Feature Selection via Block Regularized Regression Seyoung Kim

Feature Selection via Block Regularized Regression Seyoung Kim and Eric Xing Carnegie Mellon University

Block-Regularized Regression • Select a small number of covariates relevant to the output variable in the presence of a large number of irrelevant covariates • Exploit prior knowledge on stochastic block structure in covariates when the covariates are linearly ordered, and look for blocks of relevant covariates
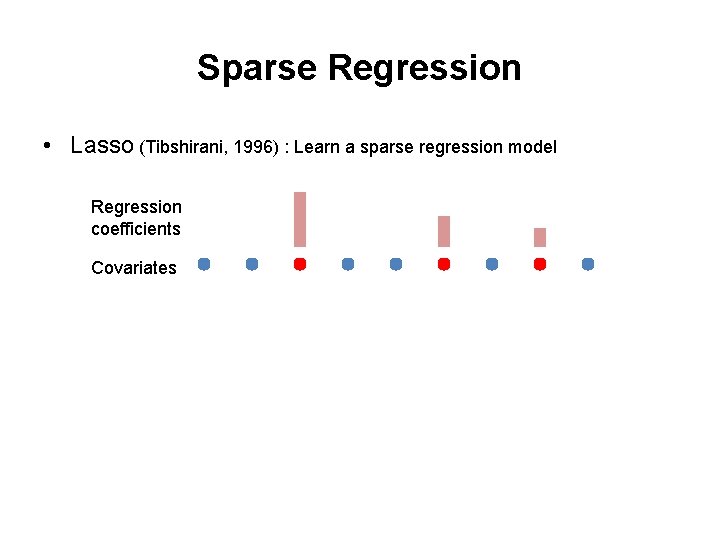
Sparse Regression • Lasso (Tibshirani, 1996) : Learn a sparse regression model Regression coefficients Covariates
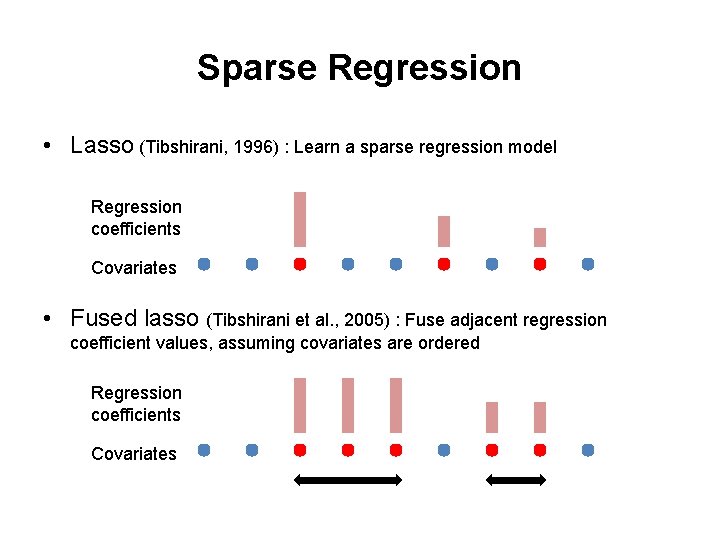
Sparse Regression • Lasso (Tibshirani, 1996) : Learn a sparse regression model Regression coefficients Covariates • Fused lasso (Tibshirani et al. , 2005) : Fuse adjacent regression coefficient values, assuming covariates are ordered Regression coefficients Covariates
