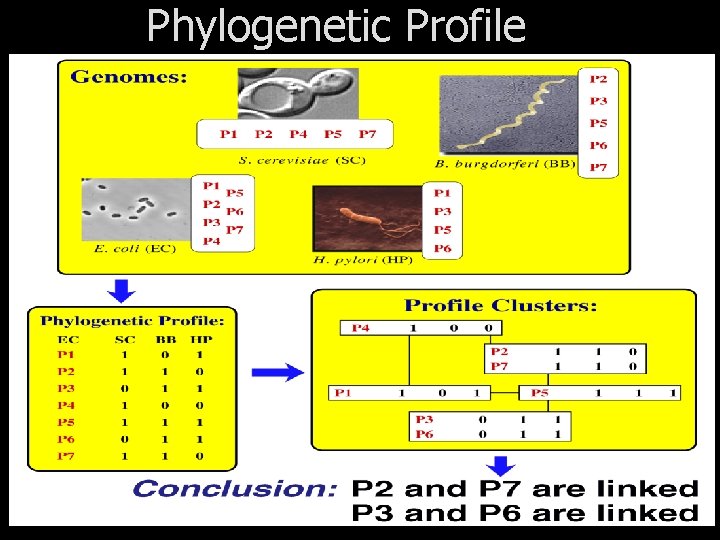
Phylogenetic Profile

Microarray Co-expression Analysis Profile Clusters P 4 Expression Profile P 1 P 2 P 3 P 4 P 5 P 6 P 7 1 1 0 1 o Pr n tei ch Ri ed m 0 1 1 1 ve r ta S 1 0 1 1 0 0 P 2 P 7 P 1 1 0 1 1 1 1 P 5 P 3 P 6 0 0 1 1 1 Conclusion: P 2 and P 7 are functionally linked P 3 and P 6 are functionally linked 0 0
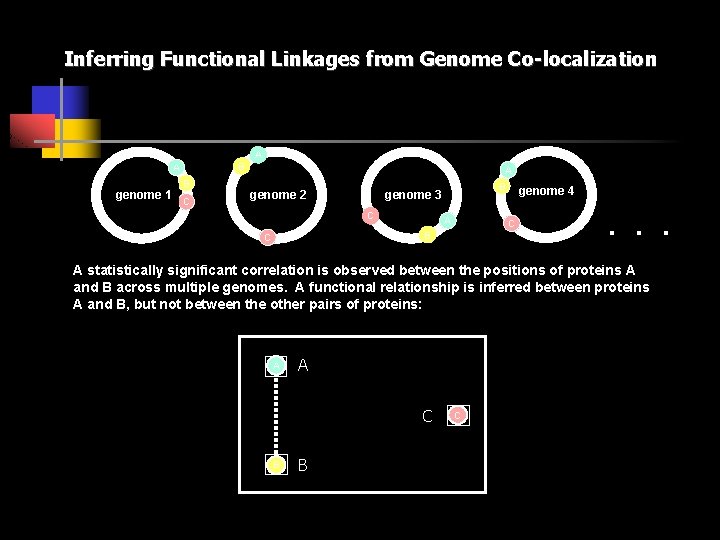
Inferring Functional Linkages from Genome Co-localization A B A genome 1 B C A genome 2 B genome 3 C A C B C genome 4 . . . A statistically significant correlation is observed between the positions of proteins A and B across multiple genomes. A functional relationship is inferred between proteins A and B, but not between the other pairs of proteins: A A C B B C

Protein interaction prediction by single domain method
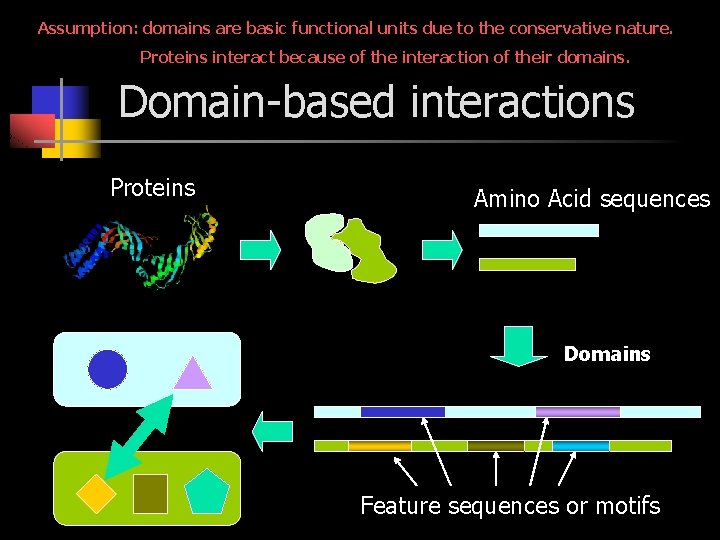
Assumption: domains are basic functional units due to the conservative nature. Proteins interact because of the interaction of their domains. Domain-based interactions Proteins Amino Acid sequences Domains Feature sequences or motifs
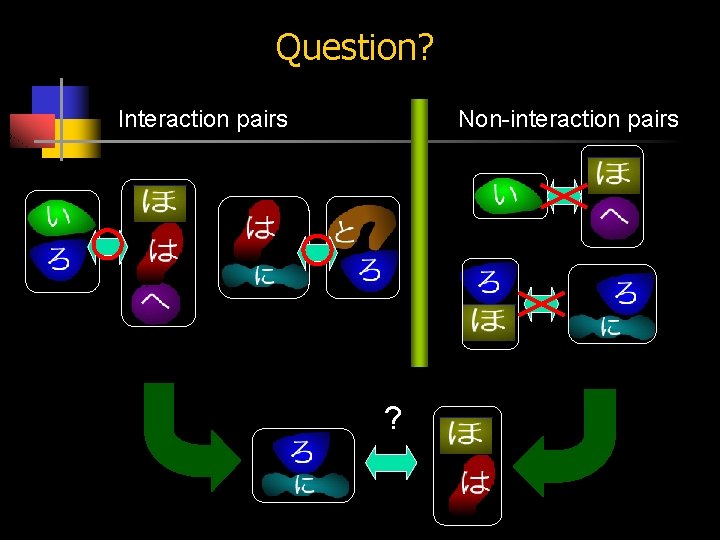
Question? Interaction pairs Non-interaction pairs ?
Inference
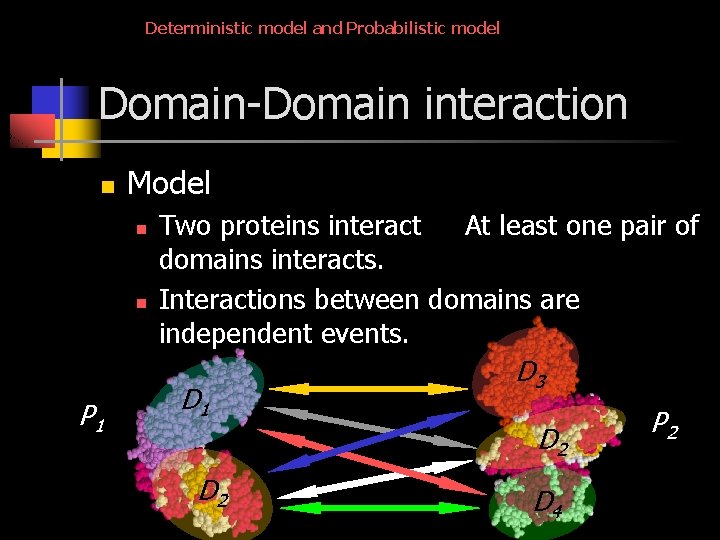
Deterministic model and Probabilistic model Domain-Domain interaction n Model n n P 1 Two proteins interact At least one pair of domains interacts. Interactions between domains are independent events. D 1 D 2 D 3 D 2 D 4 P 2

Probabilistic model for single domain pair n n n : Proteins Pi and Pj interact : Domains Dm and Dn interact : Domain pair (Dm , Dn) is included in protein pair Pi. X Pj Assumption: Independent interactions for each pair domains Proteins interacts if at least one pair of domains interacts

Association method (Sprinzak et al. , 2001) n Inference of probabilities of domain interactions using ratios of frequencies n n : Number of interacting protein pairs that include (Dm, Dn) : Number of protein pairs that include (Dm, Dn)

EM method (Deng et al. , 2002) n n n Probability (likelihood L) that experimental data {Oij={0, 1}} are observed. Use EM algorithm in order to (locally) maximize L. Estimate Pr(Dmn=1)