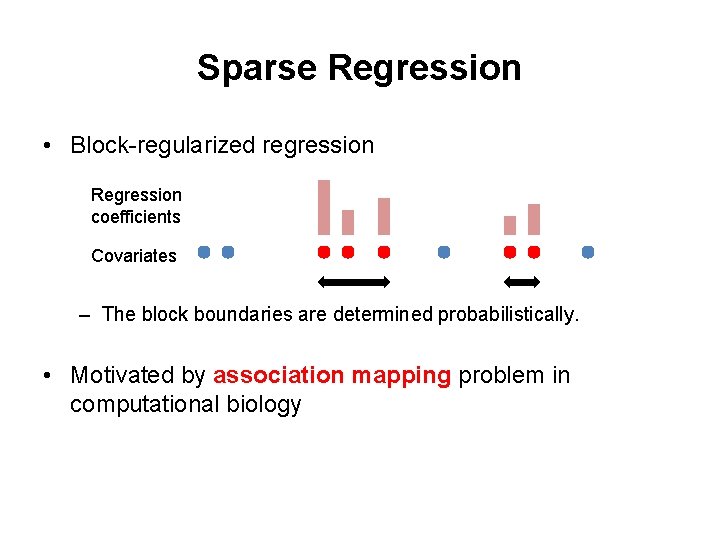
Sparse Regression • Block-regularized regression Regression coefficients Covariates – The block boundaries are determined probabilistically. • Motivated by association mapping problem in computational biology

Single Nucleotide Polymorphism (SNP) Genotypes of chromosome pair Individual 1 ATCGATCCATACAATTTACTATT ATGGATCCATAGAATTTACAATT ATCGATCCTTACAATTTACTATT … Individual 2 ATCGATCCTTACAATTTACTATT Individual N ATGGATCCTTACAATTTACTATT ATGGATCCTTAGAATTTACTATT
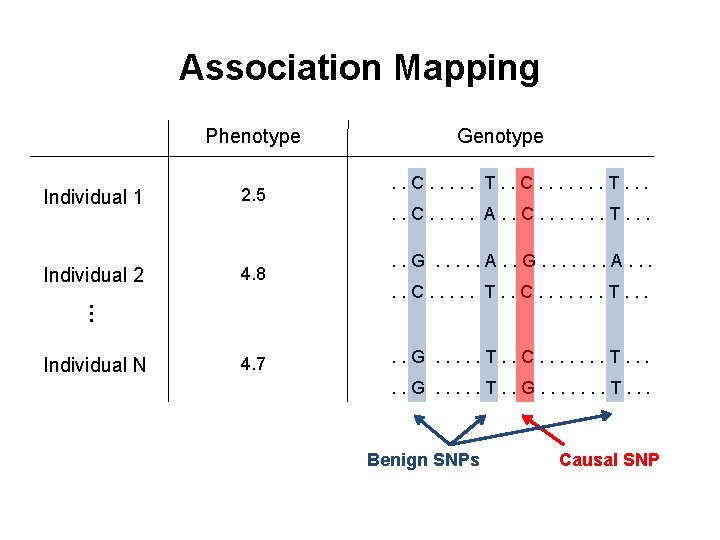
Association Mapping Phenotype 2. 5 Individual 2 4. 8 . . C. . . T. . C. . . . T. . . C. . . A. . C. . . . T. . . G. . . A. . G. . . . A. . . C. . . T. . C. . . . T. . . … Individual 1 Genotype Individual N 4. 7 . . G. . . T. . C. . . . T. . . G. . . T. . G. . . . T. . . Benign SNPs Causal SNP
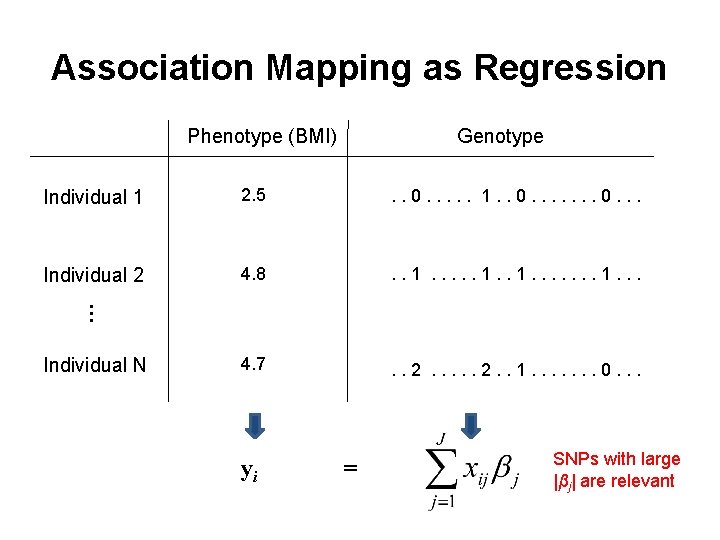