LP-based method (Hayashida, et al. ) n It seems difficult to modify EM method for numerical data. n For binary data n n LPBN Combined methods n n Linear Programming n n n LPEM EMLP SVM-based method For numerical data n n ASNM LPNM

Probabilistic model for single domain pair Variables where Constants

Predicting protein interaction n Learning process: for all protein pairs with experiment results, based on obtain domain interaction n Prediction process: for a pair of protein sequences, obtain all domains Dmn. Based on predict protein interaction

Simple SVM-based method n Feature vector Test data n Simple linear kernel with n n Interacting pairs = Positive Non-interacting pairs = Negative Margin

Overview n n Background Probabilistic model Biological experimental data Proposed methods n n APM (single domain pair) MDCinfer (multi domain pair): APMM, LPMM Results of computational experiments Conclusion

Available data Biological experimental data n n Binary data (interact or not). Experimental data using Yeast 2 hybrid n n n Ito et al. (2000, 2001) Uetz et al. (2001) For many protein pairs, different results (Oij = {0, 1}) were observed. raw numerical data. ipfam, pfam

Databases (ipfam) n A model based on domain-domain interactions has been proposed. n Use domains defined by databases like Inter. Pro or Pfam. Domain

Numerical data n Ito et al. (2000, 2001) n n For each protein pair, experiments were performed multiple times. IST (Interaction Sequence Tag) n n Number of observed interactions By using a threshold, we obtain binary data.

Overview n n n Background Probabilistic model Related work Biological experimental data Proposed methods n n APM (single domain pair) MDCinfer (multi domain pair) : APMM, LPMM Results of computational experiments Conclusion

Problems: Independent interactions for each pair domains, Proteins interacts if at least one pair of domains interacts, Single datasets with insufficient data Overview n n n Background Probabilistic model Related work Biological experimental data Proposed methods n n APM (single domain pair) MDCinfer (multi domain pair): APMM, LPMM Results of computational experiments Conclusion

BMC Bioinformatics, 8: 391, 2007 Multiple Domain Interactions and Diverse Organisms n n Multiple domain pair interactions Datasets with different reliabilities Sparse structure Negative samples Cooperativity of domains Multiple datasets
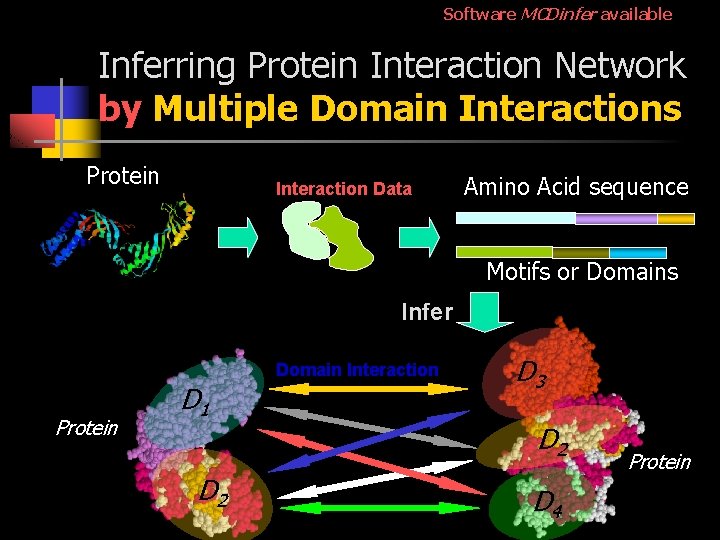
Software MCDinfer available Inferring Protein Interaction Network by Multiple Domain Interactions Protein Interaction Data Amino Acid sequence Motifs or Domains Infer Protein D 1 D 2 Domain Interaction D 3 D 2 D 4 Protein
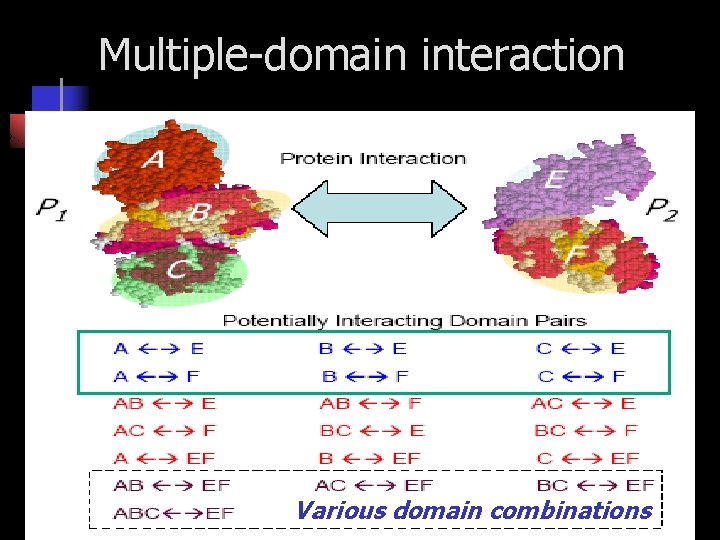
Multiple-domain interaction Various domain combinations
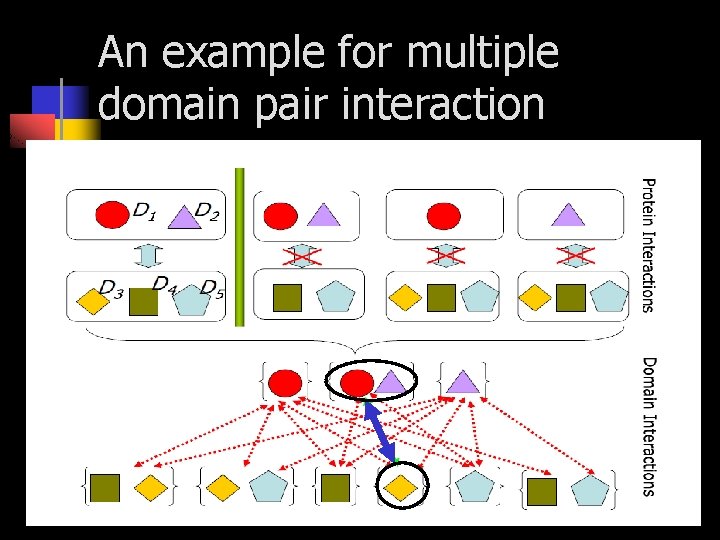
An example for multiple domain pair interaction
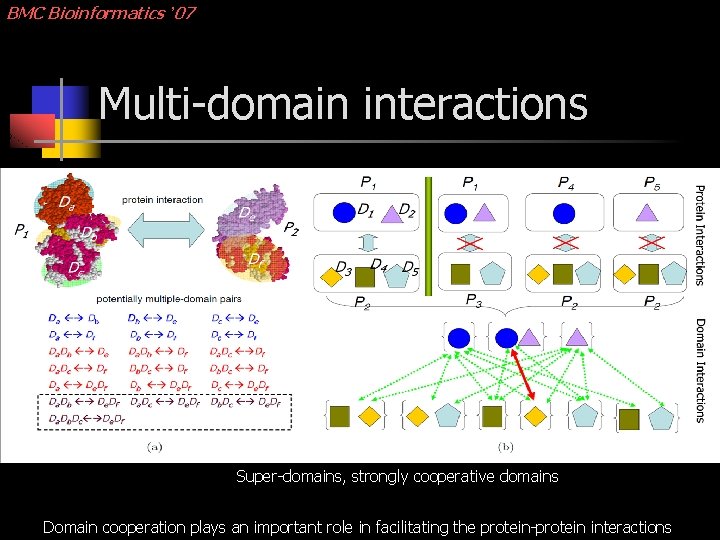
BMC Bioinformatics ’ 07 Multi-domain interactions Super-domains, strongly cooperative domains Domain cooperation plays an important role in facilitating the protein-protein interactions
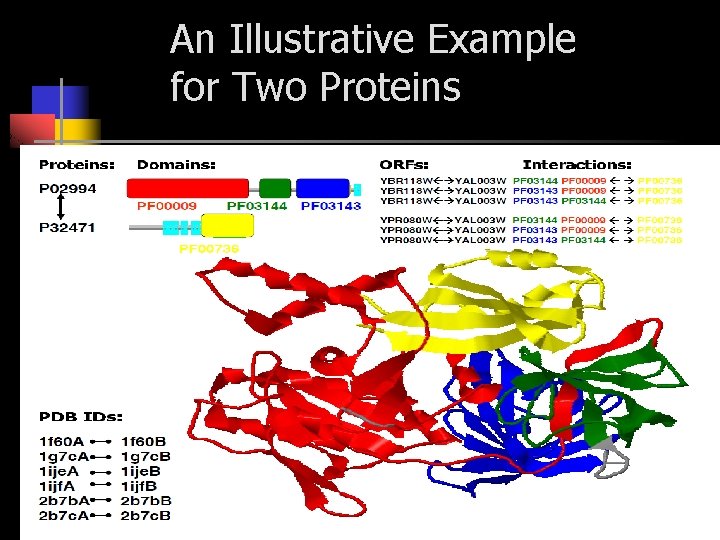
An Illustrative Example for Two Proteins

Probabilistic Model for Multiple Domain Interactions up to three domains. All sets have no overlap.