Association Mapping as Regression Phenotype (BMI) Genotype 2. 5 . . 0. . . 1. . 0. . . Individual 2 4. 8 . . 1. . . 4. 7 . . 2. . 1. . . . 0. . . … Individual 1 Individual N yi = SNPs with large |βj| are relevant

Sparse Regression • Ridge regression, lasso with L 1 penalty, etc. • Problem: block structure in genome arising from a nonrandom recombination process
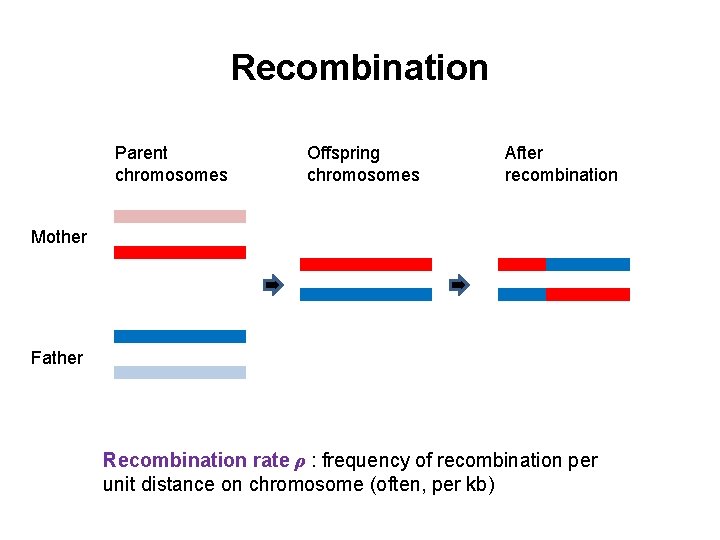
Recombination Parent chromosomes Offspring chromosomes After recombination Mother Father Recombination rate ρ : frequency of recombination per unit distance on chromosome (often, per kb)
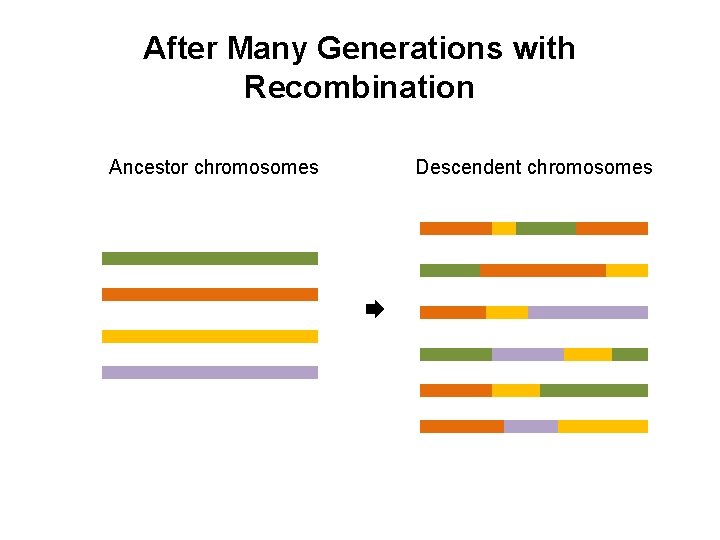
After Many Generations with Recombination Ancestor chromosomes Descendent chromosomes
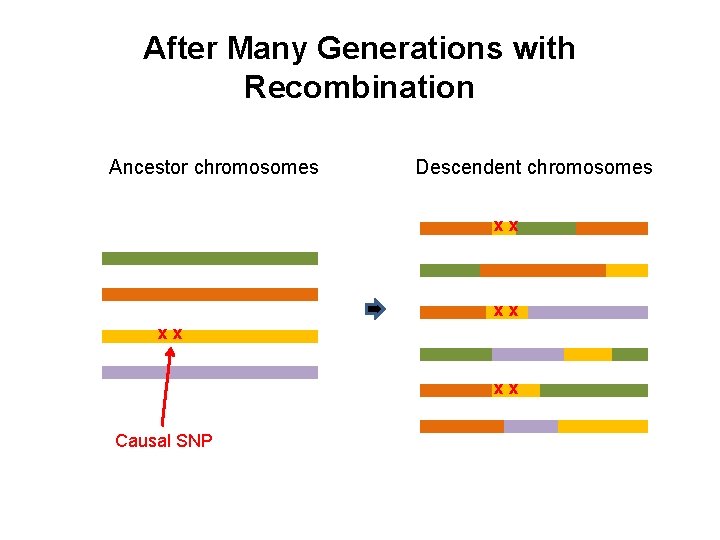
After Many Generations with Recombination Ancestor chromosomes Descendent chromosomes xx xx Causal SNP