Data integration Datasets with different reliabilities and various species To explore available information fnk: rate of false negative sample for the k-th dataset or experiment fpk: rate of false positive sample for the k-th dataset or experiment

Equivalently transform nonlinear expression to a linear form Formulation in linear form or

Linear Programming with Sparse Structure

Overview n n n Background Probabilistic model Related work Biological experimental data Proposed methods n n APM (single domain pair) MDCinfer (multi domain pair): APMM, LPMM Results of computational experiments Conclusion

Multi-Domain Cooperation n Cooperative domain pair: it has stronger interaction effect than the corresponding two-domain pairs Strong cooperative domain pair: there is an interaction effect only if the domain pair appear together Super domain pair: the two domains always appear together in individual proteins

Identification of Superdomain pairs

Strong Cooperative Domain Pairs

Cooperative Domain Pairs

Verification by Crystal Structures

PPI Prediction by Multi-domains Cooperation




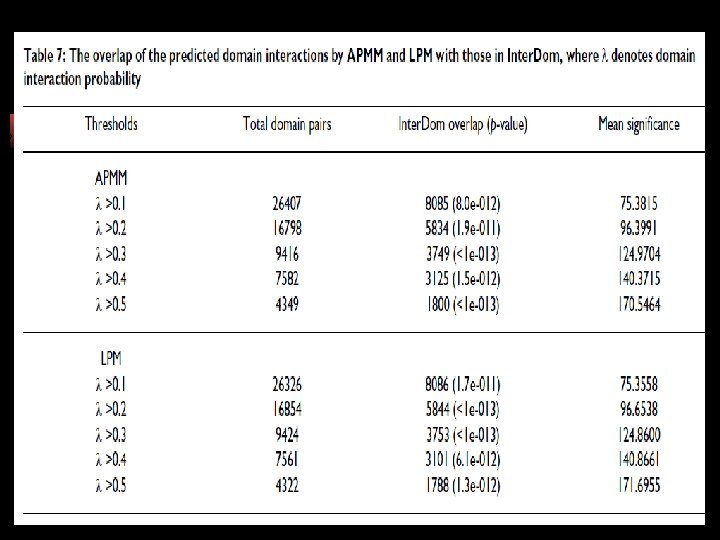
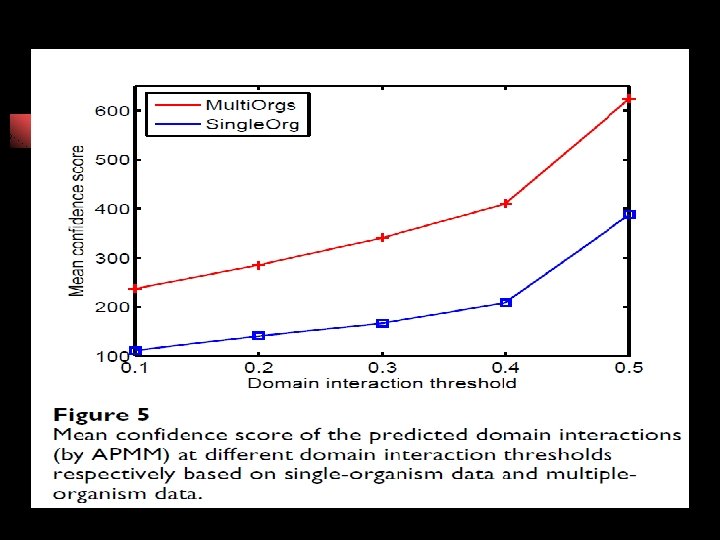
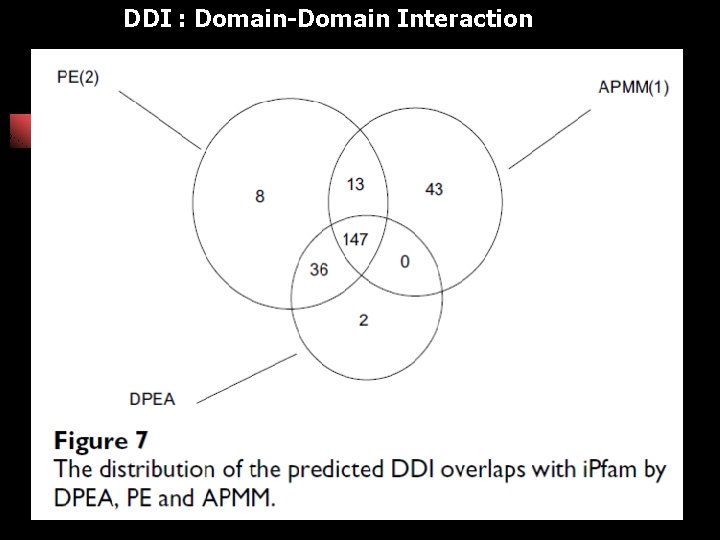
DDI : Domain-Domain Interaction