Variable Selection Methods for Association Mapping • Bayesian variable selection (George and Mc. Culloch, 1993, Ishwaran and Rao, 2005) – Without the block structure • Block-regularized regression – Explicitly model the dependencies in SNPs (covariates)
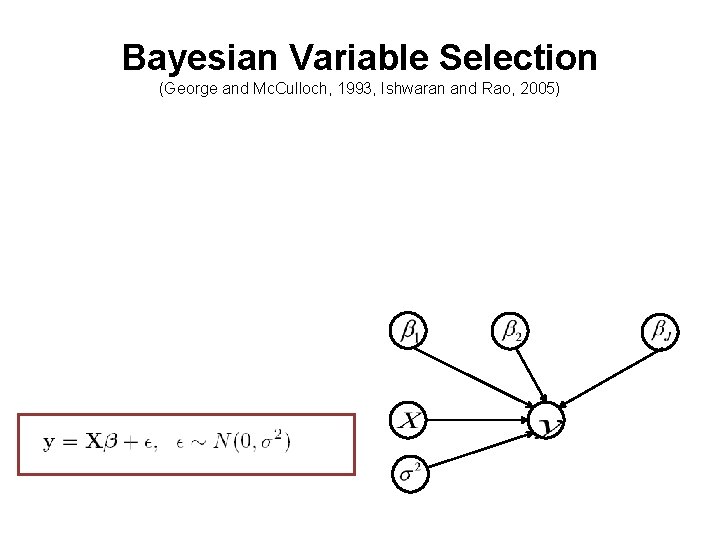
Bayesian Variable Selection (George and Mc. Culloch, 1993, Ishwaran and Rao, 2005)
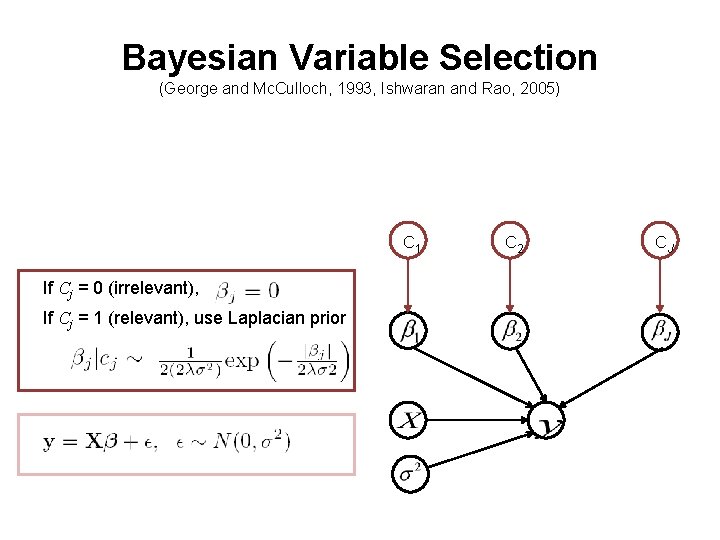
Bayesian Variable Selection (George and Mc. Culloch, 1993, Ishwaran and Rao, 2005) C 1 If Cj = 0 (irrelevant), If Cj = 1 (relevant), use Laplacian prior C 2 CJ
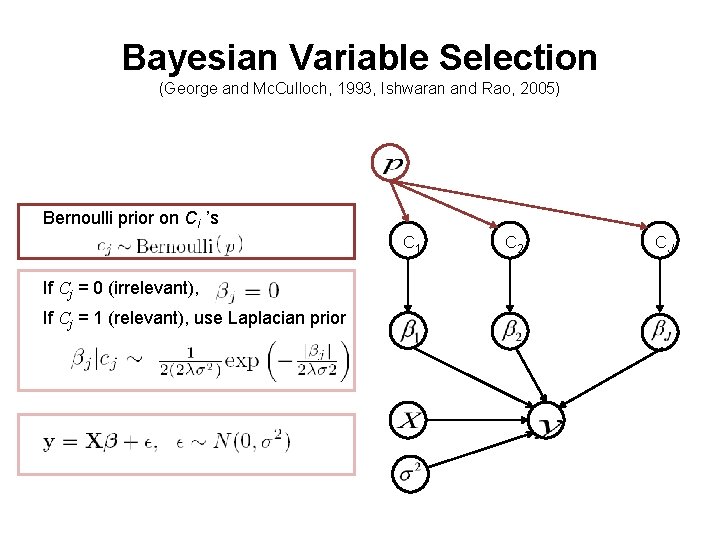
Bayesian Variable Selection (George and Mc. Culloch, 1993, Ishwaran and Rao, 2005) Bernoulli prior on Cj ’s C 1 If Cj = 0 (irrelevant), If Cj = 1 (relevant), use Laplacian prior C 2 CJ

Block-regularized Regression with Markov Chain Prior
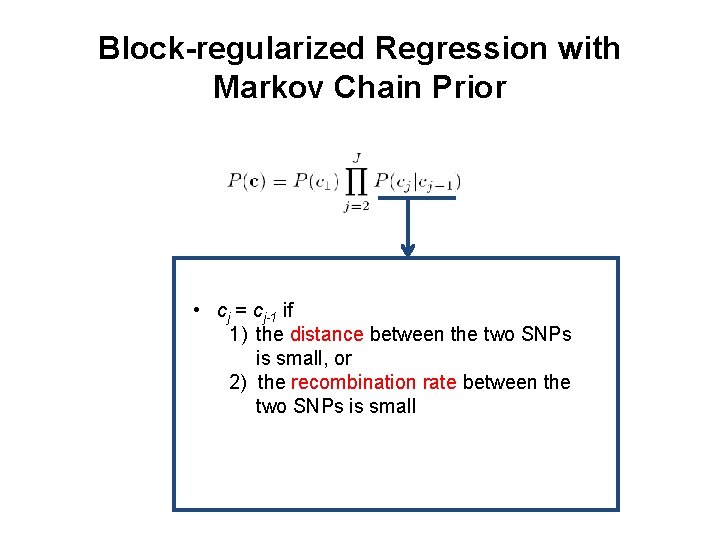
Block-regularized Regression with Markov Chain Prior • cj = cj-1 if 1) the distance between the two SNPs is small, or 2) the recombination rate between the two SNPs is small
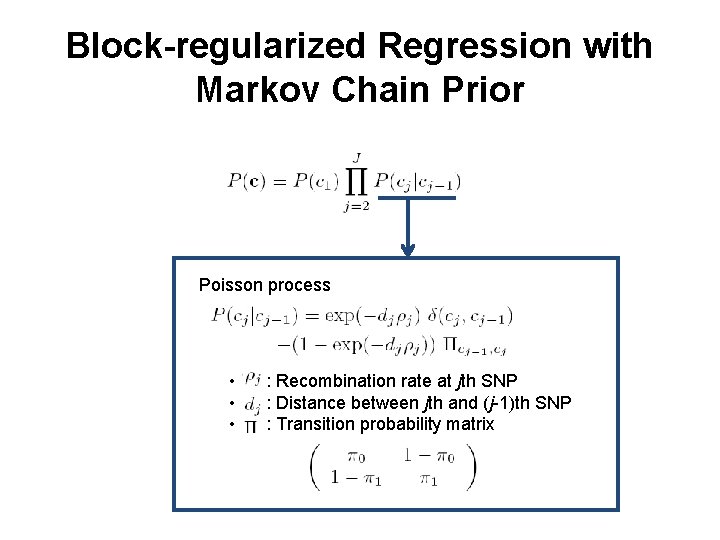
Block-regularized Regression with Markov Chain Prior Poisson process • • • : Recombination rate at jth SNP : Distance between jth and (j-1)th SNP : Transition probability matrix
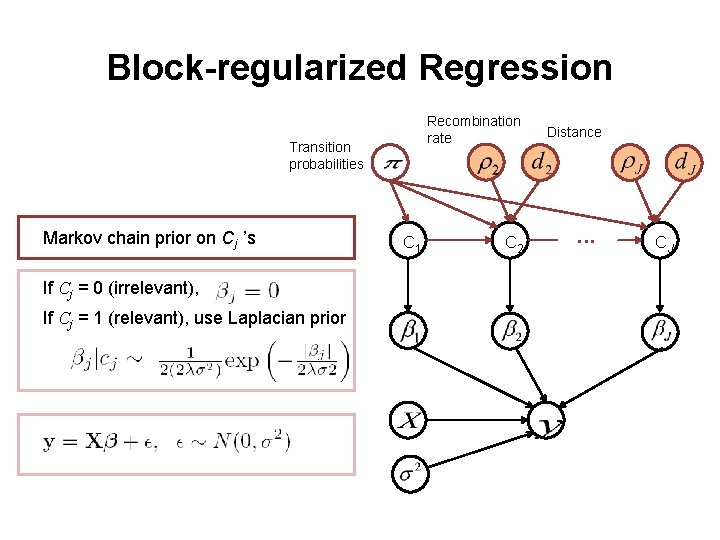
Block-regularized Regression Recombination rate Transition probabilities Markov chain prior on Cj ’s If Cj = 0 (irrelevant), If Cj = 1 (relevant), use Laplacian prior C 1 C 2 Distance . . . CJ
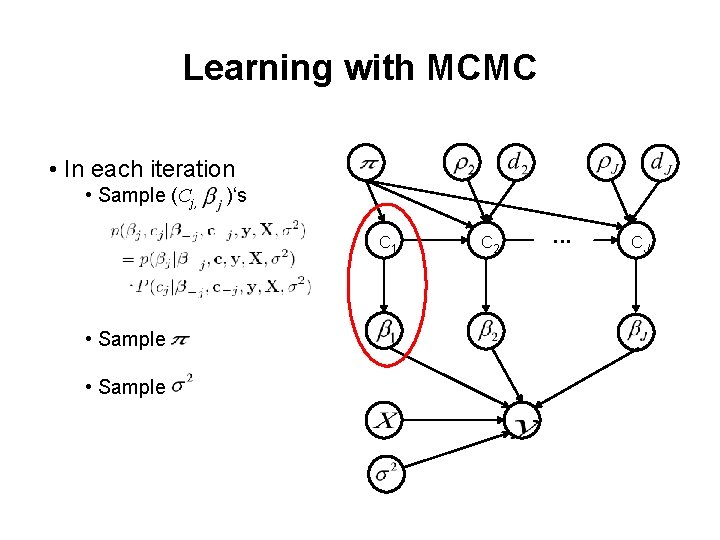
Learning with MCMC • In each iteration • Sample (Cj, )‘s C 1 • Sample C 2 . . . CJ

Experiments • Simulation study – Comparison with • Bayesian variable selection with independent Bernoulli prior • Lasso • Ridge regression – Simulate covariates from ms (Hudson, 2002) – Estimate recombination