Reconstructing Complexes, Pathways 1. From TAP-MS, identify total proteins in the complex 2. according to Pfam and protein sequences, all possible domains for each protein in terms of Pfam architectures can be obtained. 3. cooperative domain interactions are obtained by performing our approach on protein interaction data. 4. physical interactions of those proteins in the complex can be predicted at the protein level. 5. the interactions between protein pairs are further examined at the domain levels based on complex structure information. By further combining a protein docking procedure, the interactions at atomic level can be identified, which makes it possible to construct the coherent crystal structure of a complex.
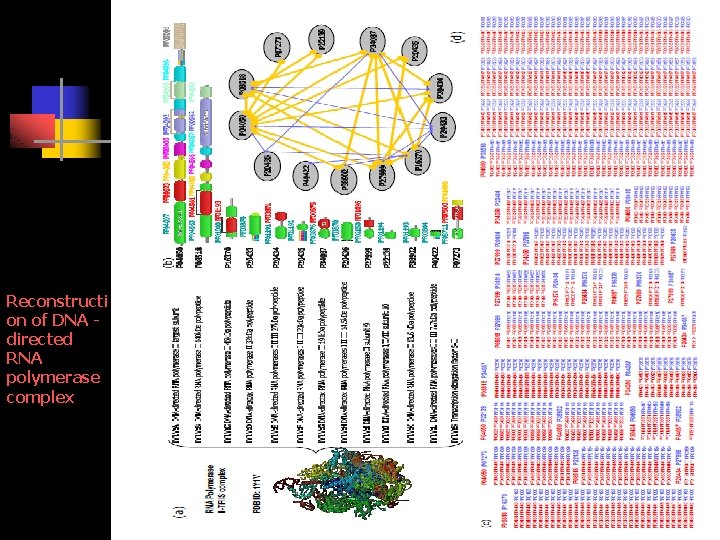
Reconstructi on of DNA directed RNA polymerase complex

Advantages for New Method n n n Multiple domain pair interactions Datasets with different reliabilities Sparse structure Negative samples Various domains
References n n n R. Wang, Y. Wang, L-Y. Wu, X-S. Zhang, L. Chen. BMC Bioinformatics, 8: 391, 2007. L. Chen, L-Y. Wu, Y. Wang, X-S. Zhang. Proteins, 62, 833 -837, 2006. X. Zhao, L. Chen, K. Aihara. Proteins, 10. 1002/prot. 21870, 2007. L. Chen, L-Y. Wu, Y. Wang, S. Zhang, X-S. Zhang, BMC Structural Biology, doi: 10. 1186/1472 -6807 -618, 2006. X. Zhao, Y. Wang, L. Chen, K. Aihara. Proteins, 2008. Z-P. Liu, L. -Y. Wu, Y. Wang, L. Chen, X. S. Zhang. BMC Bioinformatics, 8: 475, 2007.

Acknowledgment n n Chinese Academy of Sciences: Prof. XS. Zhang, Drs. Y. Wang, L-Y. Wu, etc. University of Tokyo: Prof. K. Aihara, Dr. XM. Zhao, etc. Shanghai University: Dr. R-Q. Wang, etc. Osaka Sangyo University: Dr. R-S. Wang, Dr. Zhiping Liu
- Slides: 70