rates using PHASE (Li and Stephens, 2004) – 10 relevant SNPs out of 100 -250 SNPs – 180 individuals – MCMC sampling for 5000 iterations after 2000 burn-in • Mouse dataset
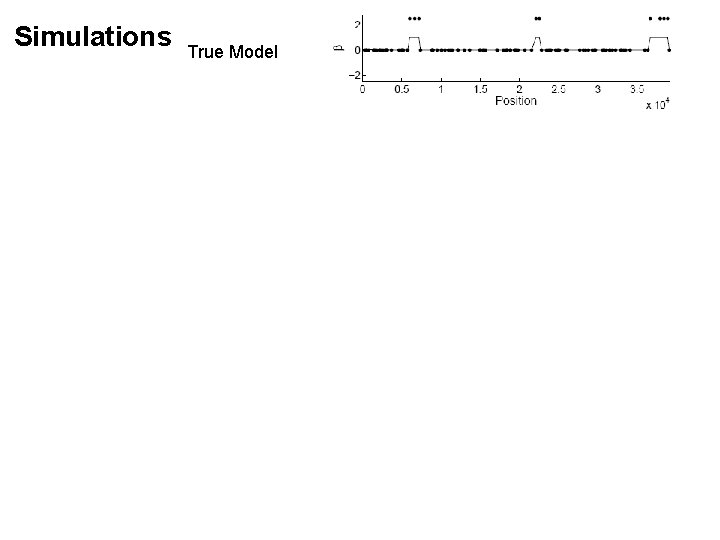
Simulations True Model
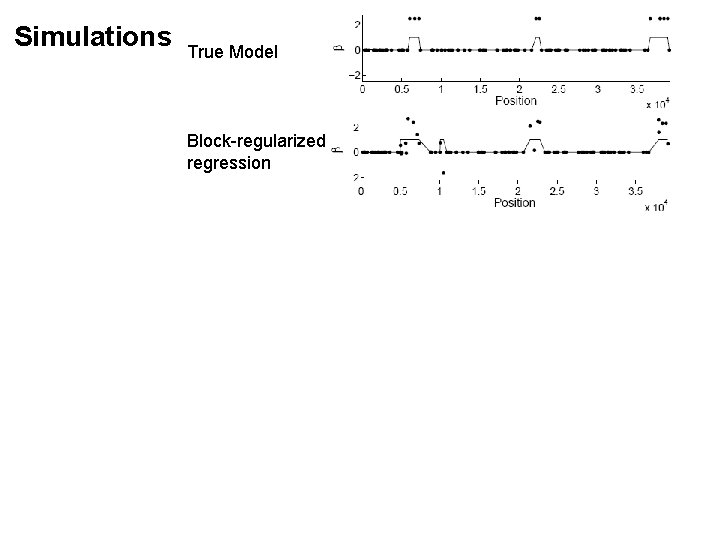
Simulations True Model Block-regularized regression
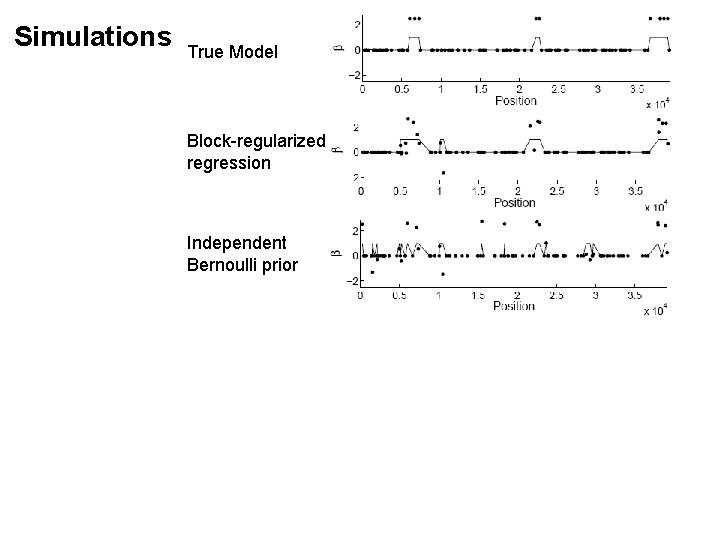
Simulations True Model Block-regularized regression Independent Bernoulli prior
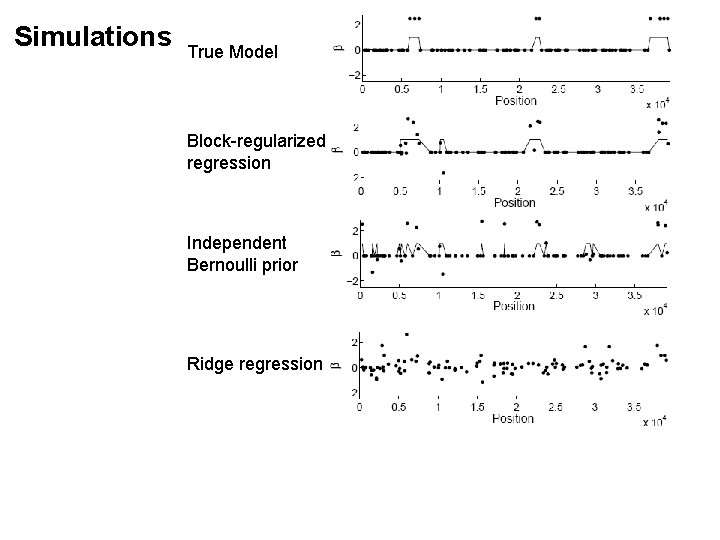
Simulations True Model Block-regularized regression Independent Bernoulli prior Ridge regression
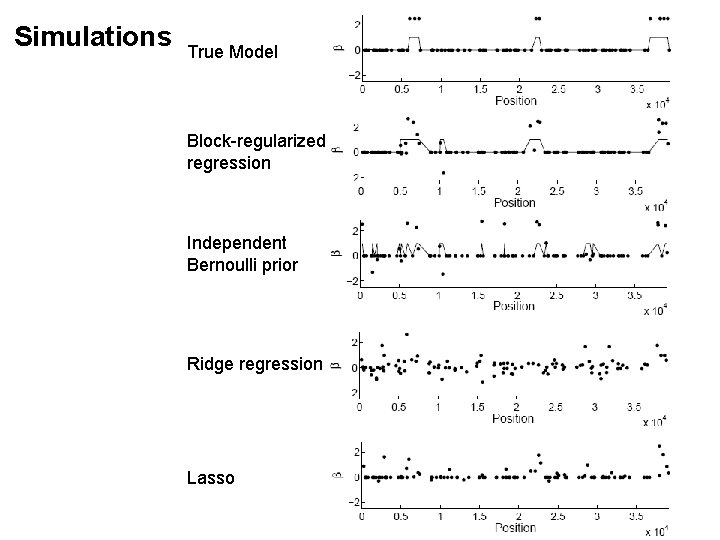
Simulations True Model Block-regularized regression Independent Bernoulli prior Ridge regression Lasso
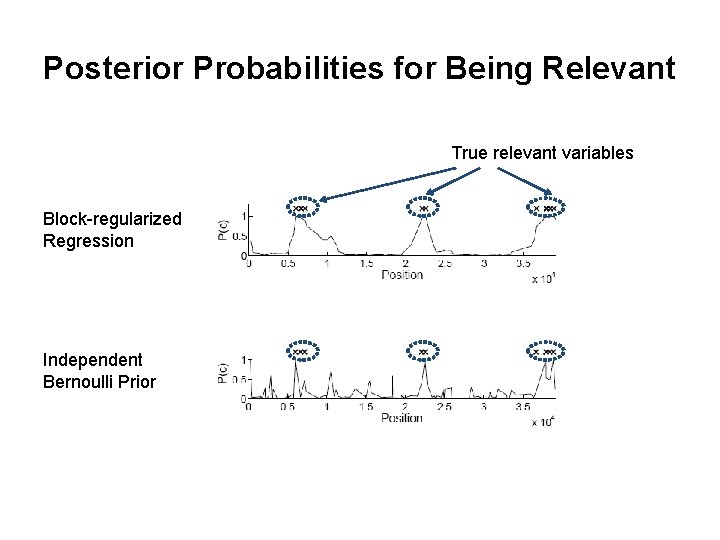
Posterior Probabilities for Being Relevant True relevant variables Block-regularized Regression Independent Bernoulli Prior
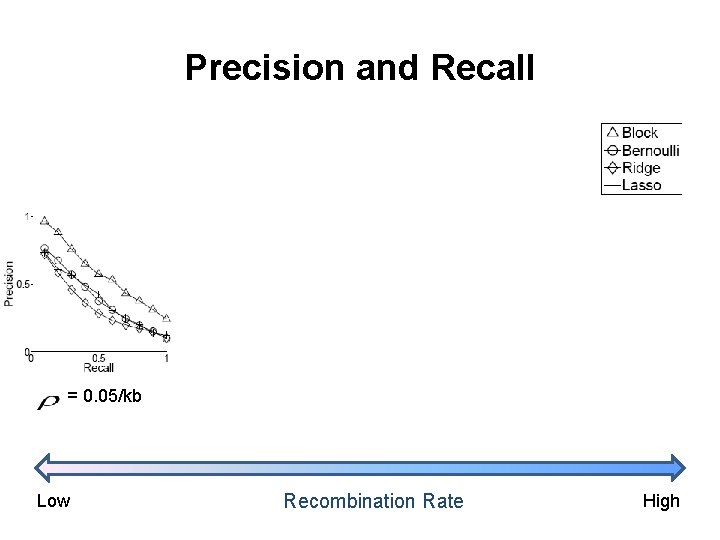
Precision and Recall = 0. 05/kb Low Recombination Rate High
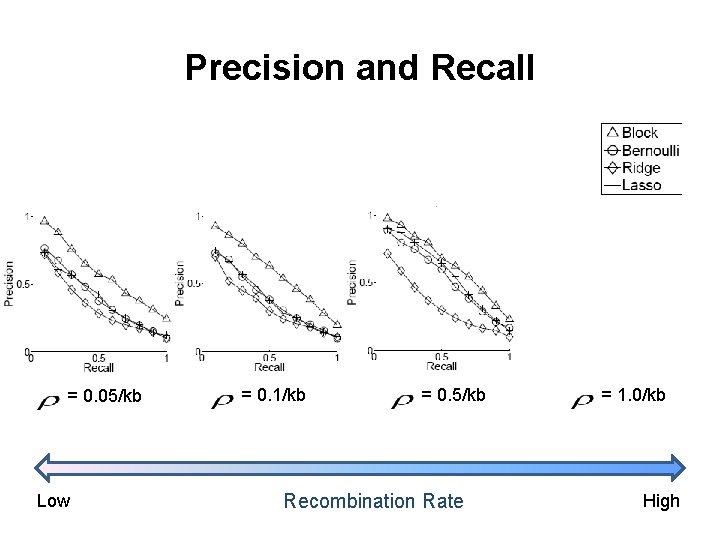
Precision and Recall = 0. 05/kb Low = 0. 1/kb = 0. 5/kb Recombination Rate = 1. 0/kb High
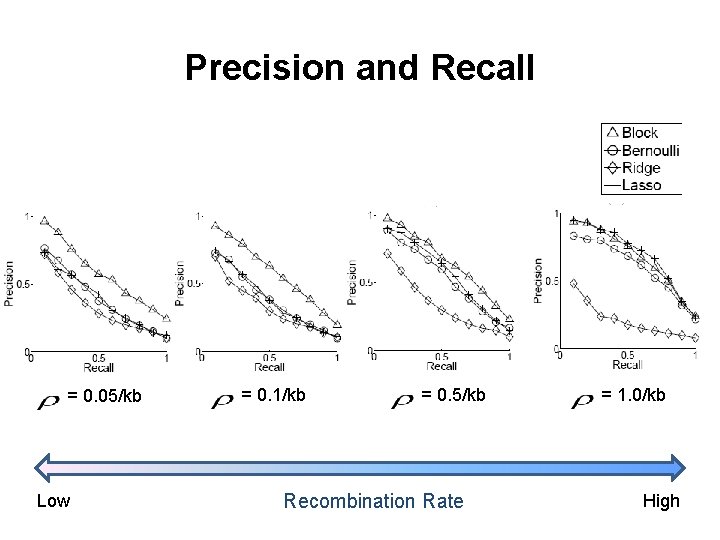
Precision and Recall = 0. 05/kb Low = 0. 1/kb = 0. 5/kb Recombination Rate = 1. 0/kb High
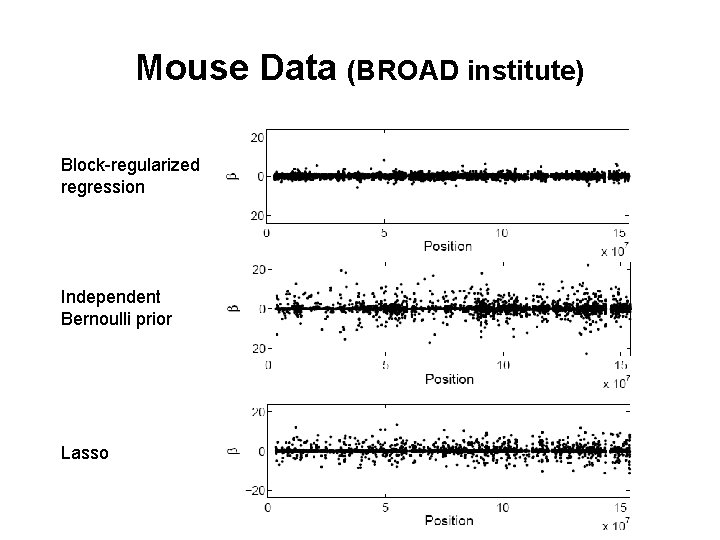
Mouse Data (BROAD institute) Block-regularized regression Independent Bernoulli prior Lasso

Conclusions • Summary – Block-regularized regression makes use of the prior knowledge on the block structure such as distance and recombination rate between adjacent SNPs. – Block-regularized regression finds blocks of relevant SNPs in association mapping. • Future Work – Generalize the Markov chain prior to a Markov random field with an arbitrary structure to model long-term dependencies in relevant SNPs

References • • • E. I. George and R. E. Mc. Culloch (1993). Variable selection via Gibbs sampling. Journal of the American Statistical Association 88: 881 -889. H. Ishwaran and J. S. Rao (2005). Spike and slab variable selection: frequentist and Bayesian strategies. The Annals of Statistics 33(2): 730773. R. Tibshirani (1996). Regression shrinkage and selection via the lasso. Journal of Royal Statistical Society, Series B 58(1): 267 -288. R. Tibshirani, M. Saunders, S. Rosset, J. Zhu, and K. Knight (2005). Sparsity and smoothness via the fused lasso. Journal of Royal Statistical Society, Series B 67(1): 91 -108. M. Yuan and Y. Lin (2006). Model selection and estimation in regression with grouped variables. Journal of Royal Statistical Society, Series B 68(1): 49 -67. N. Li and M. Stephens (2003). Modeling linkage disequilibrium, and identifying recombination hotspots using SNP data. Genetics 165: 22132233.
- Slides: 34